-
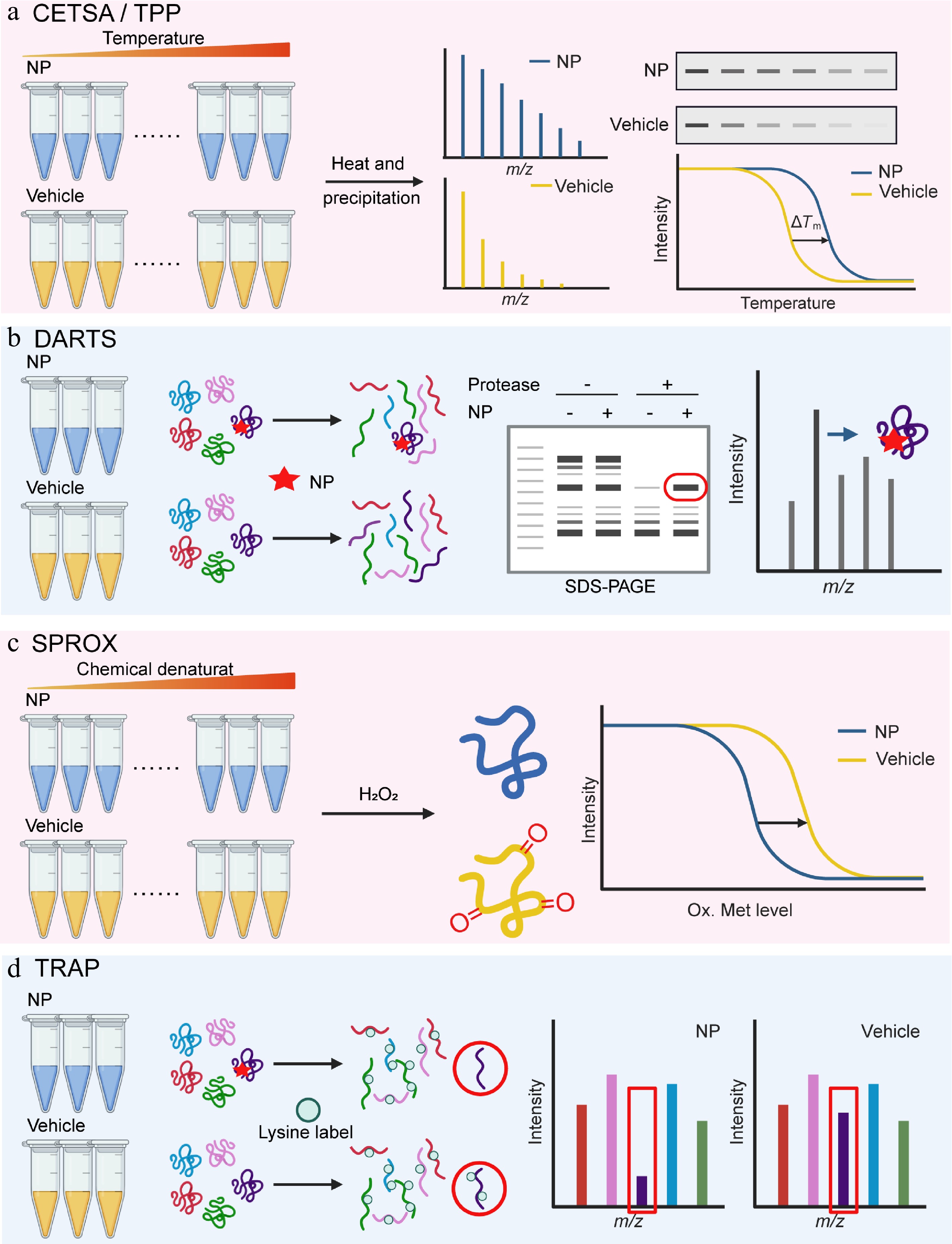
Figure 1.
Natural product target identification based on protein libraries. (a) Thermal shift assay (TSA-CETSA-TPP), (b) drug affinity responsive target stability (DARTS), (c) stability of proteins from rates of oxidation (SPROX), and (d) target-responsive accessibility profiling (TRAP).
