-
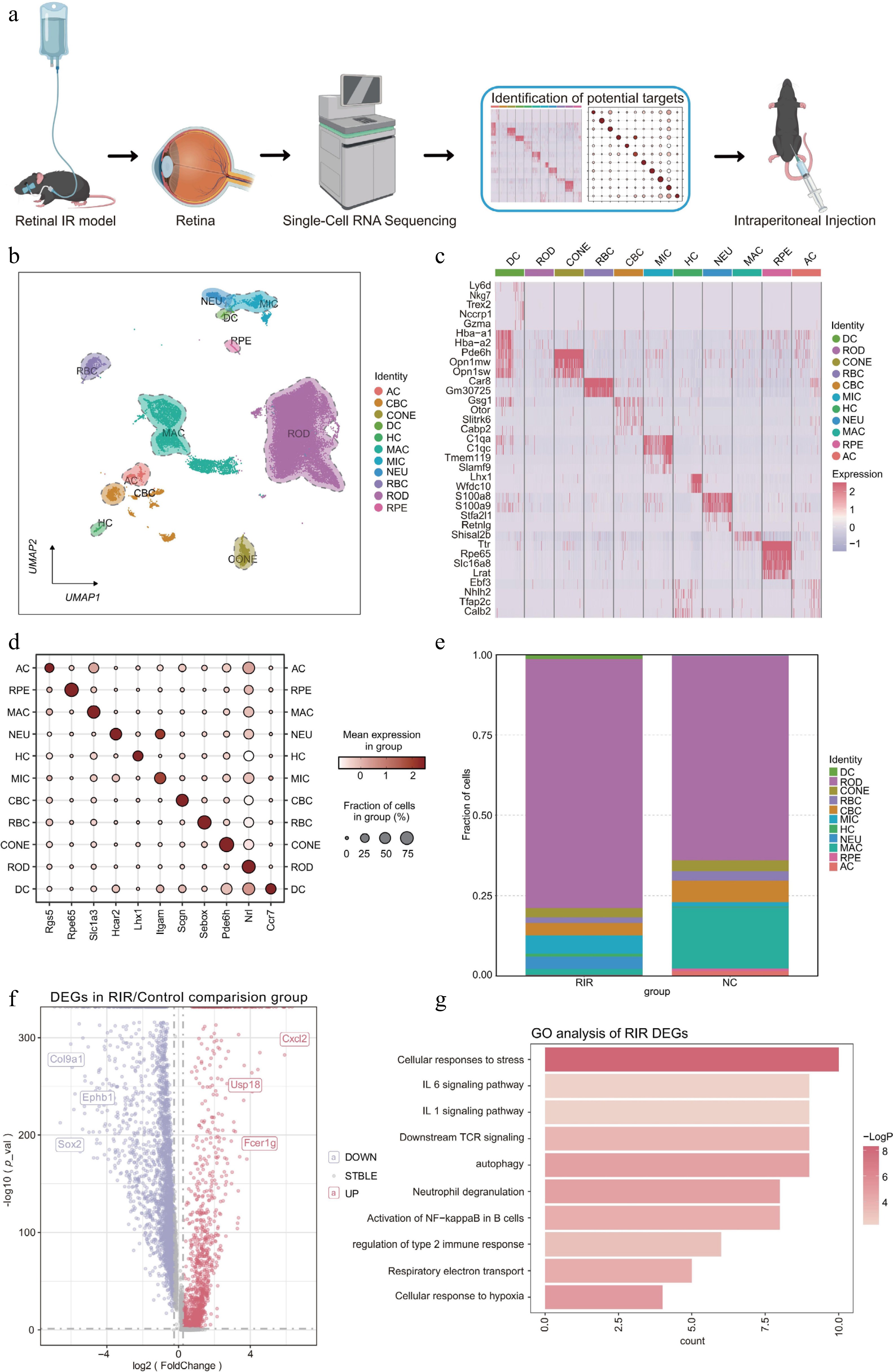
Figure 1.
