-
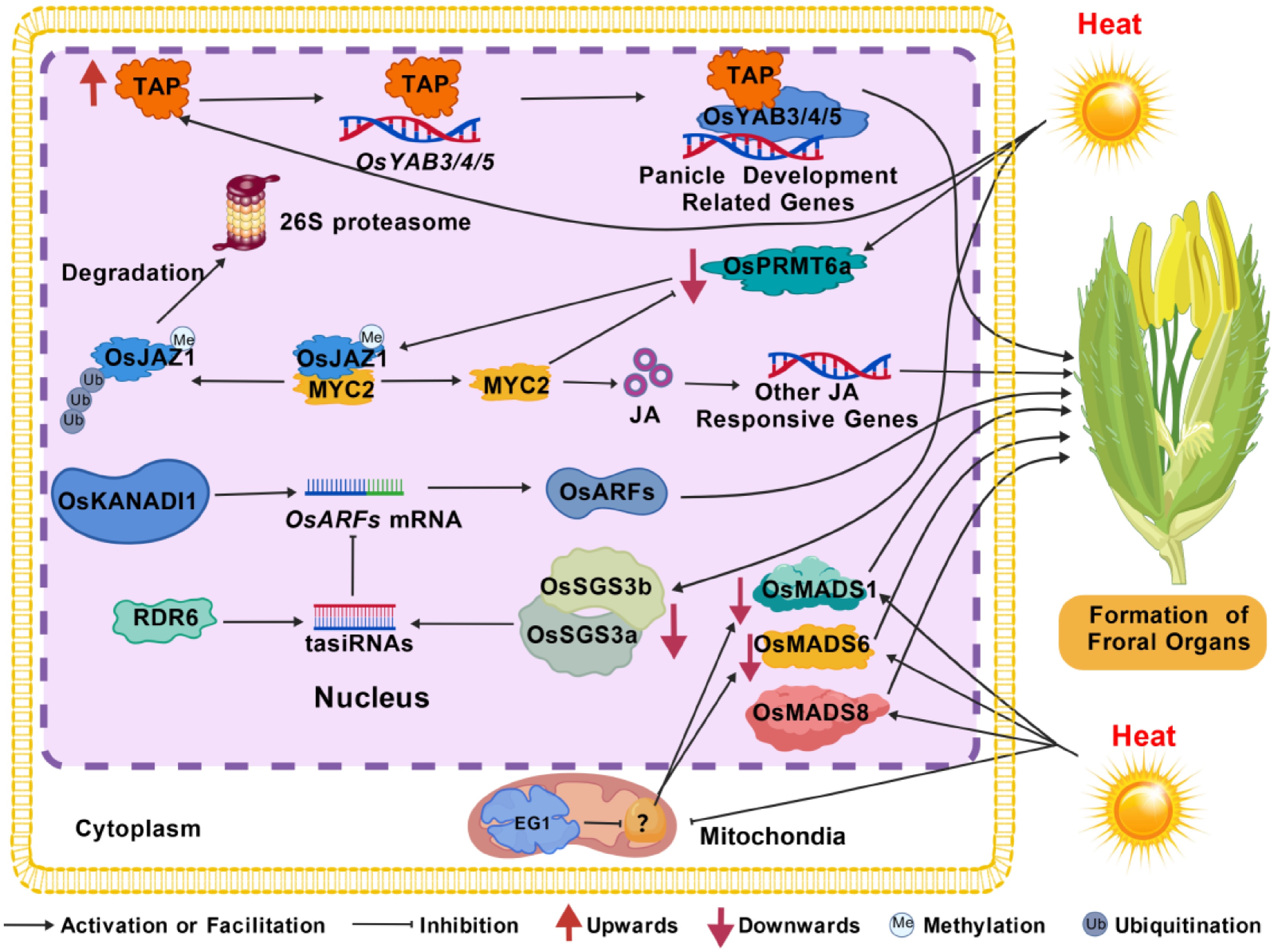
Figure 1.
