-
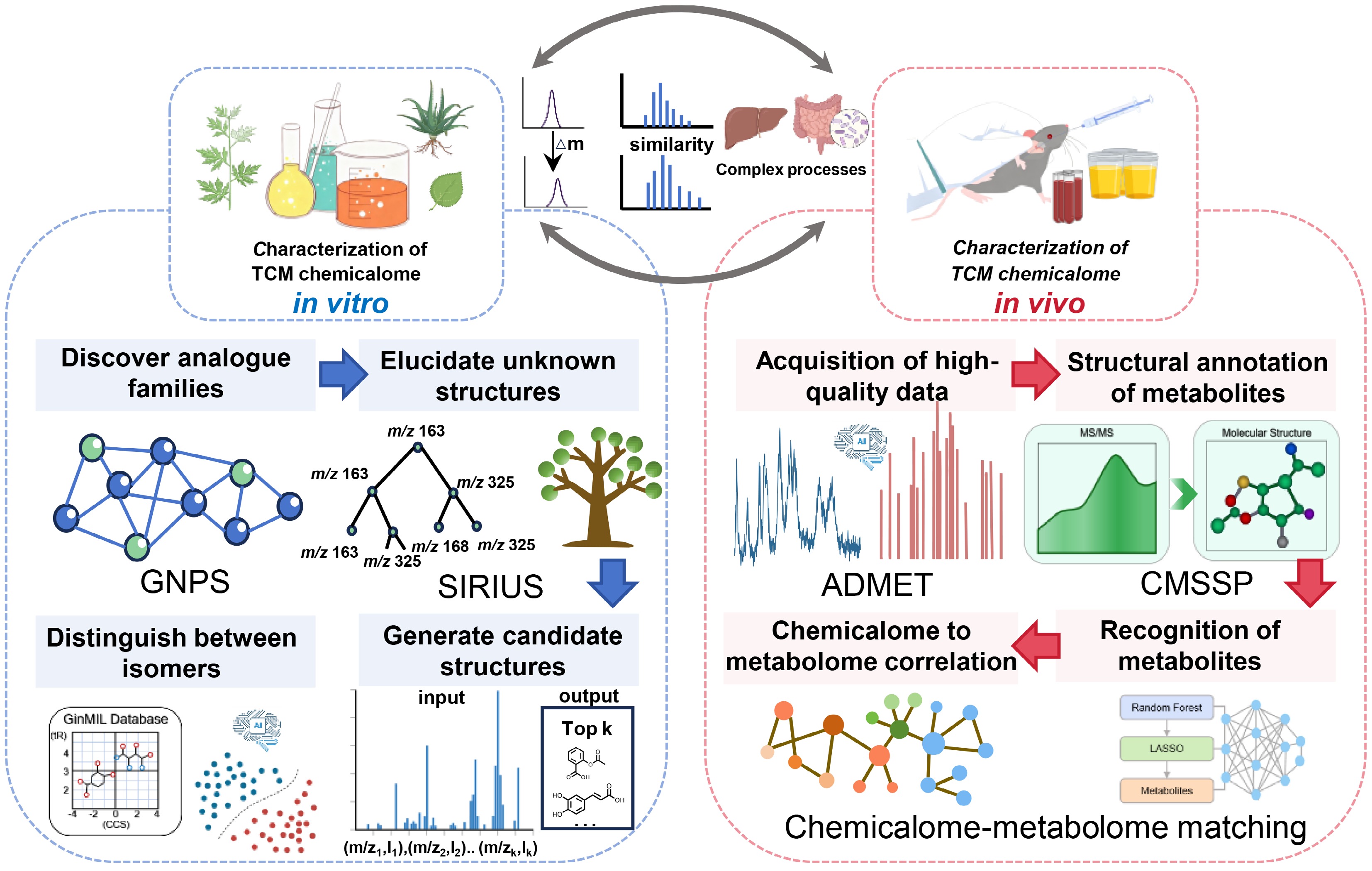
Figure 1.
Schematic diagram for AI-driven revelation of the TCM chemicalome. The diagram delineates comprehensive characterization strategies for the TCM chemicalome, categorized into in vitro (left panel) and in vivo (right panel) approaches. The in vitro workflow integrates GNPS for analog discovery via molecular networking, SIRIUS for structural elucidation using fragmentation tree analysis, and GinMIL for isomer differentiation. The in vivo workflow employs mass spectrometry for high-quality ADMET data acquisition, CMSSP for deep learning-based metabolite annotation, and a chemicalome-metabolome matching approach for the construction of metabolic networks.