-
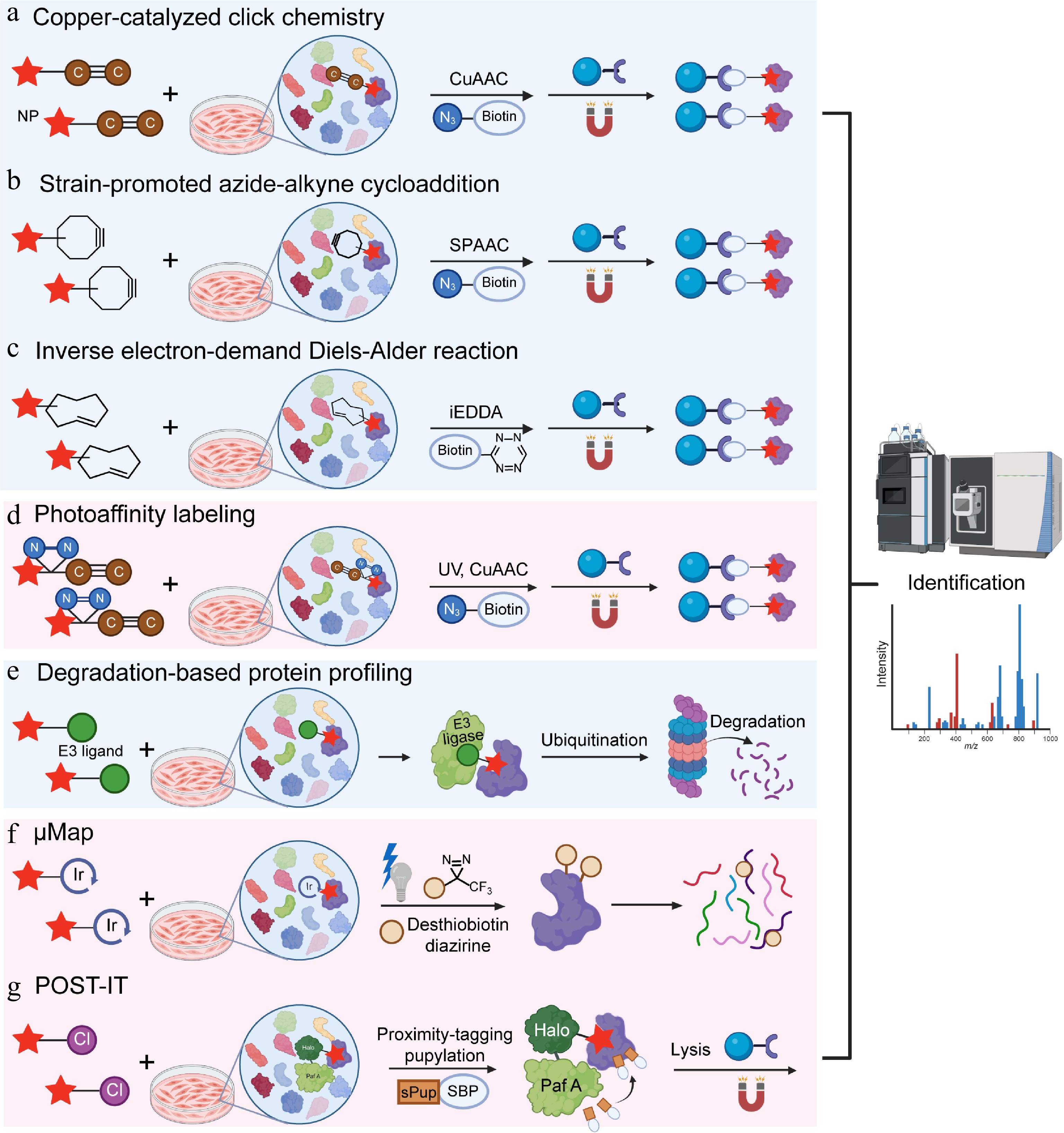
Figure 2.
Natural product target identification based on in-situ models. (a) Copper-catalyzed click chemistry (CUAAC), (b) strain-promoted azide-alkyne cycloaddition (SPAAC), (c) inverse electron-demand Diels-Alder reaction (iEDDA), (d) photoaffinity labeling (PAL), (e) degradation-based protein profiling (DBPP), (f) μMap, and (g) pup-on-target for small molecule target identification technology (POST-IT).
