-
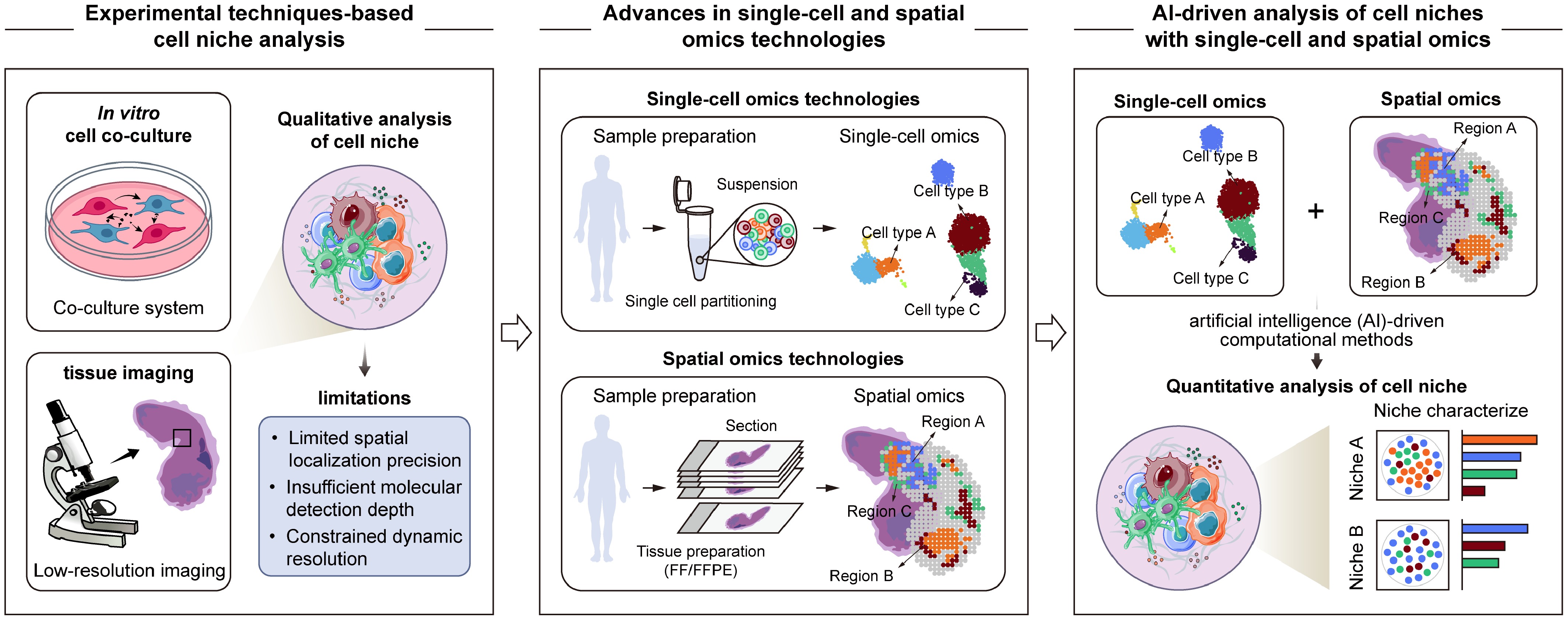
Figure 1.
Overview of advances in cell niche research. With the emergence of single-cell and spatial omics technologies, along with advances in AI-driven computational methods, cell niche research has gradually evolved from early qualitative descriptions based on experimental observations toward quantitative and systematic analysis that integrate multidimensional molecular characteristics and spatial information.