Changes in the single-cell transcriptomic landscape and cellular composition in the retina following RIR injury. (a) Schematic workflow illustrating the experimental design. (b) UMAP visualization revealing 11 distinct cell clusters in the integrated retinal dataset. (c) Heatmap displaying the expression profiles of the top marker DEGs across identified cell clusters. Rows represent genes, and columns represent individual cells sorted by cluster identity. Color intensity indicates relative gene expression levels (red: high; purple: low). (d) Dot plot showing the expression of canonical marker genes used to annotate each cell type. The size of the dot represents the percentage of cells expressing the marker, and the color intensity indicates the average expression level within that cluster. (e) Stacked bar chart illustrating the proportional shifts in cell populations between the NC and RIR groups. (f) Volcano plot of DEGs in the RIR group vs. NC group. Red and blue dots represent significantly upregulated and downregulated genes, respectively. (g) GO enrichment analysis of upregulated genes in the RIR group. Data for single-cell RNA sequencing were derived from pooled retinas of three mice per group.
-
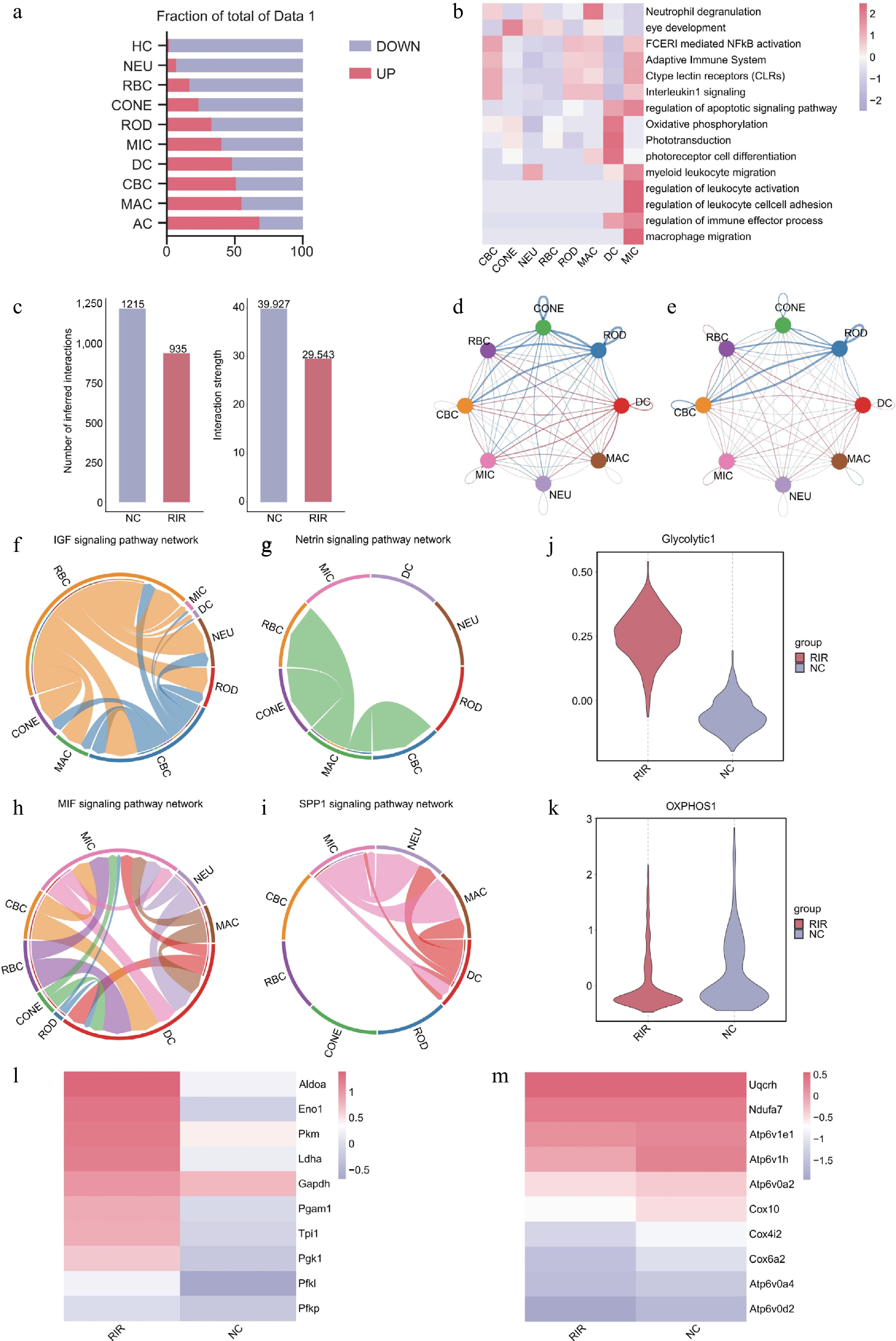
Figure 2.
