-
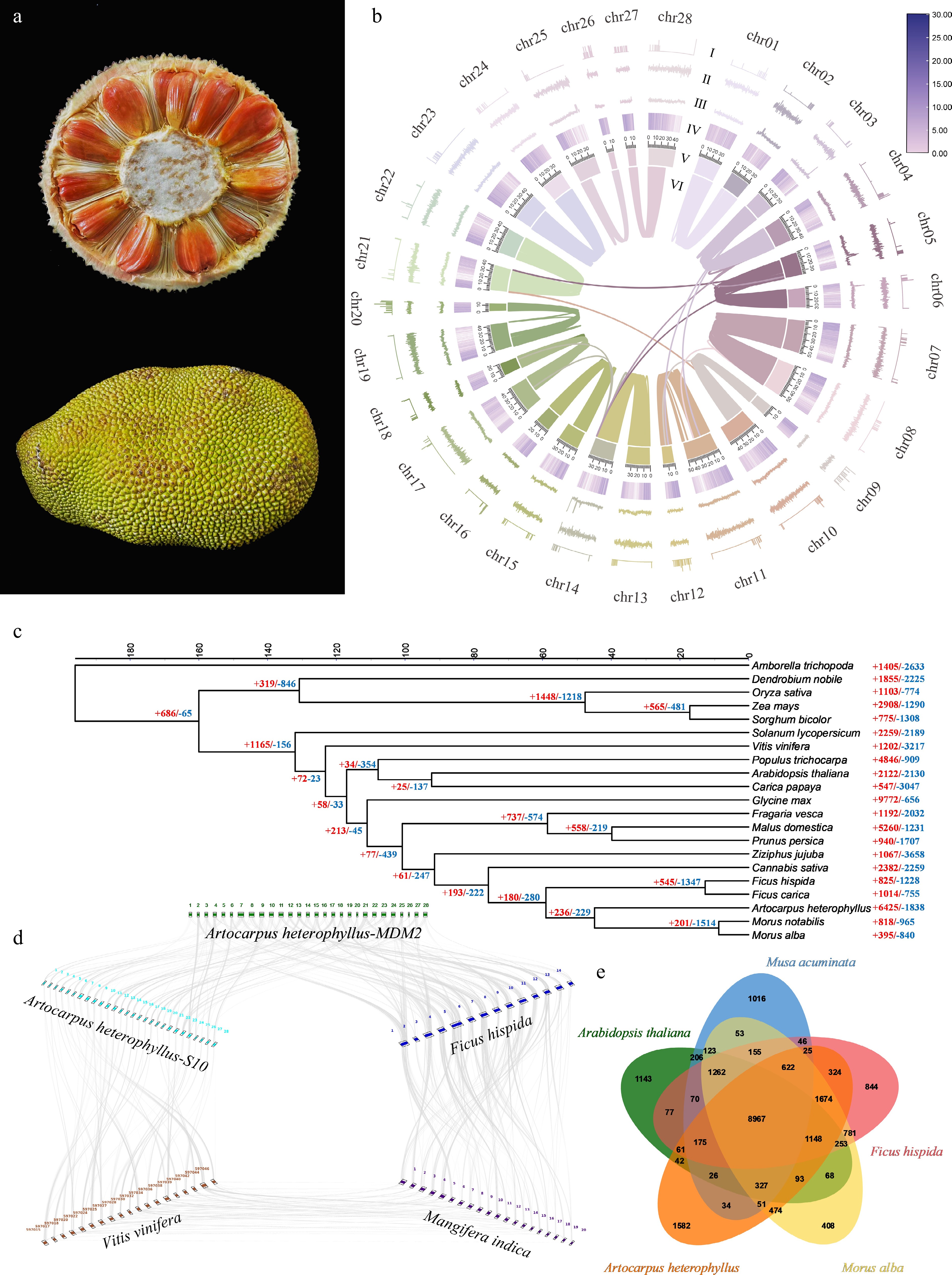
Figure 1.
(a) Morphological features of A. heterophyllus-MDM2. (b) Circos diagram of the A. heterophyllus-MDM2 genome features. (I: N [unknown bases]. II: GC skew. III: GC content. IV: gene density distribution. V: chromosome length. VI: intra-genomic chromosomal synteny among chromosomes of A. heterophyllus-MDM2 genome). (c) Phylogenetic tree and divergence time estimation of A. heterophyllus and its related species. (d) Genome collinearity analysis among A. heterophyllus-MDM2, A. heterophyllus-S10, M. alba, F. hispida, and M. indica; syntenic blocks are highlighted with grey lines connecting different chromosomes, and numbers around the rectangles indicate the chromosome IDs of each genome. (e) Venn diagram showing the shared and unique gene sets among the studied genomes.
