-
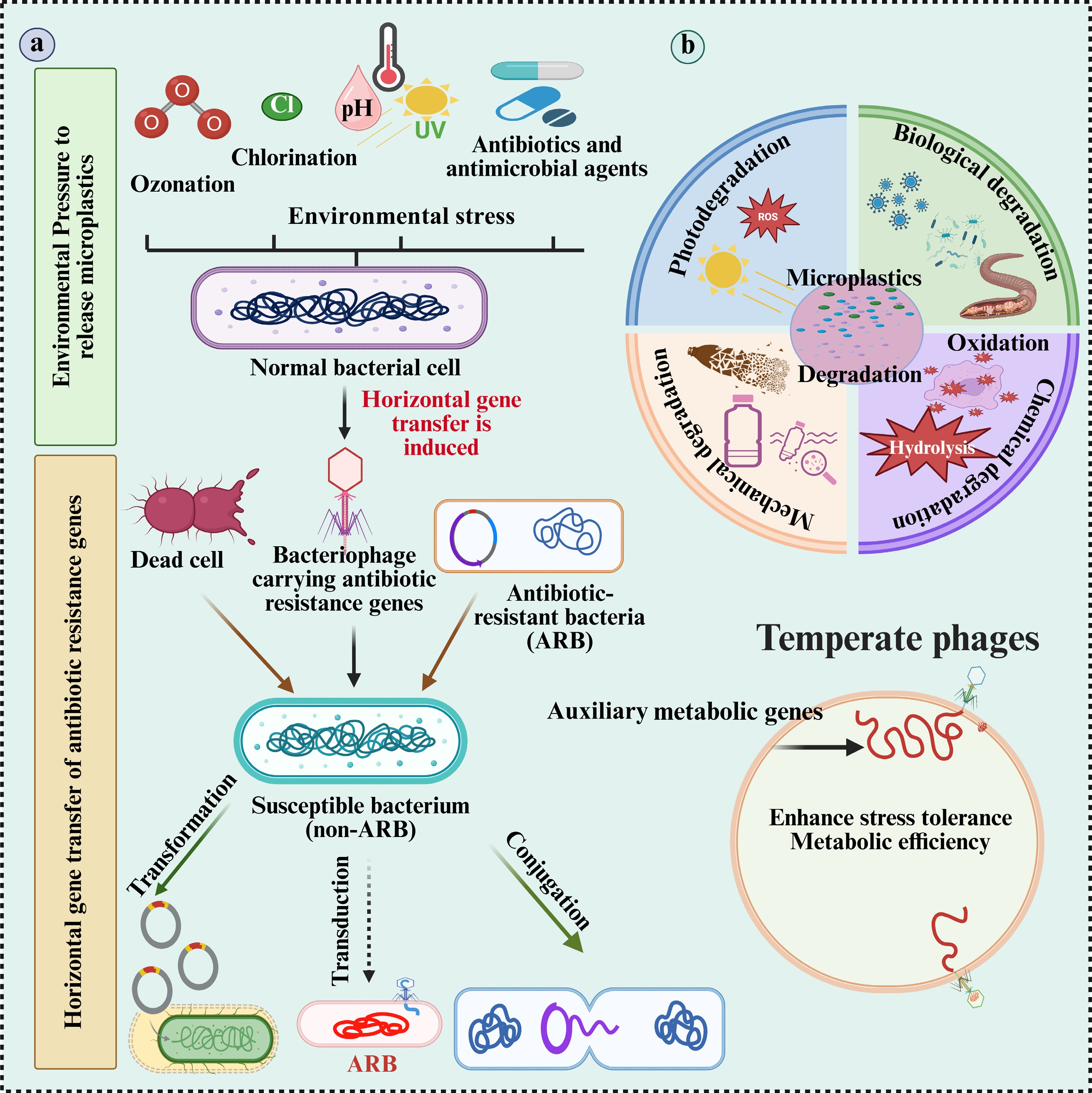
Figure 1.
