-
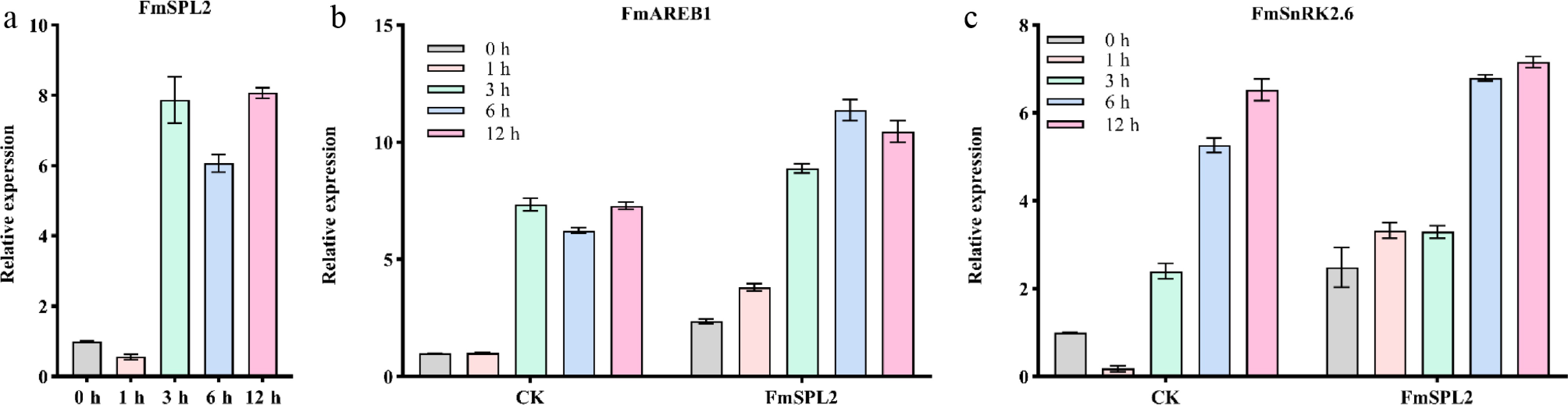
Figure 1.
The expression of the FmSPL2 gene is regulated by drought stress. (a) The expression of FmSPL2 under drought treatment at 0, 1, 3, 6, and 12 h. (b) The expression changes of FmAREB1 after transient transfection of FmSPL2 and natural drought treatment at 0, 1, 3, 6, and 12 h. (c) The expression levels of FmSnRK2.6 after transient transfection of FmSPL2 and drought treatment at 0, 1, 3, 6, and 12 h.
-
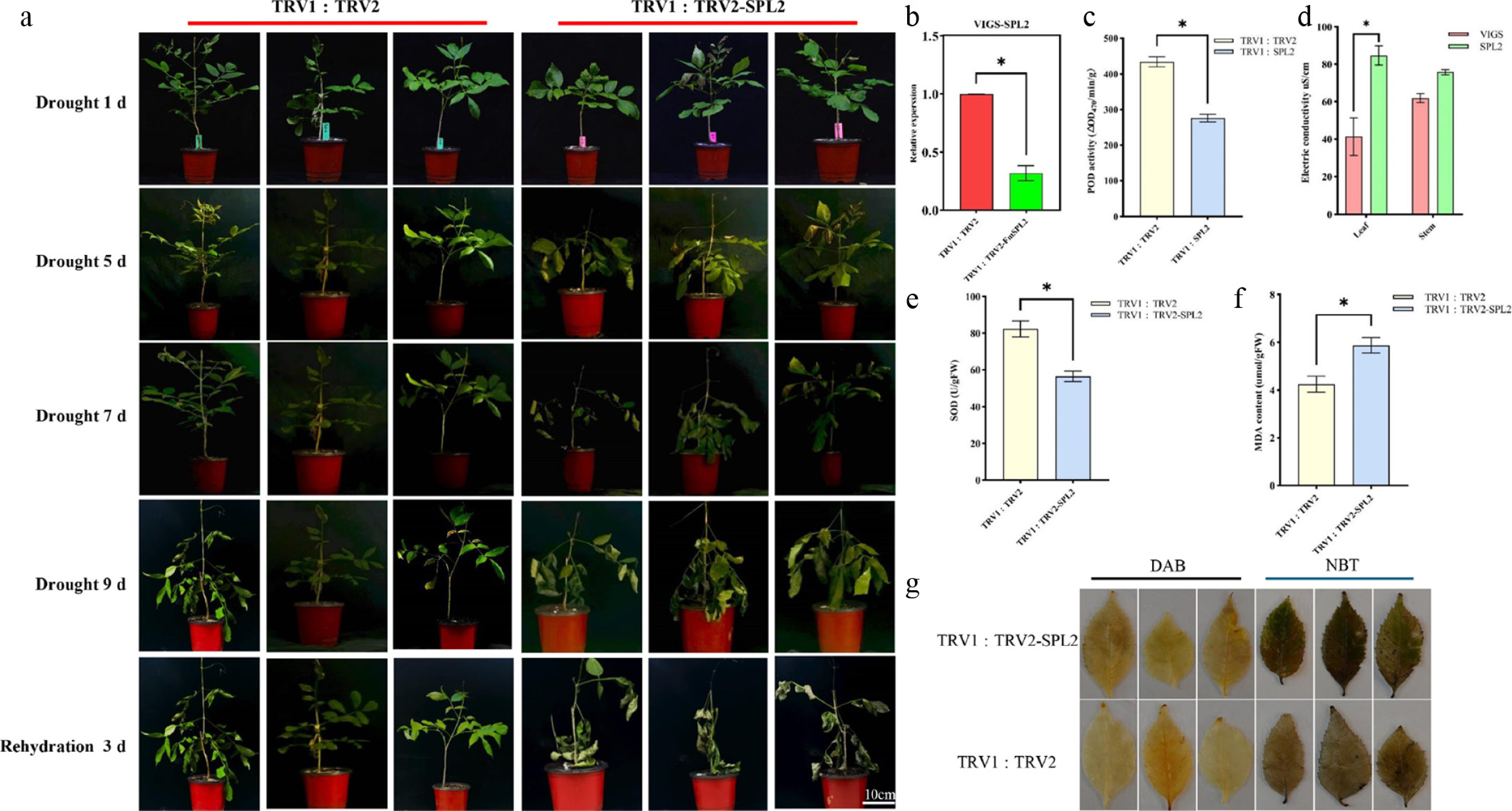
Figure 2.
