-
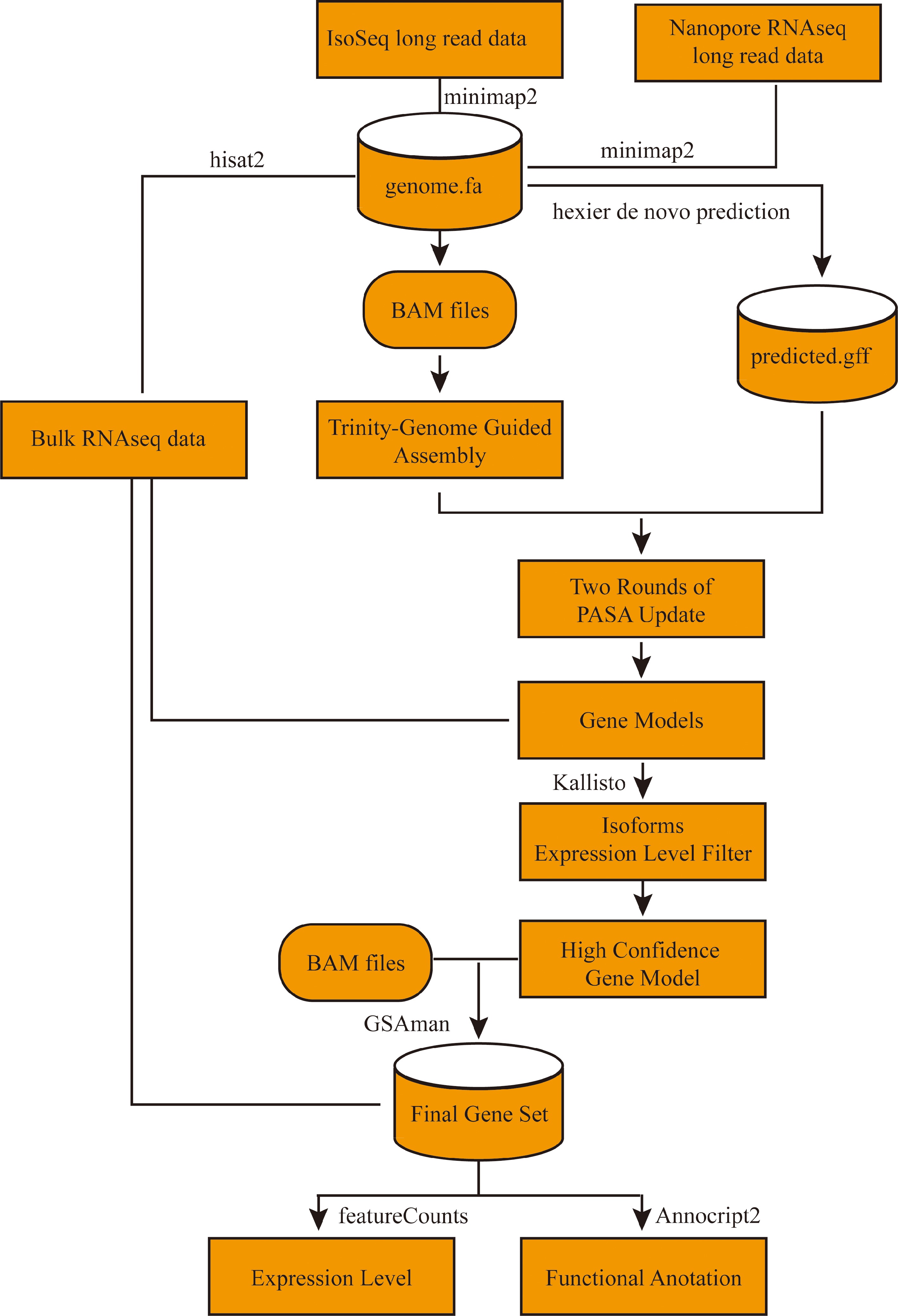
Figure 1.
Workfolw for the update annotation of octoploid strawberry 'Benihoppe'.