-
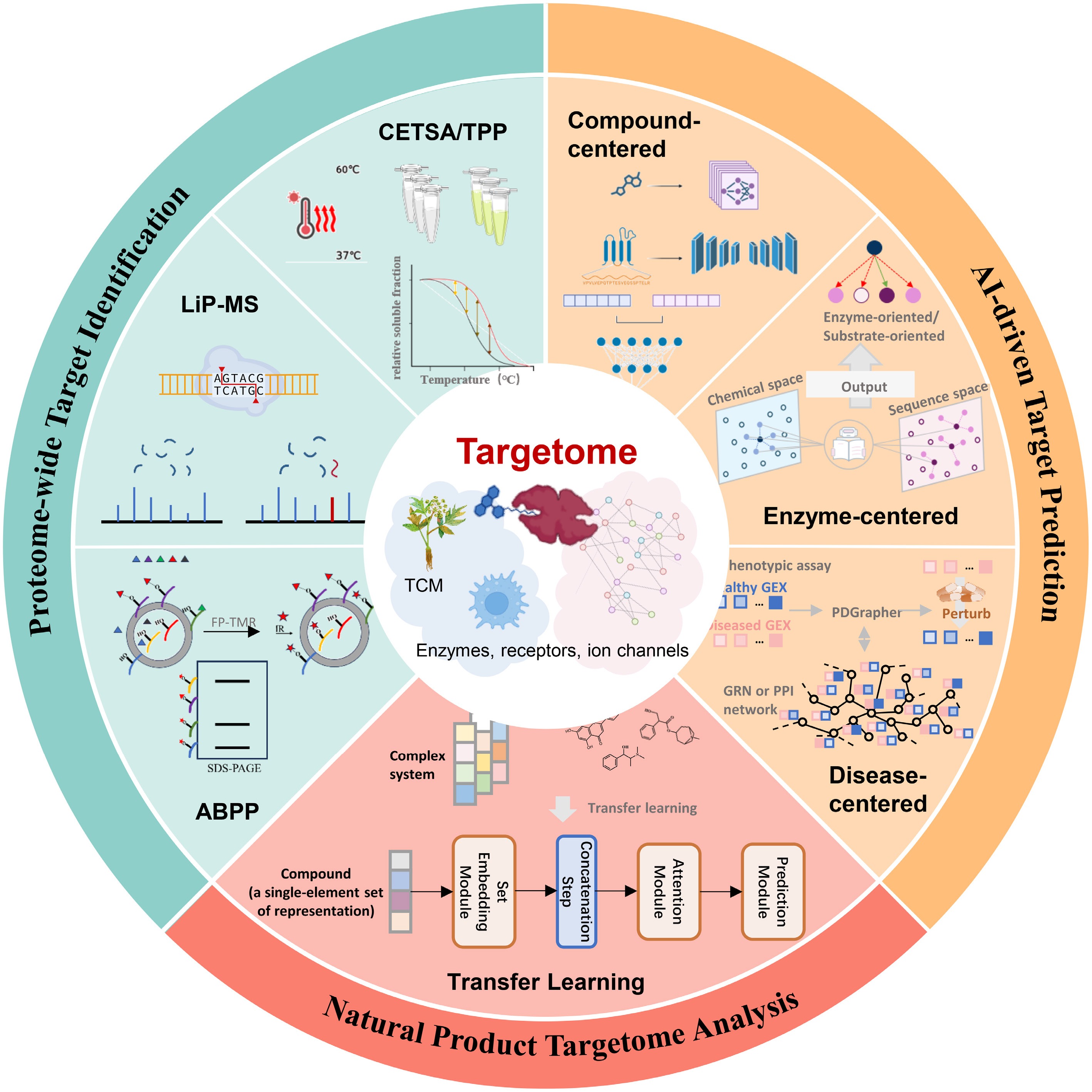
Figure 2.
Schematic diagram for AI-facilitated discovery of the TCM targetome. The diagram integrates three methodological components: AI-driven target prediction by compound-centered, enzyme-centered, and disease-centered methods; proteome-wide target identification by CETSA, TPP, LiP-MS, and ABPP; and natural product targetome analysis by transfer learning models.