Molecular regulatory networks governing rice spikelet development under heat stress. Heat stress triggers a cascade of molecular events coordinating spikelet development. Key pathways include: (1) The upregulation of THERMO-SENSITIVE BARREN PANICLE (TAP), which directly regulates and interacts with OsYABBY transcription factors (OsYAB3/4/5) to control inflorescence architecture; (2) Protein arginine methyltransferase OsPRMT6a-mediated methylation promotes the degradation of OsJAZ1, releasing OsMYC2 to activate jasmonic acid (JA)-responsive genes; (3) RNA-dependent RNA polymerase 6 (RDR6) generates trans-acting small interfering RNAs (tasiR-ARFs) that post-transcriptionally regulate AUXIN RESPONSE FACTORS (OsARF2/3), and the transcription of OsARF3A is directly promoted by OsKANADI1; (4) The dsRNA-binding protein OsSGS3a interacts with OsSGS3b to regulate spikelet development via the OsARF pathway; (5) Floral organ identity is maintained by MADS-box transcription factors (OsMADS1/6/8), with the expression of OsMADS1 and OsMADS6 being modulated by the mitochondrial lipase EXTRA GLUME1 (EG1).
-
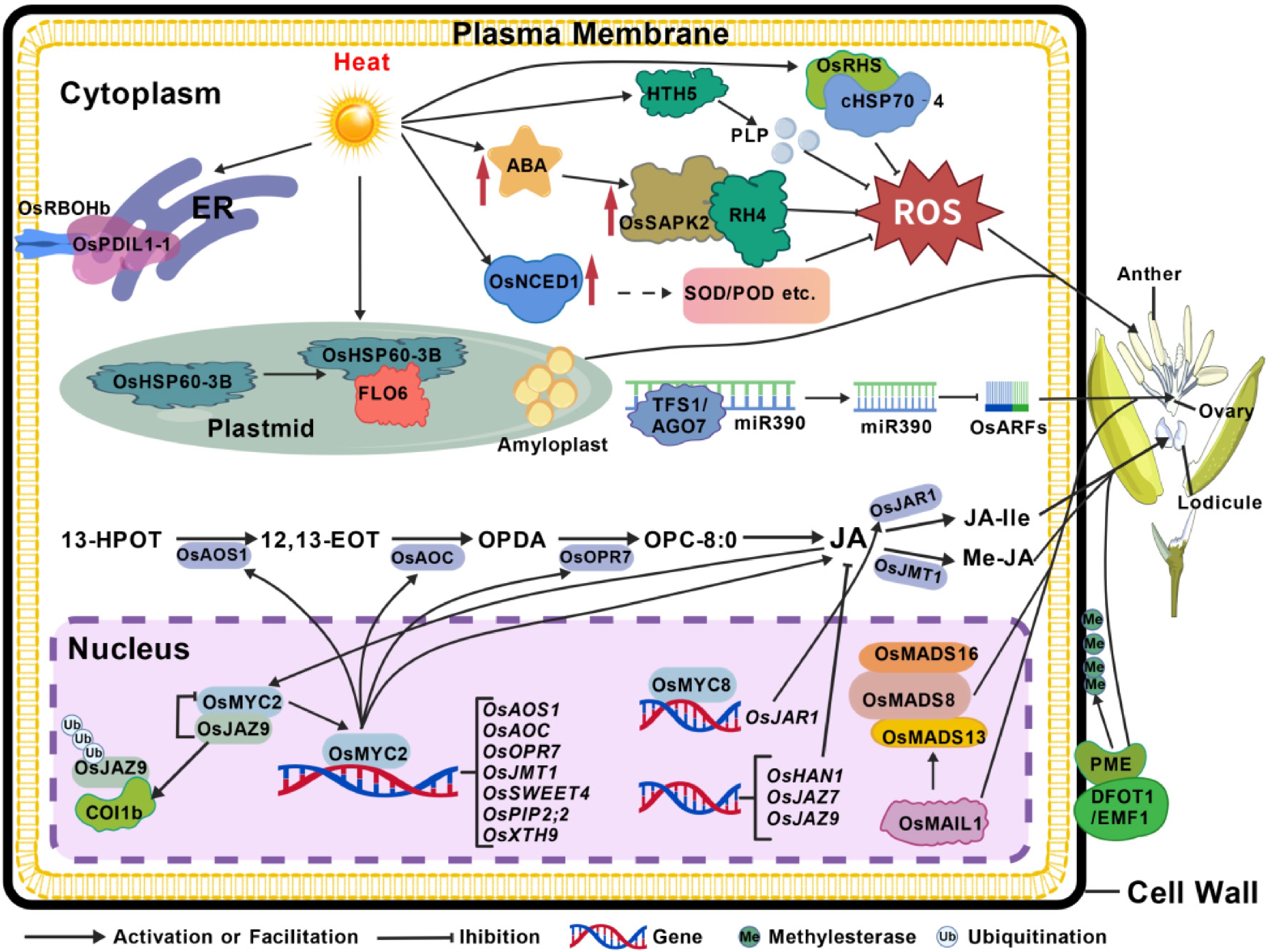
Figure 2.
Molecular networks regulating rice pollen formation, ovary development, and fertilization under heat stress. Heat stress disrupts key reproductive processes, including pollen development, ovary formation, and diurnal flower-opening timing, thereby triggering sophisticated molecular responses to mitigate these damages. The core adaptive mechanisms are outlined below: (1) Pollen development and anther dehiscence: The dynamic balance of reactive oxygen species (ROS) and starch synthesis is crucial for these