-
The discovery of plant hormones revolutionized our understanding of plant development and physiology and spurred the search for tools to modulate their levels, signaling pathways, and downstream responses. Hormones regulate a wide range of processes, from cell elongation and division, organ growth, and flower and fruit development, to tissue senescence and defense responses[1]. Hence, their pleiotropic functions are essential for nearly every aspect of plant life. Over the past century, intensive research has led to the identification and characterization of the major classes of plant hormones including auxins, gibberellins (GAs), cytokinins (CKs), ethylene (ETH), abscisic acid (ABA), jasmonic acid (JA), salicylic acid (SA), strigolactones (SLs), brassinosteroids (BRs), and peptide hormones[1]. Elucidating their biosynthesis, transport, metabolism, and signaling networks has relied heavily on genetic approaches, particularly through the isolation of hormone-deficient or constitutive mutants. However, such studies often face limitations caused by the functional redundancy in multigene families, species specificity, or the lethality of complete knockouts[2]. Although clustered regularly interspaced short palindromic repeat/CRISPR-associated protein 9 (CRISPR/Cas9) and similar technologies allow the simultaneous targeting of multiple genes, practical challenges remain[3]. A particular problem is transgene delivery and inefficient regeneration capacity in nonmodel species. Moreover, genetic approaches often also lack the temporal resolution and reversibility needed to dissect dynamic hormonal responses.
To address these challenges, small molecules have become valuable tools in plant biology. Such compounds can influence the biosynthesis, conjugation, degradation, or perception of hormones[4−6], and can even modulate specific signaling steps[7−13]. Depending on their structure, small molecules can target either specific proteins or entire protein families, thereby bypassing issues of redundancy[2]. For example, pyrabactin was useful for isolating ABA receptors, which exhibit significant functional redundancy and were unable to be identified using conventional genetic approaches[14]. Small molecules also induce reversible and dose-dependent phenotypes, allowing both temporal and spatial control[2]. This makes them especially useful for studying essential genes or stages where genetic approaches are limited. For example, using the biosynthesis inhibitor phenidone, inhibition of JA responses can be achieved without affecting fertility, which is associated with loss-of-function biosynthesis or signaling mutants[15,16]. Lastly, chemical tools also hold great promise for agriculture by enabling the manipulation of traits without the need for conventional breeding or genetic modification. In the face of global climate change and increasing food demand, the need for such tools to precisely and rapidly modulate plant traits is more urgent than ever.
However, pharmacological approaches also come with important limitations. A major challenge is the risk of low specificity and off-target effects, especially with earlier-generation compounds[17]. For instance, both auxin and ETH biosynthesis rely on pyridoxal phosphate (PLP)-dependent enzymes, and inhibitors such as aminooxyacetic acid (AOA) and aminoethoxyvinylglycine (AVG) can simultaneously block both hormone pathways[17]. Likewise, the cytochrome P450 monooxygenases form a large superfamily, with members involved in the synthesis of multiple hormones, including GAs, BRs, ABA, CKs, and JAs. Azole-type inhibitors like paclobutrazol (PAC) and uniconazole (UNI), commonly used to inhibit GA and ABA synthesis, are, in fact, broad-spectrum inhibitors with cross-pathway effects[18−20]. To address these limitations, since the early 2000s, a second wave of compounds has greatly expanded the available chemical tools through the use of chemical genetics approaches based on large-scale small-molecule screening[2]. Structurally diverse synthetic libraries combined with automated screening and high-throughput bioassays have enabled the identification of these novel compounds (see below).
In the past decade in particular, the field has shifted toward the development of more selective mechanistically defined small molecules that achieve more precise control of hormone signaling. This transition has been driven by structure-guided optimization, which has enabled receptor- and isoform-selective ligands, as well as by more systematic approaches to target validation. These advances have been supported by the increasing availability of structural information on hormone receptors and signaling components, high-throughput and artificial intelligence (AI)-assisted screening, and chemoproteomic strategies for in vivo target identification. In this review, we highlight these recent developments, with an emphasis on small-molecule tools reported within the past 5 years. We provide a broad overview of hormone modulators, and discuss the key challenges and emerging questions that are shaping the future application of chemical biology to understand and engineer the hormonal control of plant development.
-
This section outlines key concepts in chemical genomics studies and provides a framework from which the recent innovations in screening, target identification, and compound design have emerged. Central to chemical genomics studies are high-throughput screens, which can be broadly classified into forward (phenotype-driven) and reverse (target-driven) approaches (Fig. 1). Although both strategies screen chemical libraries, they differ in their goals, experimental design, and strengths, which are summarized below.
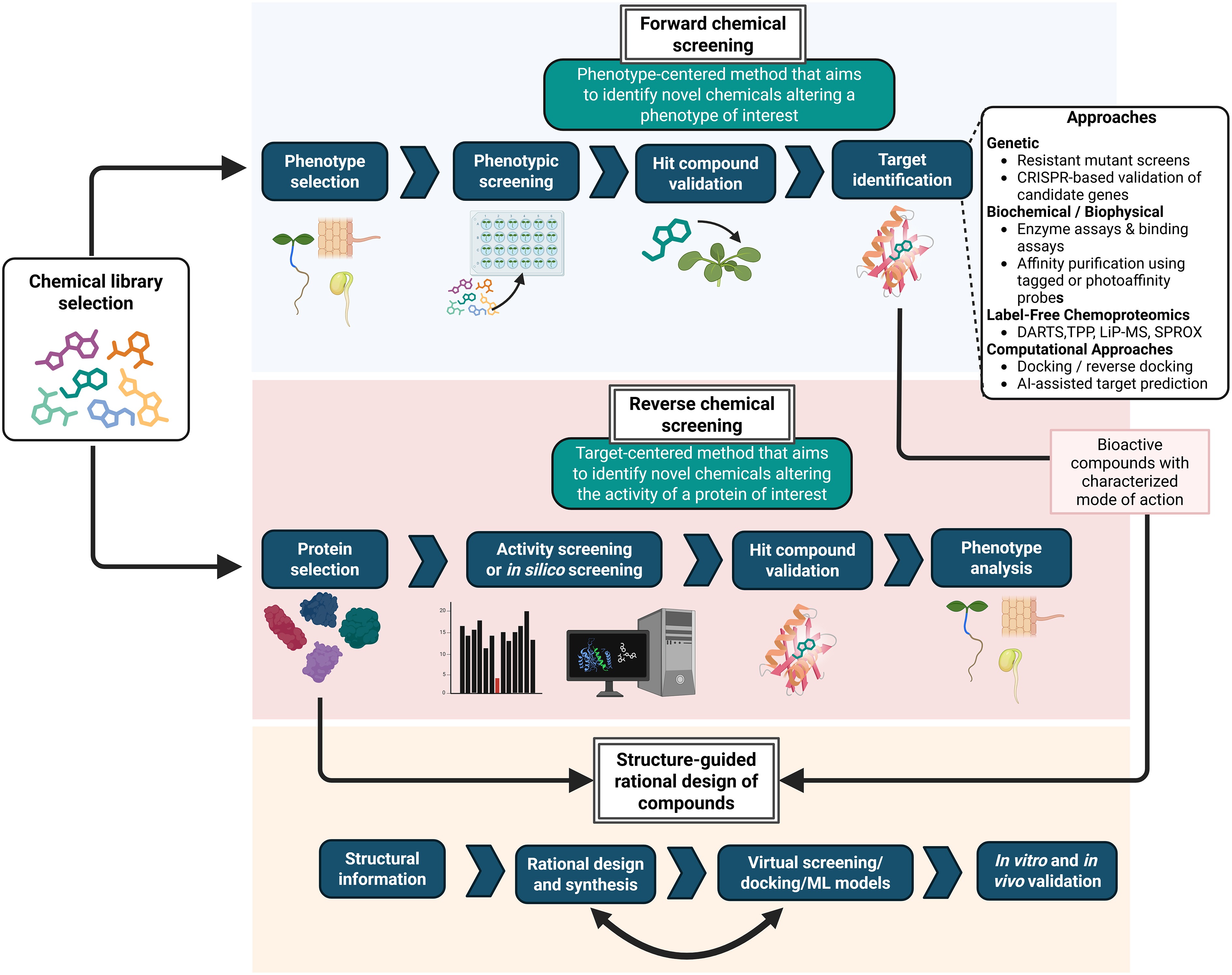
Figure 1.
Screening strategies in chemical genetics research. Two major large-scale screening approaches underpin the discovery of small molecules in plant chemical genetics. Forward chemical screening is phenotype-centered: Compounds from chemical libraries are tested for their ability to modify a developmental or physiological output, followed by extensive validation. Reverse chemical screening is target-centered and aims to identify ligands that interact with a specific protein or enzyme. Each strategy follows a distinct workflow and provides different types of information. In forward screening, a principal challenge is identification of the target compounds, which requires genetic, biochemical, and chemoproteomic methods (see box). Insights from both forward and reverse approaches can subsequently inform the rational design of novel, more selective, or more potent compounds based on known scaffolds and structural data. DARTS: drug affinity responsive target stability; LiP-MS: limited proteolysis coupled with mass spectrometry; SPROX: stability of proteins from rates of oxidation, TPP: thermal proteome profiling. Figure prepared with Biorender.
Forward chemical genetic screening starts from a phenotype or biological process and aims to identify compounds that alter it (Fig. 1)[21]. This phenotype-first approach allows the discovery of novel regulators without prior knowledge of their targets. Such screens often rely on high-throughput assays, e.g., seedlings' growth, germination, or root architecture in 96- or 384-well plates, sometimes coupled with reporter lines like pARR5::GUS for CKs to monitor pathway specificity[12,22−24]. However, after the validation of positive candidates, the major bottleneck is the identification of molecular targets (Fig. 1). This can be addressed using genetic approaches (e.g., screening for ethyl methanesulfonate (EMS)-induced resistant mutants followed by next-generation sequencing), or biochemical strategies, including tagged analogs, photoaffinity labelling, or label-free proteomics techniques (Fig. 1)[14,25,26].
Reverse chemical genetic screening, by contrast, starts with a known gene or protein of interest and seeks small molecules that modulate its activity (Fig. 1)[2]. This target-first strategy can reveal new functional roles or phenotypes and is especially powerful when detailed structural or biochemical information is available. Screens may focus on enzymatic activity, disruption of protein–protein interactions, transcriptional activation, or in silico docking. Once validated, compounds can be used to study the functional role of their in vivo target(s) (Fig. 1). Although more labor-intensive and less scalable in plants than in animal systems, reverse screening is a valuable approach for identifying specific protein modulators.
Both forward and reverse screening also support the optimization of known compounds, improving specificity and reducing off-target effects (Fig. 1). This approach has been widely used in recent years, aided by the previously mentioned technological developments. Structure-guided design has, for example, led to the identification of an agonist selective for the GA receptor GIBBERELLIN INSENSITIVE DWARF 1c (GID1c)[7], in contrast to the broadly acting GA3. Rational design can also convert agonists to antagonists, as demonstrated for opabactin and antabactin[27,28]. These advances relied on structural biology tools such as X-ray crystallography to model the compound–receptor interactions, combined with chemoproteomic strategies using click chemistry to generate and test derivative libraries in vitro. Such integrated approaches highlight the power of modern chemical genomics to refine small-molecule modulators and examine hormone signaling with high precision[27].