-
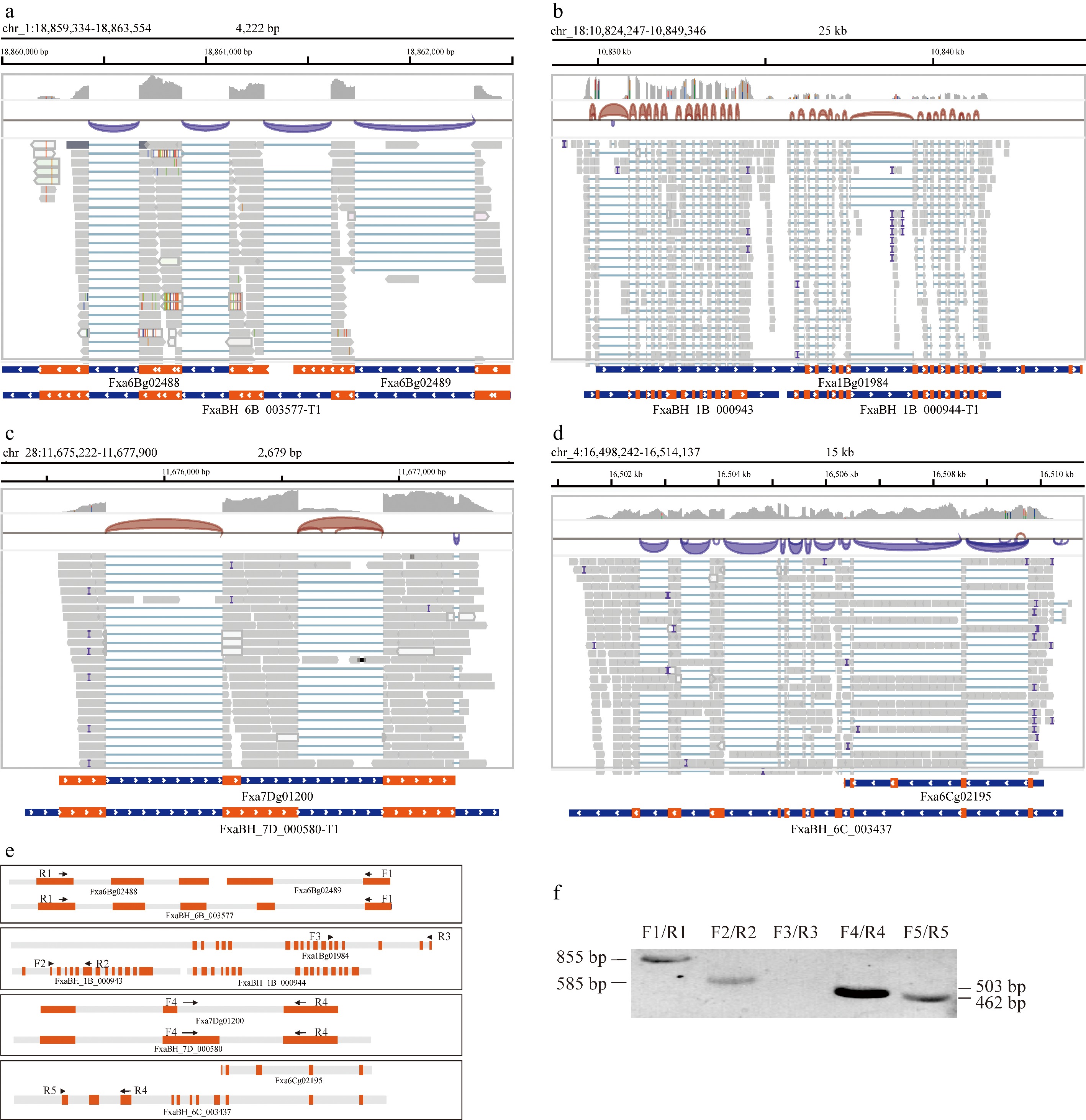

Figure 2.
Examples of modified gene structures. (a) IGV view of genes Fxa6B02488 and Fxa6B02489 merged into one gene, FxaBH_6B_003577. (b) IGV view of the gene structure for Fxa1Bg01984, split into two neighboring independent genes, each with markedly distinct read coverage. (c) IGV view of the modified FxaBH_7D_000580 gene, updated by extension of the second exon of the original Fxa7Dg01200 gene. (d) Correction of defects associated with incomplete 3' ends of the Fxa6Cg02195 gene. (e) Schematic representation of primer binding sites and expected product sizes. (f) Agarose gel electrophoresis of locus-specific PCR products.