processes. Multiple mechanisms converge to limit ROS bursts: ABA-activated protein kinase 2 (OsSAPK2) directly interacts with the DEAD-box RNA helicase RH4; CIS-EPOXYCAROTENOID DIOXYGENASE (NCED) expression is upregulated, enhancing the activity of antioxidant enzymes such as SOD and POD; HTH5-encoded pyridoxal 5'-phosphate (PLP) homeostasis protein (PLPHP) increases antioxidant PLP content; REDUCED HEAT STRESS TOLERANCE 1 (OsRHS) inhibits ROS production through interaction with cHSP70-4; and disulfide isomerase-like proteins (PDIL) regulate ROS generation by interacting with RBOHb at the endoplasmic reticulum (ER)-plasma membrane junction. Additionally, rice HEAT SHOCK PROTEIN 60-3B (OsHSP60-3B) interacts with FLOURY ENDOSPERM 6 (FLO6) to ensure proper starch granule formation. (2) Female gametophyte and pistil development: This process involves transcriptional and post-transcriptional regulation of key genes. AGO7 combines with miR390 to form an RNA-induced silencing complex (RISC) that regulates female fertility by inhibiting tasiR-ARFs. MADS8 forms heterodimers with MADS13 and MADS16 to coordinately regulate target genes. Meanwhile, OsMAINTENANCE OF MERISTEM LIKE 1 (OsMAIL1) activates carpel identity genes, including OsMADS13. (3) Diurnal flower-opening timing (DFOT): DFOT1 promotes pectin methylesterase (PME) activity and regulates diurnal flower opening by reducing pectin methylesterification in lodicule cell walls. The JA pathway components OsOPR7, OsMYC2, OsMYC8, OsAOS1, and OsAOC, together with JA inactivation factor OsHAN1 and negative regulators OsJAZ7 and OsJAZ9, jointly fine-tune diurnal flower-opening timing under heat stress.