-
Natural product Activity Target Method used Mechanisms of action Ref. Acacetin Lipid metabolism regulation LAMTOR1 DARTS Modulates autophagy via the LAMTOR1-MTORC1-AMPK signaling pathway. Alleviates metabolic dysfunction-associated fatty liver disease [31] Artemisinin Against malaria Multiple parasite proteins SPAAC Targets across glycolysis, hemoglobin degradation, antioxidant defense, and protein synthesis pathways [86] Artemisinin Against malaria Targets across different stages of the parasite PAL Interferes with the protein synthesis, glycolysis, and oxidative homeostasis pathways of parasites [99] Artemisinin derivatives Antitumor PCLAF DBPP Degrades PCLAF and activates the p21/Rb axis [102] α-Mangostin Antitumor RTN4/Nogo Protein microarray Induces proteasome degradation of RTN4 through recruiting E3 ligase UBR5 and enhances the pyroptosis phenotype [72] Bufalin Antitumor ERα (ESR1) AI + Docking + MD simulation Enhance the interaction between ERα and the E3 ligase STUB1, thereby promoting proteasomal degradation of ERα [111] Celastrol Anti-inflammation CAP1 TRAP Ameliorates metabolic syndrome via cAMP-PKA-NF-κB pathway [62] Celastrol Antitumor, anti- inflammation, alleviating of metabolic disorders Multi-targets including CHK1, OGA, ERCC6L, IKKβ, PI3Kα, and CIP2A DBPP Inhibits PI3K/AKT pathway, NF-κB pathway, and induces G2/M phase arrest [100] Cycloastragenol Antitumor Cathepsin B TRAP Enhances CD8 T cell-mediated antitumor immunity [63] 12-deoxyphorbol 13-palmitate Anti-liver fibrosis APOL2 PAL Disrupts APOL2-SERCA2-PERK-HES1 signaling and alleviates fibrosis [97] 2,4-dihydroxybenzaldehyde Alleviation of acute
kidney injuryKEAP1 HiMAP-seq Inhibits KEAP1 and downstream and NRF2-GPX4 signaling axis [106] Eupalinolide B Anti-neuroinflammation USP7 Protein microarray Inhibits USP7 to cause a ubiquitination-dependent degradation of KEAP1 and further induces an NRF2-dependent transcription activation of anti-neuroinflammation genes [71] Evodiamine Antitumor REXO4 DBPP Degrades REXO4 to induce cell death through ROS [101] Forskolin Osteoporosis TGM2 CuAAC Allostericlly activates TGM2 to improve mitochondrial dynamics and ATP production for osteoblast differentiation [80] Fructose-1,6-bisphosphate (FBP) Signaling molecule ALDH2 PAL Inhibits ALDH2 activity and results in cellular ROS upregulation accompanied by mitochondrial fragmentation [96] Gambogic acid Antitumor CNPY3 TPP Facilitates the recruitment of SIRT1 to remove lysine lactylation of CNPY3 and disrupts lysosomal proteins [27] Geldanamycin Antitumor HSP90 SPROX Binds to HSP90's N-terminal ATP binding domain and its ATPase activity [43] Grincamycin B Antitumor IDH1 Protein microarray Inhibits IDH1 to disrupt the cellular redox balance and 2-oxoglutarate homeostasis, thereby triggers ROS accumulation and eventually causes cell death [69] Halorotetin B Antitumor UBE2C PELSA Induces M phase cell cycle arrest [65] Hyperforin Anti-obesity DLAT LiP-SMap Promotes thermogenesis by stimulating AMPK and PGC-1α via a Ucp1-dependent pathway [56] Kurarinone Alleviation of Parkinson's disease sEH SIP + TPP Suppresses sEH to stabilize the level of epoxyeicosatrienoic acids and inhibits GSK3β [47] Ligustroflavone Antitumor BRD4 HiMAP-seq Inhibits BRD4 and down-regulates MYC, NFKB1 and TP53 [106] Manassantin A Antitumor 28 targets (e.g., filamin A and elongation factor 1α) SPROX Induces conformational change in filamin A to interrupt filamin A-HIF1α interaction, thereby inhibits HIF1α [42] Matrine Alleviation of spinal cord injury HSP90 DARTS Enhances the chaperon activity of HSP90, leading to axonal growth [29] Naamidine J Alleviation of acute lung injury CSE1L CuAAC Inhibits SP1 nuclear translocation and suppresses macrophage inflammation [79] Neoeriocitrin Osteogenesis and bone regeneration BECLIN1 TPP Stabilizes BECLIN1 by inhibiting ubiquitination-mediated degradation, thereby increasing autophagy [26] PF403 Antitumor NAMPT TPP Inhibits the catalytic activity of NAMPT, leading to a decrease in the NAD+ concentration [25] Shikonin Antitumor IKKβ/NEMO complex TPP + SIP Destabilizes the IKKβ/NEMO complex to suppresses NF-κB signaling and impairs cell proliferation [48] Silibinin Hepatoprotection ACSL4 TRAP Inhibits ACSL4 enzymatic activity, thereby mitigating the ACSL4-mediated ferroptosis [61] Tetrandrine Antivirus and antitumor LIMP-2 PAL Binds LIMP-2's ectodomain to inhibit lysosomal cholesterol and sphingosine transport [98] Walrobsin A Alleviation of acute kidney injury GPR75 DARTS Inhibits GPR75 to alleviate macrophage oxidative stress and the inflammatory microenvironment [30] Table 1.
Representative example of natural product target Identification.