-
After nearly two decades of progress, plant chemical genomics has produced a comprehensive toolbox of hormone modulators[29]. For some hormones, a wide variety of small-molecule modulators has been identified; for others, few are available. Throughout this review, particular emphasis is placed on compounds developed or mechanistically refined within the past 5 years. Where older small molecules tools are described alongside recent advances, this is done to highlight how modern screening and target-identification strategies have improved the specificity, selectivity, and mechanistic insight. For completeness, a comprehensive overview of both old and novel compounds is presented in Supplementary Table S1.
Auxin
-
Auxin is one of the best-characterized plant hormones. It regulates nearly every aspect of plant growth and development, including cell elongation, organ patterning, and tropic responses[30]. The complexity of auxin's biology arises from its multiple biosynthetic routes, extensive metabolic control, diverse transport mechanisms, and large, redundant signaling families[30]. Auxin therefore represents an ideal target for chemical perturbation; consequently, the auxin chemical toolbox is both diverse and expansive (Supplementary Table S1, Fig. 2).
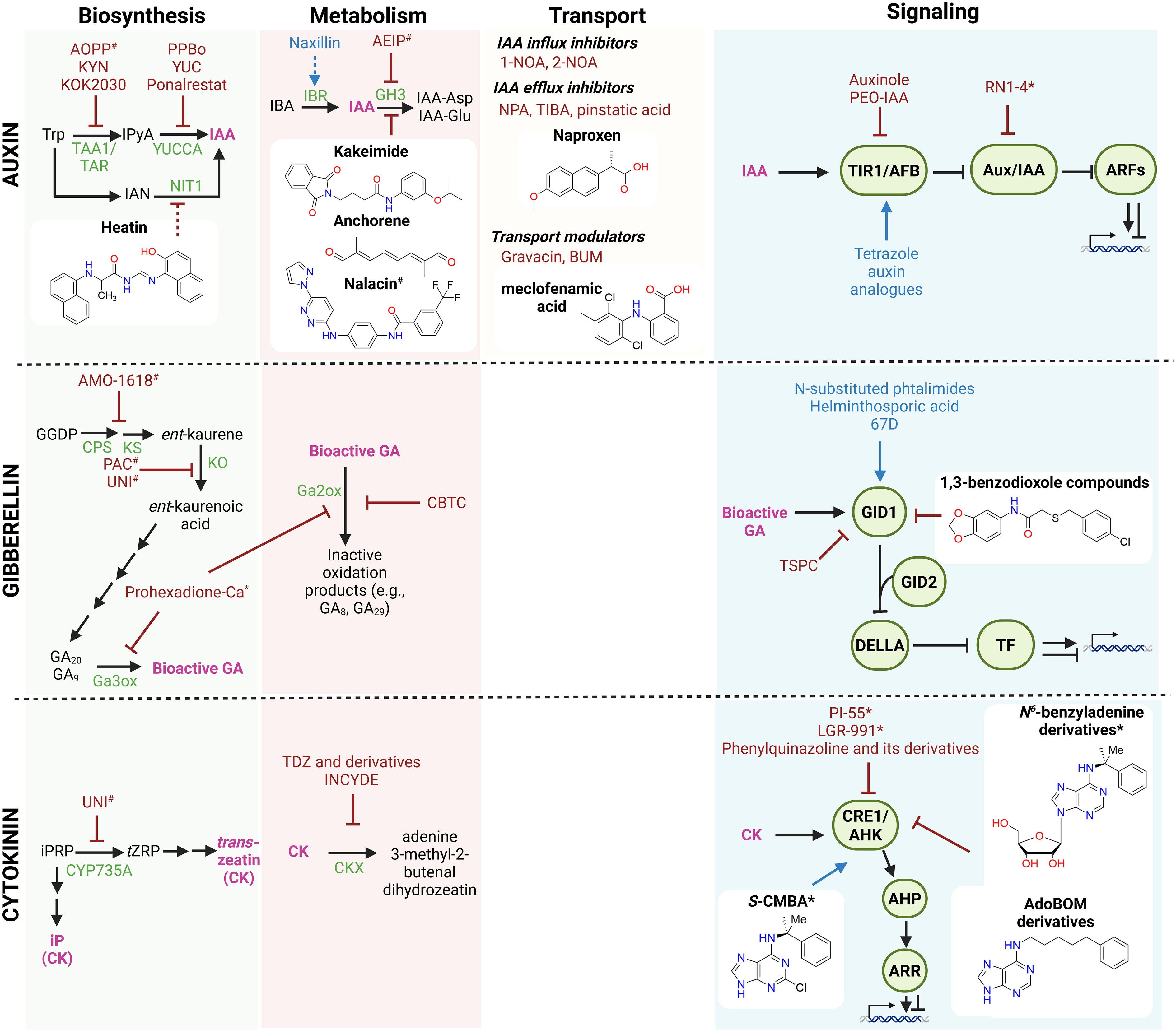
Figure 2.
Small molecules affecting the auxin, gibberellin, and cytokinin pathways. Schematic overview of small molecules that the modulate biosynthesis, conjugation, transport, and signaling of auxin, gibberellin, and cytokinin. Known small molecules with agonist or antagonist activity are indicated in blue and red, respectively. Compounds with isoform-specific effects are marked with an asterisk (*), whereas compounds with broad or off-target effects are marked with a hashtag (#). Recently discovered small molecules (in the past 5 years) are shown together with their chemical structures. Stimulatory interactions are represented by arrow-headed lines, inhibitory interactions are indicated by block-headed lines, and dashed lines indicate unproven or putative interactions. Hormones are shown in pink; precursors and intermediates are shown in black. Biosynthetic and conjugation enzymes and signaling components are depicted in green. Abbreviations: 1-NOA: 1-naphthoxyacetic acid; 2-NOA: 2-naphthoxyacetic acid; AdoBOM: N-(benzyloxymethyl)adenosine; AIEP: adenosine-5′-[2-(1H-indol-3-yl)ethyl]phosphate; AHP: ARABIDOPSIS HISTIDINE PHOSPHOTRANSFER; AMO-1618: 2-isopropyl-4-dimethylamino-5-methylphenyl-1-piperidinecarboxylate; AOPP: L-aminooxyphenylpropionic acid; ARF: AUXIN RESPONSE FACTOR; ARR: ARABIDOPSIS RESPONSE REGULATOR; Aux/IAA: auxin/indole-3-acetic acid repressors; BA: benzyladenine; BUM: 2-[4-(diethylamino)-2-hydroxybenzoyl]benzoic acid; CBTC: methyl 6-chloro-3H-1,2,3-benzodithiazole-4-carboxylate 2-oxide; CK: cytokinin; CKX: CYTOKININ OXIDASE/DEHYDROGENASE; CPS: ent-COPALYL DIPHOSPHATE SYNTHASE; CRE1/AHK: CYTOKININ RESPONSE 1/ARABIDOPSIS HISTIDINE KINASE; CYP735A: cytochrome P450 monooxygenase 735A; GA20ox: GIBBERELLIN 20-OXIDASE; GID: GIBBERELLIN INSENSITIVE DWARF; GH3: GRETCHEN HAGEN 3; IAA: indole-3-acetic acid; IAN: indole-3-acetonitrile; IBA: indole-3-butyric acid; IBR: INDOLE-3-BUTYRIC ACID RESPONSE; INCYDE: inhibitor of cytokinin degradation; iPRP: isopentenyl riboside phosphate; IPyA: indole-3-pyruvic acid; KO: ent-KAURENE OXIDASE; KS: ent-KAURENE SYNTHASE; KYN: L-kynurenine; NIT1: NITRILASE 1; NPA: naphthylphthalamic acid; PAC: paclobutrazol; PEO-IAA: phenyl-ethyl-oxyl–IAA; PPBo: p-phenoxyphenyl boronic acid; RN: RubNeddin; S-CMBA: 2-chloro-N6-(α-methylbenzyl)adenine; TAA1: TRYPTOPHAN AMINOTRANSFERASE OF ARABIDOPSIS 1; TAR: TRYPTOPHAN AMINOTRANSFERASE-RELATED; TDZ: thidiazuron; TIBA: 2,3,5-triiodobenzoic acid; TIR1/AFB: TRANSPORT INHIBITOR RESPONSE 1/AUXIN-SIGNALING F-BOX; TSPC: 3-(2-thienylsulfonyl)pyrazine-2-carbonitrile; tZRP: trans-zeatin riboside-5′-monophosphate; UNI: uniconazole; YUC: YUCCA monooxygenase. Figure prepared with Biorender. Chemical structures drawn with Chemdraw.
Classical auxin tools, developed over the past few decades, can be broadly categorized as biosynthesis and transport inhibitors (Fig. 2, indicated in red). Biosynthesis inhibitors primarily target the main indole-3-pyruvate (IPyA) pathway, and include either broad-acting PLP-dependent enzyme inhibitors, such as L-aminooxyphenylpropionic acid (AOPP), or more specific hydroxy- and methoxy-type pyruvamines, such as L-kynurenin or KOK2030, which block the first enzymatic step[4,31,32]. Inhibitors of the second step, catalyzed by YUCCA enzymes, include yucasin and p-phenoxyphenyl boronic acid (PPBo)[33,34]. These compounds were instrumental in establishing the physiological importance of the IPyA route. Similarly, transport inhibitors, including the efflux inhibitor naphthylphthalamic acid (NPA), influx inhibitors like 1-naphthoxyacetic acid (1-NOA), and compounds affecting the transporters' localization, such as 2,3,5-triiodobenzoic acid (TIBA), provided early evidence for carrier-mediated polar auxin transport and its role in development[35−37]. Synthetic auxins (e.g., 2,4-dichlorophenoxyacetic acid and naphthaleneacetic acid) and antagonists (e.g., PEO-IAA, auxinole), on the other hand, have been used to explore auxin signaling[38].
More recent advances have focused on providing novel tools to manipulate intracellular auxin levels by allowing the selective modulation of specific biosynthetic branches or conjugation pathways (Fig. 2). A notable example is the discovery of a new auxin biosynthesis regulator by van der Woude and colleagues[39]. In a chemical screen for thermomorphogenesis regulators they identified N-[(2-hydroxy-1-naphthyl)methylene]-2-(1-naphthylamino)propanohydrazide or heatin, which rescued the high temperature-induced hypocotyl elongation phenotype and was shown to impact auxin biosynthesis. Structure–activity relationship (SAR) analyses and click-chemistry-based proteomics implicated members of the NITRILASE 1 family, which convert indole-3-acetonitrile (IAN) to indole-3-acetic acid (IAA), as likely targets. Heatin therefore represents the first chemical tool enabling the study of auxin biosynthesis pathways parallel to the IPyA branch. These had previously been difficult to dissect because of the dominance and redundancy of the IPyA pathway, and may be very useful for disentangling the physiological roles of different auxin synthesis routes.
Increasing auxin levels by limiting its conjugation is also an attractive strategy, as it bypasses the previously mentioned complexity of the biosynthesis pathways. Recent efforts have focused on identifying the inhibitors of auxin-conjugating GRETCHEN HAGEN3 (GH3) proteins (Fig. 2)[40]. Kakeimide and nalacin were discovered through phenotype-based screening for root-related auxin phenotypes[6,23]. Although both compounds share a benzamide core, enzyme kinetics, molecular docking, and X-ray crystallography analyses revealed differences in their isoforms' specificity. Both compounds inhibit auxin-conjugating Type II GH3s at low micromolar concentrations. However, nalacin additionally targets the JA-conjugating AtGH3.11 through distinct binding modes within the active site of GH3. These properties therefore facilitate the selective examination of GH3 subgroups, although potential JA-related side effects must be considered when using nalacin. These studies illustrate the power of acquiring structural information to understand the precise binding modes, something that was not possible with earlier inhibitors. In addition, compared with the older inhibitor adenosine-5′-[2-(1H-indol-3-yl)ethyl]phosphate (AIEP), which was only functional in vitro, kakeimide and nalacin display robust activity in vivo and do not affect SA levels[6,23,41]. Notably, kakeimide is also active across multiple species beyond Arabidopsis. Classical genetics and physiological studies can still reveal novel regulators as well. Recently, anchorene, a naturally occurring apocarotenoid breakdown product, was shown to also inhibit GH3 activity, making it the first known endogenous auxin conjugation inhibitor[42]. Interestingly, it is structurally very distinct from kakeimide and nalacin (Fig. 2). Therefore, it can be an inexpensive alternative for its large-scale production and use in modulating auxin levels.