-
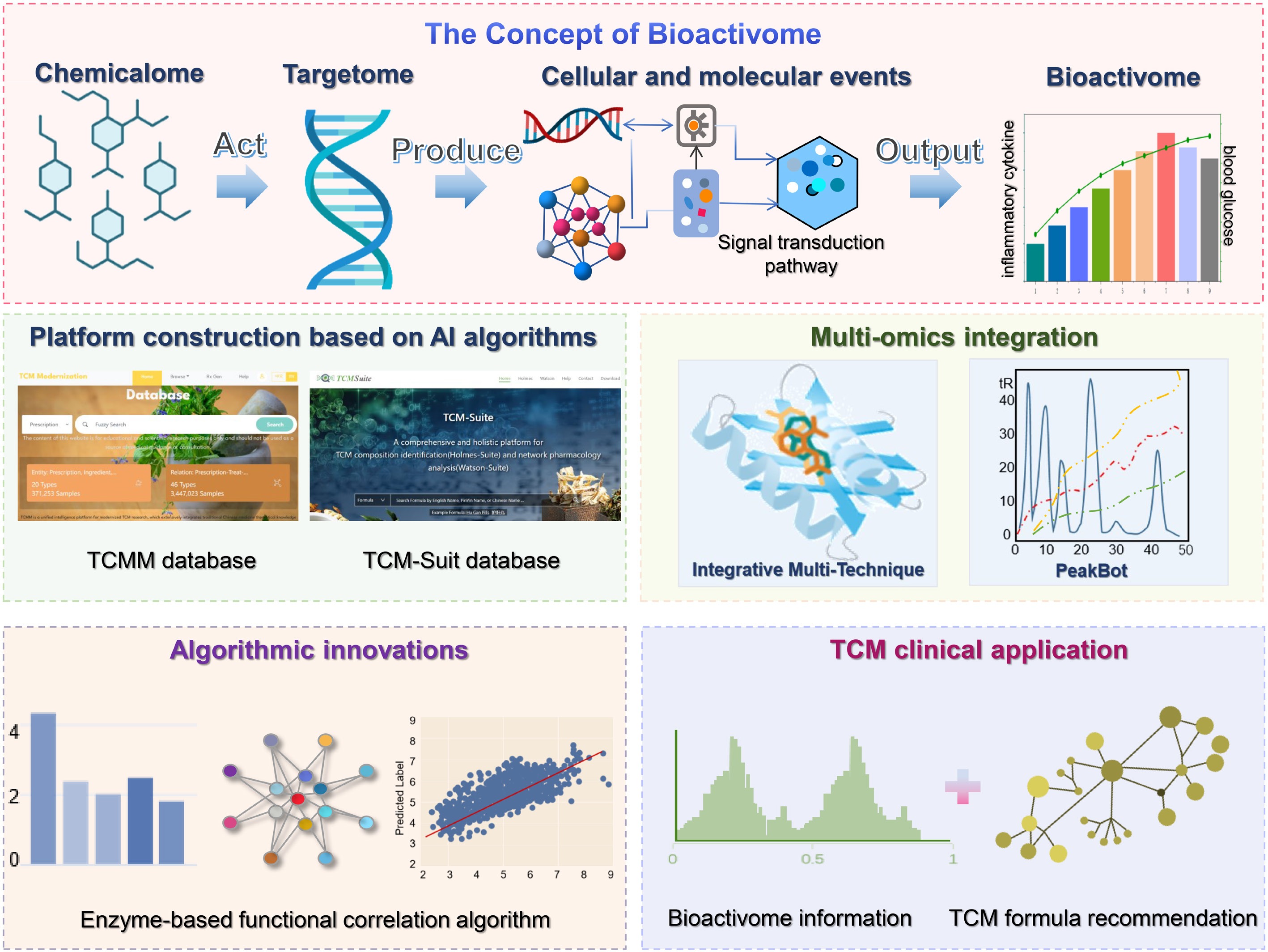
Figure 3.
Schematic diagram for AI-accelerated revelation of the TCM bioactivome. The diagram introduces the concept of the bioactivome, which refers to the set of biological activities derived from the molecular and cellular events induced by the action of the TCM chemicalome on the targetome. Research on the bioactivome mainly focuses on platform construction based on AI algorithms, multi-omics integration, algorithmic innovations, and TCM clinical application.
