-
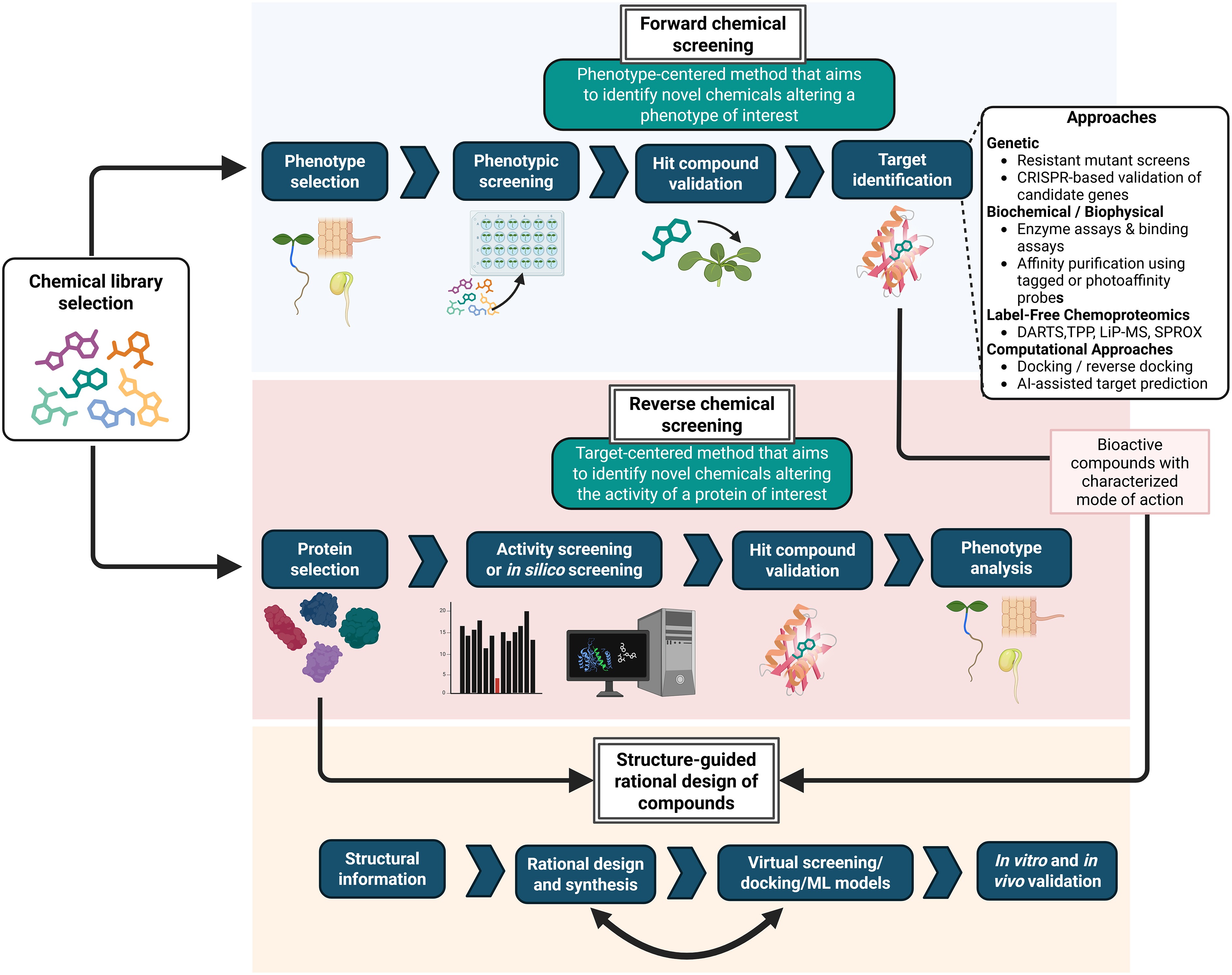
Figure 1.