Recent advances in structural biology have also refined our understanding of how auxin transport is inhibited[43]. Cryo-electron microscopy (cryo-EM) examination of the structures of PINOID (PIN) proteins, which are auxin exporters, revealed that NPA binds directly and stabilizes PINs in an inward-open conformation, preventing efflux activity[43]. Such structural insights provide a rational basis for designing next-generation inhibitors or for refining current ones. Nontraditional inhibitors have also emerged. The nonsteroidal anti-inflammatory drug (NSAID) naproxen competitively inhibits PIN-mediated auxin efflux[44], with cryo-EM revealing a distinct binding mode compared with NPA. In contrast, meclofenamic acid, another NSAID, also blocks auxin transport, but does so by interfering with TWISTED DWARF1 (TWD1), an immunophilin that regulates ABCB auxin transporters via cytoskeletal reorganization[45].
Advances in auxin signaling have been slower, although two earlier developments provide interesting avenues for future research (Fig. 2). The rational design of tetrazole-auxins has yielded an auxin agonist with selective binding to TRANSPORT INHIBITOR RESPONSE 1 (TIR1) over AUXIN-SIGNALING F-BOX 5 (AFB5), which enables the exploration of receptor-specific auxin responses[46]. Downstream of their perception, RubNeddins provide a powerful tool for dissecting the function of individual Aux/IAA family members[13], whose study has been complicated by gene family redundancy. These compounds were discovered via a genome-wide chemical screen looking for tools that selectively stabilize particular Aux/IAA proteins by preventing their neddylation and degradation.
Gibberellins
-
Gibberellins are diterpenoid plant hormones that regulate diverse developmental processes, including cell elongation, seed germination, and flowering[47]. Because GA drives shoot elongation, its modulation has long been exploited in agriculture to reduce plant height and prevent lodging in cereal crops, leading to the development of widely used GA biosynthesis inhibitors[48]. However, very little progress has been made in GA-related chemical biology in recent years, with only a few newer examples of signaling-related small molecules (Fig. 2, indicated by the molecular structures in white boxes).
Traditional GA-related small molecules mostly affect biosynthesis and can be grouped into three broad classes: Broad-spectrum cyclase inhibitors (e.g., AMO-1618 and chlormequat chloride), triazole-based P450 monooxygenase inhibitors (e.g., PAC and UNI), and 2-oxoglutarate mimics (e.g., prohexadione-Ca) (Fig. 2)[49−51]. Many of these compounds, like PAC and UNI, are widely used to control GA levels, but off-target effects (e.g., ABA biosynthesis), limit their use[18]. Prohexadione-Ca additionally blocks GA-conjugating GA2ox enzymes. More selective inhibitors, such as methyl 6-chloro-3H-1,2,3-benzodithiazole-4-carboxylate 2-oxide (CBTC), were identified via recombinant enzyme assays with radiolabeled substrates. These inhibitors enhance seed germination by limiting the inactivation of GA[52]. Despite these issues, the identification of specific inhibitors of GA biosynthesis or tools that can effectively control the oxidation and conjugation of GA has been limited in recent years. This is likely because of the complexity of the multiple redundant pathways, highlighting the need for further exploration of these metabolic routes.
In parallel, recent efforts have focused on modulating the perception of GA through the identification of receptor-selective agonists and inhibitors. Because the bioactive GAs GA1 and GA4 are difficult to synthesize at scale, several recent studies have aimed to identify low-cost synthetic GA mimics. One major advance was the identification of N-phthalimide derivatives, such as AC94377 and 67D, which activate GA signaling and promote germination (Fig. 2)[53,54]. Molecular docking and binding assays confirmed that AC94377 selectively binds GIBBERELLIN INSENSITIVE DWARF 1c (GID1c), one of the three GA receptors in Arabidopsis thaliana, without affecting the other two. This specificity provided new insights into the function of receptor subtypes, revealing that GID1c plays a distinct role in GA signaling, particularly in seed germination. This would not have been possible with the use of natural GAs, and it highlights the possibilities of designing specific inhibitors for understanding physiological function.
GA antagonists have also been developed to inhibit the perception of GA. An early example is 3-(2-thienylsulfonyl)pyrazine-2-carbonitrile (TSPC), identified in a yeast two-hybrid screen for compounds that disrupt the GID1–DELLA interaction[55]. TSPC blocks GA signaling in vivo and induces GA-deficient phenotypes. More recently, AI-driven screening approaches have enabled the rational design of novel inhibitors. Wang et al. used virtual docking to screen chemical libraries for GID1-binding compounds and identified 1,3-benzodioxole derivatives such as HTS05309[7]. HTS05309 blocks the perception of GA and induces dwarfism in treated plants. Compared with TSPC, which disrupts the formation of the GID1–DELLA complex without confirmed receptor binding, HTS05309 represents a more specific and mechanistically validated tool for studying the perception of GA. Together, these examples illustrate how recent chemical and computational approaches enable receptor- and pathway-specific modulation of GA signaling, providing precision that was previously unachievable with classical biosynthesis inhibitors.
Cytokinin
-
Cytokinins are adenine-derived plant hormones that regulate shoot apical meristem activity, branching, and leaf senescence[56]. Their roles in modulating the shoot-to-root ratio, senescence, and morphogenesis make them central in both basic research and agricultural applications. However, compared with other hormones, the availability of selective chemical tools to manipulate CK pathways has been limited, especially in terms of controlling biosynthesis. Similar to GAs, most recent developments have focused on the development of receptor-specific agonists and antagonists using structural information to guide the rational design of ligands.
Classical CK modulators act at the levels of biosynthesis, degradation, or perception. The biosynthesis of CKs involves isopentenyltransferases (IPTs) producing N6-(Δ2-isopentenyl)adenosine (iP riboside), which can be hydroxylated by CYP735A to trans-zeatin riboside, before LONELY GUY (LOG) enzymes release the bioactive free base, trans-zeatin, which is the active hormone[57]. Despite a well-characterized pathway, few inhibitors exist. The triazole-based P450 inhibitor UNI blocks CYP735A's activity and reduces trans-zeatin levels, but its broad off-target effects limit its use for examining CK biosynthesis with precision (Fig. 2)[58]. The inhibition of CK degradation, on the other hand, can be used to increase CK levels. Phenylurea-like compounds (e.g., thidiazuron) and INCYDE irreversibly block CYTOKININ OXIDASE/DEHYDROGENASE (CKX) enzymes, preventing degradation and elevating endogenous CK levels[59,60]. These compounds, while not specific to individual CKX isoforms, offer a practical route for enhancing CK in plants, particularly in biotechnological and agricultural contexts. CK perception modulators have traditionally been N6-substituted adenine analogs that mimic endogenous CKs and compete for receptor binding. In Arabidopsis, these receptors include ARABIDOPSIS HISTIDINE KINASE 2 (AHK2), AHK3, and CYTOKININ RESPONSE 1 (CRE1)/AHK4[57]. Earlier receptor antagonists, such as 2-hydroxy-3-methyl benzyl adenine (PI-55) and 6-(2,5-dihydroxybenzylamino)purine (LGR-991), have been valuable for dissecting receptors' function[61,62]. PI-55 preferentially blocks CRE1/AHK4, whereas LGR-991 has broader receptor-binding activity, highlighting the potential for the rational design of isoform-specific receptor antagonists.
More recent work has expanded N6-substituted adenine analogs to generate receptor-specific agonists and antagonists. Using structural modification, the CK reporter line pARR5::GUS, and other CK bioassays, stereochemically defined molecules were developed with striking receptor selectivity. For example, N6-((S)-α-methylbenzyl)adenosine and related molecules act as antagonists of CRE1 and AHK2, whereas the structurally similar analogs (S-MBA, S-FMBA, and 2-chloro-N6-[α-methylbenzyl]adenine) behave as agonists with a preference for AHK3 (Fig. 2, with the specificity of compounds indicated by *)[24,63]. These subtle structural changes allow the ligands to switch between agonist and antagonist roles, demonstrating the precision that is achievable through rational design. Additional advances have come from the discovery of N6-substituted adenosine and N6-(benzyloxymethyl)adenosine (AdoBOM) derivatives that appear to bind outside the canonical ligand pocket[12,63]. Docking studies suggest that these compounds interact with allosteric sites, inducing conformational shifts that alter receptor activity, similar to the mechanism of older phenylquinazoline derivatives[64].
Compared with older CK modulators, the newest small molecules thus offer enhanced receptor selectivity; the ability to act as agonists or antagonists, depending on the context; and novel allosteric modes of action. Classical compounds still have value for controlling endogenous CK levels, though novel tools are required.
Abscisic acid
-
ABA is recognized for its pivotal role in seed dormancy, drought tolerance, and other osmotic stress responses. Because of its central role in stress adaptation, ABA is a major target for chemical modulation[65]. Compared with other stress hormones ETH and JA, ABA has a relative lively field of chemical biology, with tools developed to affect different aspects of ABA biology (Fig. 3). Most older chemicals affect endogenous ABA levels by interfering with its biosynthesis, metabolism, or transport. Most recent progress, however, has focused on perception and signaling, yielding a new generation of selective receptor agonists, antagonists, and synthetic receptor–ligand systems.