Phenotypic analysis of FmSPL2 gene silencing by VIGS in Fraxinus mandshurica. (a) VIGS-FmSPL2 drought phenotype analysis; (b) VIGS-FmSPL2 qPCR analysis; (c) POD activity assay; (d) electrical conductivity measurement; (e) SOD activity assay; (f) MDA content assay; (g) DAB and NBT staining.
-
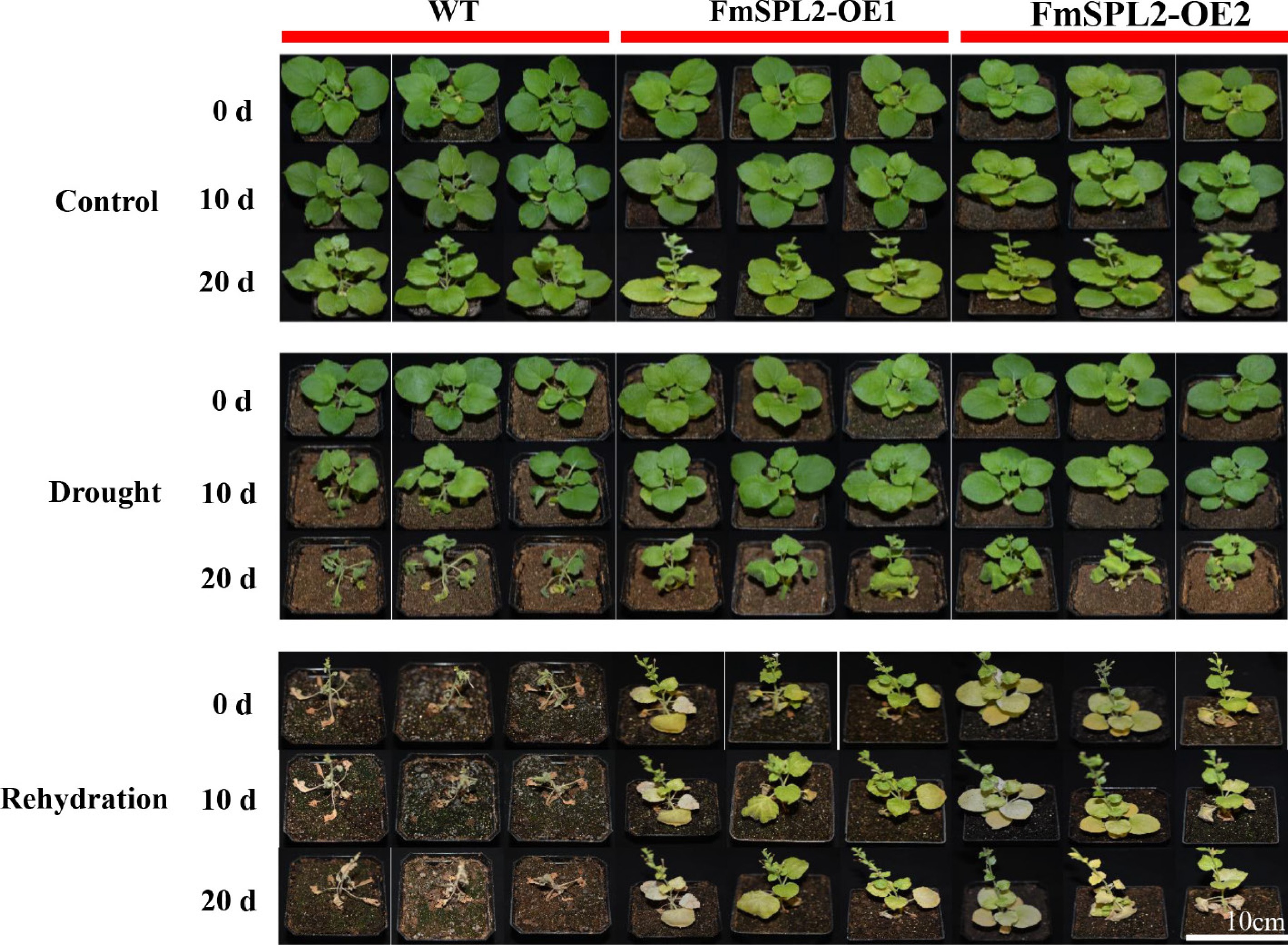

Figure 3.
FmSPL2 transgenic tobacco drought treatment phenotype image. Fifteen days of soil cultivation with drought stress and rehydration after stress, drought treatment for 20 d, followed by 15 d of rehydration.