Screening strategies in chemical genetics research. Two major large-scale screening approaches underpin the discovery of small molecules in plant chemical genetics. Forward chemical screening is phenotype-centered: Compounds from chemical libraries are tested for their ability to modify a developmental or physiological output, followed by extensive validation. Reverse chemical screening is target-centered and aims to identify ligands that interact with a specific protein or enzyme. Each strategy follows a distinct workflow and provides different types of information. In forward screening, a principal challenge is identification of the target compounds, which requires genetic, biochemical, and chemoproteomic methods (see box). Insights from both forward and reverse approaches can subsequently inform the rational design of novel, more selective, or more potent compounds based on known scaffolds and structural data. DARTS: drug affinity responsive target stability; LiP-MS: limited proteolysis coupled with mass spectrometry; SPROX: stability of proteins from rates of oxidation, TPP: thermal proteome profiling. Figure prepared with Biorender.
-
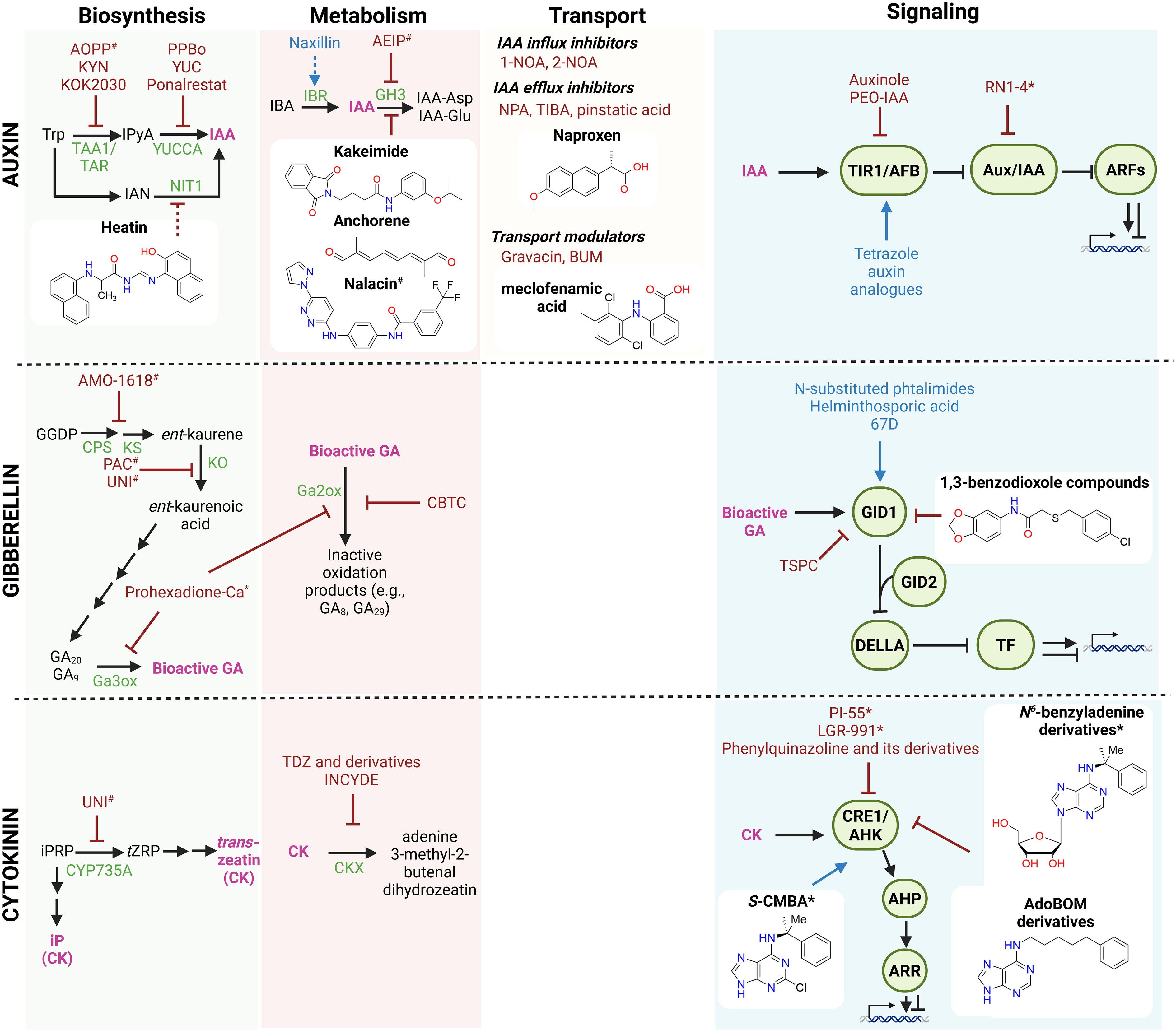
Figure 2.