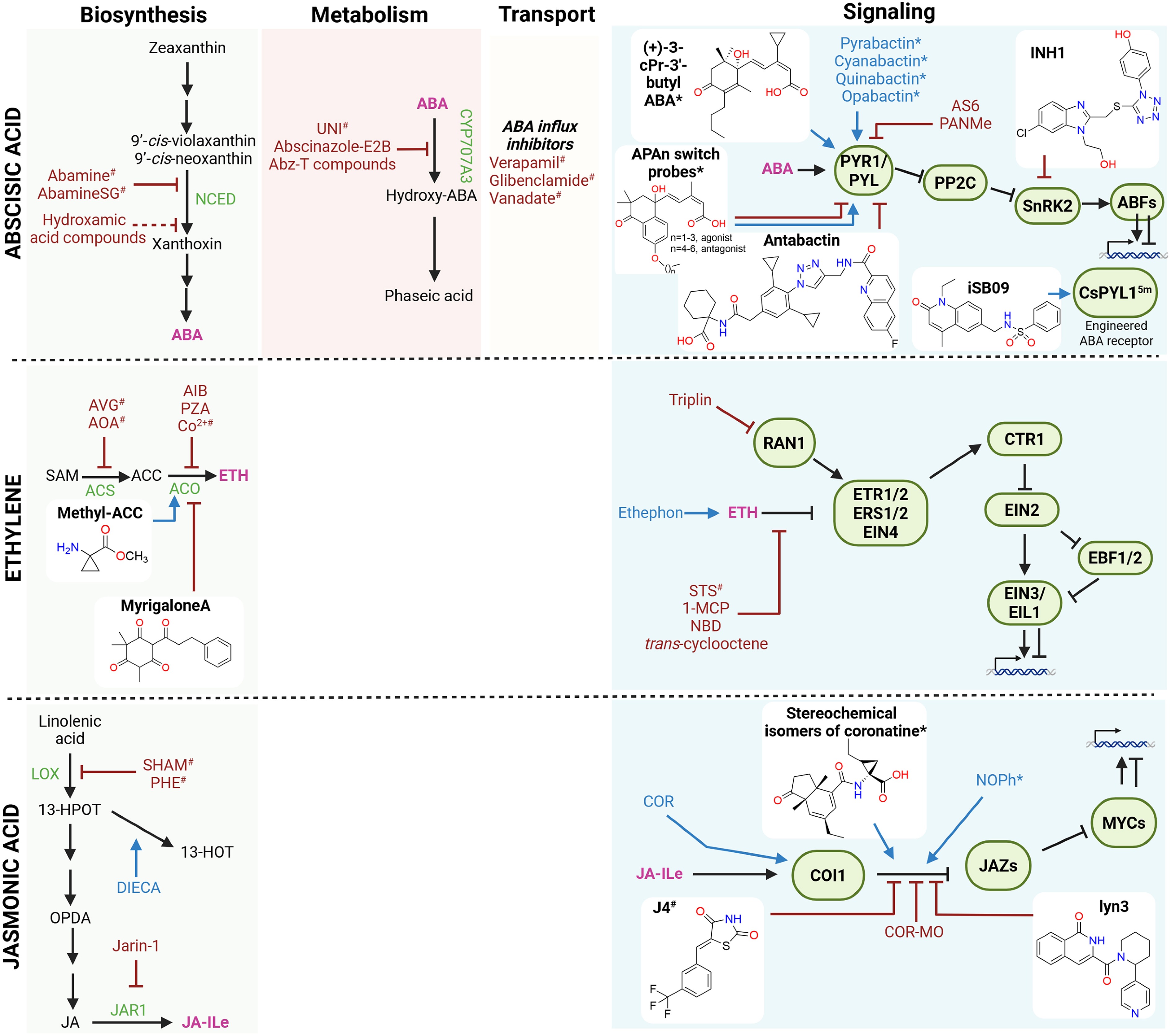
Figure 3.
Small molecules affecting the abscisic acid, ethylene, and jasmonic acid pathways. Schematic overview of small molecules that modulate the biosynthesis, conjugation, transport, and signaling of abscisic acid (ABA), ethylene (ETH), and jasmonic acid (JA). Known small molecules with agonist or antagonist activity are indicated in blue and red, respectively. Compounds with isoform-specific effects are marked with an asterisk (*), whereas compounds with broad or off-target effects are marked with a hashtag (#). Recently discovered small molecules (past 5 years) are shown together with their chemical structures. Stimulatory interactions are represented by arrow-headed lines, inhibitory interactions are indicated by block-headed lines, and dashed lines indicate unproven or putative interactions. Hormones are shown in pink; precursors and intermediates are shown in black. Biosynthetic and conjugation enzymes, as well as their signaling components, are depicted in green. Abbreviations: 1-MCP: 1-methylcyclopropene; 13-HPOT: 13-hydroperoxy-9,11,15-octadecatrienoic acid; 13-HOT: 13-hydroxy-9,11,15-octadecatrienoic acid; ABFs: ABA-RESPONSIVE ELEMENT BINDING FACTORs; Abz-T: abscinazole-T; ACC: 1-aminocyclopropane-1-carboxylic acid; ACS: ACC SYNTHASE; ACO: ACC OXIDASE; AIB: 2-aminoisobutyric acid; AOA: aminooxyacetic acid; AS6: 4′-O-phenylpropynyl ABA derivative; AVG: aminoethoxyvinylglycine; Co2+: cobalt ion; COI1: CORONATINE INSENSITIVE1; COR: coronatine; COR-MO: coronatine O-methyloxime; CTR1: CONSTITUTIVE TRIPLE RESPONSE 1; DIECA: diethyldithiocarbamic acid; EBF1/2: EIN3-BINDING F-BOX proteins 1 and 2; EIN: ETHYLENE INSENSITIVE; EIL1: EIN3-LIKE 1; ERS: ETHYLENE RESPONSE SENSOR; ETR: ETHYLENE RESPONSE; JAR1: JASMONATE RESISTANT 1; JAZ: JASMONATE ZIM-DOMAIN; LOX: lipoxygenase; NCED: 9-cis-epoxycarotenoid dioxygenase; NBD: norbornadiene; OPDA: 12-oxo-phytodienoic acid; PP2C: protein phosphatase 2C; PYR/PYL: PYRABACTIN RESISTANT/PYR-LIKE; PZA: pyrazinamide; RAN1: RESPONSIVE-TO-ANTAGONIST 1; SAM: S-adenosylmethionine; SHAM: salicylhydroxamic acid; SnRK2: SNF1-related protein kinase 2; STS: silver thiosulfate; UNI: uniconazole. Figure prepared with Biorender. Chemical structures drawn with Chemdraw.
Classical ABA modulators span biosynthesis and catabolism inhibitors, as well as receptor-targeted agonists and some antagonists. Biosynthesis inhibitors include 9-cis-epoxycarotenoid dioxygenase (NCED) inhibitors such as abamine and hydroxamic acids, which reduce ABA levels (Fig. 3)[66,67]. Triazole-based molecules like UNI, and more selective analogs such as abscinazole-E2B, on the other hand, function as competitive inhibitors of ABA catabolism and increase endogenous ABA[20,68]. Both classes, however, suffer from off-target effects, as they can bind to other CAROTENOID CLEAVAGEDEOXYGENASEs (CCDs) or P450 monooxygenases, respectively, making them less useful. These limitations motivated a second wave of ABA-related chemical biology focused on receptor-targeted tools. Sulfonamide-based agonists such as pyrabactin, quinabactin, and the highly potent opabactin, among others, can induce ABA responses with differing selectivity for specific PYRABACTIN RESISTANT (PYR)/PYR-LIKE (PYL)/REGULATORY COMPONENTS OF ABA RECEPTORS (RCAR) isoforms[14,69,70]. Opabactin, discovered through in silico docking, exhibits even greater potency than ABA itself (e.g., a lower nanomolar range)[70].
Recent advances have exploited click-chemistry to generate a large library of opabactin derivatives, leading to the discovery of antabactin[27,28]. Click-chemistry has enabled the diversification of the opabactin scaffold, allowing a broad exploration of structure–function relationships that would have been difficult to achieve with conventional synthesis. Unlike the earlier sulfonamides, antabactin functions as a broad-spectrum antagonist, resulting from the addition of a triazole residue to the ABA receptor-binding sulfonamide moiety. It therefore represents a significant advance over earlier antagonists such as AS6 or 4′-O-phenylpropynyl ABA derivatives (e.g., PanMe)[71,72], which are ABA analogs with more restricted receptor coverage or limited chemical stability.
Building on the same theme, Che et al. produced alkylated variants of phenyl-based iso-ABA, a synthetic ABA analog. These derivatives (APAn) were designed to switch between agonist and antagonist activity depending on the length of the 6′-alkoxyl chain (Fig. 2)[73]. In a separate effort, Yoshida et al., developed 3′-alkyl-ABA derivatives, which act solely as agonists, with the receptor affinity varying with the alkyl chain's length[74]. Both studies illustrate how rational modification of the side chains can fine-tune receptors' activity. The 3′-alkyl-ABA derivatives have some limitations; although chemically more stable than classical ABA or sulfonamide agonists, they are less potent, being active in the micromolar range.
Beyond receptor-level modulation, recent small molecules have also been used to probe downstream signaling. This approach is particularly noteworthy and relatively unique among plant hormone chemical biology because it enables exploration of the physiological roles of downstream components (Fig. 3). High-throughput screening identified INH1 and its related analogs as inhibitors of SNF1-related protein kinase 2 (SnRK2), the central effectors released through the repression of PROTEIN PHOSPHATASE 2C (PP2C) upon ABA perception[75]. By directly blocking SnRK2-mediated phosphorylation, INH1 suppresses ABA-induced gene expression and physiological responses. Although related compounds like INH3-4 are less specific and can affect other kinases, INH1 is relatively selective for SnRK2 and shows activity across multiple species. These inhibitors provide a new entry point into the ABA signaling cascade, complementing receptor-targeted approaches and broadening the experimental toolbox. Collectively, biosynthesis and catabolism inhibitors provide broad modulation but suffer from off-target effects, whereas receptor agonists and antagonists allow isoform-specific control. Downstream inhibitors such as INH1 or receptor-specific agonists and antagonists complement these approaches by targeting signaling effectors directly, enabling functional dissection of ABA pathways with greater precision.
A noteworthy recent innovative development is the creation of orthogonal ABA signaling systems (Fig. 3)[11]. By engineering Citrus sinensis PYL receptors with altered ligand specificity, researchers developed CsPYL15m variants that no longer recognize endogenous ABA but instead respond to synthetic ligands such as sulfobactin and iSB09. This synthetic rewiring enables the conditional activation of ABA signaling in Arabidopsis, independent of endogenous hormone fluctuations.
Ethylene
-
Ethylene is the only gaseous plant hormone, which makes direct experimental manipulation technically challenging. Nevertheless, it plays central roles in development and stress physiology, regulating processes such as fruit ripening, abscission, senescence, and responses to flooding, wounding, or salinity[76]. Therefore, because of its economic importance, numerous efforts have been made to identify chemical tools that interfere with ethylene production or perception. Unfortunately, since ethylene metabolism and transport are not well characterized, the toolkit is restricted to modulating biosynthesis and signaling, as indicated by the lack of available chemical tools (Fig. 3).
Ethylene biosynthesis proceeds via a two-step pathway: S-adenosylmethionine (SAM) is converted to 1-aminocyclopropane-1-carboxylic acid (ACC) by ACC synthases (ACS), and ACC is then oxidized by ACC oxidases (ACO) to produce ethylene. Classical modulators target these enzymes: Vinylglycine analogs such as AVG inhibit ACS, whereas cobalt ions and ACC analogs such as aminoisobutyric acid (AIB) interfere with ACO (Fig. 3)[77−79]. However, these molecules suffer from limited specificity and off-target effects[17], and AIB requires millimolar concentrations to be active. Sun et al. identified pyrazinamide (PZA) and its related analogs as more potent ACO inhibitors[22]. Compared with AIB, these compounds are approximately two orders of magnitude more potent and are therefore increasingly used as experimental tools to block ethylene responses.
Chemical interference at the receptor level is also well established. It has traditionally relied on silver ions, which replace copper cofactors in the receptors, or on strained alkenes (Fig. 3)[10,80]. The most successful receptor-targeting compound to date is 1-methylcyclopropene (1-MCP), a high-affinity ethylene competitor that effectively delays ripening and senescence and is widely applied after harvest[10]. However, because 1-MCP is gaseous as well, its agricultural use requires specialized handling and remains impractical for field applications.
Given the availability of several effective classical tools, comparatively fewer fundamentally new chemical strategies have emerged for ethylene-related chemical biology in recent years. One interesting route is the development of cheap alternatives to elevate the levels of ACC, the soluble precursor of ethylene, in plants, as ACC itself is rather expensive. A promising development is methyl-ACC (MeACC), a methylated derivative of ACC with enhanced lipophilicity and uptake (Fig. 3)[81]. In planta, MeACC is hydrolyzed to ACC, most likely by an unknown esterase, thereby releasing the immediate precursor of ethylene. Application of MeACC induces robust ethylene responses, including the triple response phenotype, and its improved membrane permeability suggests its potential as a practical ethylene-releasing compound.