-
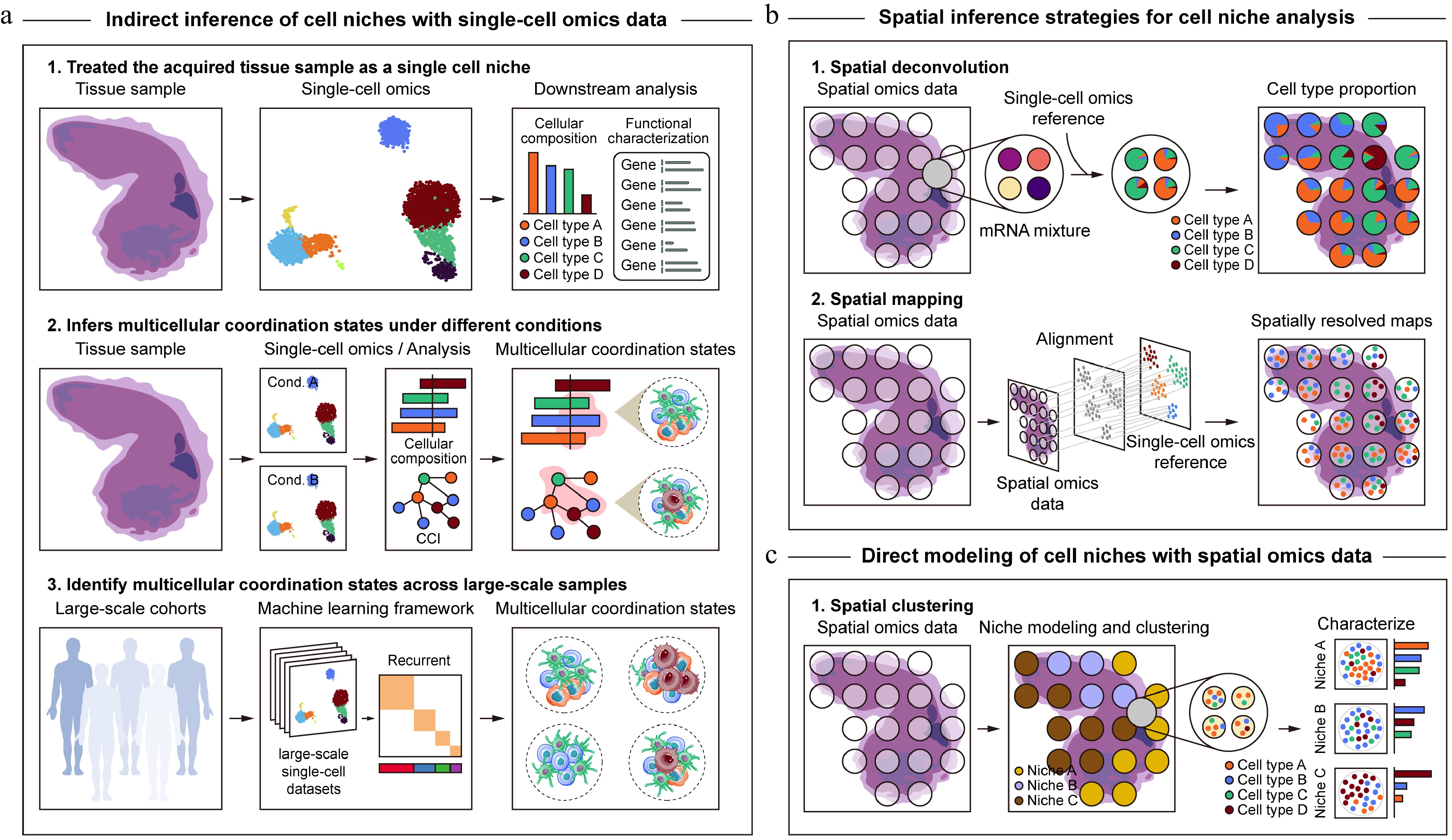
Figure 2.
Schematic pipeline for inferring cell niches from single-cell and spatial omics data. (a) Cell niche analysis based on single-cell omics data. Given the loss of spatial location information, cell niche inference from single-cell omics data mainly relies on systematic characterization of cellular composition and functional states at single-cell resolution, indirect inference of multicellular coordination states through changes in cell type proportions or CCIs, and machine learning-based identification of recurrent multicellular coordination states across large-scale single-cell cohorts. (b), (c) Cell niche analysis based on spatial omics data. In contrast to single-cell omics, spatial omics technologies explicitly preserve spatial context, enabling direct modeling of spatial proximity, cellular composition, and molecular interactions within tissues to delineate cell niches. Existing approaches can be broadly categorized into two conceptual paradigms, where the first (b), focuses on spatial inference of cell niches based on reconstructed high-resolution cellular composition and molecular features, and the second (c), emphasizes direct modeling and identification of cell niches by jointly utilizing the molecular profiles and spatial information from spatial omics data.