-
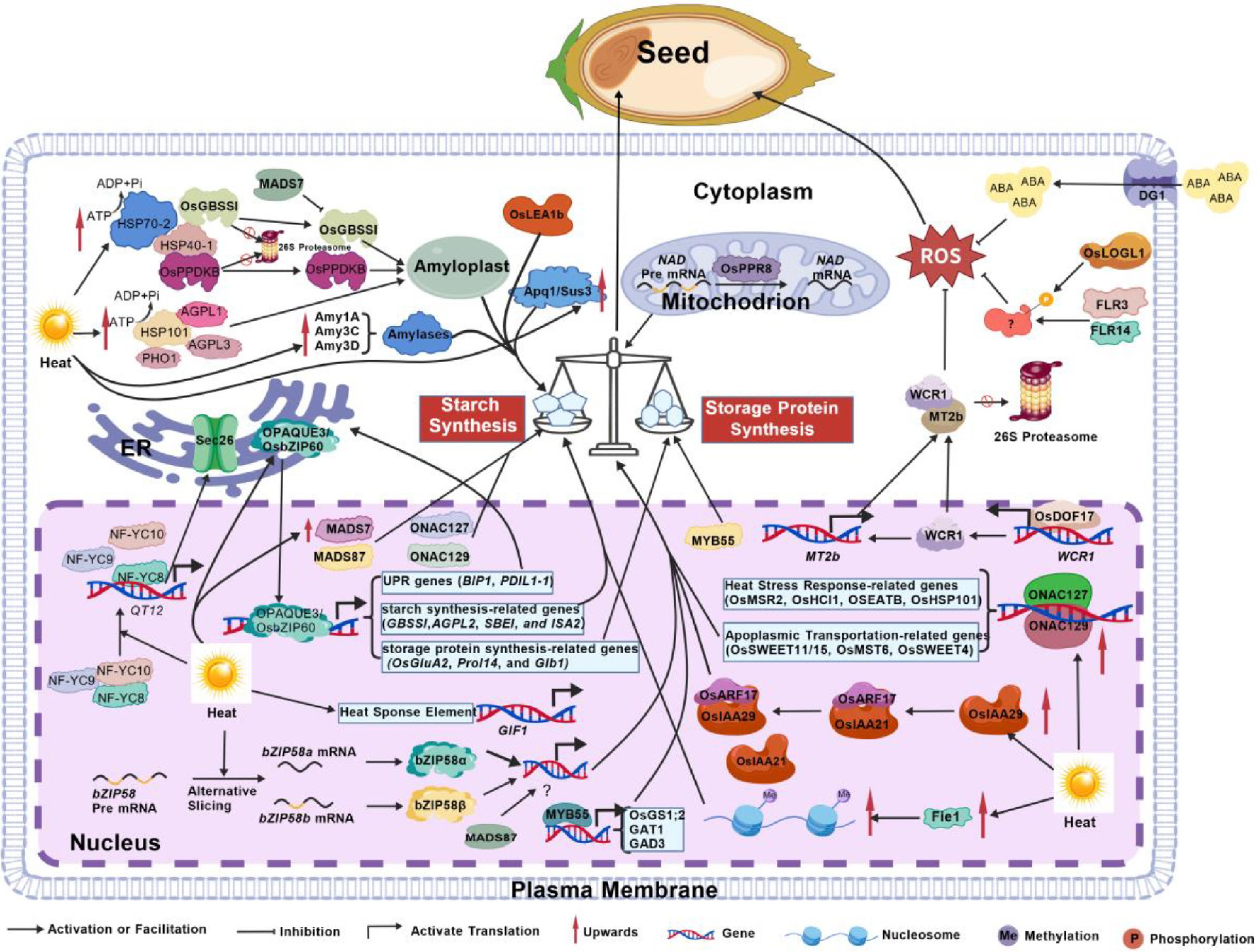
Figure 3.