Natural products are also gaining attention because of their structural diversity and regulatory advantages. Myrigalone A, a flavonoid-derived allelochemical isolated from Myrica gale, was recently shown to inhibit seed germination and seedling growth in a species-specific manner[82]. Resistance screens in Dictyostelium and subsequent docking studies identified ACO enzymes as likely targets, making Myrigalone A one of the first naturally occurring small molecules reported to block ACO in vivo. Such natural products highlight the potential of chemical ecology as a source of novel hormone pathway regulators. Nevertheless, although Myrigalone A is active in a similar concentration range to PZA, it appears to display distinct species specificity, an aspect that warrants further systematic investigation.
Overall, ethylene-related chemical biology is dominated by long-established inhibitors and receptor antagonists that remain highly effective. Recent advances, including modulators derived from natural products and ACC-increasing chemicals have expanded the toolkit incrementally rather than fundamentally. These developments nonetheless highlight opportunities for future efforts aimed at improving their specificity, tunability, and applicability across species and experimental contexts.
Jasmonates
-
Jasmonates are lipid-derived hormones that are central to plant defense, as well as to developmental processes such as root elongation, flower development, and senescence[83]. Certain classical tools affect early JA biosynthesis (Fig. 3), targeting 13-lipoxygenase (LOX) activity (e.g., phenidone, SHAM), and are therefore not fully specific (Fig. 3)[15,84]. Conversion of JA to its bioactive form, JA-isoleucine (JA-Ile), can be prevented by jarin-1[5]. However, a recent study in 2023 showed that jarin-1's action is species-specific, as it blocks root elongation in Arabidopsis and Medicago truncatula but fails to suppress JA responses in Solanum lycopersicum and Brassica nigra[85]. These findings suggest either differential uptake or the presence of alternative bioactive JAs in some species, highlighting the need for more versatile inhibitors of JA-Ile formation.
Major progress has been made in JA signaling (Fig. 3). These advances were largely driven by the structure-guided derivatization of coronatine, a phytotoxin produced by Pseudomonas syringae that acts as a JA agonist[86], and by screening-based identification of receptor antagonists. Perception of JA relies on CORONATINE INSENSITIVE 1 (COI1), an F-box protein that, upon binding of JA-Ile, promotes the interaction between COI1 and JAZ repressors, leading to subsequent JAZ degradation and activation of JA genes' expression[83]. Recent structure-guided derivatization of coronatine has produced novel derivatives, such as its phenyloxime derivative NOPh[87]. Compared with coronatine, which broadly promotes the degradation of JAZ proteins, NOPh preferentially destabilizes JAZ9 and JAZ10, allowing a partial uncoupling of different JA responses. Interestingly, even stereochemical isomers of coronatine itself were able to differentially affect COI1–JAZ interactions, specifically acting on JAZ9[88].
High-throughput screening strategies have also yielded important discoveries, identifying antagonists that are structurally different from coronatine. Chini et al. combined yeast two-hybrid assays with a JAZ1-GUS reporter to identify novel antagonists, leading to the discovery of J4 (Fig. 3)[89]. J4 broadly blocked JA responses in planta but also interfered with certain TIR1–IAA interactions, suggesting potential crosstalk between jasmonate and auxin signaling pathways. In parallel, taking advantage of novel computational strategies, structure-based in silico screening of large compound libraries has emerged as a complementary approach to identify JA inhibitors[90]. Lyn3, identified by this strategy, inhibits JA responses by binding COI1 directly, further demonstrating the feasibility of computational design in plant hormone research. Both J4 and Lyn3 provide novel chemical scaffolds for developing JA antagonists, though they act at low-to-mid micromolar concentrations, whereas COR-Mo is active in the submicromolar range[89−91].
Together, these recent advances provide highly selective, mechanistically precise tools that enable the functional exploration of JA signaling at multiple levels, overcoming the limitations of classical biosynthesis inhibitors.
Brassinosteroids
-
Brassinosteroids (BRs) are steroidal hormones that are essential for cell expansion, vascular differentiation, reproductive development, and stress responses[92]. The biosynthesis of BRs originates from the mevalonate pathway, and the production of bioactive brassinolide proceeds through a complex metabolic grid of hydroxylation, oxidation, and epimerization reactions[93]. Two partially redundant routes, the early and late C-6 oxidation pathways, generate multiple intermediates, complicating genetic analyses and creating opportunities for chemical intervention (Fig. 4). Key enzymes include cytochrome P450 monooxygenases such as DWARFED4 (DWF4) and CONSTITUTIVE PHOTOMORPOGENIC AND DWARFISM (CPD), which catalyze the rate-limiting hydroxylation steps. Because of this, common nonspecific P450 inhibitors like UNI can induce BR-deficient phenotypes[94]. The development of brassinazole (Brz), a triazole that specifically inhibits DWF4-mediated C-22 hydroxylation, was a breakthrough (Fig. 4)[95]. Brz became the gold-standard tool for probing BRs' function in planta. Unlike Brz, which has been in use for over two decades, the yucaizol (Ycz) series of compounds represents a more recent generation of biosynthesis inhibitors that selectively block the late biosynthetic steps of BR catalyzed by CYP90D1[96]. Such tools are instrumental for specific exploration of the BR pathway and are complements to earlier chemicals like Brz.
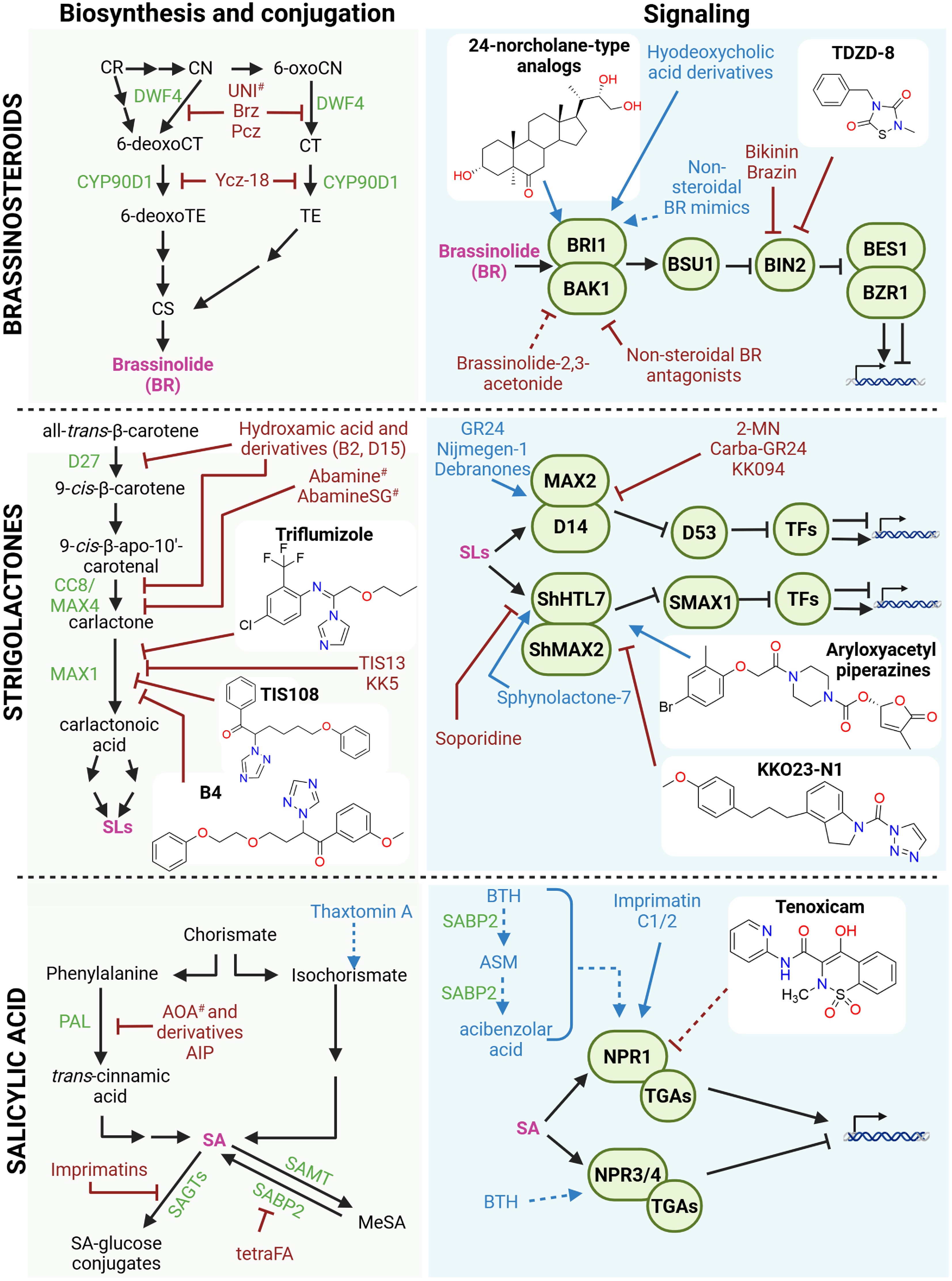
Figure 4.
Small molecules affecting the brassinosteroid (BR), strigolactone (SL), and salicylic acid (SA) pathways. Schematic overview of small molecules that modulate the biosynthesis, conjugation, transport, and signaling of BR, SL, and SA. Known small molecules with agonist or antagonist activity are indicated in blue and red, respectively. Compounds with isoform-specific effects are marked with an asterisk (*), whereas compounds with broad or off-target effects are marked with a hashtag (#). Recently discovered small molecules (past 5 years) are shown together with their chemical structures. Stimulatory interactions are represented by arrow-headed lines, inhibitory interactions are indicated by block-headed lines, and dashed lines indicate unproven or putative interactions. Hormones are shown in pink; precursors and intermediates are shown in black. Biosynthetic and conjugation enzymes, as well as their signaling components, are depicted in green. Abbreviations: 2-MN: 2-methoxy-1-naphthaldehyde; AIP: 2-aminoindan-2-phosphonic acid; AOA: 2-aminooxyacetate; ASM: acibenzolar-S-methyl; BAK1: BRASSINOSTEROID INSENSITIVE 1-ASSOCIATED KINASE 1; BES1: BRI1-EMS-SUPPRESSOR 1; BTH: benzo(1,2,3)thiadiazole-7-carbothioic acid S-methyl ester; BZR1: BRASSINAZOLE RESISTANT 1; BIN2: BRASSINOSTEROID INSENSITIVE 2; Brz: brassinazole; BRI1: BRASSINOSTEROID INSENSITIVE 1; BSU1: BRI1 SUPPRESSOR 1; CCB/MAX4: CAROTENOID CLEAVAGE DIOXYGENASE 8/MORE AXILLARY GROWTH 4; CN: campestanol; CR: campesterol; CS: castasterone; CT: cathasterone; D14: DWARF14; D27: DWARF27; DWF4: DWARF4; MeSA: methyl-salicylic acid; NPR: NONEXPRESSOR OF PATHOGENESIS-RELATED; PAL: PHENYLALANINE AMMONIA-LYASE; Pcz: propiconazole; SABP2: SA-binding protein 2; SAGT: SA glucosyltransferase; SAMT: SA carboxyl methyltransferase; ShHTL7: Striga hermonthica HYPOSENSITIVE TO LIGHT 7; SMAX1: SUPPRESSOR OF MAX1; TE: teasterone; tetraFA: tetra-fluoroacetophenone; TGA: TGACG-BINDING FACTOR. Figure prepared with Biorender. Chemical structures drawn with Chemdraw.