-
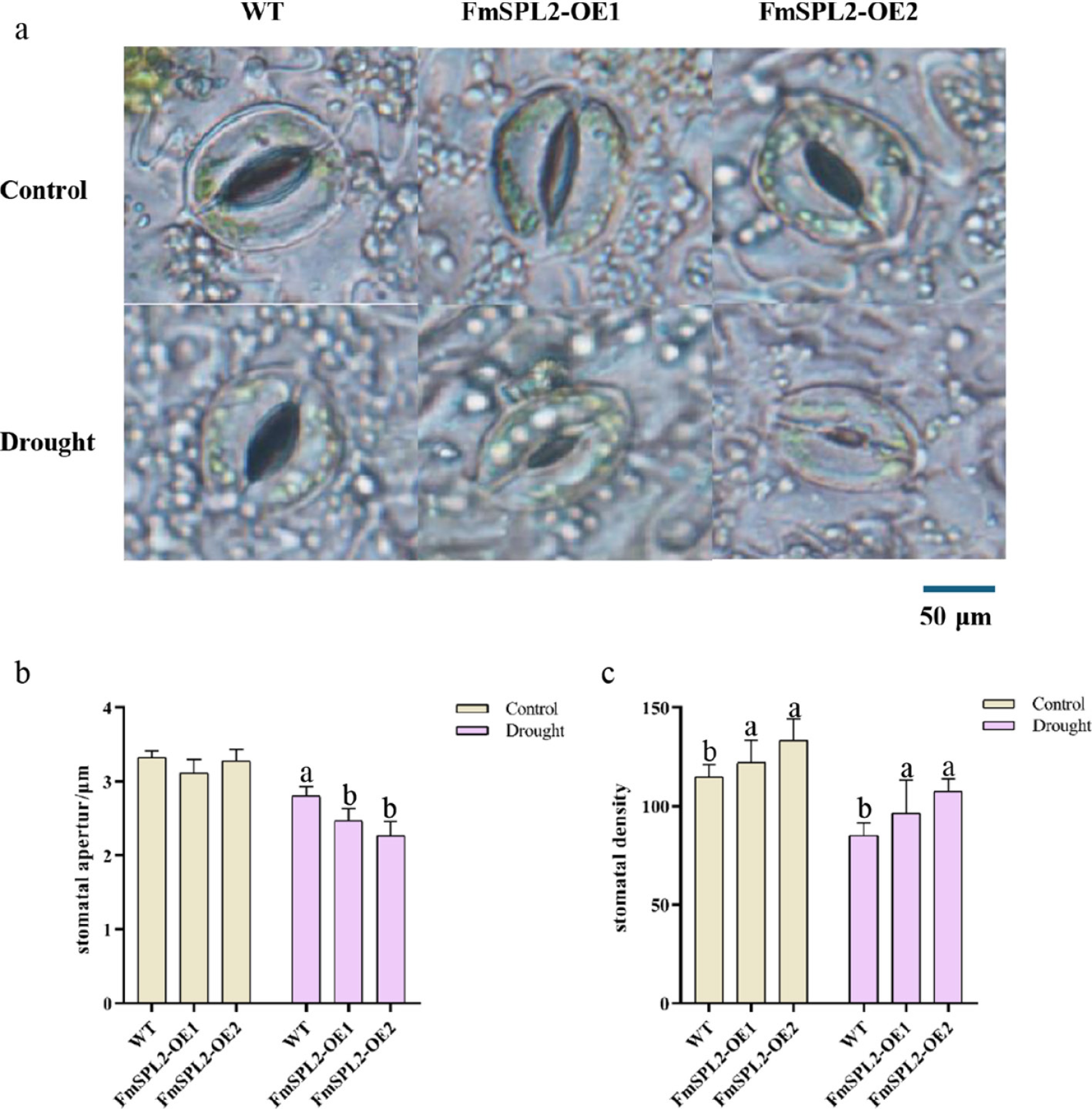
Figure 4.
Effect of FmSPL2 on stomata under drought treatment. (a) Stomatal state after drought treatment; (b) stomatal aperture; (c) stomatal density.
-
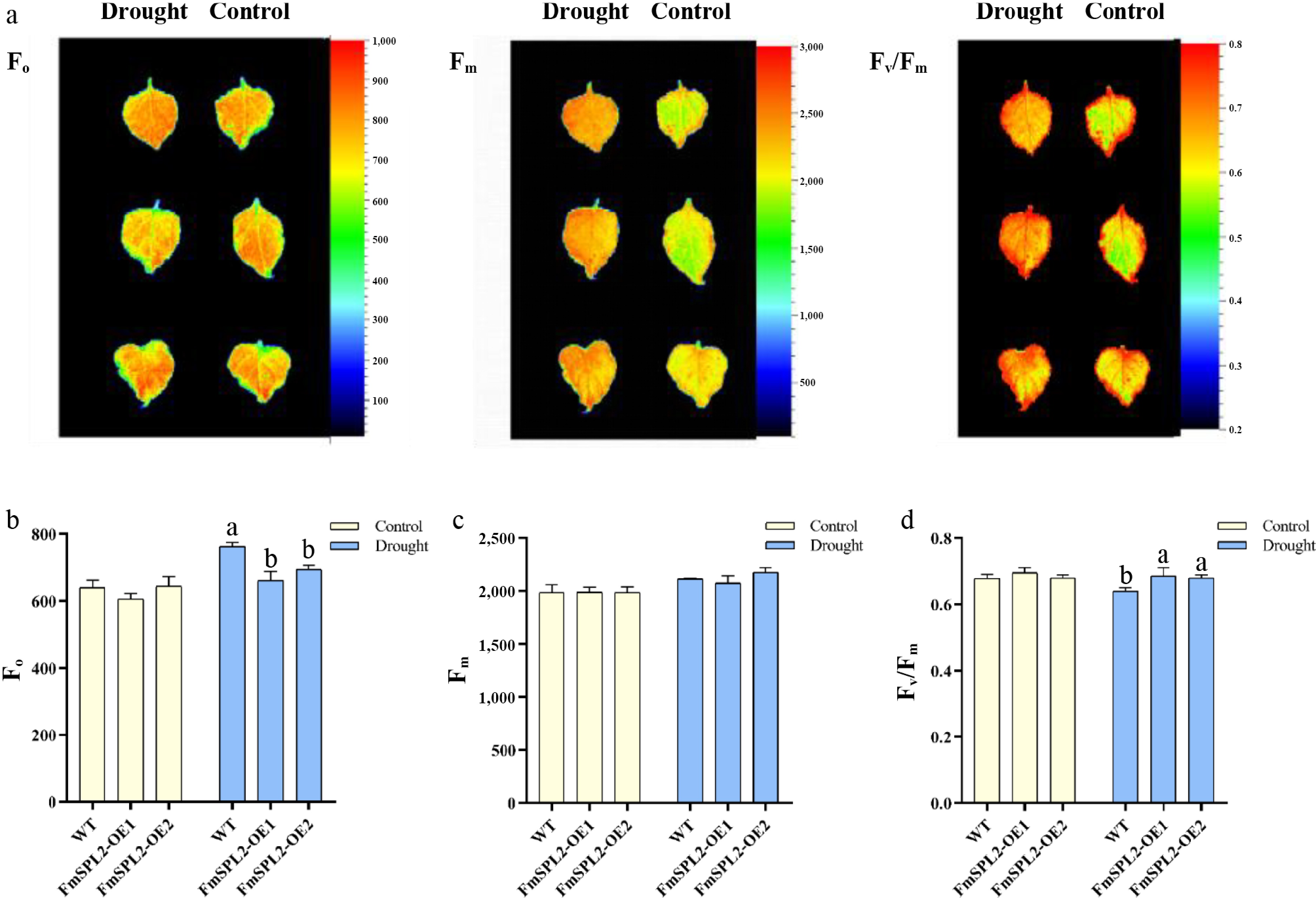
Figure 5.