Integrated analysis reveals inflammatory signaling networks and metabolic reprogramming in RIR injury. (a) Bar graph quantifying upregulated (red) and downregulated (blue) DEGs across retinal cell types. (b) Heatmap of pathway enrichment analysis of significantly upregulated DEGs in the RIR group across eight major cell clusters. (c) Comparison of total cell–cell interactions and interaction strength between NC and RIR groups. Chord diagrams visualizing the differential number of interactions (d) and interaction strength (e). Circle plots of specific signaling pathways. Homeostatic pathways like IGF (f) and netrin (g) characterize the NC group, whereas inflammatory pathways like MIF (h) and SPP1 (i) dominate the RIR landscape. Box plots showing single-cell scores for glycolysis (j) and oxidative phosphorylation (OXPHOS) (k). Heatmaps displaying the expression of key genes involved in glycolysis (l) and OXPHOS (m).
-
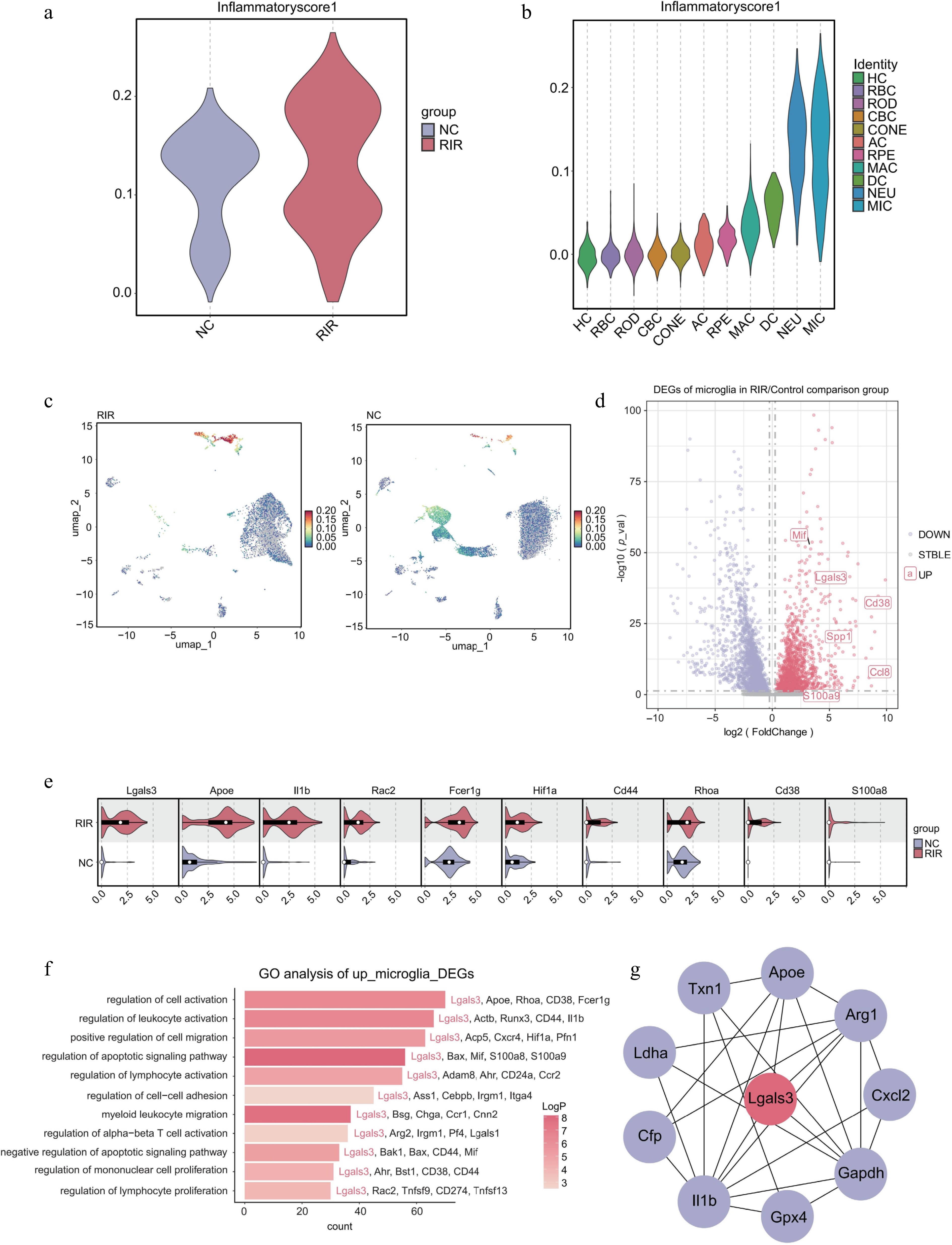
Figure 3.