-
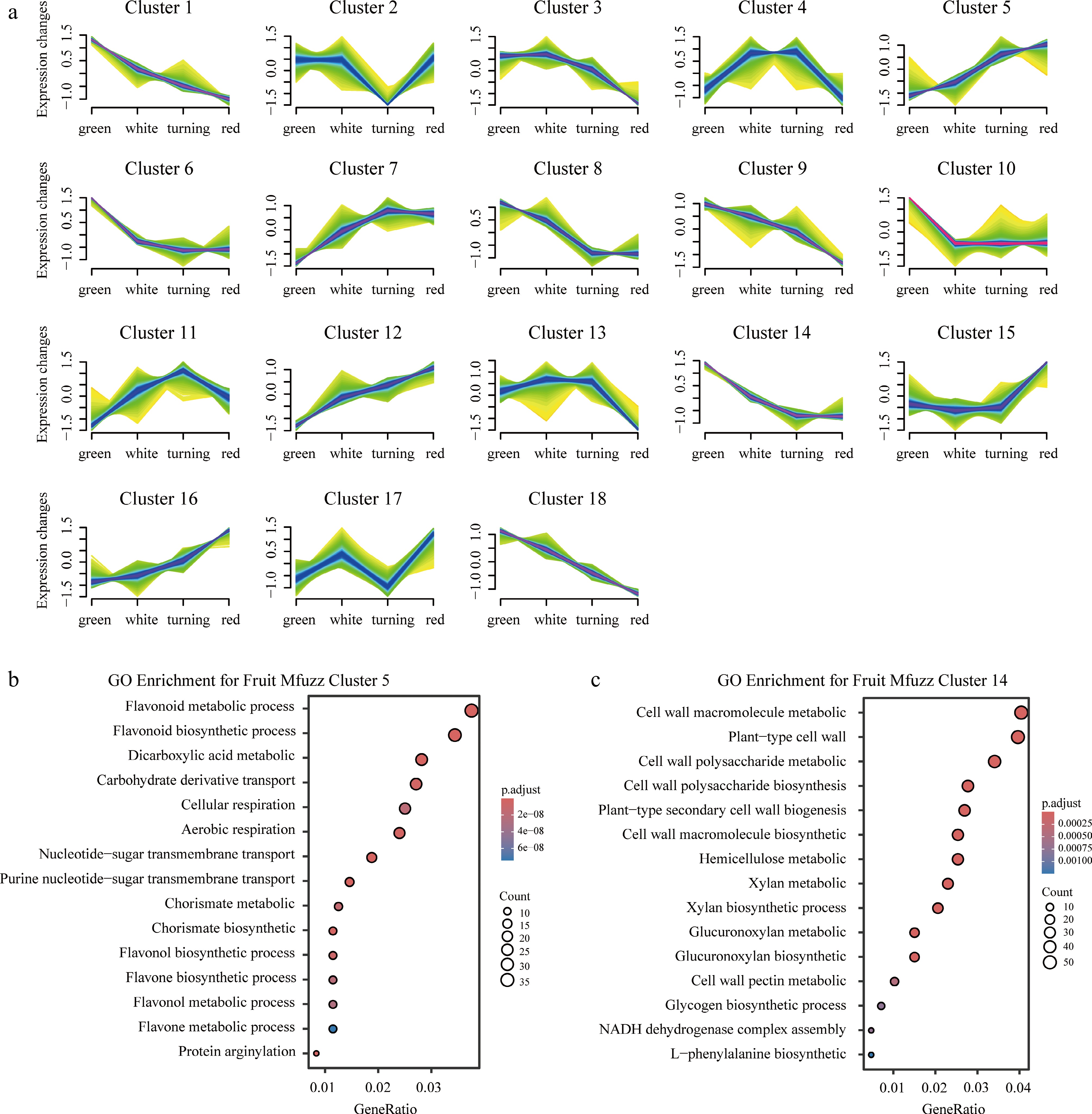
Figure 3.
