-
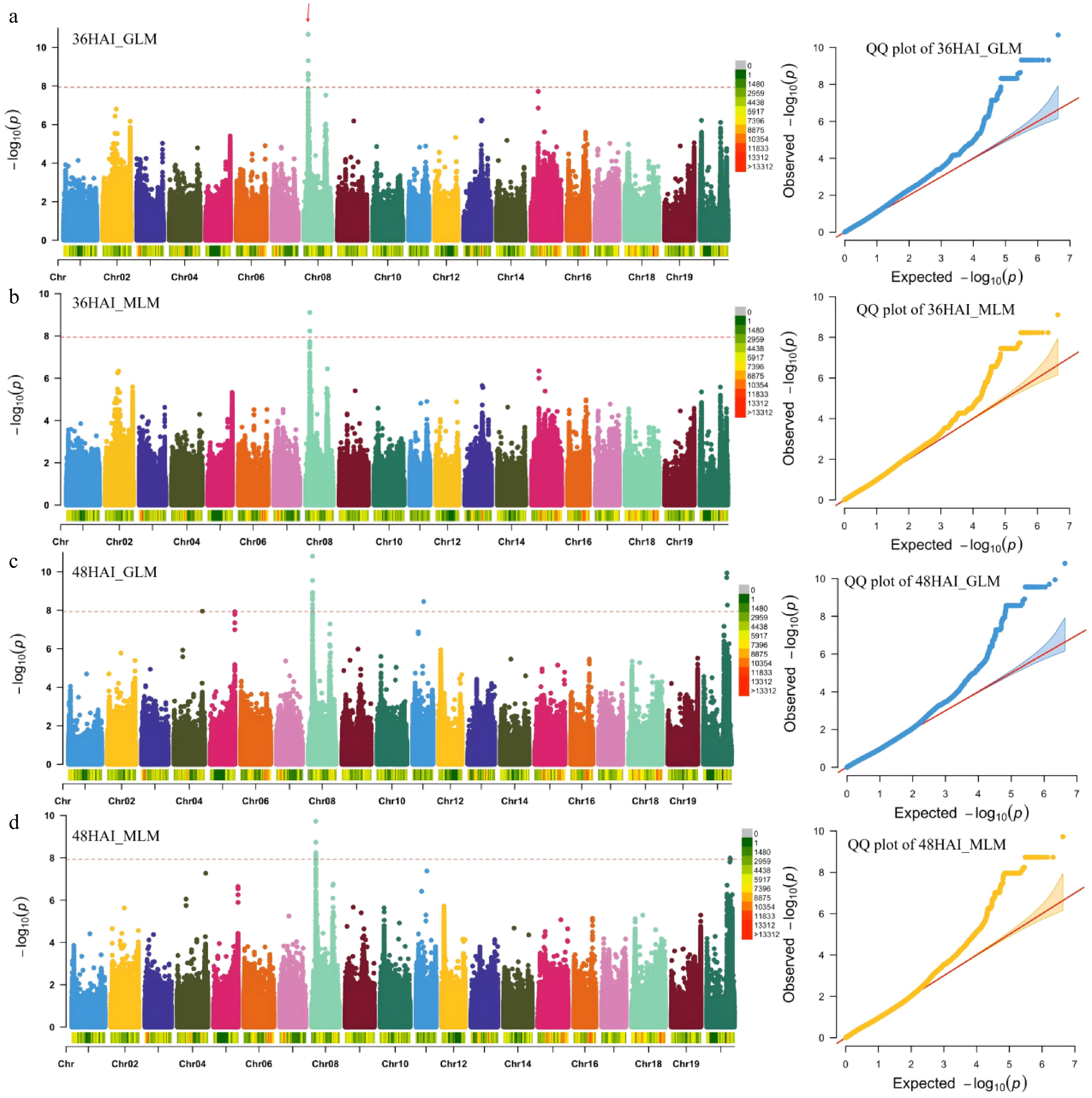
Figure 1.