Small molecules affecting the auxin, gibberellin, and cytokinin pathways. Schematic overview of small molecules that the modulate biosynthesis, conjugation, transport, and signaling of auxin, gibberellin, and cytokinin. Known small molecules with agonist or antagonist activity are indicated in blue and red, respectively. Compounds with isoform-specific effects are marked with an asterisk (*), whereas compounds with broad or off-target effects are marked with a hashtag (#). Recently discovered small molecules (in the past 5 years) are shown together with their chemical structures. Stimulatory interactions are represented by arrow-headed lines, inhibitory interactions are indicated by block-headed lines, and dashed lines indicate unproven or putative interactions. Hormones are shown in pink; precursors and intermediates are shown in black. Biosynthetic and conjugation enzymes and signaling components are depicted in green. Abbreviations: 1-NOA: 1-naphthoxyacetic acid; 2-NOA: 2-naphthoxyacetic acid; AdoBOM: N-(benzyloxymethyl)adenosine; AIEP: adenosine-5′-[2-(1H-indol-3-yl)ethyl]phosphate; AHP: ARABIDOPSIS HISTIDINE PHOSPHOTRANSFER; AMO-1618: 2-isopropyl-4-dimethylamino-5-methylphenyl-1-piperidinecarboxylate; AOPP: L-aminooxyphenylpropionic acid; ARF: AUXIN RESPONSE FACTOR; ARR: ARABIDOPSIS RESPONSE REGULATOR; Aux/IAA: auxin/indole-3-acetic acid repressors; BA: benzyladenine; BUM: 2-[4-(diethylamino)-2-hydroxybenzoyl]benzoic acid; CBTC: methyl 6-chloro-3H-1,2,3-benzodithiazole-4-carboxylate 2-oxide; CK: cytokinin; CKX: CYTOKININ OXIDASE/DEHYDROGENASE; CPS: ent-COPALYL DIPHOSPHATE SYNTHASE; CRE1/AHK: CYTOKININ RESPONSE 1/ARABIDOPSIS HISTIDINE KINASE; CYP735A: cytochrome P450 monooxygenase 735A; GA20ox: GIBBERELLIN 20-OXIDASE; GID: GIBBERELLIN INSENSITIVE DWARF; GH3: GRETCHEN HAGEN 3; IAA: indole-3-acetic acid; IAN: indole-3-acetonitrile; IBA: indole-3-butyric acid; IBR: INDOLE-3-BUTYRIC ACID RESPONSE; INCYDE: inhibitor of cytokinin degradation; iPRP: isopentenyl riboside phosphate; IPyA: indole-3-pyruvic acid; KO: ent-KAURENE OXIDASE; KS: ent-KAURENE SYNTHASE; KYN: L-kynurenine; NIT1: NITRILASE 1; NPA: naphthylphthalamic acid; PAC: paclobutrazol; PEO-IAA: phenyl-ethyl-oxyl–IAA; PPBo: p-phenoxyphenyl boronic acid; RN: RubNeddin; S-CMBA: 2-chloro-N6-(α-methylbenzyl)adenine; TAA1: TRYPTOPHAN AMINOTRANSFERASE OF ARABIDOPSIS 1; TAR: TRYPTOPHAN AMINOTRANSFERASE-RELATED; TDZ: thidiazuron; TIBA: 2,3,5-triiodobenzoic acid; TIR1/AFB: TRANSPORT INHIBITOR RESPONSE 1/AUXIN-SIGNALING F-BOX; TSPC: 3-(2-thienylsulfonyl)pyrazine-2-carbonitrile; tZRP: trans-zeatin riboside-5′-monophosphate; UNI: uniconazole; YUC: YUCCA monooxygenase. Figure prepared with Biorender. Chemical structures drawn with Chemdraw.