Molecular mechanisms regulating of rice grain development and quality under heat stress. High temperature affects rice grain development by disrupting the balance between starch and storage protein synthesis and by inducing reactive oxygen species (ROS) accumulation. Key molecular pathways involved in this process are outlined as follows. (1) Starch metabolism and chalkiness formation: α-amylases (including Amy1A, Amy3C, and Amy3D) influence chalkiness formation under thermal stress. Heat shock proteins HSP70-2 and HSP40-1 interact with each other and directly bind key starch synthesis enzymes—GRANULE-BOUND STARCH SYNTHASE (OsGBSSI) and PYRUVATE ORTHOPHOSPHATE DIKINASE (OsPPDKB)—to maintain their stability and activity. HSP101 interacts with starch biosynthesis enzymes such as AGPL1, AGPL3, and PHO1 to regulate starch synthesis and endosperm development. Appearance Quality of Brown Rice 1 (Apq1)/Sucrose Synthase 3 (Sus3) participates in sucrose synthesis, thereby influencing final grain quality. (2) Hormone signaling and redox homeostasis: The transporter DG1 exports abscisic acid (ABA) from vascular tissues into developing caryopses, contributing to stress adaptation. Cytokinin-activating enzyme LONELY GUY-Like 1 (LOGL1), along with FERONIA-LIKE RECEPTORS FLR3 and FLR14, helps maintain redox homeostasis in the endosperm, potentially through phosphorylation of unknown target proteins. (3) Transcriptional and post-transcriptional regulation: OsIAA29 competes with OsIAA21 for binding to OsARF17, thus enhancing the transcriptional activity of OsARF17 and subsequently regulating genes involved in starch and protein synthesis. In addition, transcription factors from multiple families—including MADS-box, MYB, NAC, and bZIP—serve as core regulators of grain quality under high temperature by directly controlling the expression of starch/protein synthesis genes or other heat-responsive genes.
-
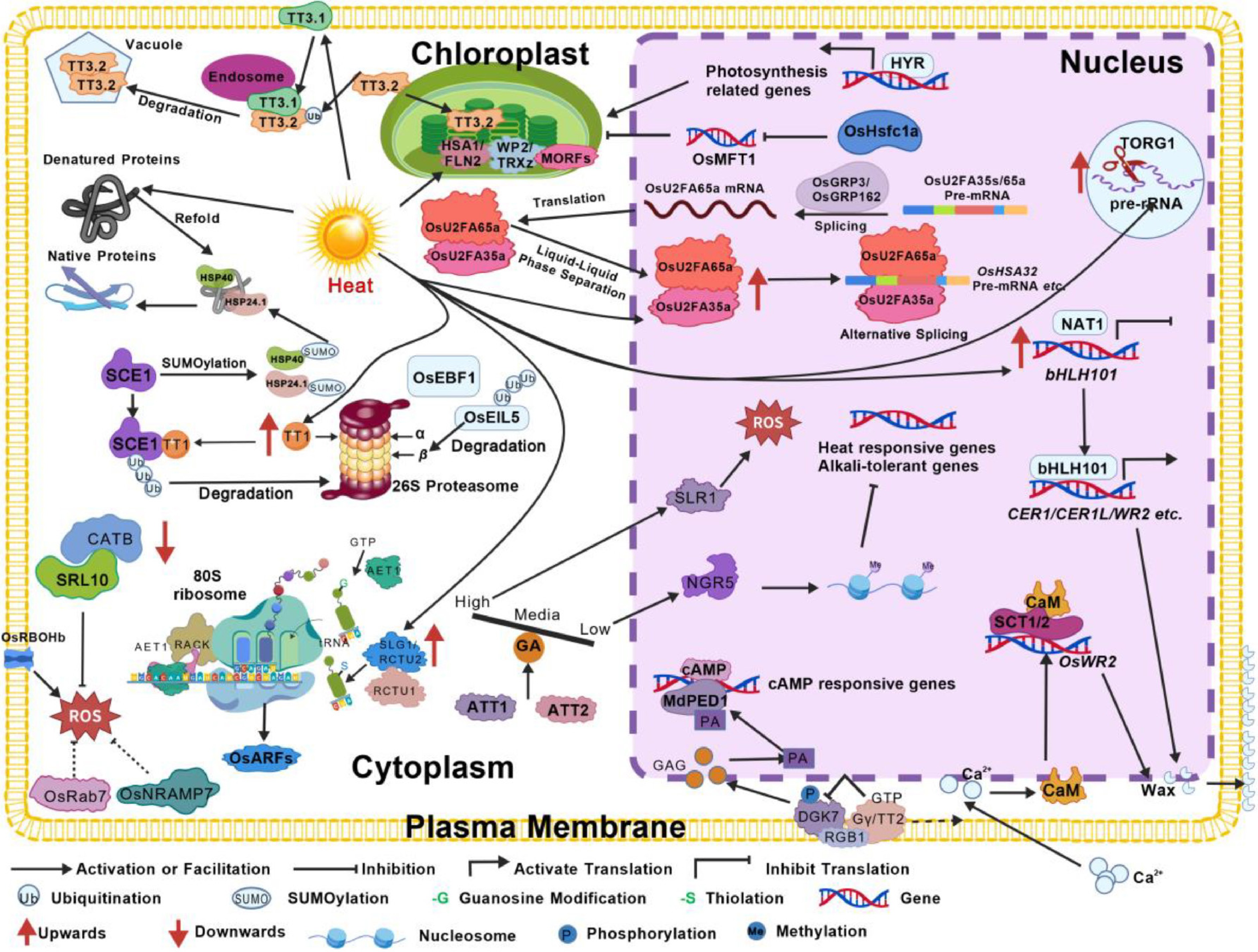
Figure 4.