Microglia drive the neuroinflammatory response via a Galectin-3-centered regulatory network. (a) Violin plot comparing the global inflammatory score between NC and RIR groups. (b) Stratified violin plot displaying inflammatory scores across different retinal cell clusters. (c) Feature plots visualizing the distribution of inflammatory scores on the UMAP. (d) Volcano plot of DEGs in the microglia (RIR vs. NC). (e) Split violin plots validating the upregulation of specific genes in RIR-associated microglia. (f) GO enrichment analysis of upregulated DEGs in RIR microglia. (g) PPI network analysis of microglial DEGs.
-
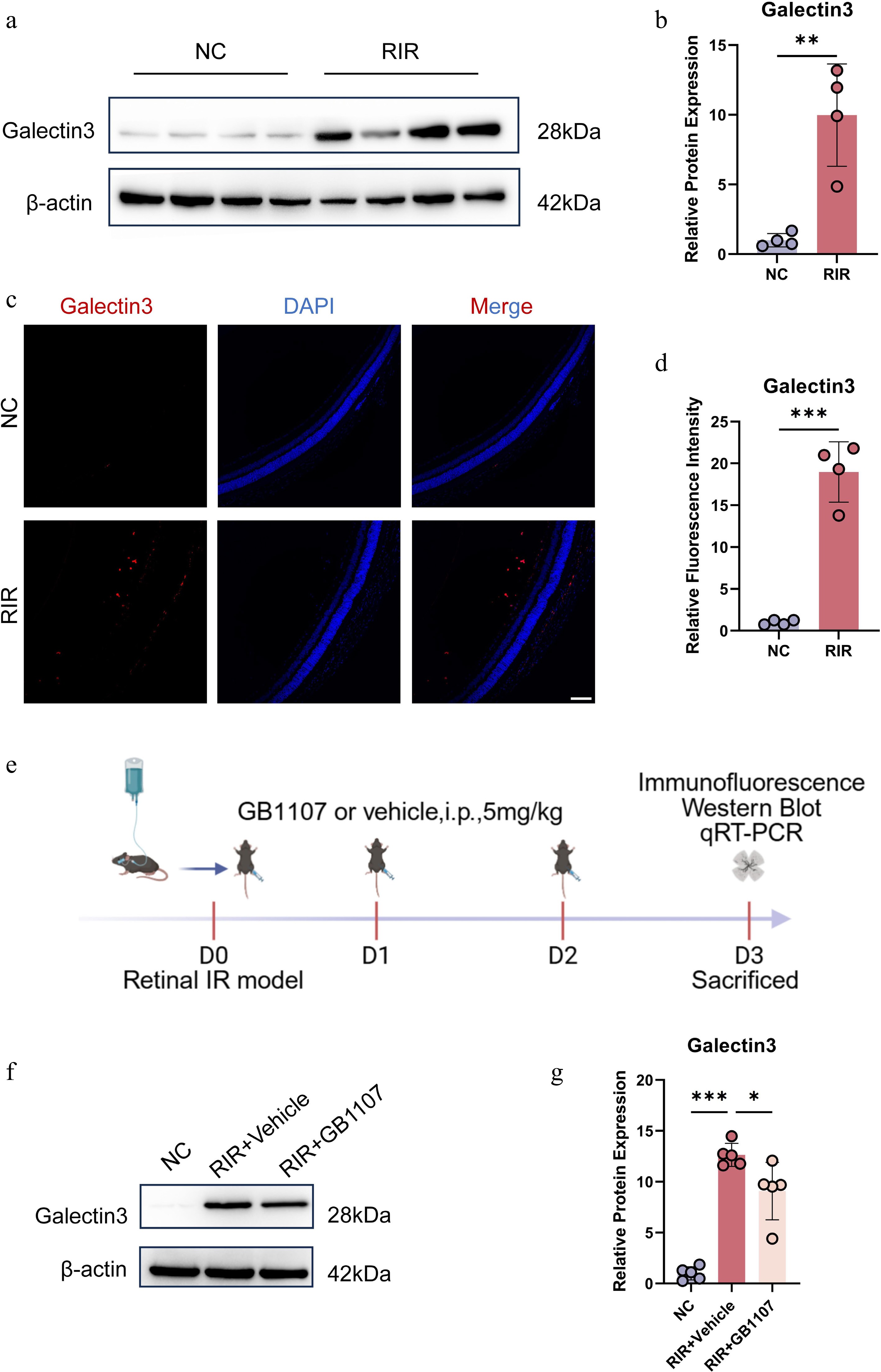
Figure 4.
