-
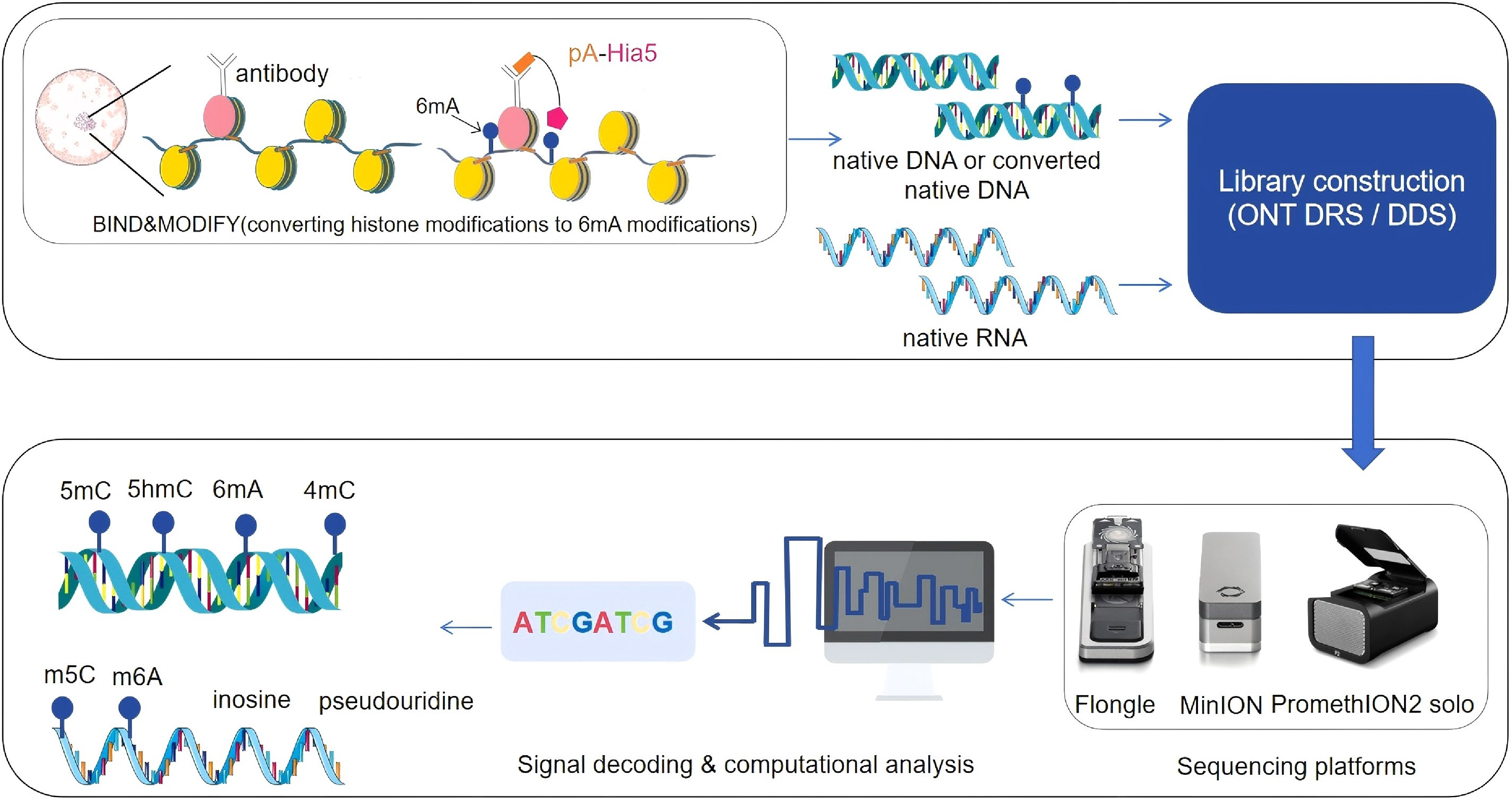
Figure 1.