GWAS of the rate of seed germination in soybean. (a)−(d) Manhattan plots and quantile–quantile (Q–Q) plots for the whole population of soybean accessions. The red arrows indicate the quantitative trait loci (QTLs) identified. GLM, general linear model; MLM, mixed linear model; HAI, hours after imbibition.
-
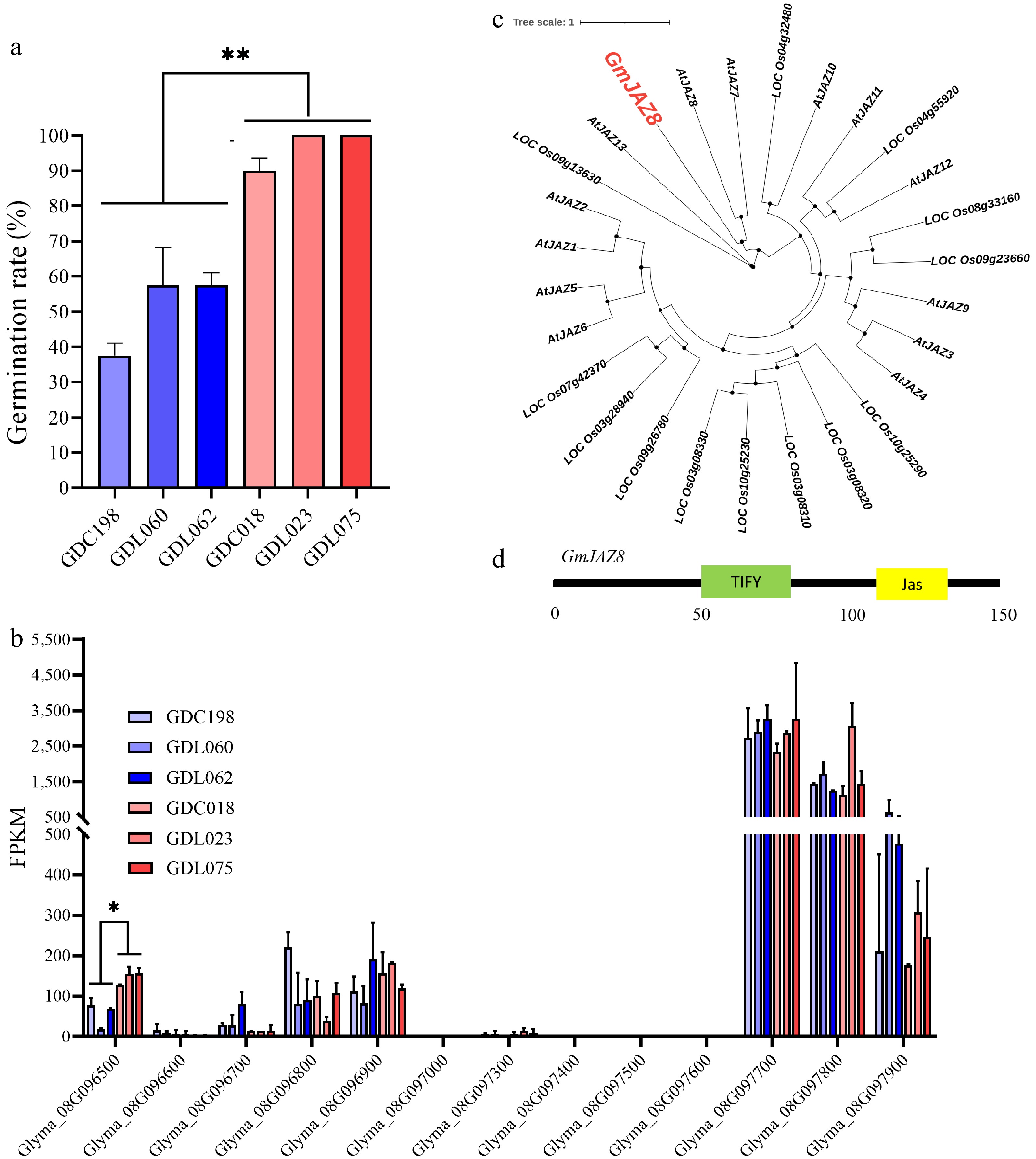
Figure 2.