Since the elucidation of BR signaling, major efforts have focused on finding BR mimetics and receptor modulators. Natural BRs are structurally complex, costly to produce, and chemically unstable, motivating the search for synthetic alternatives that can activate BR signaling at a lower cost by activating the BRASSINOSTEROID INSENSITIVE 1 (BRI1) receptor[93]. Early synthetic BR receptor agonists included hyodeoxycholic acid and its derivatives, which partially mimic BR's activity but require relatively high concentrations and show limited specificity compared with natural ligands[97]. A second group of BR agonists are nonsteroidal mimics of BR (Fig. 4)[98]. The latter were developed to reduce synthetic complexity and improve stability, although they are generally less bioactive than natural BRs. Antagonists of BR perception have also been identified in the past and include some nonsteroidal molecules as well as brassinolide-2,3-acetonide[99,100]. Beyond receptor perception, inhibitors targeting downstream signaling also exist. Bikinin, a small molecule identified through chemical genetics, directly inhibits the GSK3-like kinase BRASSINOSTEROID INSENSITIVE2 (BIN2), which is a negative regulator of BR signaling. Bikinin and its analogs induce strong BR responses by competing with adenosine triphosphate (ATP) at the kinase's active site[101].
Recent studies have continued efforts to extend the current set of BR perception and signaling modulators. For example, 24-norcholane-type BRs are rationally designed BR analogs with varying alkyl side chains (Fig. 4)[102]. These confer increased chemical stability compared with natural BRs, facilitating their application. In addition, compared with older tools like hyodeoxycholic acid derivatives, they are more active towards the BRI1 receptor. Nevertheless, they are more expensive to synthesize and are more metabolized in planta[97,102]. Given the chemical complexity of most BRI1 ligands, a recent study set out to modulate BR signaling downstream from perception. Because BIN2 belongs to the broader GSK3 family, several mammalian GSK3 inhibitors, such as TDZD-8, are possible candidates for inhibiting BIN2. Indeed, TDZD-8 is able to activate BR signaling in diverse plants, thereby promoting somatic embryogenesis; this highlights the conserved structural features across kingdoms (Fig. 4)[9]. However, TDZD-8 is two to four times less active then bikinin and bikinin-like molecules and is less specific for BIN2, making it a less interesting tool for exploring BR signaling.
Strigolactones
-
Strigolactones (SLs) are carotenoid-derived hormones with dual roles as endogenous regulators of shoot branching, root architecture, and mycorrhizal symbioses, and as exogenous signals that trigger the germination of parasitic weeds such as Striga and Phelipanche[103]. Their developmental and agricultural importance has driven intensive efforts to develop chemical tools that probe the biosynthesis, perception, and signaling of SLs, as evident from the number of tools available (Fig. 4).
Three types of classical SL biosynthesis inhibitors exist: Amino acid-derived (e.g., abamine and abamineSG) and hydroxamic acid-derived (e.g., D15 and B2) inhibitors of CCDs, and triazole-based inhibitors of MORE AXILLARY GROWTH 1 (MAX1)/CYP711A (e.g., TIS13 and KK5) (Fig. 4)[103−105]. Because CCD inhibitors generally produce pleiotropic effects, inhibitors of later biosynthetic steps are crucial for specifically examining the functionof SL. Nevertheless, the first generation of triazole-based inhibitors had either lower efficacy in planta or were prone to off-target effects on other P450 monooxygenases[105,106]. Through SAR analyses, a more potent and specific inhibitor was developed, namely TIS108[107]. Recently, it was proven, using recombinant enzyme assays, that TIS108 can effectively and directly block the conversion of carlactone to the canonical SL 4-deoxyorobanchol (4DO) in rice (Oryza sativa)[108]. This study showed that TIS108 can lower Striga infestation without affecting growth and highlighted a possible divergence between canonical and noncanonical SLs.
Using TIS108 as a lead, novel derivatives were also designed using a backbone modification strategy and they have been tested in vitro and in vivo[109]. One such compound, B4, induces clear SL-deficient phenotypes in both Arabidopsis and rice, likely related to a reduction in endogenous SL levels (Fig. 4). Another triazole-based compound was recently identified in a screen of N-containing heterocyclic P450 inhibitors[106]. From this screen, triflumizole was identified as an Os900 inhibitor in rice, and it also blocked the carlactone–4DO conversion. Although triflumizole is highly stable and effective, it inhibits other P450 monooxygenases, in contrast to TIS108, which is highly specific for MAX1[106−108].
In terms of SL perception, butenolide-based agonists (e.g., GR24), noncanonical SL agonists (e.g., sphynolactone-7), and receptor antagonists (e.g., KK094) can be distinguished (Fig. 4)[110−113]. Natural SLs are structurally complex and costly to synthesize, motivating the widespread use of simplified analogs such as GR24 that activate the SL receptor DWARF14 (D14)[111,114]. However, these analogs are generally chemically unstable. In addition, GR24 acts on both the canonical D14 receptor as well as on the noncanonical SL receptor HYPOSENSITIVE TO LIGHT 7 (ShHTL7), which is an SL receptor in parasitic weed species[111,114]. For selective activation of these parasitic weed receptors, noncanonical SL agonists like sphynolactone-7 are more suitable[110]. Sphynolactone-7 specifically and potently activates ShHTL7 at femtomolar concentrations. Since it does not induce SL in the host plants, it is considered to be a powerful germination stimulant of Striga hermonthica that can be used for suicidal germination strategies.
More recently, two-dimensional (2D) molecular similarity-based virtual screening using parasitic seed germination stimulants as bait and subsequent structure-guided modification yielded a series of aryloxyacetyl piperazines as potent germination stimulants (Fig. 4)[110,115]. In particular, the addition of a carbamate-D-ring, similar to sphynolactone-7, significantly increased the compound's activity to subnanomolar concentrations. Moreover, while sphynolactone-7 is highly specific for ShHTL7, most active aryloxyacetyl piperazine derivatives likely target other HTLs as well, and are also functional in Phelipanche aegyptiaca. Conversely, some recent compounds preferentially bind and induce SL responses via the canonical SL receptor D14, and have minor effects on ShHTL7. The debranone derivative 2-nitrodebranone was recently designed and showed strong SL responses in Arabidopsis in terms of growth and development[8]. Moreover, it was almost 10 times more active than GR24, unlike older debranones.
In parallel, receptor antagonists have provided complementary approaches to disrupt SL signaling. For canonical D14 receptors, biochemical assays such as the yoshimulactone green (YLG) competition assay have enabled the identification of inhibitors. These include the triazole urea derivative KK094, which blocks the interaction between D14 receptor and the D53 repressor protein, effectively inhibiting the SL signaling pathway and promoting branching in rice[112,116]. For parasitic weed receptors, selective inhibitors of ShHTL7 have also been described, including sporodine derivatives and the recently developed KKO23-N1/N2 compounds (Fig. 4)[117,118]. The latter were designed by combining the structural features of known inhibitory scaffolds with ShHTL modeling, and their specificity was validated by docking, enzyme kinetics, and germination assays.
Salicylic acid
-
Salicylic acid is a central regulator of plant immunity, particularly systemic acquired resistance, but it also influences growth and senescence[119]. Despite its long-recognized role, several aspects of SA's biosynthesis and signaling remain incompletely understood[119]. Because SA strongly enhances defense against pathogens, chemical modulation of SA pathways holds great promise for protecting crops from pathogens and other diseases. In contrast to other hormone pathways, SA's chemical biology has seen relatively few genuinely new small-molecule probes in recent years. Most progress has come from incremental improvements or repurposing of existing compounds rather than the discovery of pathway-specific biosynthesis or perception modulators
The biosynthesis of SA occurs through two routes: the ISOCHORISMATE SYNTHASE (ICS) pathway and the PHENYLALANINE AMMONIA-LYASE (PAL) pathway. Most available biosynthesis inhibitors act on PAL. Transaminase inhibitors such as 2-aminooxyacetic acid (AOA) and α-aminooxy-β-phenylpropionic acid (AOPP) block PAL's activity but are not specific[120]. Further research showed that the phosphonic analog 2-aminoindan-2-phosphonic acid (AIP) provides stronger and more selective competitive inhibition[121]. Conversely, thaxtomin A, a Streptomyces-derived phytotoxin, induces the accumulation of SA upstream from isochorismate in an ENHANCED DISEASE SUSCEPTIBILITY1- (EDS1-) and PHYTOALEXIN DEFICIENT4 (PAD4)-dependent manner[122]. SA homeostasis is also controlled by reversible methylation (MeSA) and glucosylation[119]. A chemical genetic screen identified imprimatins, which inhibit SA glucosyltransferases (SAGTs), increasing active SA and enhancing disease resistance[123]. In contrast, the synthetic SA analog tetrafluoroacetophenone blocked the release of SA from MeSA, which helped to clarify the contribution of MeSA as a mobile SAR signal[124].
The perception of SA is mediated by NONEXPRESSOR OF PATHOGENESIS-RELATED GENES (NPR) receptors that activate transcription with TGACG-BINDING FACTOR factors[119]. Only a few direct modulators of SA perception are known. The widely used analog benzothiadiazole (BTH) induces immune responses in an NPR1-dependent manner and is metabolized to acibenzolar-S-methyl in planta[125]. Although BTH has become an important research and field tool, its exact receptor-binding mechanism remains to be fully resolved. More recently, oxicam-type NSAIDs including tenoxicam and meloxicam were identified as suppressors of SA-mediated immunity[126]. These compounds inhibit defense responses in an NPR1-dependent but NPR3/4-independent manner, suggesting the modulation of transcriptional output rather than biosynthesis or perception. Direct physical interaction with NPR1, however, remains unconfirmed.
In conclusion, compared with other hormone pathways, few selective SA-related chemical probes exist. Therefore, future developments, including ICS-targeting inhibitors or NPR-specific agonists and antagonists, will be key to unlocking the unresolved questions of SA's biology and advancing chemical immune-priming strategies in crops.
Peptide hormones
-
Peptide hormones form the most recently established class of plant hormones, and their recognition as bona fide phytohormones was debated for many years[1]. They are structurally diverse signaling molecules, typically 5 to 100 amino acids in length, that are derived from larger precursor prepropeptides through precise proteolytic processing and post-translational modification. To date, more than 30 peptide hormone families have been identified in plants, including SYSTEMINS, CLAVATA3/EMBRYO SURROUNDING REGION-RELATED peptides (CLEs), and INFLORESCENCE DEFICIENT IN ABSCISSION (IDA)[1]. Characterization of their cognate receptors has greatly expanded our understanding of peptide hormones' function. Functionally, they regulate a wide range of developmental and physiological processes, including meristem maintenance, organ abscission, reproduction, growth, and responses to biotic and abiotic stress, to name a few[1]. Despite their biological significance, chemical tools for modulating peptide hormone pathways remain scarce. This lack largely reflects their structural complexity and relatively recent discovery. Nevertheless, a few pioneering studies have demonstrated that peptide signaling pathways are amenable to pharmacological intervention. Given the diversity of peptide hormones, only selected representative strategies will be discussed below.