A New paradigm shift for epigenetic profiling with ONT DDS/DRS, for simultanoeusly profiling DNA methylation and histone modifications at single molecular level, antibodies against histone modifications and proteinA fused with 6 mA methyltransferase to convert histone modifications to 6 mA marked genomic DNA, then the extracted DNA with BIND&MODIFY treated or without treated native DNA or native RNA can be subjected to library preparation, following sequencing platform (Flongle, MinION or PromethION) sequencing, the acquired ionic current signals can be decoded to the sequences of DNA or RNA with native modifications. The drawing for the figure got the support from BioIcons techniques and the images of sequencers are from Oxford Nanopore Technologies.
-
Epigenetic parameter ONT direct sequencing (DDS/DRS) Traditional technologies
(WGBS/ChIP-seq/meRNA-IP, etc.)DNA methylation Advantages • Simultaneous base-resolution detection of 5 mC, 5 hmC, 6 mA[40] • Gold standard for 5mC (bisulfite sequencing) • Resolves repetitive regions (e.g., centromeres)[22] • Lower cost per sample • Single-molecule resolution[7,40,41] • Mature bioinformatics pipelines Disadvantages • Requires high-input DNA quality • Limited to 5 mC (requires additional assays for
5 hmC/6 mA)• Basecalling accuracy for modifications needs improvement[16,17] • Poor performance in repetitive regions • Computationally intensive (GPU-dependent)[16] • Bulk averaging masks single-molecule heterogeneity RNA base modifications Advantages • Direct detection without enrichment/conversion[18] • Lower cost • Multi-modification profiling in single reads[16,19] • Tolerates moderate RNA degradation • Single-molecule resolution[18,31] • Established protocols Disadvantages • Technical expertise required • Antibody-dependent (bias risk) • Higher cost per run • Single-modification per experiment • Underdeveloped modification-detection algorithms[16] • Labor-intensive workflow Histone modifications Advantages • BIND&MODIFY: Simultaneous DNA methylation + histone mark profiling[24] • Field standard for histone marks • Single-cell epigenetic state resolution[23] • Lower cost • Direct correlation of histone-DNA crosstalk[23,24] • Mature analysis tools[13] Disadvantages • Requires nuclear isolation & protein engineering • GC bias in sequencing depth • Needs dual-modification algorithm development • Bulk averaging obscures cellular heterogeneity • Histone-to-6 mA conversion step adds complexity • No DNA methylation co-profiling Alternative splicing Advantages • Full-length transcript isoforms[18,31] • Lower cost • Single-molecule splicing patterns • Abundant third-party software • Multi-process integration (splicing + modifications)[31] Disadvantages • Higher cost than short-read RNA-seq • Assembly errors in isoform reconstruction • Data quality-dependent • No direct modification/splicing linkage • Limited statistical tools • Population-averaged data Poly(A) tail length Advantages • Multi-parameter profiling (tail length + splicing/modifications)[18] • Lower cost • Isoform-specific tail length detection • Detects tail heterogeneity • No additional experimental steps • Simple analysis Disadvantages • Higher cost than PAL-seq[42] • Isolated measurement (no multi-process context) • Insensitive to tail heterogeneity • Population-averaged data • Algorithmic limitations in estimation • Isoform-agnostic quantification Non-coding
RNA (ncRNA)Advantages • Full-length lncRNA characterization • Universal ncRNA compatibility • ncRNA processing/modification detection[31] • Standardized protocols Disadvantages • Poly(A)-tail dependency limits non-polyadenylated ncRNAs • Fragmented long ncRNA assembly • Poor small ncRNA performance without adaptations • No direct modification detection • Higher cost • No single-molecule resolution Table 1.
Comparative performance of ONT direct sequencing vs traditional epigenetic technologies.
Figures
(1)
Tables
(1)