Identification of GmJAZ8 for soybean the rate of seed germination. (a) Germination rate of six soybean accessions. **, p < 0.01. (b) FPKM of nine genes. *, p < 0.05. ns, no significance. Only genes with FPKM values greater than 0 are displayed. (c) Phylogenetic analysis of the GmJAZ8 and JAZ family from Arabidopsis and rice. (d) Domain organization of GmJAZ8. The numbers under the black line indicate the position of the amino acids.
-
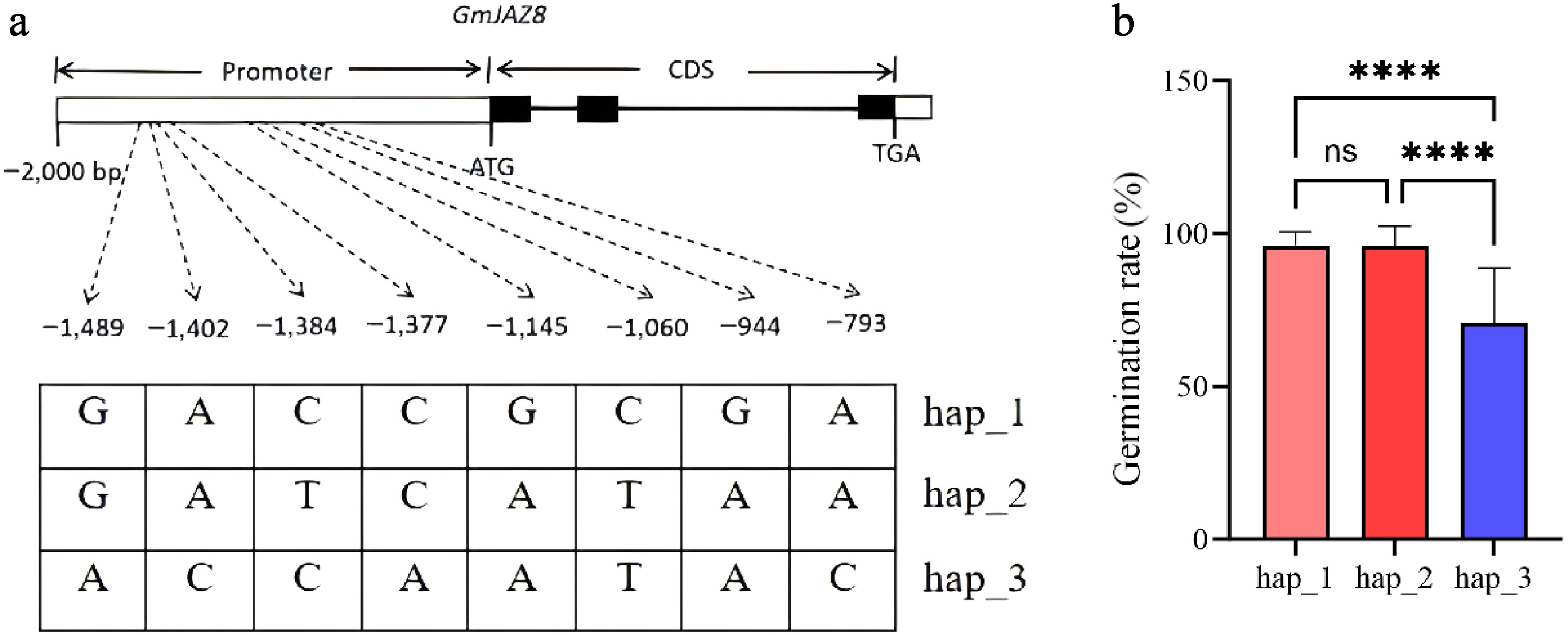
Figure 3.