Changes of chlorophyll fluorescence parameters after drought stress. (a) Chlorophyll fluorescence imaging; (b) F0 value of chlorophyll fluorescence parameter; (c) chlorophyll fluorescence parameter Fm value; (d) chlorophyll fluorescence parameter Fv/Fm value.
-
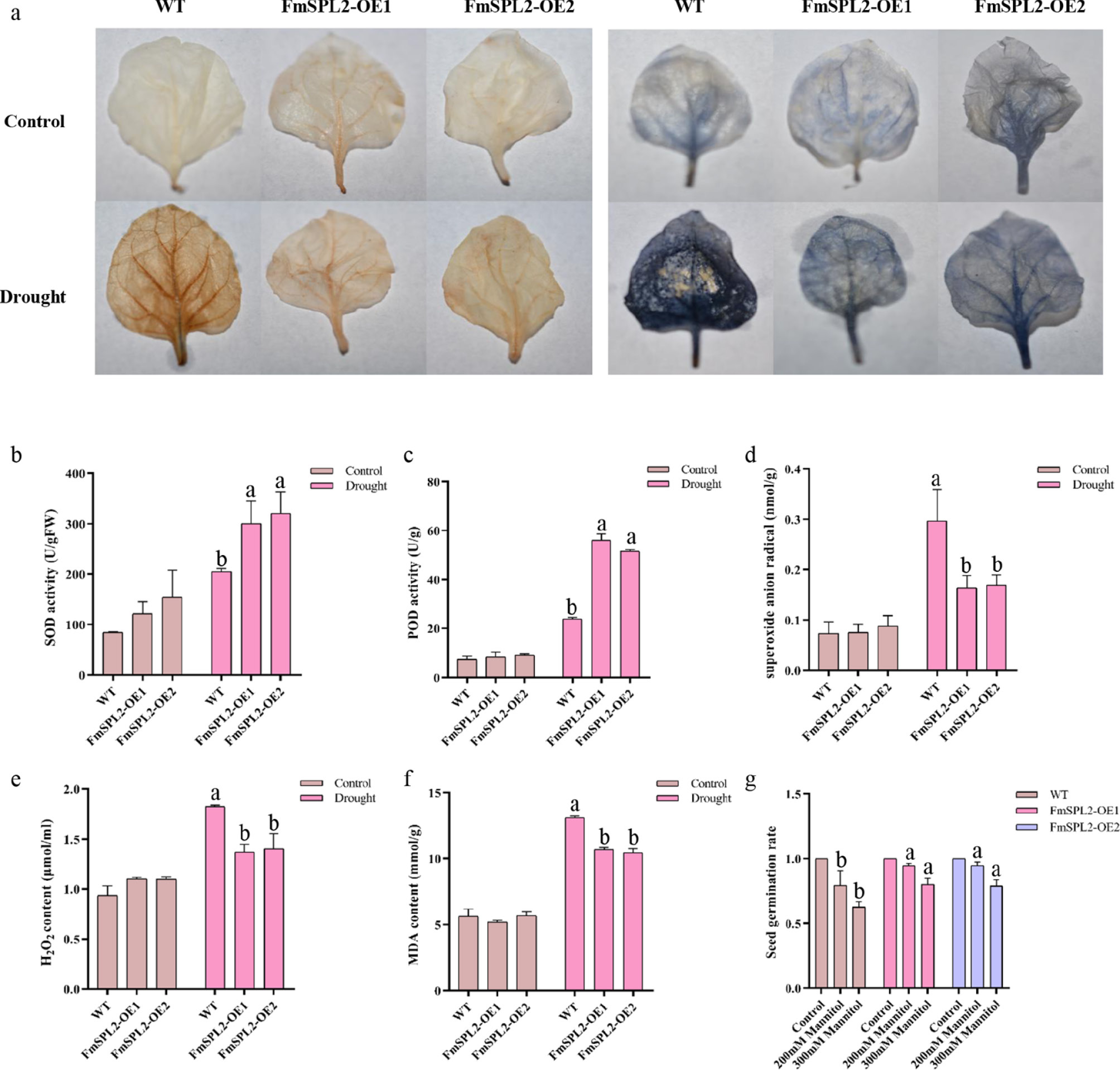
Figure 6.