Schematic representation of the proposed role of microplastics (MPs) in microbial adaptation under environmental stress. (a) Environmental stressors, such as ozonation, chlorination, UV radiation, pH changes, and antibiotics which accelerate the oxidation, hydrolysis, and ionization of MPs. These processes promote the degradation of MPs and the generation of reactive oxygen species (ROS). (b) The same stressors also induce horizontal gene transfer (HGT), particularly transduction, wherein bacteriophages carrying antibiotic resistance genes (ARGs) transfer these genes to susceptible bacteria, converting them into antibiotic-resistant bacteria (ARB). Additionally, dead cells may release ARGs that are acquired via transformation. MP degradation enhances microbial stress tolerance and metabolic efficiency, creating a feedback loop that supports the survival and proliferation of ARB under persistent environmental stress. ARB: antibiotic-resistant bacteria; ARGs: antibiotic resistance genes.
-
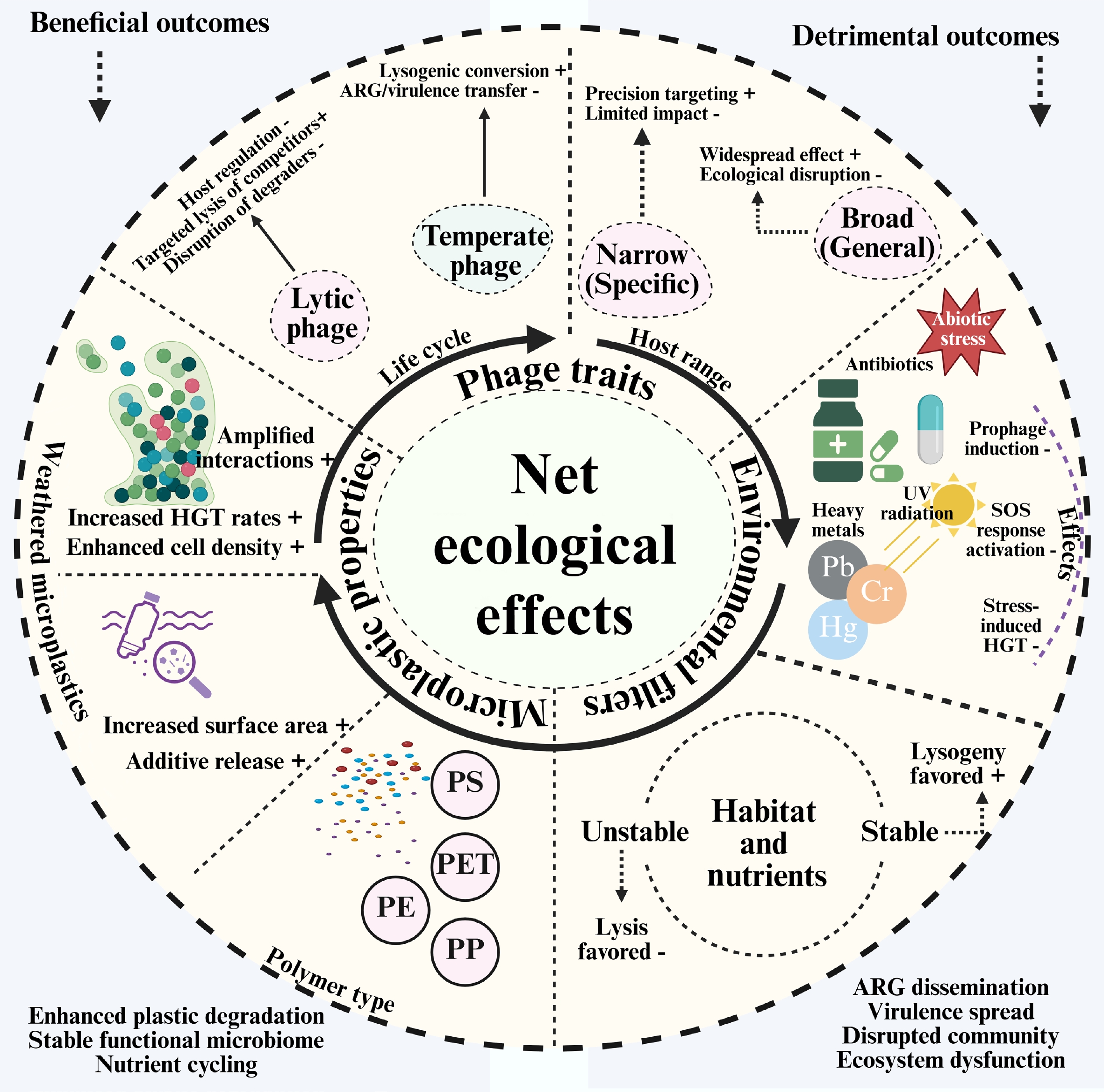
Figure 2.