Molecular networks of entire growth period of rice under high temperature. High-temperature stress triggers multi-level molecular responses that coordinately regulate rice development during both vegetative and reproductive phases. The key mechanisms can be summarized as follows: (1) Phytohormones and signal transduction: ATT1/2 encode GA20-oxidases that fine-tune gibberellin levels, balancing ROS scavenging and epigenetic regulation via the SLR1-NGR5-LC2 complex. TT2 promotes CaM-SCT1 interaction, inhibiting wax synthesis gene OsWR2 expression. TT2 inhibits diacylglycerol kinase 7 (DGK7) activity by dephosphorylation, locking metal-dependent phosphodiesterase (MdPDE1) activity and nuclear translocation, regulating the expression of the second messenger cyclic adenosine monophosphate (cAMP)-responsive genes and enhancing the heat tolerance of rice. NEGATIVE REGULATOR OF THERMOTOLERANCE 1 (NAT1) inhibits bHLH110 expression that promotes the expression of genes such as ECERIFERUM1 (CER1) and CER1L, regulating rice wax synthesis and positively enhancing rice heat resistance; (2) Chloroplast development and photosynthesis: TT3.1 ubiquitinates the chloroplast precursor protein TT3.2 for degradation, and YIELD RICE (HYR) is a key regulator of rice photosynthesis. Heat shock transcription factor OsHsfc1a enhances thermotolerance by regulating MOTHER OF FT AND TFL1 (OsMFT1) and preserving chloroplast structure under heat stress; (3) RNA processing: OsU2AF35a interacts with OsU2AF65a and undergoes liquid-liquid phase separation to promote correct pre-mRNA splicing. OsGRP3 and OsGRP162 interact with spliceosome components to regulate alternative splicing, while TOGR1 maintains rRNA homeostasis. HEAT-STRESS SENSITIVE ALBINO 1 (HSA1)/fructokinase-like protein 2 (FLN2) interacts with thioredoxin z (TRXz) regulates plant chloroplast RNA editing by controlling the redox states of MORFs, thereby contributing to the accurate translation of chloroplast-encoded proteins; (4) Protein synthesis, modification and degradation: AET1 and SLG1/RCTU2 coordinate tRNA modification and translation efficiency. TT1 encodes the α2 subunit of the 26S proteasome, clearing cytotoxic denatured proteins under heat stress and promoting degradation of SCE1 to regulate HSP accumulation by mediating their SUMOylation; (5) ROS and antioxidant regulation: OsRbohB positively regulates ROS generation, while OsRab7, OsNRAMP7, and the SRL10-CATB complex enhance antioxidant capacity and inhibit ROS production.
Figures
(4)
Tables
(0)