Physiological indexes determination after drought treatment. (a) Hydrogen peroxide (DAB), and superoxide anion (NBT) staining; (b) superoxide dismutase (SOD) activity; (c) peroxidases (POD) activity; (d) superoxide anion (DFR) content; (e) hydrogen peroxide (H2O2) content; (f) Malondialdehyde (MDA) content, and (g) germination rate.
-
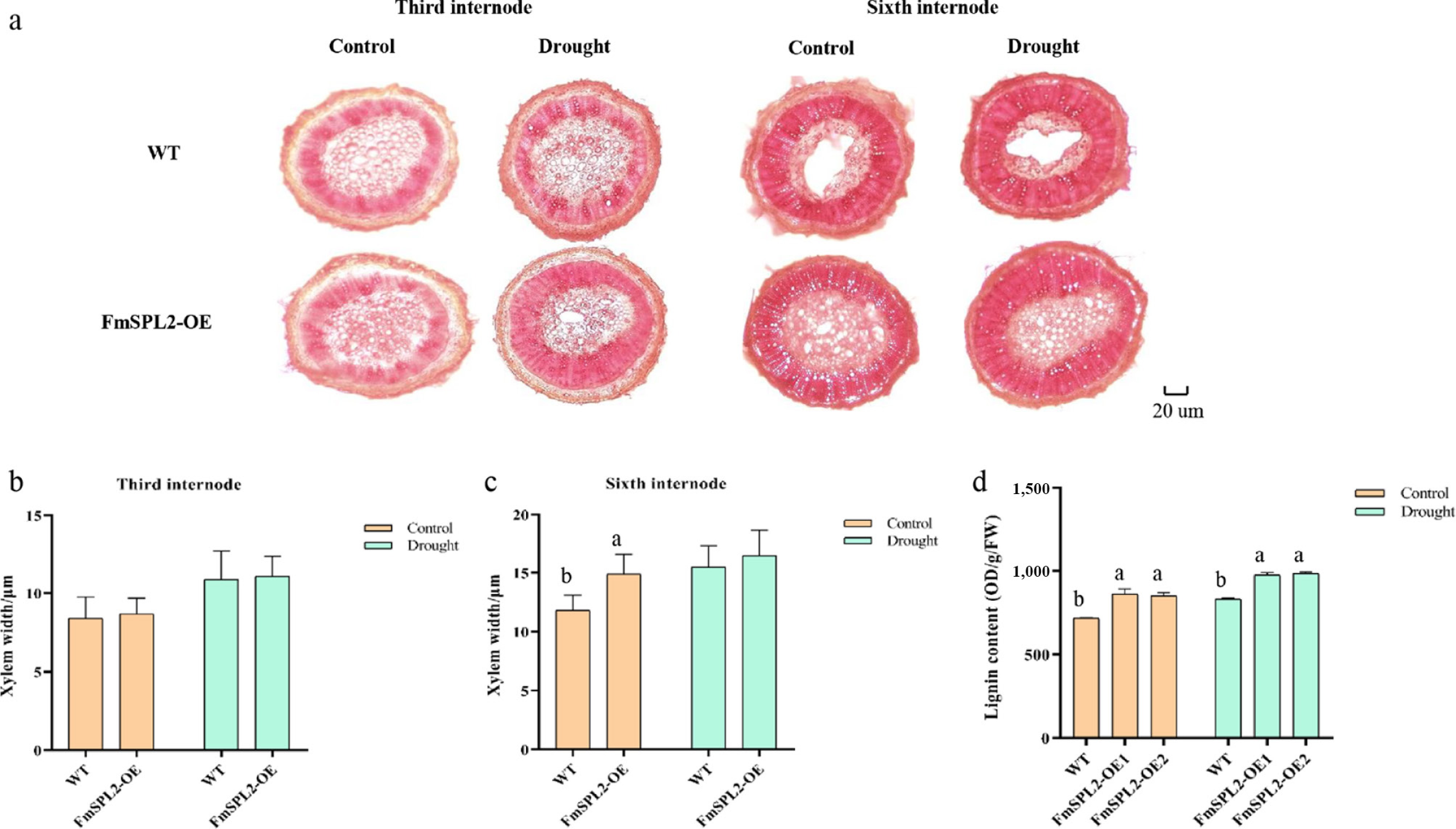
Figure 7.
FmSPL2 gene affects xylem development. (a) Xylem staining after drought treatment; (b) xylem thickness of the third internode; (c) xylem thickness of the sixth internode, and (d) lignin content.