Allelic variations in the GmJAZ8 gene (promoter and intragenic regions). (a) Haplotypes of GmJAZ8. (b) Box plots of the germination rate of the accessions of different haplotypes. Different colors indicate the different haplotypes. ****, p < 0.0001; ns, no significance.
-
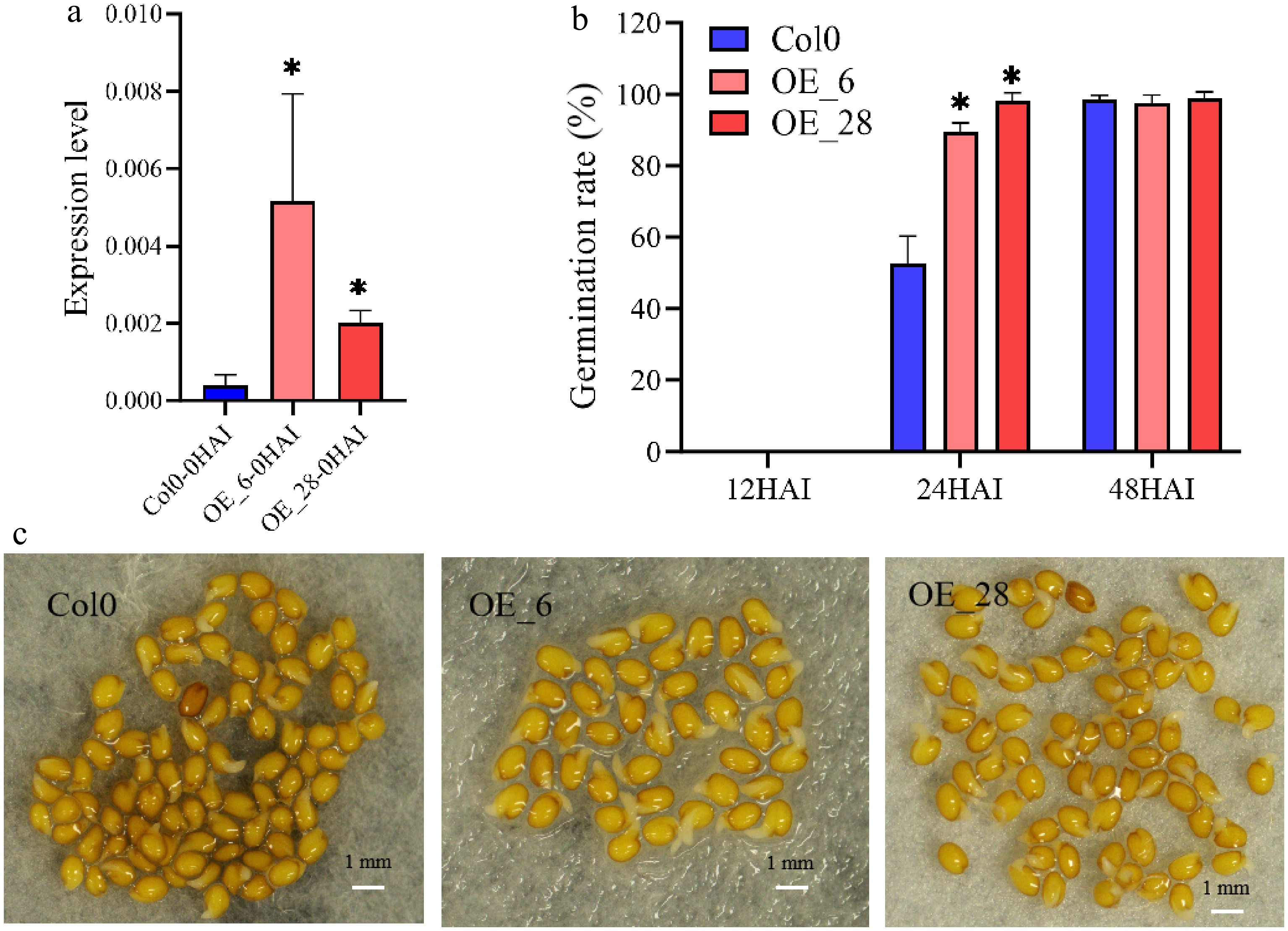
Figure 4.