(a) Mfuzz soft-cluster heat map, and (b), (c) GO/KEGG enrichment bubble plots for clusters 5 and 14 of the receptacle. Color intensity indicates relative expression; bubble size represents gene ratio; color scale denotes adjusted p-value.
-
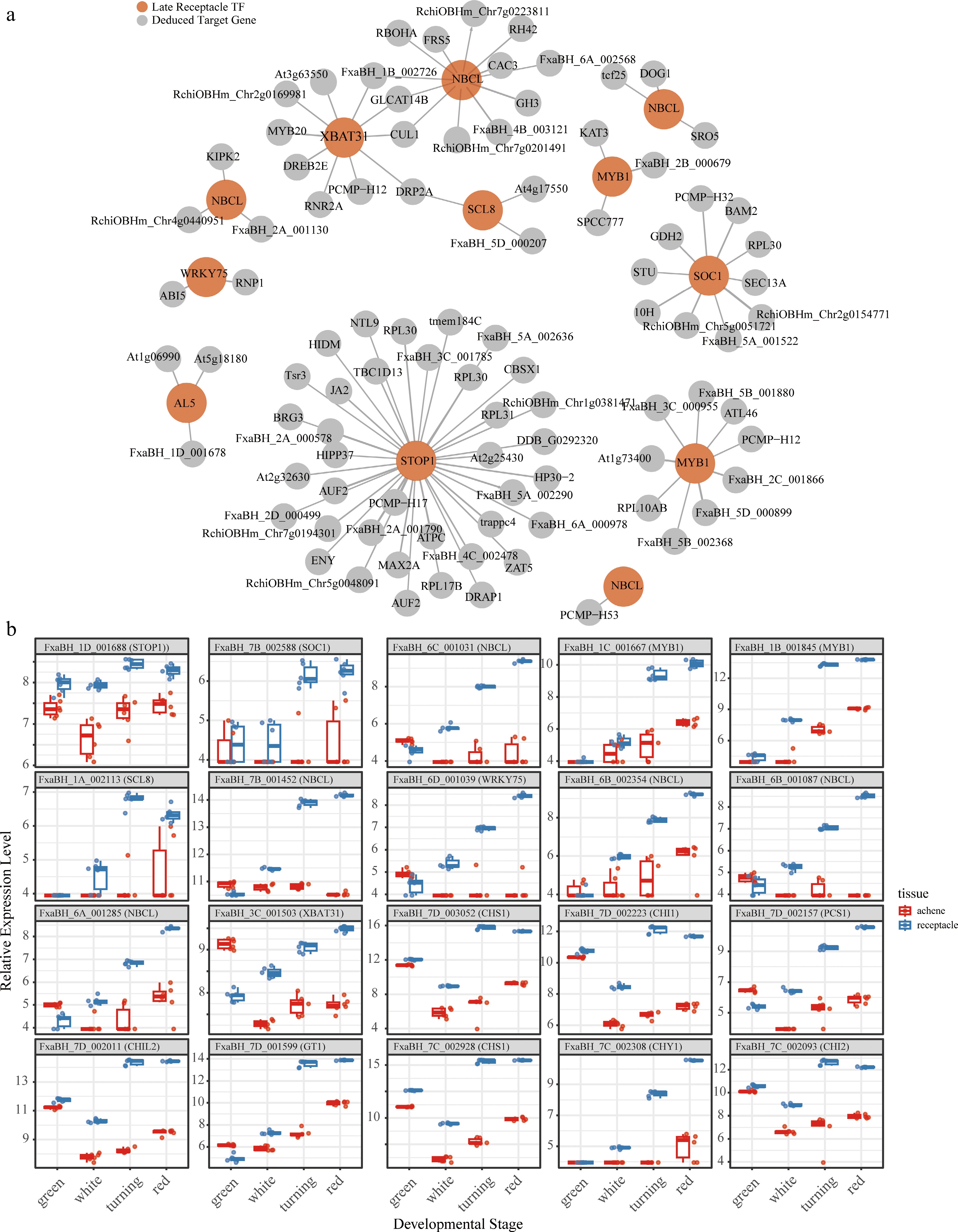
Figure 4.
(a) The top 100 regulator-target interactions unveiled in the preent study, and (b) relative expression patterns of the top 20 in receptacle and achene tissues from different developmental stages.