-
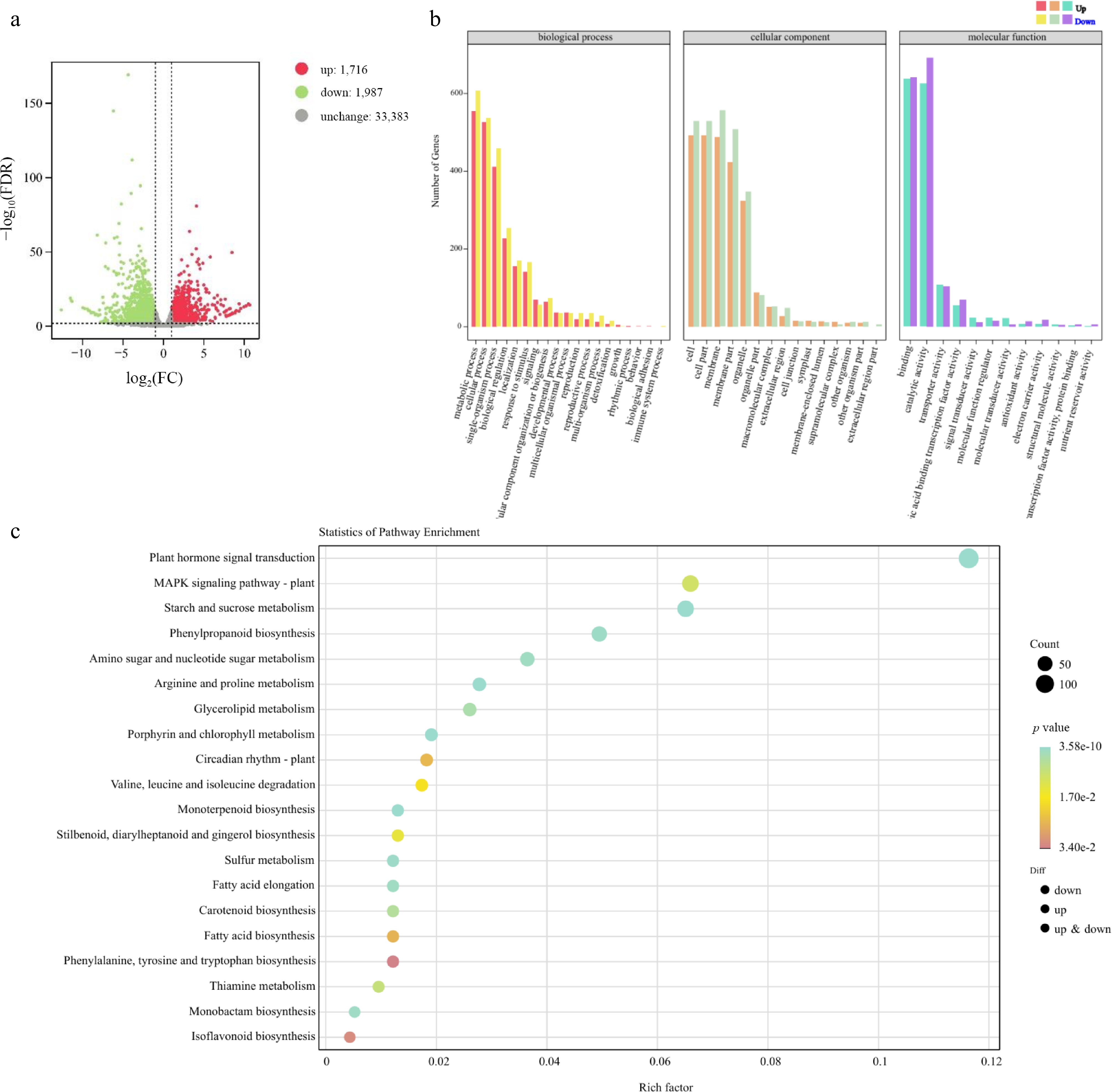
Figure 8.
FmSPL2 affects drought resistance by regulating the expression of lignin biosynthesis genes.
-
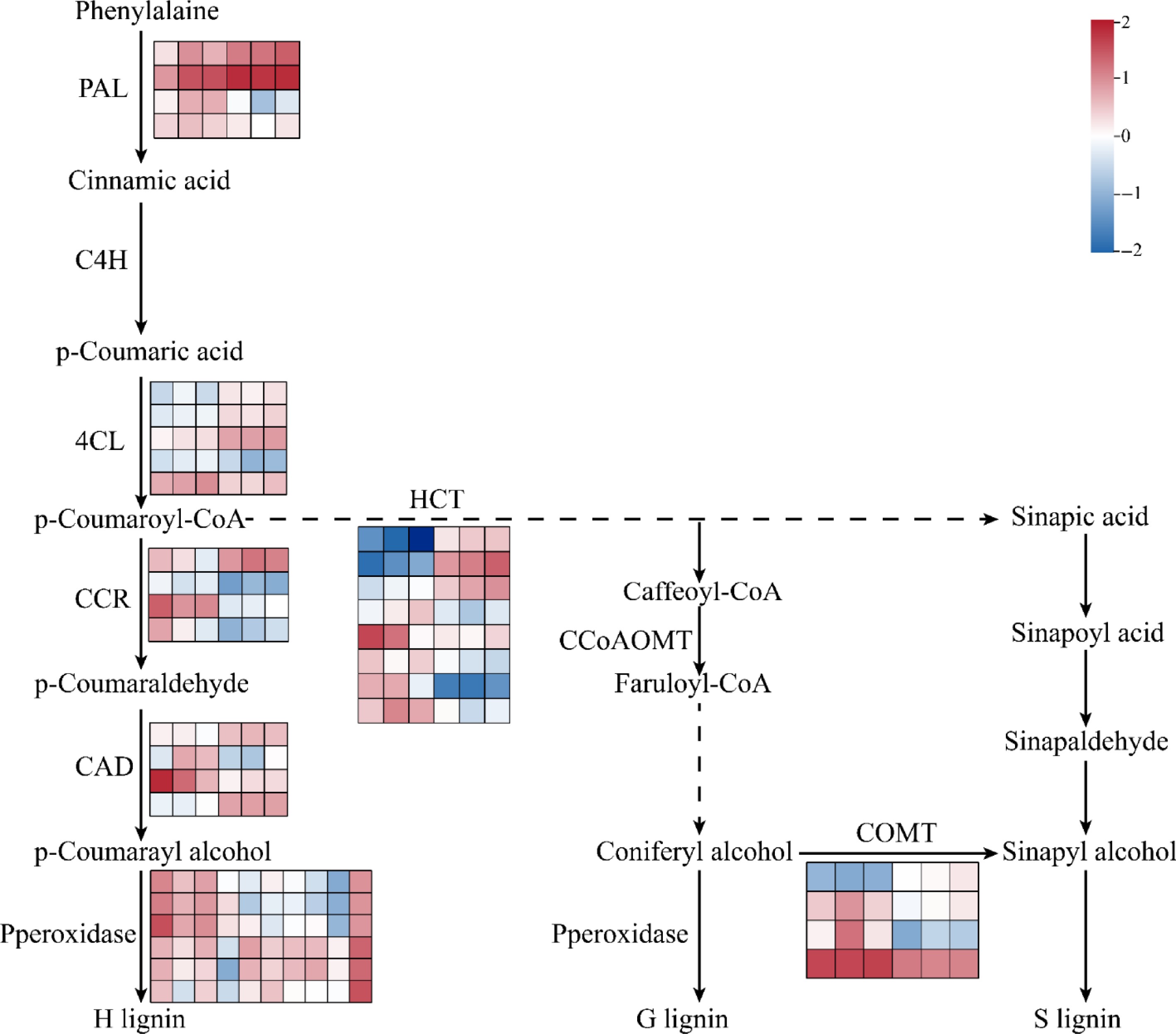
Figure 9.
Phenylpropanoid and flavonoid biosynthesis network diagram. Gene expression is shown according to the mean FPKM of three biological replicates; blue indicates low expression and