GmJAZ8 promotes seed germination. (a) Relative expression of GmJAZ8 in Col0 and transgenic lines (OE_6, and OE_28) at 0 HAI. (b) The rate of germination of Col0 and the transgenic lines (OE_6, and OE_28). (c) Seedlings of the wild-type, OE_6, and OE_28 lines at 24 HAI. *, p < 0.05.
-
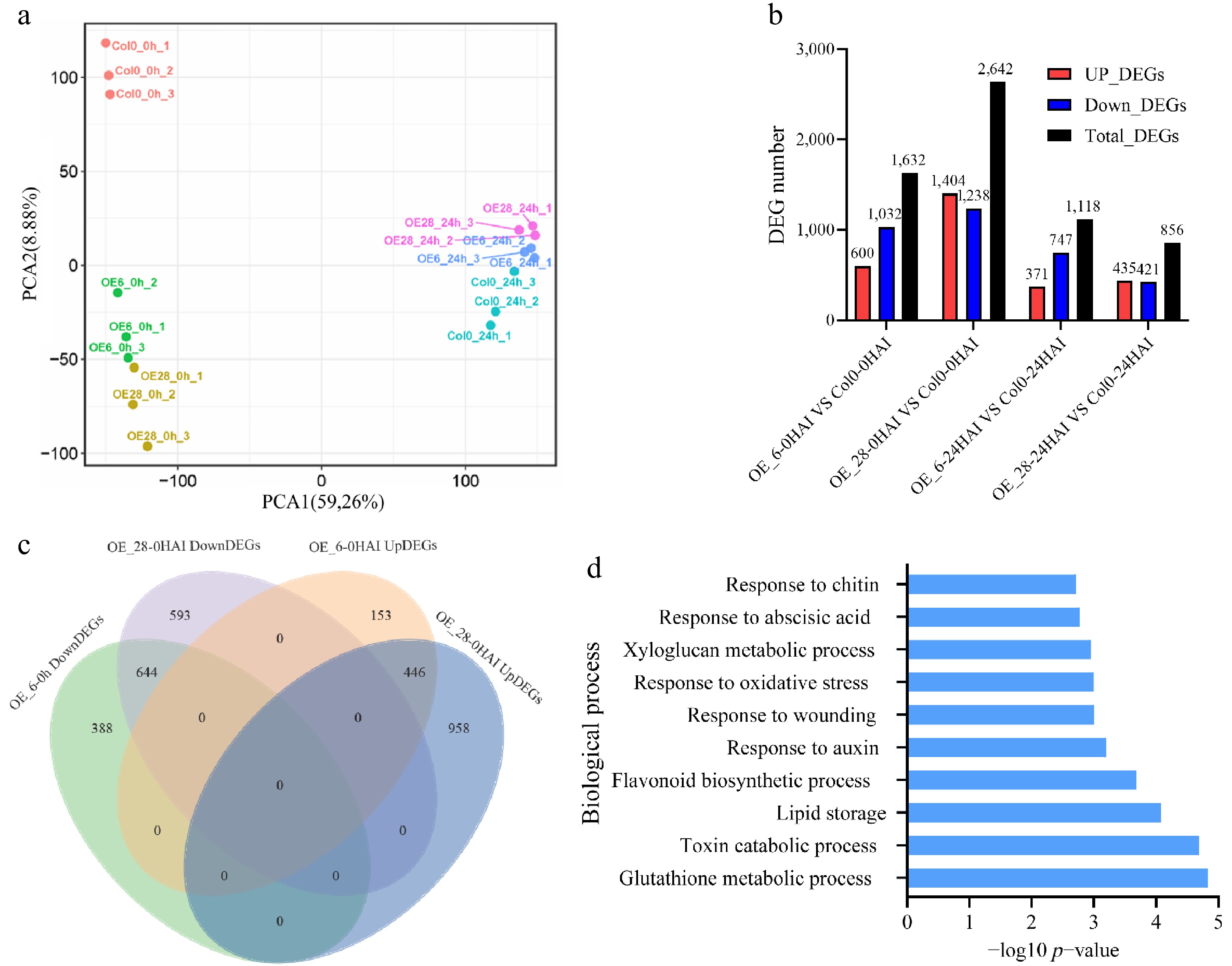
Figure 5.