Two principal approaches have emerged: Small-molecule modulators targeting peptide receptors or their downstream components, and synthetic peptide agonists or antagonists. A recent example of the first category is NPD12704, identified from a 30,000-compound screen for competitors of CLE9 binding to BARELY ANY MERISTEM1 (BAM1)[127]. Importantly, NPD12704 selectively antagonizes BAM1 without affecting other closely related receptors such as CLAVATA1, illustrating how receptor-specific screening can overcome the redundancy limiting peptide hormone studies. Similarly, Liu et al. conducted a chemical genetic screen using kinase inhibitor libraries to identify the modulators of FERONIA (FER), a RAPID ALKALINIZATION FACTOR (RALF) receptor involved in immunity and growth[128]. Several known kinase inhibitors blocked the phosphorylation of FER in vitro and enhanced disease resistance in vivo by preventing ATP binding. Although most of these inhibitors lack complete specificity, reversine and lavendustin A exhibit preferential FER inhibition and are useful tools for disentangling FER's function. Together, these studies reveal trends similar to those observed for other hormone pathways. Approaches that exploit the conserved features of receptor kinases and repurpose compounds with known activity against related protein families can efficiently yield functional inhibitors, but these often suffer from limited specificity. In contrast, unbiased large-scale screening of diverse synthetic libraries, as well as structure-guided and virtual screening strategies, show greater potential to deliver receptor-selective chemical probes. As information on the structure of peptide hormone receptors continues to accumulate, structure-based ligand design is likely to become an increasingly powerful strategy in this field.
A second strategy for modulating responses to peptide hormones is the use of synthetic peptide analogs, an approach which is well established in medicine with the introduction of therapeutic peptides. In plants, rational modification of peptide sequences, although relatively unexplored, has enabled the creation of certain variants with altered receptor specificity and bioactivity. For example, Pro9-modified CLV3 peptides retain their binding to CLV receptors but function as antagonists in vivo[129]. Similarly, certain endogenous peptide antagonists also exist, such as AntiSYS in tomato, which specifically blocks the systemin receptor and counterbalances systemin-mediated defense responses[130]. Such natural antagonists can provide valuable templates for the rational design of synthetic peptides. Nevertheless, the design and synthesis of such synthetic peptides that bind to specific receptors in vivo remains challenging, primarily because of post-translational modifications and the high specificity of the binding pocket. In addition, custom synthesis is still fairly costly and needs to be improved if this would be a viable strategy.
Both strategies, small-molecule modulators and synthetic peptides, illustrate the potential for examining peptide hormone pathways, but they differ in feasibility and scalability. Small molecules can be rapidly screened and repurposed but often suffer from lower specificity and redundancy issues. Synthetic peptides offer higher specificity and the ability to precisely mimic or block natural signaling, yet are currently limited by the cost, delivery, and structural complexity. Future directions likely involve integrating structure-guided ligand design, virtual screening, and peptidomimetics to balance specificity, stability, and cost. Continued accumulation of the receptors' structural data, along with advanced screening platforms and rational design strategies, will increase the efficiency of generating selective and practical chemical tools for peptide hormone research.
Across hormone classes, recent chemical biology efforts reveal a common trade-off between breadth and precision. Classical biosynthesis and metabolism inhibitors often provide strong and robust phenotypes but are limited by off-target effects and pathway crosstalk. In contrast, newer receptor- and signaling-directed ligands, frequently enabled by structural and computational approaches, offer higher specificity, isoform selectivity, and mechanistic clarity, albeit with narrower applicability and higher development cost. Together, these complementary tool classes form a layered chemical toolbox that enables both global pathway perturbation and fine-scale exploration of hormone signaling networks.
-
In fundamental research, confident target identification is essential for using a compound to probe specific biosynthetic steps or signaling nodes. In applied contexts, full molecular annotation of in vivo targets may be less critical, yet an understanding the mode of action and likely off-targets remains invaluable, particularly to assess risks in crop applications. To that end, several technologies, both old and new, aim to identify in planta targets (Fig. 5).
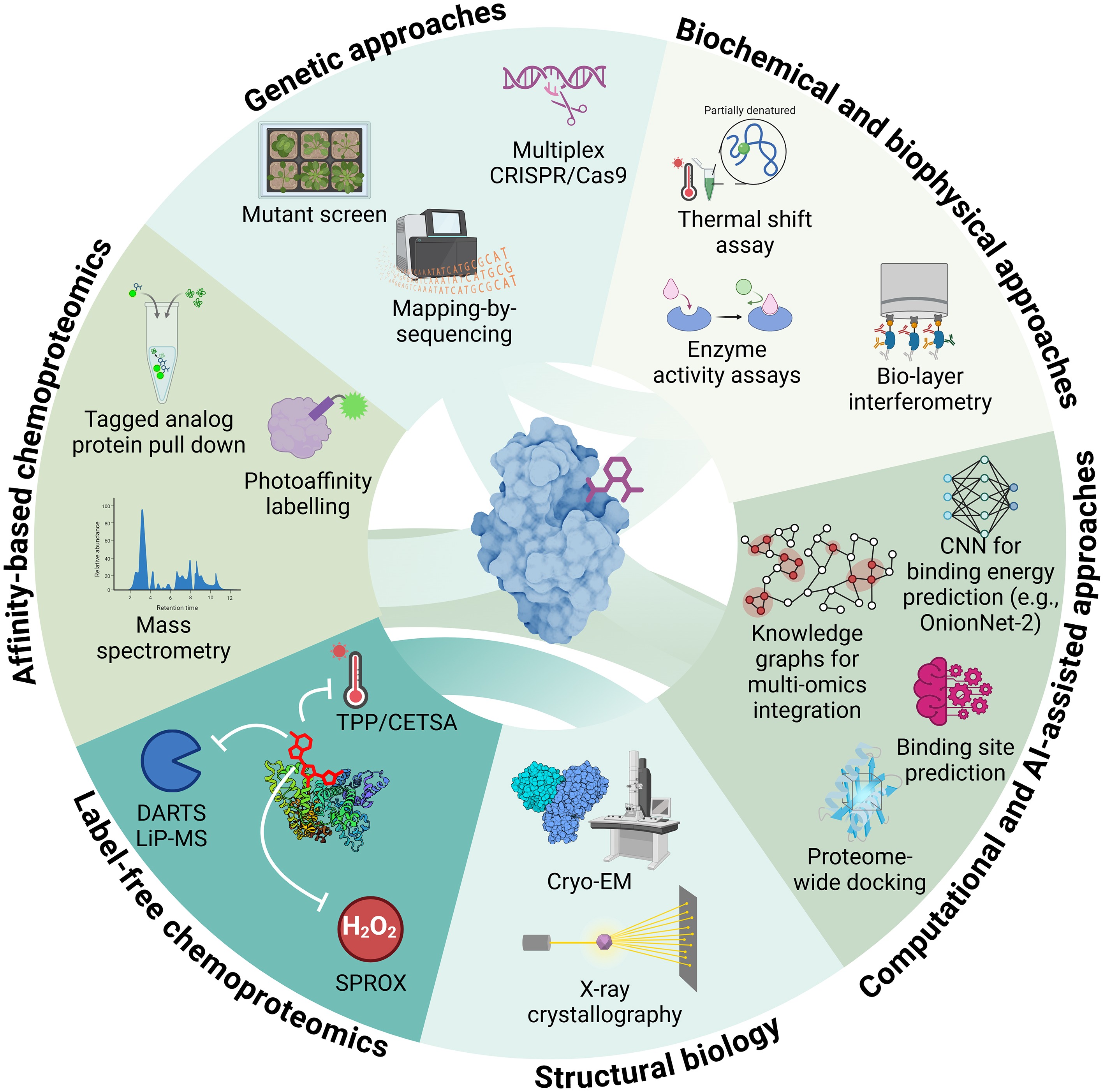
Figure 5.
Integrated strategies for small-molecule target identification. Overview of complementary genetic, biochemical, chemoproteomic, structural, and computational approaches for identifying molecular targets of bioactive compounds in plants. Abbreviations: CETSA, Cellular Thermal Shift Assay; CNN, Convolutional Neural Network; CRISPR/Cas9, Clustered Regularly Interspaced Short Palindromic Repeats/CRISPR-associated protein 9; DARTS, Drug Affinity Responsive Target Stability; LiP-MS, Limited Proteolysis–Mass Spectrometry; SPROX, Stability of Proteins from Rates of Oxidation; TPP, Thermal Proteome Profiling. Figure prepared in Biorender.
Traditionally, target identification in plants have relied on a genetic approach, combining mechanistic studies with mutant analyses or forward genetic screens for resistant mutants, followed by mapping-by-sequencing to pinpoint causal mutations (Fig. 5)[14,86]. Though powerful, these strategies face limitations, including broad compound activity, redundancy in large gene families, and lethality when complete loss of function is required. Multiplex CRISPR/Cas can help overcome redundancy but only when candidate gene families have already been implicated by mechanistic studies[3]. Complementary biochemical approaches are, however, usually required for validation, as they provide the most direct and accessible means to confirm putative protein targets. Enzyme kinetics or biophysical assays such as biolayer interferometry (BLI) and isothermal titration calorimetry (ITC) can establish direct interactions, although detailed molecular information on binding often requires more advanced techniques (Fig. 5)[6,70]. Structural biology tools, including X-ray crystallography, and cryo-EM, provide insights into ligand binding at the atomic level and can structurally explain ligand specificity[131]. Such high-resolution information has guided the rational design of ligands, as exemplified by antabactin, cyanabactin, and COR-Mo[27,91,132]. Nevertheless, like biochemical approaches, these methods depend on successful protein purification, may not fully reflect in vivo protein behavior, and are costly. They are typically used for validation rather than initial discovery.
Chemoproteomic strategies offer an unbiased alternative for target discovery. Affinity-based approaches use tagged analogs (e.g., biotinylated derivatives) or photoaffinity probes to covalently trap binding partners, followed by affinity purification and mass spectrometry (MS)[133]. Click-chemistry is one such approach that can both facilitate the generation of chemical libraries (see ABA section) and the tagging of ligands for purifying the protein targets[134]. This approach has enabled the identification of NIT1, revealing an unexpected auxin biosynthesis pathway as target of heatin, which would have been difficult to uncover with targeted methods[39]. However, chemical tags can impair uptake or binding, complicating in planta applications. Label-free methods avoid this derivatization by exploiting ligand-induced changes in protein stability and sensitivity to proteases or to other environmental changes. Drug affinity responsive target stability (DARTS) detects protection from proteolysis upon ligand binding[25], whereas thermal proteome profiling (TPP) measures ligand-induced protein melting shifts coupled to multiplexed mass spectroscopy[26]. TPP has successfully retrieved targets in vivo, including bikinin-binding proteins, and can also reveal interacting partners within complexes[135]. Despite these strengths, these methods can be constrained by incomplete proteome coverage, low sensitivity for weak or low-abundance targets, and the prerequisite that ligand binding alters the protein's stability.
An alternative route is using in silico tools to validate putative protein targets, using docking experiments and molecular dynamics studies. However, although these tools have been used for years, recent computational advances are now complementing experimental strategies and even offering strategies for unbiased large-scale target identification (Fig. 5). For example, high-quality structural predictions from AlphaFold have enabled proteome-scale docking[136]. In forward screening, this could aid the high-throughput docking of virtual libraries to candidate proteins; however, reverse screening could assist in docking a single ligand across the proteome. Moreover, virtual screening and docking can be assisted by machine learning-based rescoring to improve hit enrichment. For instance, combining docking with OnionNet-SFCT, a deep-learning model, retrieved 4 of 14 known ABA-binding proteins in the top 10 predictions, whereas docking alone retrieved none[137]. Therefore, such AI-augmented pipelines can clearly increase confidence in identifying the true targets and reducing off-target effects. AI methods are also being applied to predict binding sites, refine scoring functions, and accelerate large-scale virtual screening in pharmacological studies[138]. Though not yet applied to plant research, it offers an interesting route for further study.