Conceptual framework determining the net ecological effect of bacteriophages in the plastisphere.
-
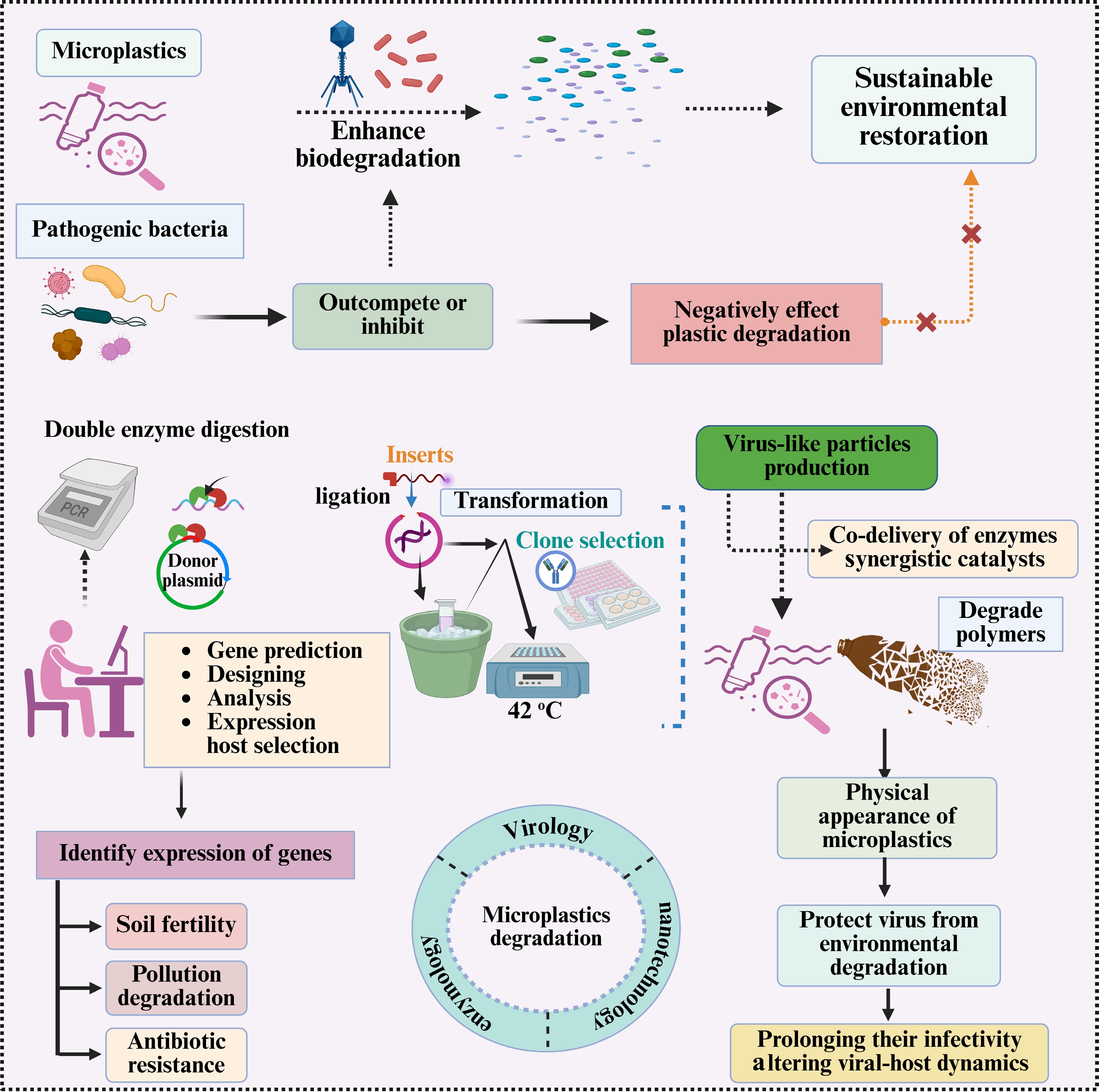
Figure 3.