-
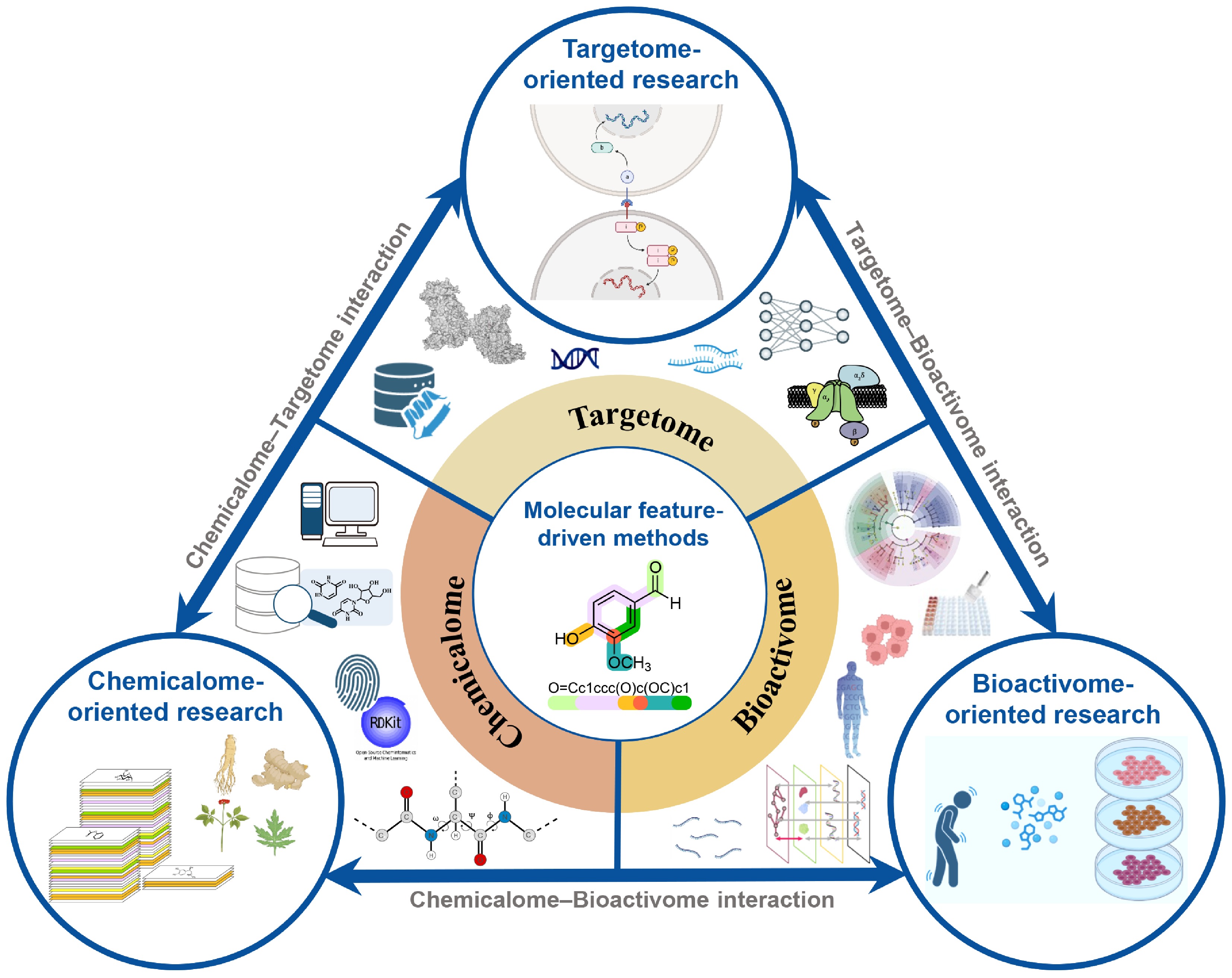
Figure 4.
Schematic diagram for interactions among the chemicalome, targetome, and bioactivome. The framework includes chemicalome-oriented, targetome-oriented, and bioactivome-oriented research to construct their interactions. Molecular feature-driven methods play an important role in this process.