-
v1.0.a1 v1.0H
(haplotype)v1.0.a2
(this study)Number of genes 109,320 116,232 103,316 Number of mRNAs 109,320 175,313 113,259 Mean length of genomic loci 2,462 2,823 3,211 Mean exon number 4.5 5.5 5.6 Mean CDS length 1,049 1,232 1,169 Single-exon gene number 31,703 29,344 17,048 Multi-exon gene number 77,617 86,888 86,268 Genes with 5'UTR 37,673 45,601 102,967 Genes with 3'UTR 42,605 46,873 101,449 Genes with both 5'UTR and 3'UTR 31,198 40,542 101,404 Mean 5'UTR length 173 371 248 Mean 3'UTR length 350 631 414 Shortest gene length 228 155 104 Longest gene length 151,412 808,995 67,370 Shortest intron length 4 20 20 Longest intron length 50,784 483,249 29,051 Complete BUSCOs (%) 96.2 98.7 99.5 Fragmented BUSCOs (%) 1.1 0.3 0 Missing BUSCOs (%) 2.7 1.0 0.5 Genes with GO terms 54,718 NA 64,266 Genes with KEGG assignment 20,318 NA 43,732 Table 1.
Annotation feature comparison among the initial release, haplotype release and the updated version in this study.
-
GeneID TotalScore Functional description ShortName FxaBH_1D_001688 7.5 Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 STOP1 FxaBH_7B_002588 7.5 MADS-box protein SOC1 SOC1 FxaBH_6C_001031 7.5 BTB/POZ domain and ankyrin repeat-containing protein NBCL NBCL FxaBH_1C_001667 7.5 Transcription factor MYB1 MYB1 FxaBH_1B_001845 7.5 Transcription factor MYB1 MYB1 FxaBH_1A_002113 7.5 Transcription factor MYB1 MYB1 FxaBH_7B_001452 6.5 Scarecrow-like protein 8 SCL8 FxaBH_6D_001039 6.5 BTB/POZ domain and ankyrin repeat-containing protein NBCL NBCL FxaBH_6B_002354 6.5 Probable WRKY transcription factor 75 WRKY75 FxaBH_6B_001087 6.5 BTB/POZ domain and ankyrin repeat-containing protein NBCL NBCL FxaBH_6A_001285 6.5 BTB/POZ domain and ankyrin repeat-containing protein NBCL NBCL FxaBH_3C_001503 6.5 Putative E3 ubiquitin-protein ligase XBAT31 XBAT31 FxaBH_7D_003052 4.5 Chalcone synthase CHS1 FxaBH_7D_002223 4.5 Chalcone--flavanone isomerase 1 CHI1 FxaBH_7D_002157 4.5 Aspartic proteinase PCS1 PCS1 FxaBH_7D_002011 4.5 Chalcone isomerase-like protein 2 CHIL2 FxaBH_7D_001599 4.5 Anthocyanidin 3-O-glucosyltransferase 1 GT1 FxaBH_7C_002928 4.5 Chalcone synthase CHS1 FxaBH_7C_002308 4.5 3-hydroxyisobutyryl-CoA hydrolase 1 CHY1 FxaBH_7C_002093 4.5 Chalcone--flavanone isomerase 2 CHI2 FxaBH_7C_001900 4.5 Chalcone isomerase-like protein 2 CHIL2 FxaBH_7C_001470 4.5 Anthocyanidin 3-O-glucosyltransferase 1 GT1 FxaBH_7B_003068 4.5 Chalcone synthase CHS1 FxaBH_7B_002249 4.5 Chalcone -flavanone isomerase 2 CHI2 FxaBH_7B_002068 4.5 Chalcone isomerase-like protein 2 CHIL2 FxaBH_7B_000048 4.5 Polygalacturonase FxaBH_7A_003557 4.5 Chalcone synthase CHS1 FxaBH_7A_003556 4.5 Chalcone synthase CHS1 FxaBH_7A_003555 4.5 Chalcone synthase CHS1 FxaBH_7A_002558 4.5 Chalcone--flavanone isomerase 1 CHI1 Table 2.
TOP candidate genes in regulating strawberry fruit maturation.
Figures
(4)
Tables
(2)