A multi-step approach combining genetic engineering and viral-like particles to improve microplastic degradation and soil bioremediation. It includes: (1) double enzyme digestion for plasmid preparation, gene prediction, and expression host selection; (2) ligation and transformation for clone selection (42 °C); and (3) virus-like particle production to co-deliver synergistic enzymes while protecting viral infectivity. It also highlights the challenges posed by pathogenic bacteria that may inhibit plastic degradation, as well as strategies to monitor the impacts of gene expression on soil fertility, pollution degradation, and antibiotic resistance.
-
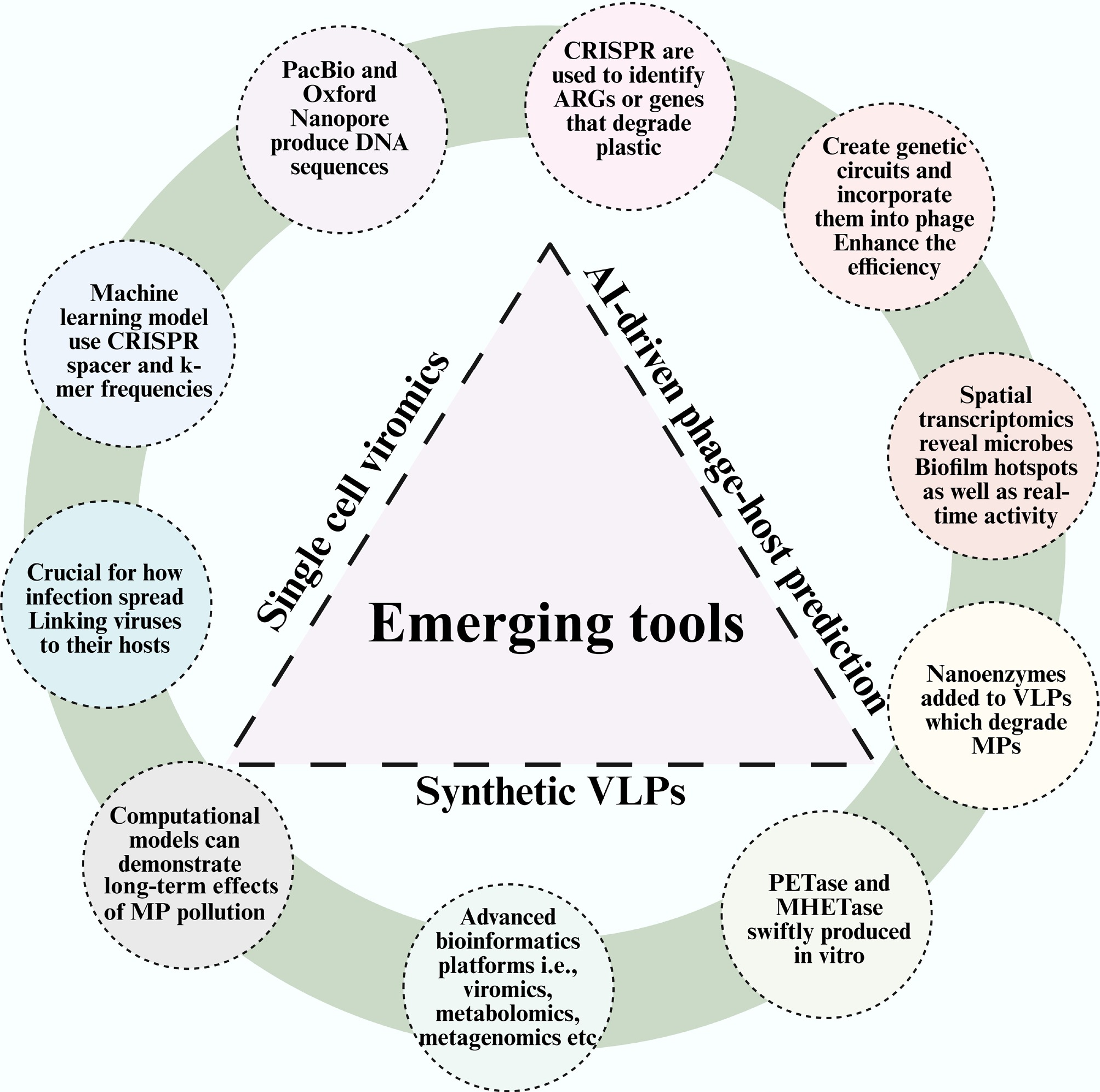
Figure 4.