Another interesting avenue is the use knowledge-graph-based approaches that can integrate multiple sources and layers of information (e.g., chemical, genetic, proteomic, and metabolomic data) and process them into relational networks. By embedding small molecules, proteins, and phenotypes in a shared graph, machine learning can infer new links between hormone modulators and biological processes that would be difficult to derive from any single dataset[139]. Large-scale omics studies and meta-analyses feed these frameworks, enabling prediction of the secondary targets and broader traits that may be influenced by a compound[140]. Such integrative strategies are particularly promising in plant systems, where redundancy and pathway crosstalk often obscure direct mechanisms.
In conclusion, each approach has strengths and drawbacks. Classical genetics provides clear functional connections but struggles with redundancy; chemoproteomics is unbiased but limited by proteome coverage; computational tools offer speed and scale but require experimental validation. The most convincing studies therefore combine multiple, complementary strategies that are used in parallel to converge on the robust identification of small-molecule targets (Fig. 5).
-
Chemical biology is entering a phase characterized by increased precision, scalable screening platforms, and integrative discovery pipelines that are reshaping how plant hormones can be manipulated and studied. These advances hold promise for resolving some long-standing gaps in hormone biology and for translating applications to crop improvement. However, their practical impact will depend on overcoming some constraints, including compound delivery, metabolic stability, species specificity, and even regulatory hurdles, which can influence their adoption. In this section, we highlight key areas where next-generation chemical tools are most likely to provide conceptual and practical advances.
Towards a next generation of chemical tools?
-
A defining trend in plant-focused chemical genomics is the move toward greater selectivity, including isoform- or tissue-, and context-specific regulation. Advances in cryo-EM, crystallography, and protein structure prediction using AlphaFold/Multimer have substantially expanded opportunities for structure-guided ligand design. These approaches have already enabled the development of isoform-specific regulators of CK and auxin[24,46]. They may also help to better understand other long-standing questions in hormone biology, such as the role of the membrane-bound auxin receptor AUXIN BINDING PROTEIN 1 (ABP1) in auxin signaling by isolating ABP1-binding ligands[141]. Nevertheless, the predictive power of structural approaches is constrained by their limitations.
Screening strategies are also diversifying. In that regard, the screening of chemical libraries is likely to shift from single-phenotype bioassays, to high-throughput and integrative approaches. Platforms such as Microphenotron enable, for example, chemical libraries to be screened for root growth regulators[142]. Other systems could also be repurposed to allow chemical screening, but can take advantage of high-throughput multispectral imaging to, for example, monitor ABA-dependent stomatal conductance and isolate ABA inhibitors[143]. When combined with hormone-responsive reporter lines, these systems facilitate the identification of compounds with pathway- or tissue-specific activity, reducing possible off-targets. Nevertheless, to further propel advances in the discovery of small molecules, collaborative efforts such as data sharing may significantly benefit the field. For example, open access to chemical screening data and standardized screening platforms would allow researchers to compare results across different systems and species, revealing conserved and species-specific regulatory mechanisms. Such collaborative efforts are essential for overcoming the current challenges in data integration and interpretation.
Another emerging theme is the engineering of synthetic hormone circuits. Orthogonal receptor–ligand pairs, exemplified by ABA systems such as the CsPYL15m receptor–iSB09 pair[11], could be extended to other protein targets and hormones. They enable the conditional activation of hormone response pathways without interfering with endogenous hormone signals. These systems are elegant and offer temporal precision, but their deployment beyond laboratory settings faces obstacles, including the need for targeted mutagenesis and regulatory acceptance, which currently limit their agricultural scalability.
A more speculative but possibly powerful strategy to selectively target hormone pathways is the application of proteolysis-targeting chimeras (PROTAC)-like approaches[144]. PROTAC-like strategies induce selective degradation of the target proteins by recruiting endogenous E3 ligases. Though they have been successful in mammalian systems, in plants, their adoption remains rare. This is largely because of the difficulties in the optimal design of PROTACs that target specific E3 ligases for selective degradation of the target proteins. In plants, this is challenging, given the lack of structural information on proteins[145]. Apart from this, plants also face practical challenges related to the foliar uptake, bioavailability, and unintended degradation of proteins[146]. Finally, scaling PROTAC applications to crops would require cost-effective synthesis, delivery methods that are compatible with field conditions, and regulatory approval, all of which currently pose major bottlenecks. Nevertheless, PROTAC could offer a powerful way to deplete hormone receptors or signaling regulators with high temporal precision.
Finally, future pipelines for target identification are likely to be hybrid and iterative (Fig. 5). In such a view, AI-assisted reverse docking could generate prioritized target lists, followed by label-free engagement assays (e.g., TPP, CETSA, LiP-MS) in relevant tissues, targeted biophysics for validation, and precise genetic confirmation via base editing to install binding pocket mutations. Adding spatial omics, such as cell-type-resolved proteomics or transcriptomics, would ensure that the candidate targets are present where the compound acts. Moreover, these novel strategies might offer renewed insights into the modes of action of several previously identified small molecules that remain without proven in vivo protein targets but produce phenotypes of interest[147−149].
Opportunities for resolving questions in hormone biology
-
Despite the breadth of available chemical tools, many aspects of hormonal pathways remain underexplored, which represent lost opportunities to gain more insights into the function of molecular players. In auxin biology, the recent discovery of GH3 inhibitors has opened avenues to study auxin metabolism[6,23], but other conjugation and oxidation pathways lack specific pharmacological probes. This imbalance limits the ability to dissect how distinct metabolic branches contribute to spatial and temporal auxin gradients in vivo. In addition, beyond the well-studied IPyA route, compounds like heatin illustrate how chemical tools can reveal overlooked biosynthetic branches[39]. A similar situation exists in CK biology. Although the perception and degradation of CK are supported by well-established chemical tools, the biosynthetic steps remain comparatively underexplored, largely because of the absence of selective inhibitors for key enzymes such as IPTs and CYP735A monooxygenases. This lack of specificity hampers efforts to temporally or tissue-specifically modulate cytokinin production and obscures the relative contributions of de novo biosynthesis versus interconversion pathways.
Increasing evidence suggests that certain hormone precursors themselves can act as signaling molecules[150]. An ETH precursor, ACC, has been reported to trigger physiological responses independent of ethylene production. However, the lack of selective chemical probes has hindered a mechanistic exploration of these precursor-mediated effects. Developing ligands for putative ACC receptors, or inhibitors that selectively block its synthesis or turnover, would allow better separation of precursor signaling from downstream hormone action and would clarify how these parallel signaling layers contribute to stress and developmental responses
The functional redundancy in large receptor families has also hindered genetic studies; receptor-specific ligands and isoform-selective inhibitors can now resolve these redundancies and uncover hidden diversity. Another complexity of hormone biology is the subcellular compartmentalization of precursors, hormones, and their conjugates. Small molecules targeted to specific compartments, using peptide inhibitors with targeting motifs or prodrugs that are released by organelle-specific conditions, could clarify where within the cell these hormones act.
Chemical tools for agriculture: avenues and limitations
-
Finally, in applied contexts, small molecules hold great promise. Currently, plant growth regulators account for approximately 2.5% of the global crop protection market, dominated by GA inhibitors and GAs themselves[151]. Nevertheless, climate change and sustainable intensification demand more versatile tools. Selective modulators could fine-tune stress tolerance, flowering time, or root architecture, which are controlled by hormones, without genetic modification. Orthogonal circuits could enable crops that respond only to the application of synthetic ligands, decoupling yield or defense traits from endogenous trade-offs.
However, their translation faces hurdles, and synthetic molecules raise regulatory concerns over environmental persistence. Despite major progress, chemical tools are not without important constraints. As mentioned previously, specificity remains a bottleneck, which could be resolved using structure-guided rational design of ligands. That would reduce the risk of unintended developmental trade-offs in crops[17]. Second, plant-related delivery and metabolic stability are difficult to predict. The cuticle and cell wall, vascular trafficking, and rapid metabolic detoxification can prevent compounds from reaching their targets or shorten their effective lifetime. Together with the means of application, these remain major obstacles for field performance, where environmental exposure further alters their uptake and persistence. Third, ecological and regulatory concerns limit deployment. Some classical growth regulators are poorly biodegradable or affect nontarget organisms, and new synthetic chemistries require a stringent evaluation of their environmental fate and food safety[152]. Even when effective biologically, translation into registered agricultural products can take a decade or more. Natural product derivatives or microbiome-derived compounds may face fewer barriers. Many microbes produce hormone mimics or antagonists, such as auxin analogs[153]. Natural products like coronatine and myrigalone already serve as powerful chemical tools[82,86]. Exploring microbial metabolites and screening natural product libraries in vivo, in vitro, or in silico offers alternatives to synthetic compounds[154]. Finally, hormone responses are deeply context-dependent, varying by species, genotype, microbiome, and environment. A chemical that performs well in Arabidopsis or under controlled conditions may fail in heterogeneous field settings.
Overcoming these limitations will require hybrid target validation pipelines, combining chemical proteomics with structural modeling and precise genetics, early metabolite profiling to assess in-plant stability, and greater use of natural product-inspired or microbe-derived scaffolds that align with regulatory and sustainability goals. Moreover, cross-species comparisons and data sharing are crucial for accelerating the development of effective, broadly applicable chemical tools. By establishing standardized screening platforms and sharing datasets, researchers can better understand how chemical tools perform across different species, ensuring faster translation from model organisms to crops.
-
The future of chemical tools for plant hormones lies in greater precision, integrative discovery pipelines, and translational applications. By combining synthetic chemistry, structural biology, AI, and high-throughput phenotyping, researchers will gain unprecedented control over hormone pathways. This will clarify long-standing questions in hormone biology while opening sustainable avenues to engineer crop performance in a changing world.
Thomas Depaepe would like to thank Sergi Munné-Bosch and the three anonymous reviewers for their useful insights and comments during the writing and revision of this article. Thomas Depaepe was employed under a postdoctoral fellowship from the European Union's Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement No. 10120950 during the preparation of this manuscript.
-
The author confirms sole responsibility for the following: study conception and design, data collection, analysis and interpretation of results, manuscript preparation, and approved the final version of the manuscript.
-
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
-
The author declares that there is no conflict of interest.
- Supplementary Table S1 Overview of small molecules in plant hormone research.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of Chongqing University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Depaepe T. 2026. Recent advances in the chemical control of plant hormone responses. Plant Hormones 2: e004 doi: 10.48130/ph-0026-0002
Recent advances in the chemical control of plant hormone responses
- Received: 02 December 2025
- Revised: 21 January 2026
- Accepted: 02 February 2026
- Published online: 28 February 2026
Abstract: Chemical modulation of plant hormones has emerged as a powerful complement to genetic approaches. It enables the precise interrogation of complex and often redundant regulatory networks. Beyond their value for basic research, small-molecule tools offer reversible and tunable control of hormone responses, with growing relevance for sustainable agriculture. Although chemical regulators have been used since the earliest studies of plant hormones, recent methodological advances have substantially reshaped the field. Advances over the past 5 years in chemical optimization, target identification strategies, structural biology and artificial intelligence (AI)-assisted approaches have driven a field-wide shift. This shift favors highly specific, tunable ligands with defined protein targets. Along with a broad overview of small-molecule regulators, we integrate these discoveries across hormone classes and synthesize the emerging principles, challenges, and strategies in the chemical manipulation of plant hormones. We discuss how these advances are expanding both the scope and translational potential of these tools to support crops' performance under climate stress.