red indicates high expression. PAL, phenylalanine ammonialyase; C4H, cinnamate 4-hydroxylase; 4CL, 4-coumaroyl CoA ligase; CCR, cinnamoyl-CoA reductase; HCT, shikimate O-hydroxycinnamoyltransferase; CAD, cinnamyl-alcohol dehydrogenase; CCoAOMT, caffeoyl-CoA O-methyltransferase; COMT caffeic acid 3-O-methyltransferase.
-
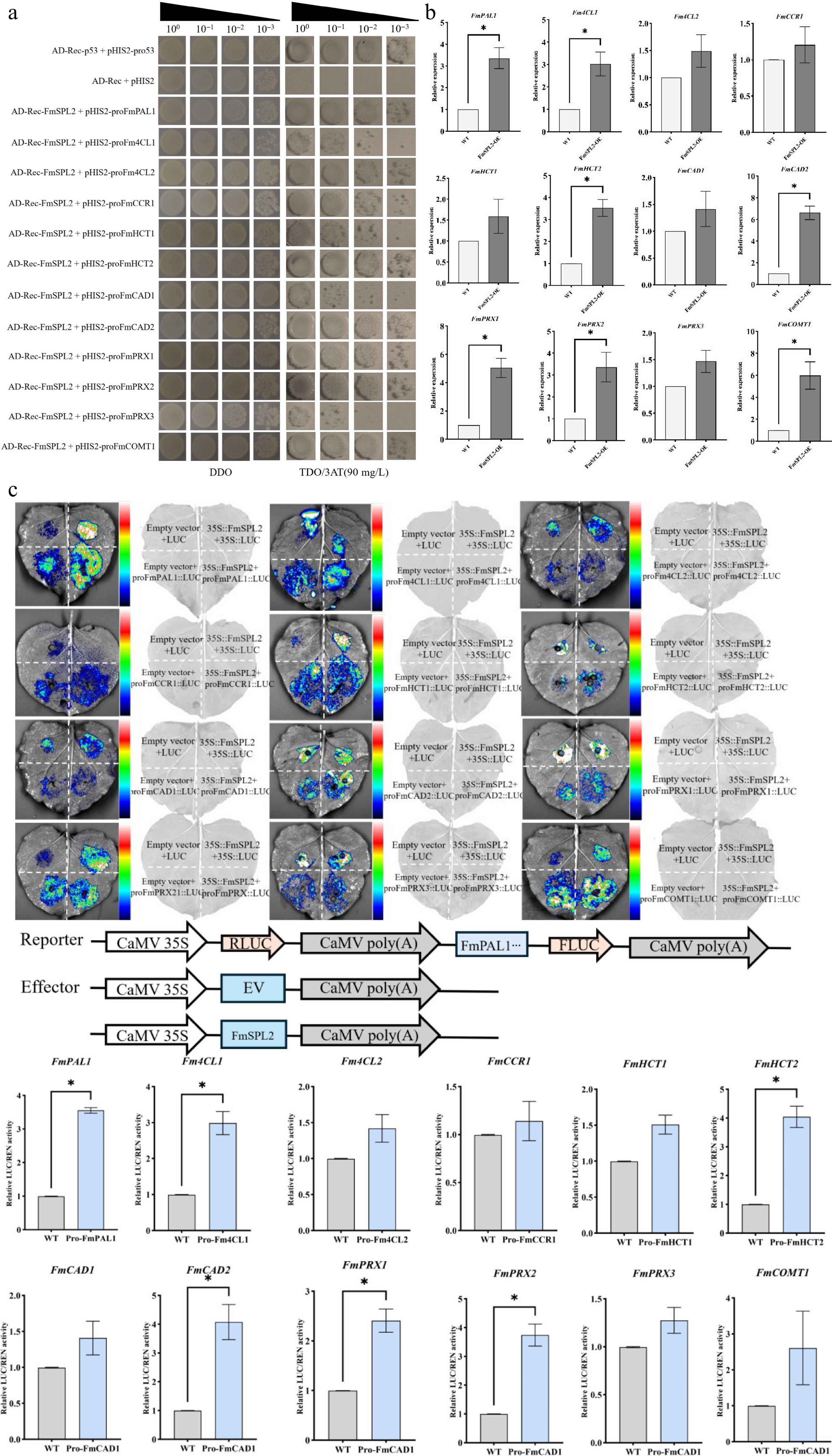
Figure 10.