-
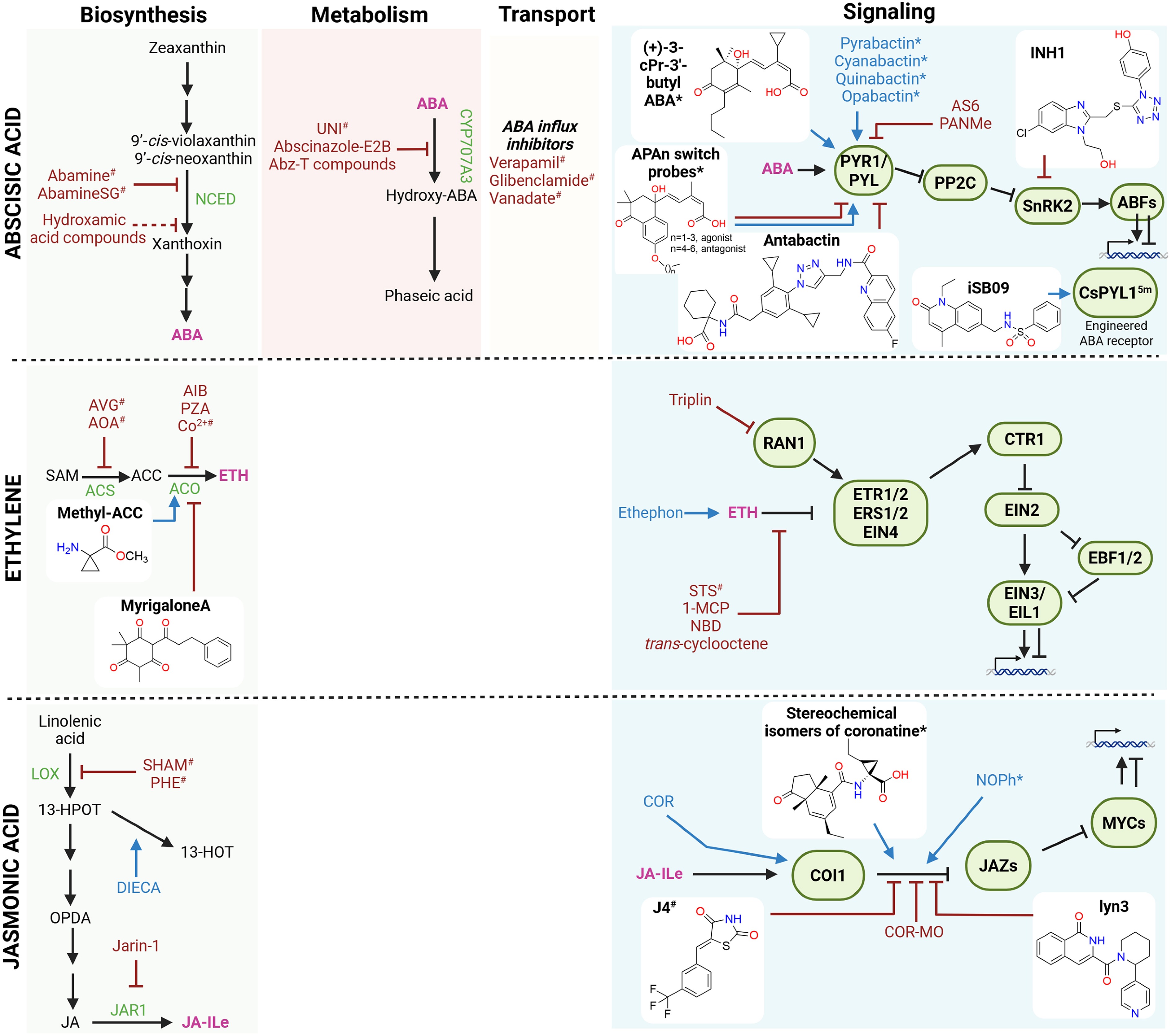
Figure 3.