-
Method Principle Key features Advantages Limitations TSA/CETSA/TPP Ligand binding alters protein thermal stability
Tm shift detected via fluorescence (TSA), immunoblot (CETSA), or MS (TPP)– Label-free
– TSA: purified proteins
– CETSA: cells/lysates
– TPP: proteome-wide MS– No modification
– CETSA: live-cell compatible
– TPP: high-throughput, unbiased– TSA: pure protein only
– CETSA: low throughput
– TPP: costly, long runtime, membrane protein under-representationDARTS Drug binding confers resistance to proteolytic digestion – Label-free
– SDS-PAGE/MS readout– No probe synthesis
– Works with crude extracts
– Detects low-affinity interactions– False positives
– Low-abundance proteins missed
– Protease choice criticalSPROX Ligand stabilizes protein, reducing methionine oxidation rates; quantitated by MS – Chemical denaturation gradient
– MS detection of oxidized/non-oxidized Met peptides– Quantitative affinity data – Requires Met-containing peptides
– No binding site informationSIP Organic solvents precipitate free proteins; ligand-bound proteins remain soluble. – Mixed organic solvent
– MS readout– Good proteome coverage
– No probe modification– In vitro only
– Organic solvent may disrupt some interactionsLiP-MS Ligand-induced conformational changes alter protease accessibility; MS quantifies differential peptides. – Two-step proteolysis
– Peptide-level resolution– Binding site mapping (~12 aa)
– Proteome-wide
– Near-physiological conditions– Requires high sequence coverage
– Cannot localize distal allosteric sitesTRAP Ligand binding changes lysine accessibility to reductive dimethylation – Isotope-coded formaldehyde labeling
– MS readoutNo protein modification – Requires reactive Lys near binding site PELSA Local stabilization upon ligand binding hinders tryptic cleavage; reduced peptide abundance pinpoints binding region – High-concentration trypsin digestion – No modification
Direct binding site information
– High sensitivity– May need optimization for each protein
– Sequence coverage dependentProtein microarrays Immobilized recombinant proteins probed with labeled or unlabeled NP – High-density chip
– Fluorescence/MSI/DESI-MS readout– Ultra-high throughput
– Parallel screening of thousands of proteins– Improper protein folding and truncated fragments
– Non-physiological in vitro environment
– High experimental costCuAAC Azide/alkyne-modified NP; Cu(I) catalyzes cycloaddition with biotin tag for pulldown – Bioorthogonal probe
– Affinity enrichment– Fast, specific, sensitive – Copper cytotoxicity
– Chelation inhibits catalysisSPAAC Strain-promoted cycloaddition; no catalyst – Copper-free
– Ring-strain driven– No metal toxicity
– Live-cell compatible– Slower kinetics (~0.1 M−1·s−1)
– Larger functional groupsiEDDA Tetrazine (Tz) + trans-cyclooctene (TCO) inverse electron-demand Diels-Alder – Extremely fast – Ultrafast kinetics
Bioorthogonal– Bulky Tz/TCO may alter NP properties PAL UV irradiation activates photoreactive group; covalent crosslinking to target – Probe: NP + photoreporter + tag
– UV-induced carbene/nitrene– Captures transient/weak interactions
– Membrane and low-abundance proteins– Probe synthesis may alter activity
– UV damageDBPP NP-PROTAC conjugate recruits E3 ligase; target degradation – Ternary complex formation
– Quantitative proteomics– Detects moderate/weak binders
– Direct pulldown evidence– Complex design
– Costly, platform-dependentμMap Visible-light photocatalyst (Ir) conjugated to NP; Dexter energy transfer activates diazirine probe nearby – Photocatalytic proximity labeling
– 450 nm blue light– No UV damage
- High membrane protein enrichment– Poor tissue penetration (blue light) POST-IT HaloTag-PafA fusion recruited by HTL-NP; pupylation of proximal lysines in live cells/animals – Prokaryotic pupylation system
– Non-diffusive labeling– Live-cell and in vivo
– Preserves transient interactions– Requires genetic fusion
– Engineering of PafA ligaseHiMAP-seq Multiplexed RNA-seq with UMIs; generates Chemical-Induced Gene Signatures (CIGS) for mechanism inference – Pooled sample barcoding
– No mRNA purification– Ultra-high throughput
– Cost-effective, reproducible– Indirect target inference
– Requires validationAI/
BioinformaticsIntegrates multi-omics, network models, machine learning; predicts targets from chemical structure and expression data – Computational pipelines
– Deep learning, network pharmacology– Accelerates hypothesis generation
– Reveals polypharmacology– Prediction accuracy depends on training data
– Requires experimental confirmationTable 2.
Comparative summary of natural product target identification methods.
Figures
(2)
Tables
(2)