-
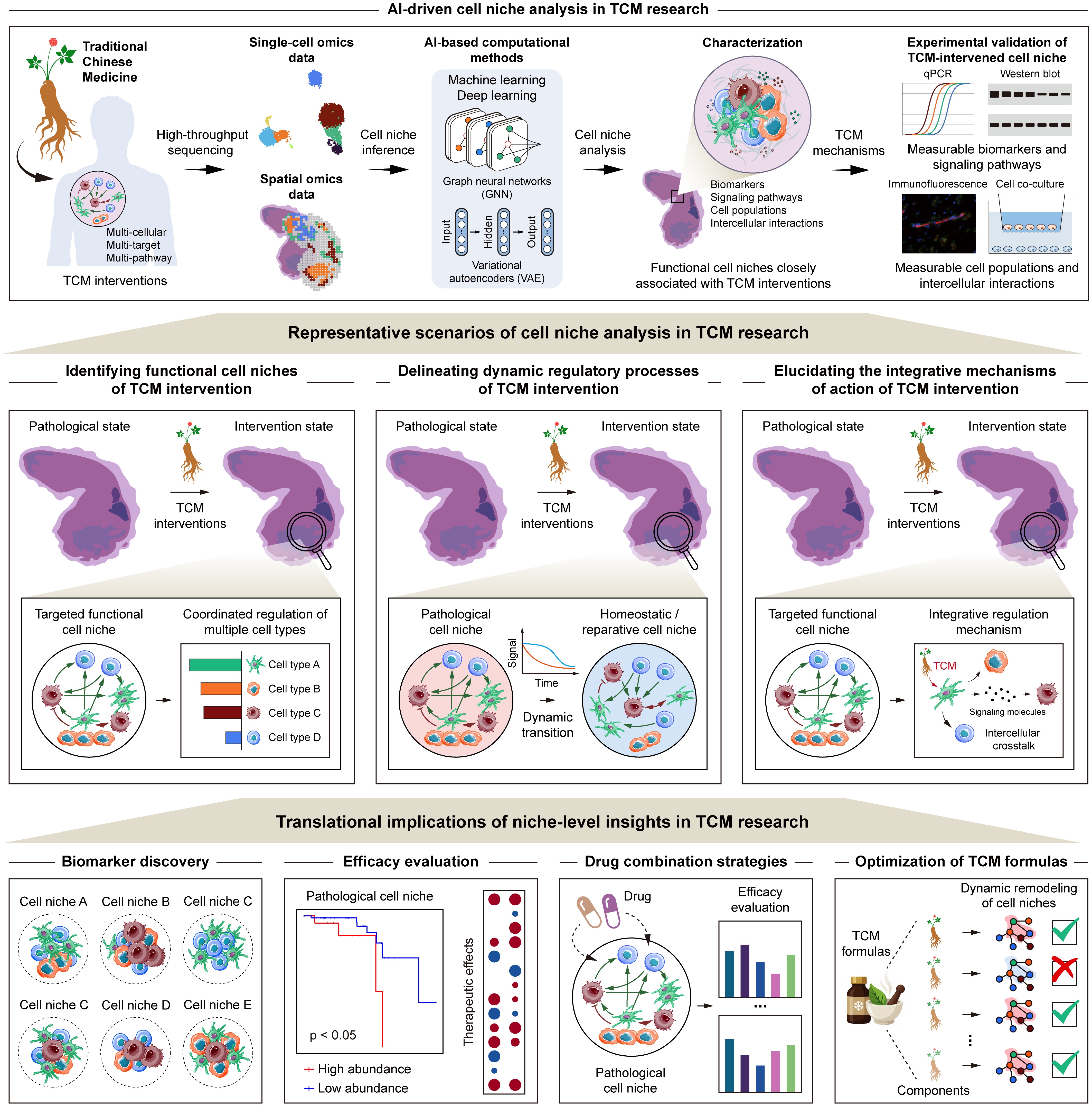
Figure 3.