-
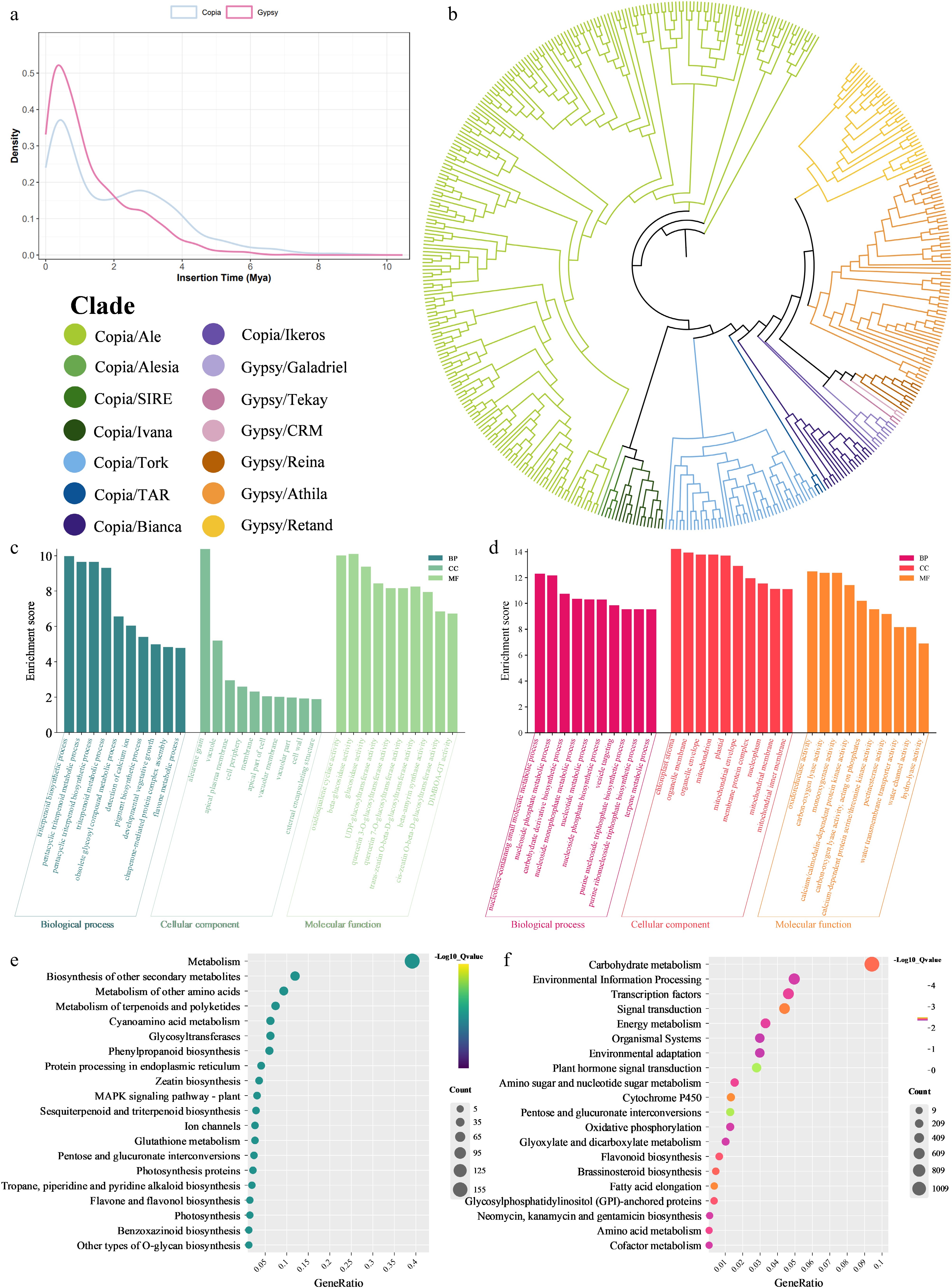
Figure 2.
(a) Density plot of LTR retrotransposon insertion times. (b) Phylogenetic clustering of LTR-RTs. (c) GO enrichment analysis plots of contracted gene families. (d) GO enrichment analysis plots of expanded gene families. (e) KEGG enrichment analysis plots of contracted gene families. (f) KEGG enrichment analysis plots of expanded gene families.