Small molecules affecting the abscisic acid, ethylene, and jasmonic acid pathways. Schematic overview of small molecules that modulate the biosynthesis, conjugation, transport, and signaling of abscisic acid (ABA), ethylene (ETH), and jasmonic acid (JA). Known small molecules with agonist or antagonist activity are indicated in blue and red, respectively. Compounds with isoform-specific effects are marked with an asterisk (*), whereas compounds with broad or off-target effects are marked with a hashtag (#). Recently discovered small molecules (past 5 years) are shown together with their chemical structures. Stimulatory interactions are represented by arrow-headed lines, inhibitory interactions are indicated by block-headed lines, and dashed lines indicate unproven or putative interactions. Hormones are shown in pink; precursors and intermediates are shown in black. Biosynthetic and conjugation enzymes, as well as their signaling components, are depicted in green. Abbreviations: 1-MCP: 1-methylcyclopropene; 13-HPOT: 13-hydroperoxy-9,11,15-octadecatrienoic acid; 13-HOT: 13-hydroxy-9,11,15-octadecatrienoic acid; ABFs: ABA-RESPONSIVE ELEMENT BINDING FACTORs; Abz-T: abscinazole-T; ACC: 1-aminocyclopropane-1-carboxylic acid; ACS: ACC SYNTHASE; ACO: ACC OXIDASE; AIB: 2-aminoisobutyric acid; AOA: aminooxyacetic acid; AS6: 4′-O-phenylpropynyl ABA derivative; AVG: aminoethoxyvinylglycine; Co2+: cobalt ion; COI1: CORONATINE INSENSITIVE1; COR: coronatine; COR-MO: coronatine O-methyloxime; CTR1: CONSTITUTIVE TRIPLE RESPONSE 1; DIECA: diethyldithiocarbamic acid; EBF1/2: EIN3-BINDING F-BOX proteins 1 and 2; EIN: ETHYLENE INSENSITIVE; EIL1: EIN3-LIKE 1; ERS: ETHYLENE RESPONSE SENSOR; ETR: ETHYLENE RESPONSE; JAR1: JASMONATE RESISTANT 1; JAZ: JASMONATE ZIM-DOMAIN; LOX: lipoxygenase; NCED: 9-cis-epoxycarotenoid dioxygenase; NBD: norbornadiene; OPDA: 12-oxo-phytodienoic acid; PP2C: protein phosphatase 2C; PYR/PYL: PYRABACTIN RESISTANT/PYR-LIKE; PZA: pyrazinamide; RAN1: RESPONSIVE-TO-ANTAGONIST 1; SAM: S-adenosylmethionine; SHAM: salicylhydroxamic acid; SnRK2: SNF1-related protein kinase 2; STS: silver thiosulfate; UNI: uniconazole. Figure prepared with Biorender. Chemical structures drawn with Chemdraw.