Pharmacological inhibition of Galectin-3 with GB1107 attenuates its ischemia-induced upregulation. Representative Western blot images (a) and quantitative analysis (b) of Galectin-3 protein expression in retinal tissues. (c) Immunofluorescence image labeling Galectin-3 with red is shown. Cell nucleuses were stained with DAPI (blue). Scale bar: 100 µm. (d) Quantification of Galectin-3 fluorescence intensity. (e) Experimental timeline. Representative Western blots (f) and quantification (g) demonstrating that GB1107 treatment suppresses RIR-induced Galectin-3 protein elevation. Data are mean ± SD (n = 5 per group). One-way ANOVA with Bonferroni's post hoc test was used. *p < 0.05, **p < 0.01, ***p < 0.001.
-
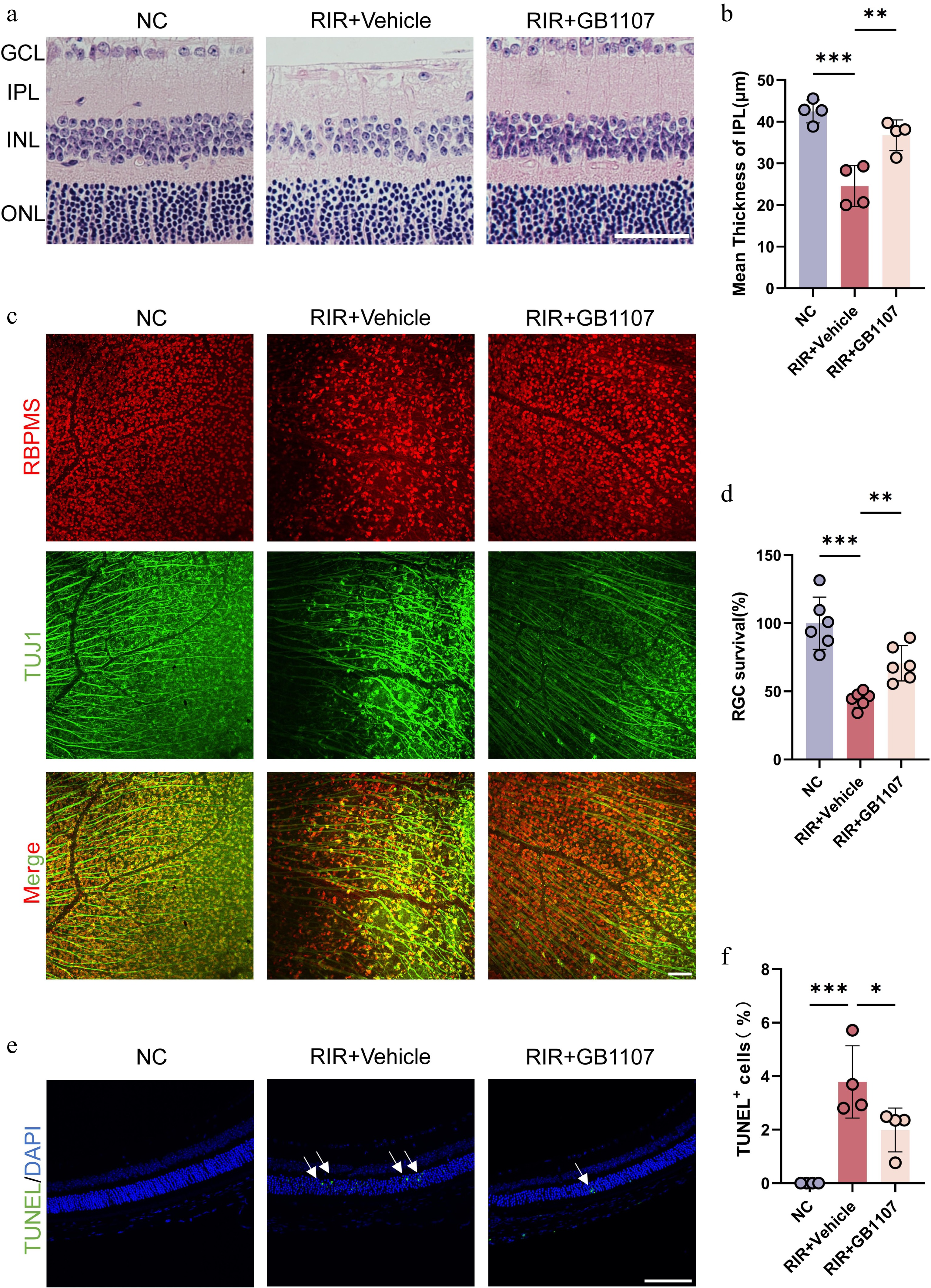
Figure 5.
GB1107 alleviates RIR-induced retinal structural damage and confers neuroprotection. (a) Representative H&E staining of retinal sections. Scale bar: 50 µm. (b) Quantification of IPL thickness across groups (n = 4 per group). (c) Immunofluorescence image labeling RGCs in the retina with RBPMS (red) and TUJ1 (green), and the merged image of two channels is shown above. Scale bar: 100 µm. (d) Quantification of RBPMS-positive RGC density (n = 6 per group). (e) Representative TUNEL staining (green) assessing retinal apoptosis. Nuclei are stained with DAPI (blue). Scale bar: 100 µm. (f) Quantification of TUNEL-positive cells (n = 4 per group). All data are presented as the mean ± SD. One-way ANOVA with Bonferroni's post hoc test was used for statistical analyses. * p < 0.05, ** p < 0.01, *** p < 0.001.