Schematic representation of various emerging tools that elucidate the interaction of viruses, bacteria, and MPs.
-
Database Primary focus areas Main features Limitations specific to general virology Soil MP research limitations Ref. NCBI RefSeq Viral
(viral genomes; proteins; transcripts)High-quality reference sequences; Taxonomic breadth; Data standardization; Comprehensive metadata Standardized annotation framework; Integration with NCBI resources; Facilitation of viral research; Cross-disciplinary utility Representation bias; Homology-dependent curation; Limited environmental virome data; Genomic diversity challenges Incomplete coverage of environmental viruses; Difficulty in novel viral discovery; Annotation limitation; Host-assignment complexity [70,71] IMG/VR (Integrated Microbial Genomes/Virus)
(Studying viral genomes and their integration with microbial genomes in various environments)Viral genetic diversity; Host-virus interactions; Functional annotation; Ecosystem-level implications Scalability and dataset size; Integrated analysis tools; Ecological context mapping; Host prediction frameworks; Downloadable and visualizable data; Continuous updates ensure that researchers have access to the latest insights Sampling biases; Error-prone predictions (inaccuracies stemming from incomplete genome assemblies); Data resolution gaps (insufficient in cases where a detailed functional role, life cycle, or ecological impacts of specific virus need further exploration) Underrepresentation of soil viral sequences (lag behind other environments like marine and human gut ecosystems); Challenges in linking the virus to microbial hosts (dense and diverse with intertwined tropic network); Depth and assembly gaps (low-abundance and challenging to resolve due to sequencing depth limitation) [72,73] GVD (Giant virus database) Access to genomics, proteomics, and phylogenetic data of Nucleocytoviricota, which are notable for their genome sizes reaching megabases. (Supporting taxonomic classification and evolutionary studies; Enabling exploration of interactions between giant viruses and their host organisms; Investigating the role in terrestrial and aquatic ecosystems) Advanced data integration (functional dynamics of giant viruses); Search capabilities (access tailored datasets); Visualization tools (improve accessibility and interpretation of data); Global coverage (aquatic systems, forest soils, wastewater, and human-associated habitats); Interoperability with other databases Narrow scope (limits studying RNA viruses, bacteriophages, or other small genome DNA viruses); Limited clinical relevance (i.e., studying human pathogenic viruses or animal-associated viromes); Bias in sampling (difficult to detect viruses in environmental samples); Data completeness Lack of direct linkages (do not explicitly focus on giant virus activity in relation to MP particles); Limited environmental metadata (lack of information specific to soil ecosystems impacted by MPs); Neglect of abiotic factors (integration in hydrophobicity, chemical sorption, and mechanical effects of MPs on microbial communities); Host specificity constraints (giant viruses infect protists and algae while MP-associated microbiomes may involve bacteria, archaea, and other small organisms) [74] Table 1.
An overview of various viral databases and their potential use in soil microplastic studies
Figures
(4)
Tables
(1)