-
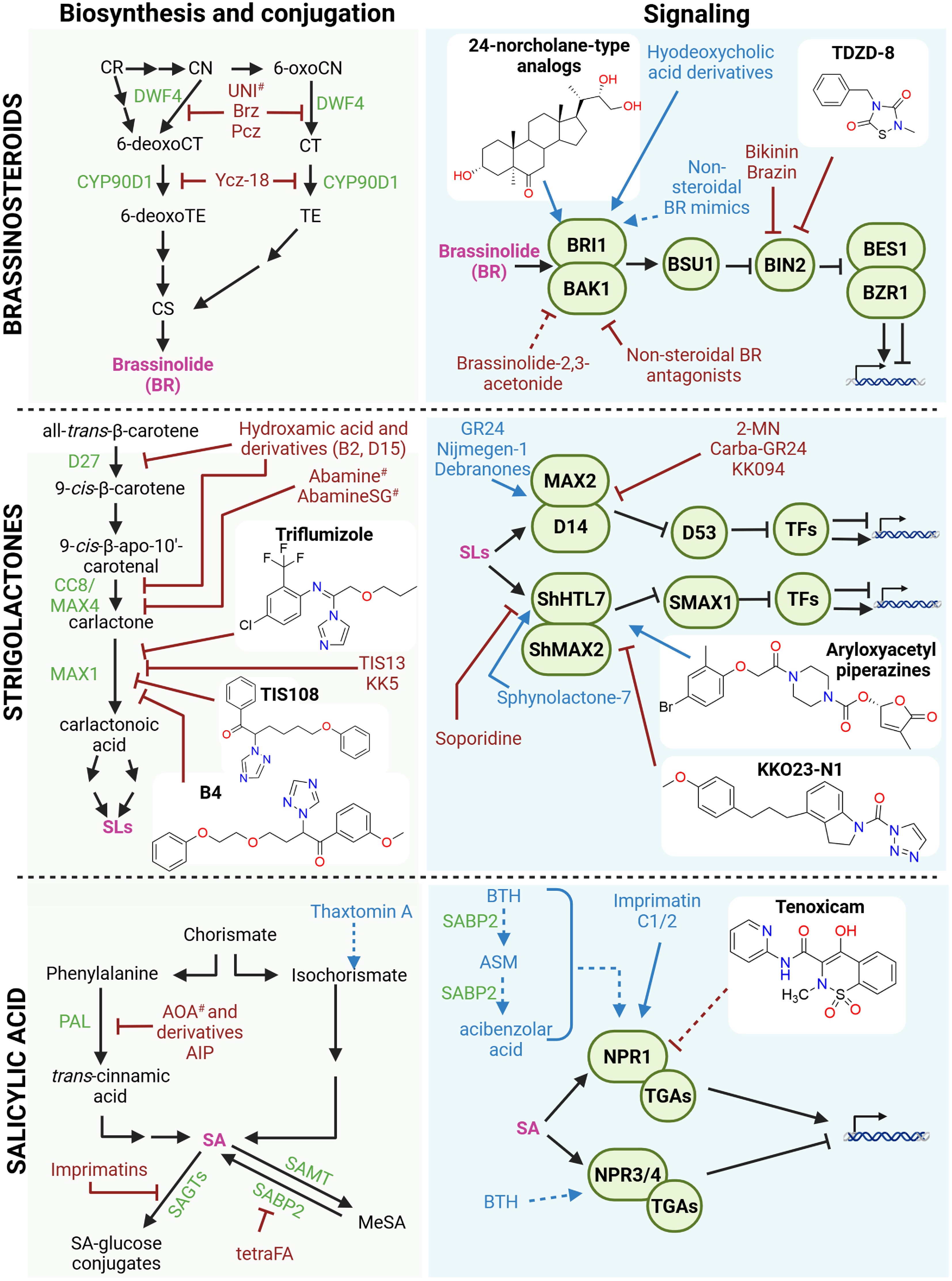
Figure 4.
Small molecules affecting the brassinosteroid (BR), strigolactone (SL), and salicylic acid (SA) pathways. Schematic overview of small molecules that modulate the biosynthesis, conjugation, transport, and signaling of BR, SL, and SA. Known small molecules with agonist or antagonist activity are indicated in blue and red, respectively. Compounds with isoform-specific effects are marked with an asterisk (*), whereas compounds with broad or off-target effects are marked with a hashtag (#). Recently discovered small molecules (past 5 years) are shown together with their chemical structures. Stimulatory interactions are represented by arrow-headed lines, inhibitory interactions are indicated by block-headed lines, and dashed lines indicate unproven or putative interactions. Hormones are shown in pink; precursors and intermediates are shown in black. Biosynthetic and conjugation enzymes, as well as their signaling components, are depicted in green. Abbreviations: 2-MN: 2-methoxy-1-naphthaldehyde; AIP: 2-aminoindan-2-phosphonic acid; AOA: 2-aminooxyacetate; ASM: acibenzolar-S-methyl; BAK1: BRASSINOSTEROID INSENSITIVE 1-ASSOCIATED KINASE 1; BES1: BRI1-EMS-SUPPRESSOR 1; BTH: benzo(1,2,3)thiadiazole-7-carbothioic acid S-methyl ester; BZR1: BRASSINAZOLE RESISTANT 1; BIN2: BRASSINOSTEROID INSENSITIVE 2; Brz: brassinazole; BRI1: BRASSINOSTEROID INSENSITIVE 1; BSU1: BRI1 SUPPRESSOR 1; CCB/MAX4: CAROTENOID CLEAVAGE DIOXYGENASE 8/MORE AXILLARY GROWTH 4; CN: campestanol; CR: campesterol; CS: castasterone; CT: cathasterone; D14: DWARF14; D27: DWARF27; DWF4: DWARF4; MeSA: methyl-salicylic acid; NPR: NONEXPRESSOR OF PATHOGENESIS-RELATED; PAL: PHENYLALANINE AMMONIA-LYASE; Pcz: propiconazole; SABP2: SA-binding protein 2; SAGT: SA glucosyltransferase; SAMT: SA carboxyl methyltransferase; ShHTL7: Striga hermonthica HYPOSENSITIVE TO LIGHT 7; SMAX1: SUPPRESSOR OF MAX1; TE: teasterone; tetraFA: tetra-fluoroacetophenone; TGA: TGACG-BINDING FACTOR. Figure prepared with Biorender. Chemical structures drawn with Chemdraw.