Differentially expressed genes (DEGs) with p < 0.05 between the wild-type (Col0) and GmJAZ8 overexpressing lines. (a) PCA of all samples. (b) The number of DEGs. (c) Overlapping DEGs between OE_6 and OE_28 at 0 HAI. (d) GO assay of downregulated overlapping DEGs at 0 HAI.
-
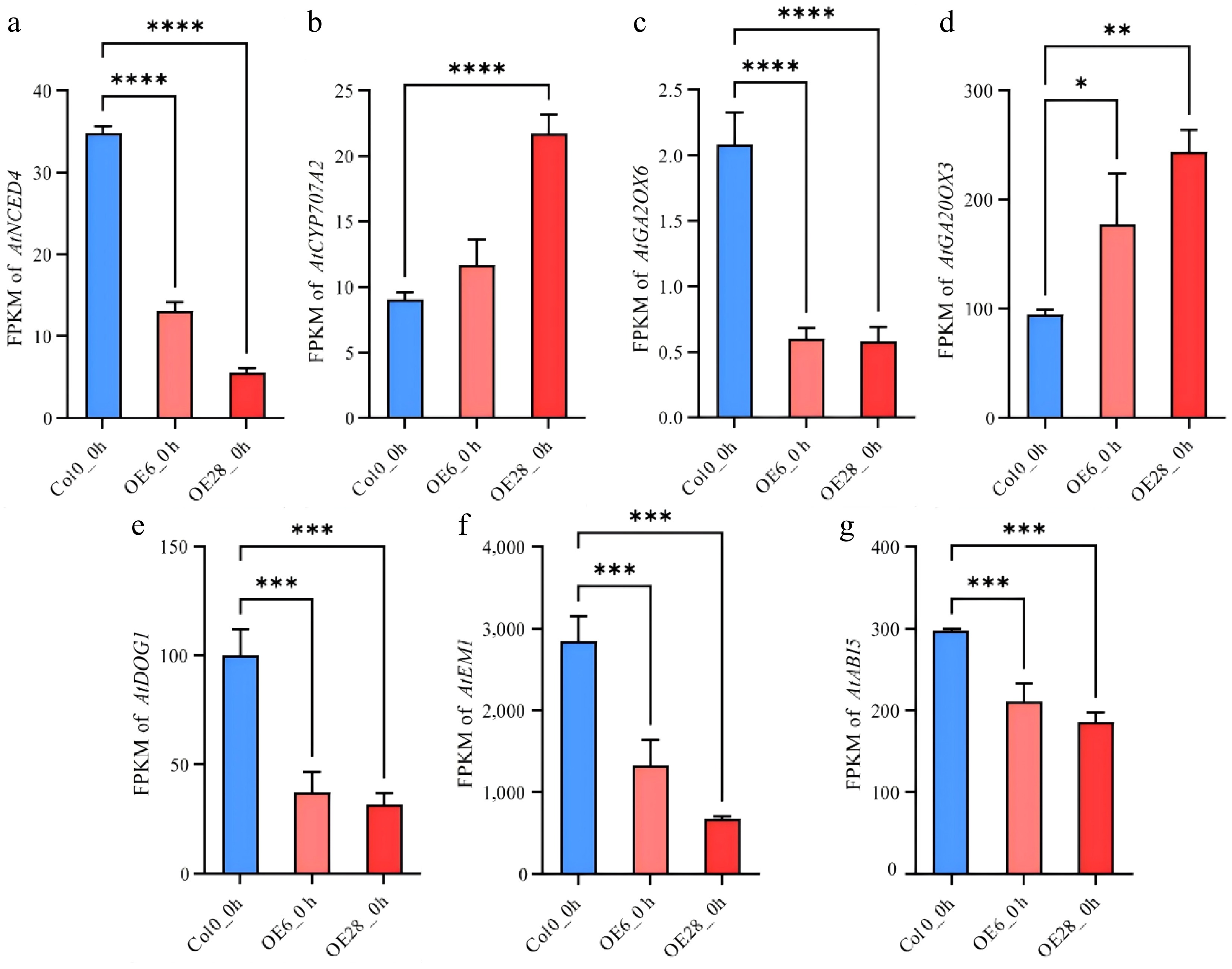
Figure 6.