-
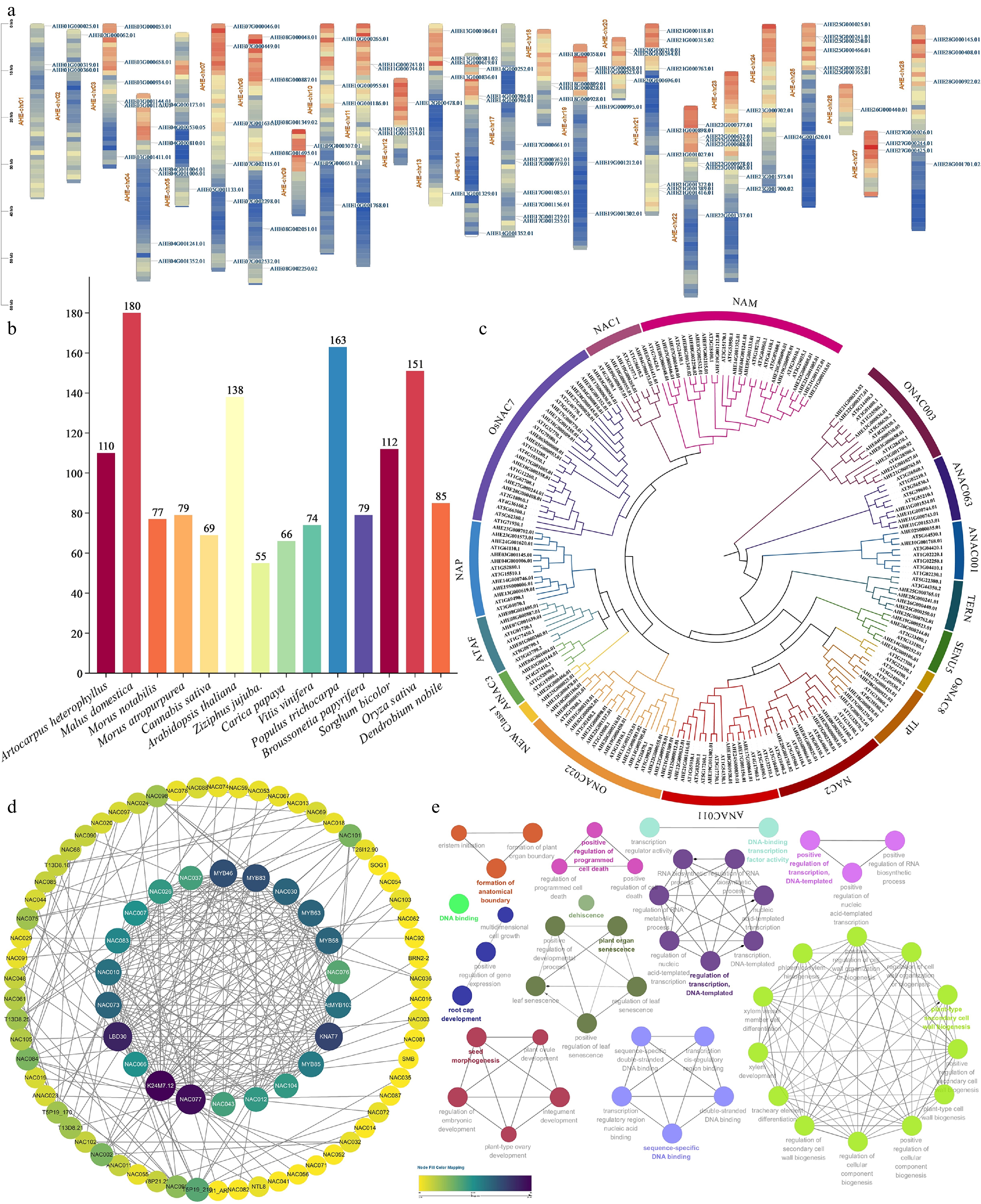
Figure 3.
(a) Schematic presentations for the distribution of AhNAC genes in A. heterophyllus chromosomes. (b) Comparison diagram of NAC gene families of various species. (c) Phylogenetic analysis of NAC proteins from A. heterophyllus and A. thaliana; different subfamilies are highlighted with specific colors. (d) Protein-protein interaction (PPI) network of AtNAC proteins. Nodes represent AtNAC proteins. (e) ClueGO functional enrichment analysis bubble plot.
-
Assembly feature MDM2 Total sequence length (bp) 1,027,292,696 Number of chromosomes 28 Number of scaffolds 60 Scaffolds N50 (bp) 39,361,022 Scaffolds L50 12 GC content (%) 34.82 Number of genes 45,366 Total TE length (bp) 605,212,121 LAI 26.88 BUSCO (%) 93.8 QV 76.11 Table 1.
Statistics of the A. heterophyllus-MDM2 genome assembly.
-
Type Number of elements Length (bp) In genome
(%)DNA transposons 514,818 109,019,163 10.61 Long interspersed nuclear elements (LINEs) 8,203 2,163,258 0.21 Rolling-circles 1,646 1,315,557 0.13 Long terminal repeats (LTR) 627,910 459,633,274 44.74 Simple repeats 90 5,989 < 0.01 Total − 605,212,121 58.91 Table 2.
Statistics of repeat elements in the A. heterophyllus-MDM2 genome.
-
Type MDM2 Gene density (gene/Mb) 44.16 Gene number 45,366 Average gene length (bp) 3,985.64 Average CDS length (bp) 1,983.49 Average exon per gene 6.82 Average exon length (bp) 293.65 Average intron length (bp) 456.40 Table 3.
Gene structural characteristics of the A. heterophyllus-MDM2 genome.
Figures
(3)
Tables
(3)