-
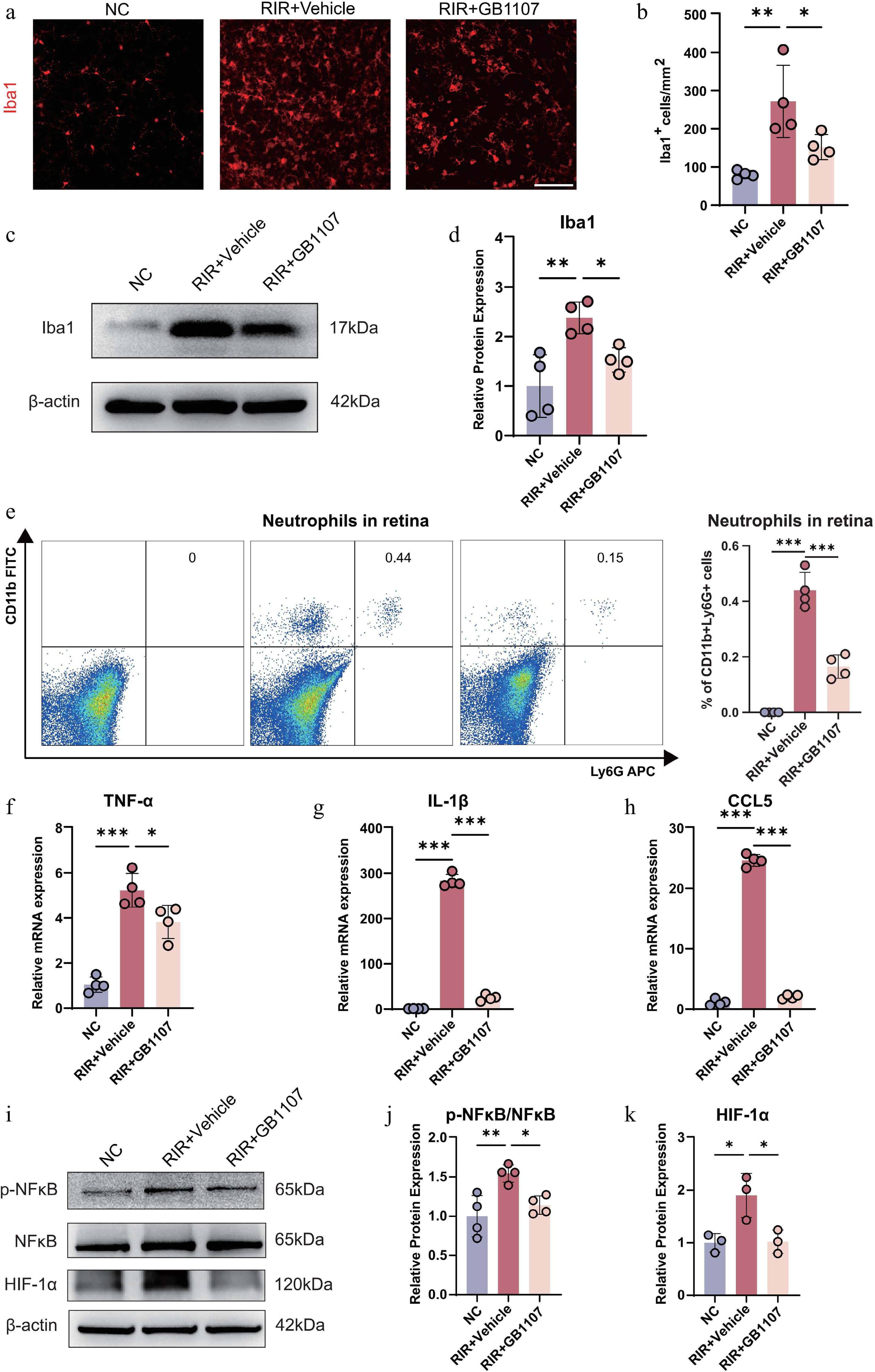
Figure 6.
GB1107 suppresses microglial activation and neuroinflammation associated with the downregulation of NF-κB and HIF-1α signaling. (a) Immunofluorescence showing microglial morphology (Iba1, red). Scale bar: 100 µm. (b) Quantification of Iba1-positive cell density. (n = 4 per group). (c) Western blot analysis and (d) quantification of Iba1 protein levels, confirming reduced microglial activation with GB1107 treatment. (n = 4 per group). (e) Representative flow cytometry plots and quantification of retinal neutrophils (CD11b+Ly6G+). (n = 4 per group). The mRNA expression levels of pro-inflammatory cytokines TNF-α (f) and IL-1β (g), and chemokine CCL5 (h) were measured with qRT-PCR (n = 4 per group). (i) Western blot analysis of p-NF-κB, NF-κB, and HIF-1α, with (j) ratio quantification of p-NF-κB/NF-κB (p65) (n = 4 per group) and (k) protein quantification of HIF-1α (n = 3 per group). All data are presented as the mean ± SD. One-way ANOVA with Bonferroni's post hoc test was used for statistical analyses. * p < 0.05, ** p < 0.01, *** p < 0.001.
Figures
(6)
Tables
(0)