Verification of the binding and regulatory relationship between FmSPL2 and downstream lignin biosynthetic genes. (a) Yeast one-hybrid (Y1H) assay showing the physical binding of FmSPL2 to the promoters of genes related to the phenylpropanoid biosynthetic pathway. (b) Relative expression levels (qRT-PCR) of the target genes in FmSPL2 overexpressing lines. (c) Dual-luciferase reporter assay in N. benthamiana leaves demonstrating the transcriptional activation of downstream gene promoters by FmSPL2.
-
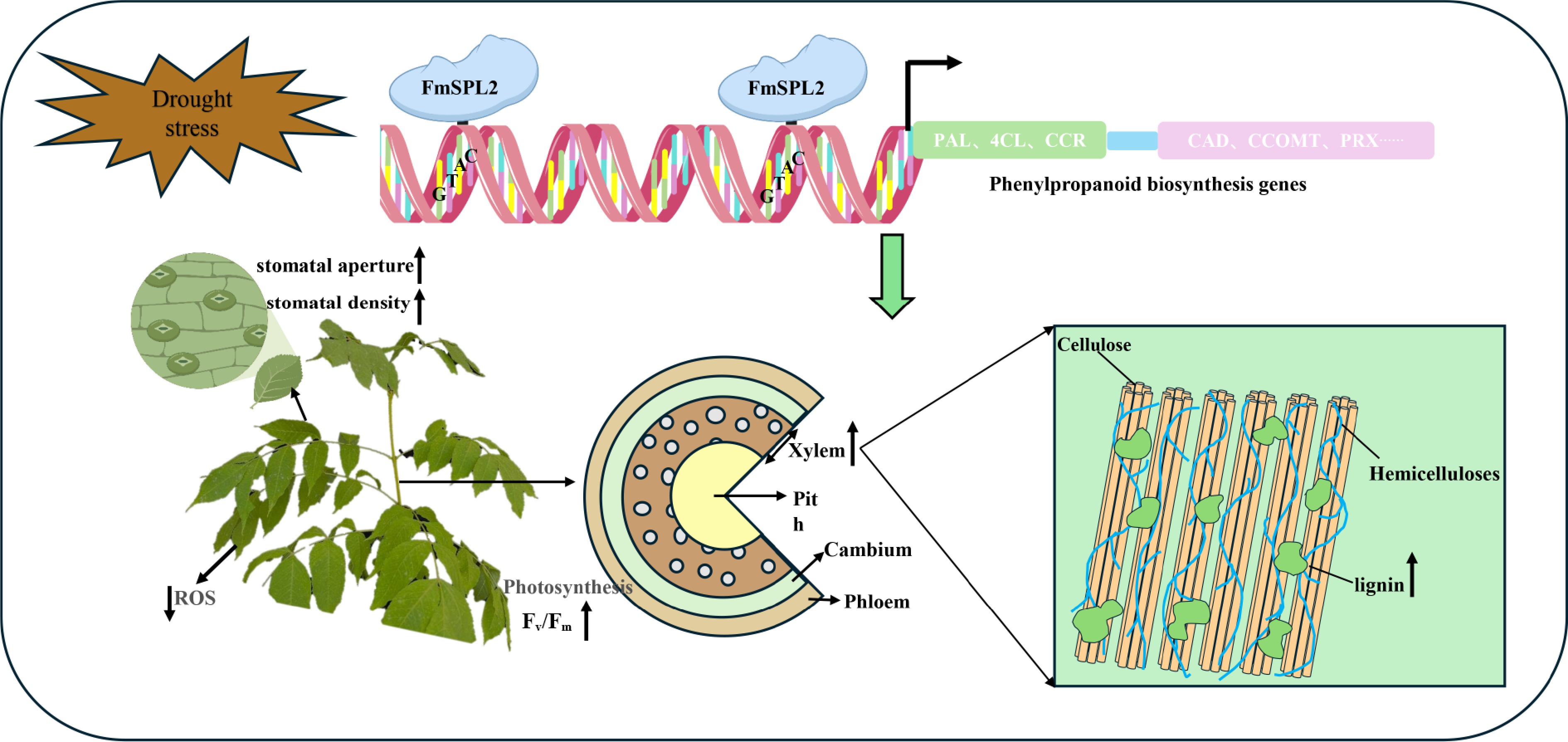
Figure 11.
Proposed molecular model demonstrating that FmSPL2 regulates lignin biosynthesis and drought adaptation in Fraxinus mandshurica. FmSPL2 is rapidly expressed under drought stress, regulating the expression of phenylpropanoid metabolism-related genes to promote lignin biosynthesis. At the same time, it enhances drought tolerance in Fraxinus mandshurica by modulating stomatal aperture, improving photosynthetic efficiency, and increasing ROS scavenging capacity.
Figures
(11)
Tables
(0)