Representative scenarios and translational implications of AI-driven cell niche analysis in TCM research. AI-driven cell niche analysis provides a systematic framework for investigating the mechanisms of TCM interventions, with three representative scenarios, including identifying functional multi-cellular targets associated with TCM interventions by resolving coordinated cellular architectures within specific spatial contexts, characterizing the dynamic remodeling of pathological cell niches toward homeostatic or reparative states during TCM treatment, and elucidating the integrative mechanisms of action of TCM from a multi-cellular, multi-target, and multi-pathway perspective. The niche-level insights further demonstrate translational implications, offering guidance for the discovery of functional biomarkers, efficacy evaluation, the design of drug combination strategies, and the optimization of TCM formulas.
-
Method Strategy Underlying AI algorithm Language Ref. NMFreg Regression-based NNLS MATLAB [90] SPOTlight Regression-based NNLS R [91] Redeconve Regression-based Regularized NNLS R [92] spatialDWLS Regression-based DWLS R [93] SpatialDecon Regression-based log-normal regression R [94] SONAR Regression-based Poisson-Gamma regression + GWR R [95] Cottrazm Regression-based DWLS R [96] SpaCET Regression-based NNLS R [97] RCTD Probabilistic modeling Poisson distribution with MLE R [98] stereoscope Probabilistic modeling NB distribution with MAP Python [99] cell2location Probabilistic modeling NB distribution with variational Bayesian inference Python [100] Starfysh Probabilistic modeling Poisson-Gamma distribution with variational inference Python [101] STdeconvolve Probabilistic modeling LDA R [102] STRIDE Probabilistic modeling LDA Python [103] Bulk2Space Deep learning-based VAE Python [104] DestVI Deep learning-based VAE Python [105] SPACEL Deep learning-based VAE Python [106] DSTG Deep learning-based GCN R and Python [107] STdGCN Deep learning-based GCN Python [108] SD2 Deep learning-based GCN + Autoencoder R and Python [109] CellDART Deep learning-based ADDA Python [110] spSeudoMap Deep learning-based ADDA Python [111] TransformerST Deep learning-based Transformer Python [112] AntiSplodge Deep learning-based Feedforward neural network Python [113] UCDbase Deep learning-based Feedforward neural network Python [114] LETSmix Deep learning-based Feedforward neural network Python [115] Tangram Deep learning-based Nonconvex optimization Python [116] CARD Other NMF + CAR R [117] CellPie Other NMF + factor analysis Python [118] Seurat Other Enrichment score R [119] CellTrek Other Random forest R [120] SpaOTsc Other OT Python [121] NovoSpaRc Other OT Python [122] TACCO Other OT Python [123] DOT Other OT R [124] Table 1.
AI methods for spatial deconvolution.
-
Method Strategy Underlying AI algorithm Language Ref. Seurat Joint dimension reduction-based CCA + KNN MATLAB [119] LIGER Joint dimension reduction-based NMF + KNN R [128] Harmony Joint dimension reduction-based PCA + KNN R [129] SpaGE Joint dimension reduction-based SVD + KNN Python [130] gimVI Joint dimension reduction-based Joint VAE Python [131] stPlus Joint dimension reduction-based Autoencoder + KNN Python [132] SpatialScope Joint dimension reduction-based Deep generative model Python [133] SpaTrio Mapping matrix-based OT R and Python [134] SIMO Mapping matrix-based OT Python [135] Tangram Mapping matrix-based Nonconvex optimization Python [116] CellTrek Mapping matrix-based Random forest R [120] SpaOTsc Mapping matrix-based OT Python [121] novoSpaRc Mapping matrix-based OT Python [122] CytoSPACE Mapping matrix-based Convex optimization Python [136] Table 2.
AI methods for spatial mapping.