-
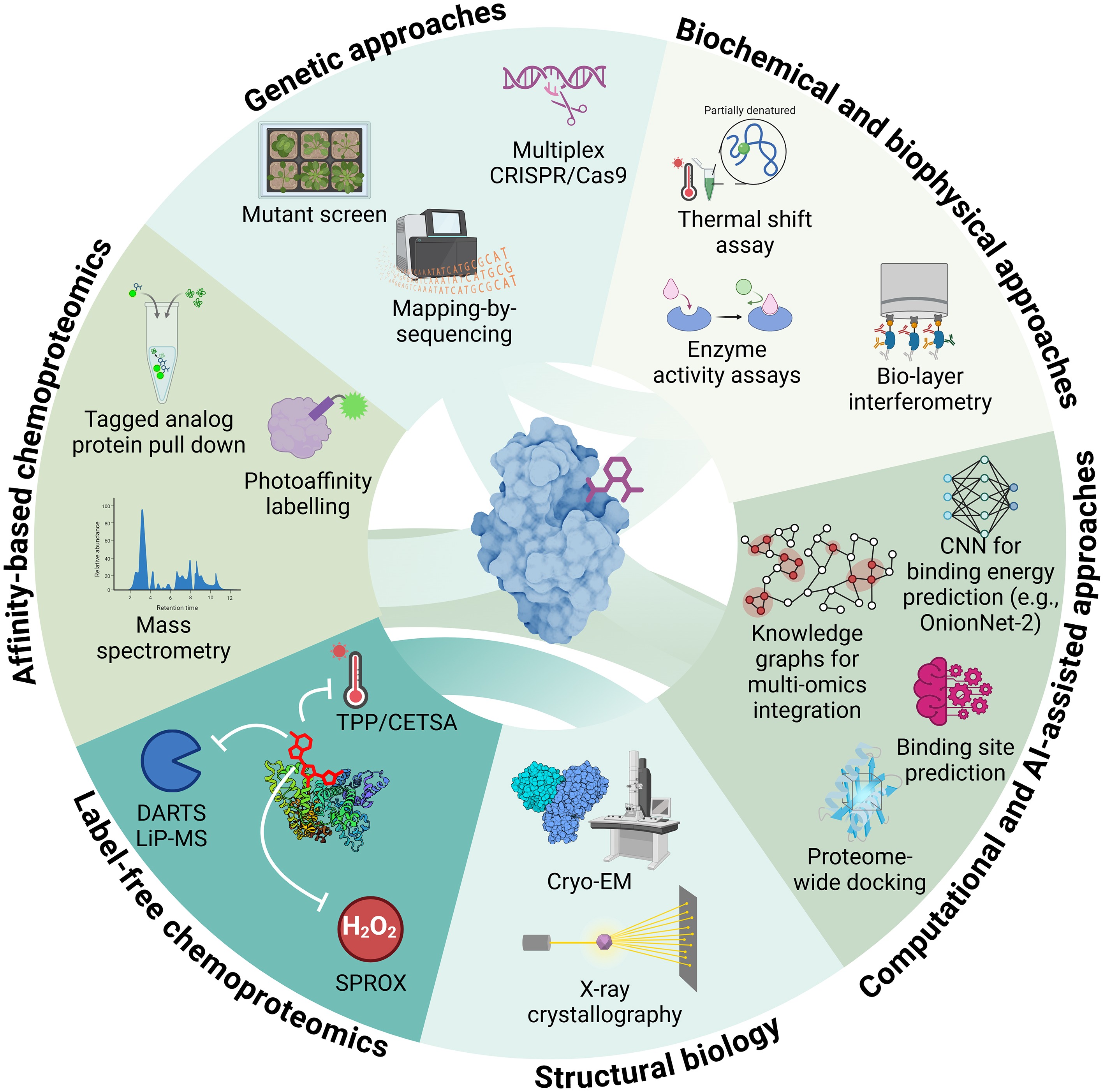
Figure 5.
Integrated strategies for small-molecule target identification. Overview of complementary genetic, biochemical, chemoproteomic, structural, and computational approaches for identifying molecular targets of bioactive compounds in plants. Abbreviations: CETSA, Cellular Thermal Shift Assay; CNN, Convolutional Neural Network; CRISPR/Cas9, Clustered Regularly Interspaced Short Palindromic Repeats/CRISPR-associated protein 9; DARTS, Drug Affinity Responsive Target Stability; LiP-MS, Limited Proteolysis–Mass Spectrometry; SPROX, Stability of Proteins from Rates of Oxidation; TPP, Thermal Proteome Profiling. Figure prepared in Biorender.
Figures
(5)
Tables
(0)