GmJAZ8 altered expression of hormonal signaling genes. the FPKM values of (a) AtNCED4, (b) AtCYP707A2, (c) AtGA2OX6, (d) AtGA20OX3, (e) AtDOG1, (f) AtEM1 and (g) AtABI5. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001.
-
SNP Alleles 36HAI_GLM
p-value36HAI_MLM
p-value48HAI_GLM
p-value48HAI_MLM
p-valueChr08_7394400 C/A 2.12E-11 7.74E-10 2.72E-09 1.90E-10 Chr08_7400080 C/T 4.86E-10 5.82E-09 1.57E-11 1.86E-09 Chr08_7416347 C/T 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7427722 G/T 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7459314 T/C 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7464017 G/A 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7465749 G/A 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7470131 A/G 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7476997 C/A 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7490743 T/A 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7490778 T/C 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7500442 T/C 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7500556 T/G 4.86E-10 5.82E-09 2.82E-10 1.86E-09 Chr08_7507912 A/T 4.86E-10 5.82E-09 2.82E-10 1.86E-09 HAI, hours after imbibition; GLM, general linear model; MLM, mixed linear model. Table 1.
Information of 14 significant SNPs within the 113.5-kb interval block (7,394,400–7,507,912 bp).
-
Gene ID Position (bp) Functional annotation Homologous Arabidopsis genes Glyma.08G096500 7,396,887–7,399,006 Jasmonate–ZIM domain protein 8 AT1G30135 Glyma.08G096600 7,404,774–7,409,459 (S)-2-hydroxy-acid oxidase AT4G18360 Glyma.08G096700 7,412,769–7,417,800 (S)-2-hydroxy-acid oxidase AT4G18360 Glyma.08G096800 7,418,970–7,424,007 (S)-2-hydroxy-acid oxidase AT3G14420 Glyma.08G096900 7,424,892–7,426,785 Tetratricopeptide repeat (TPR)-like superfamily protein AT4G21065 Glyma.08G097000 7,429,593–7,429,941 AA_trans domain-containing protein AT5G19875 Glyma.08G097100 7,432,417–7,434,453 (S)-2-hydroxy-acid oxidase AT3G14420 Glyma.08G097200 7,435,704–7,438,009 (S)-2-hydroxy-acid oxidase AT4G18360 Glyma.08G097300 7,438,009–7,444,035 (S)-2-hydroxy-acid oxidase AT4G18360 Glyma.08G097400 7,447,072–7,448,398 Leucine-rich repeat receptor-like protein kinase AT2G42800 Glyma.08G097500 7,457,157–7,457,877 Cotton fiber expressed protein AT2G34610 Glyma.08G097600 7,460,358–7,463,786 BED zinc finger AT3G42170 Glyma.08G097700 7,468,455–7,476,125 RNA-binding KH domain-containing protein AT5G46190 Glyma.08G097800 7,478,544–7,486,117 Ornithine aminotransferase AT5G46180 Glyma.08G097900 7,489,719–7,494,791 TCP2 family transcription factor AT4G18390 Table 2.
Fifteen potential candidate genes within the 113.5-kb interval block (7,394,400–7,507,912 bp).
Figures
(6)
Tables
(2)