-
Tool name Core technologies Key functionalities Advantages Limitations GNPS Molecular network algorithm, MS/MS fragment clustering, unsupervised learning 1. MS/MS fragment similarity visualization;
2. Structural homology mining;
3. Known compound family screening1. Intuitive MS data visualization;
2. Enables rapid natural product tracing;
3. Accelerates discovery of known TCM compound families1. Reliance on existing MS databases;
2. Low structural annotation accuracy for rare TCM components with unique fragments;
3. Clustering affected by low-quality MS dataSIRIUS Deep learning, fragmentation tree analysis, tandem MS database matching, de novo structural analysis 1. Tandem MS-based unknown structure identification;
2. MS fragment decomposition;
3. Structural similarity search1. Enables computational de novo prediction for unknown TCM components;
2. Integrated strategies enable higher annotation accuracy than single-algorithm tools;
3. Supports batch processing of large-scale TCM MS data1. Reduced de novo accuracy for TCM components with multiple chiral centers;
2. Relies on high-quality tandem MS data;
3. Slow analysis for high-diversity TCM samplesMSNovelist MS-driven de novo molecular structure generation, deep neural network, single MS/MS spectrum interpretation 1. Direct molecular structure deduction from single MS/MS spectrum;
2. Reference-free unknown structure generation1. Breaks dependence on physical references;
2. Requires only a single MS/MS spectrum;
3. Improves novel TCM compound discovery efficiency1. Diverse structural candidates require manual verification;
2. Low accuracy for TCM macromolecules;
3. Poor adaptability for rare TCM componentsGinMIL Machine learning, ion mobility separation, multi-dimensional database construction 1. Comprehensive ginsenoside database construction;
2. Ginsenoside isomer differentiation1. Solves the traditional MS bottleneck in TCM isomer distinction (ginsenosides);
2. Multi-dimensional data fusion enables high component identification accuracy;
3. Specialized for saponin-rich TCM targeted analysis1. Ginsenoside-focused;
2. Relies on ion mobility separation equipment;
3. Unable to analyze unknown saponin isomersSwissADME Machine learning, QSAR model, molecular descriptor calculation 1. Rapid ADME indicator assessment for small molecules 1. Fast prediction for high-throughput TCM active component screening;
2. Predicted indicators align with TCM small-molecule in vivo metabolic characteristics;
3. Visualizable results1. Applicable only to small molecules;
2. Low prediction accuracy for rare TCM components with unique structures;
3. Lacks prediction of in vivo metabolic pathways for TCM componentsADMETlab 2.0 Integrated machine learning, multi-task feature learning, large-scale ADMET database 1. Comprehensive prediction of ADMET indicators;
2. Toxicity/metabolic stability prediction;
3. Drug-drug interaction prediction1. Provides extensive ADMET indicators relevant to TCM research;
2. High accuracy;
3. Supports batch/customized prediction;
4. Predicts TCM compound prescription drug-drug interactions1. Basic molecular descriptor knowledge required;
2. TCM metabolite ADMET prediction needs metabolic pathway tools;
3. Lacks specific guidance for lead optimization of TCM componentsCMSSP Contrastive learning model, molecular graph convolution,
MS-structure unified representation space construction1. Metabolite structural annotation;
2. Low-abundance metabolite identification in complex biological matrices;
3. In vivo MS data interpretation1. The MS-structure unified representation space significantly enhances annotation accuracy;
2. Strong anti-interference ability for complex biological matrices;
3. Reduces dependence on in vivo metabolite databases1. Requires large-scale natural product MS-structure paired training data;
2. Slow analysis for TCM in vivo samples with massive metabolites;
3. Sensitive to high-abundance endogenous biological matrix interferencesChemicalome-Metabolome Matching Platform Machine learning, metabolic network construction, chemicalome-metabolome correlation 1. In vivo MS raw data processing;
2. Endogenous background elimination;
3. Parent compound and metabolite distinction;
4. Screening of xenobiotic metabolites1. Integrates in vitro-in vivo matching;
2. Time-series filtering outlines TCM component temporal metabolic characteristics;
3. Constructs complex TCM metabolic networks;
4. Adapts to dynamic biological matrix processing1. High in vitro and in vivo MS data quality requirements;
2. Large-scale metabolic network construction needs high computing resources;
3. Weak automatic identification for rare TCM metabolic pathwaysTable 1.
Comparative analysis of AI tools for TCM in vitro and in vivo chemicalome characterization.
Figures
(4)
Tables
(1)