-
Method Strategy Underlying AI algorithm Language Ref. Giotto PGM-based HMRF R and Python [137] DR-SC PGM-based HMRF R [138] BayesSpace PGM-based Fully Bayesian model R [139] BASS PGM-based Potts model with additional hierarchical
modeling structuresR [140] PRECAST PGM-based Potts model + CAR R [141] SPICEMIX PGM-based NMF-HMRF Python [142] UTAG Feature augmentation Linear weighting Python [145] CellCharter Feature augmentation Neighborhood aggregation Python [32] ClusterMap Feature augmentation Neighborhood aggregation Python [146] BANKSY Feature augmentation Linear combination R and Python [147] SpaGCN GNN-based GCN + DEC Python [148] Spatial-MGCN GNN-based Multi-view GCN Python [149] STMGCN GNN-based Multi-view GCN Python [150] STAGATE GNN-based GATE Python [151] STAligner GNN-based GATE Python [152] DSSC GNN-based GATE Python [153] CCST GNN-based DGI Python [154] SpaceFlow GNN-based Spatially regularized DGI Python [155] SPIRAL GNN-based GraphSAGE Python [156] SEDR GNN-based VGAE + DEC Python [157] scNiche GNN-based M-GAE + GFN Python [36] SPACEL GNN-based GCN + DANN Python [106] DeepST GNN-based Denoising autoencoder + VGAE + DANN Python [158] SpaDAC GNN-based graph attention network (GAT) + CNN Python [159] stMDA GNN-based GCN + CNN + VAE Python [160] GraphST GNN-based GCN with SCL Python [161] MERINGUE Improved community detection algorithm-based Expression graph with spatial weighting + Louvain R [164] SCGP Improved community detection algorithm-based Spatial graph with expression weighting + Leiden Python [165] SpatialLeiden Improved community detection algorithm-based Multiple graph + Leiden Python [166] Schürch's method Neighborhood phenotype-driven Neighborhood cell type frequency matrix + k-means Python [167] Spatial-LDA Neighborhood phenotype-driven Neighborhood cell type frequency matrix + Spatial-LDA + k-means Python [168] CytoCommunity Neighborhood phenotype-driven Neighborhood cell type frequency matrix + GNN Python [169] Table 3.
AI methods for spatial clustering.
-
Method Data requirement Inference strategy Interpretability Scalability Robustness Typical application scenario Single-cell-based indirect inference Single-cell data Cell niches are inferred based on multicellular coordination states, without explicit spatial context High; results are directly linked to cell types, conditions, or samples High; scalable to large cohorts High; relatively stable across datasets and parameter settings Preliminary characterization of cell niches, large-scale cohort studies, and cross-condition comparisons Spatial inference strategy: Spatial deconvolution/spatial mapping Single-cell/low-quality spatial data Cell niches are inferred based on reconstructed high-resolution cellular composition and molecular features High; results are driven by biological priors and directly linked to cell types Moderate; depends on inference strategy and
data sizeModerate; affected by batch effects, reference quality, and parameter settings Intuitive and refined descriptions of composition and molecular features of cell niches at the spatial level PGM-based spatial clustering methods Spatial data Moderate to high; clear modeling assumptions and the strong statistical foundation Low to moderate; generally computationally intensive Moderate to high; relatively stable across parameter settings Feature augmentation-based spatial clustering methods Spatial data Moderate to high; interpretability depends on feature design High; generally computationally efficient Moderate to high; simplified modeling assumptions with few parameters, but sensitive to feature design GNN-based spatial clustering methods Spatial data Cell niches are inferred based on direct modeling and unsupervised partitioning by utilizing the molecular profiles as well as spatial information of spatial omics data jointly Low to moderate; results are often difficult to interpret Low to moderate; generally limited by graph size and
GPU memoryLow to moderate; greater modeling flexibility but typically involve numerous parameters and are more prone to overfitting Discovery of previously uncharacterized cell niches
in complex tissues in a
data-driven mannerImproved community detection algorithm-based spatial clustering methods Spatial data Moderate to high; results are interpretable at the community structure level High; generally computationally efficient High; simplified modeling assumptions with few parameters Neighborhood phenotype-driven spatial clustering methods Spatial data High; results are directly linked to cell types and biologically intuitive Moderate to high; depends on inference strategy High; simplified modeling assumptions with few parameters Table 4.
Comparison of AI methods for cell niche analysis.
Figures
(3)
Tables
(4)