-
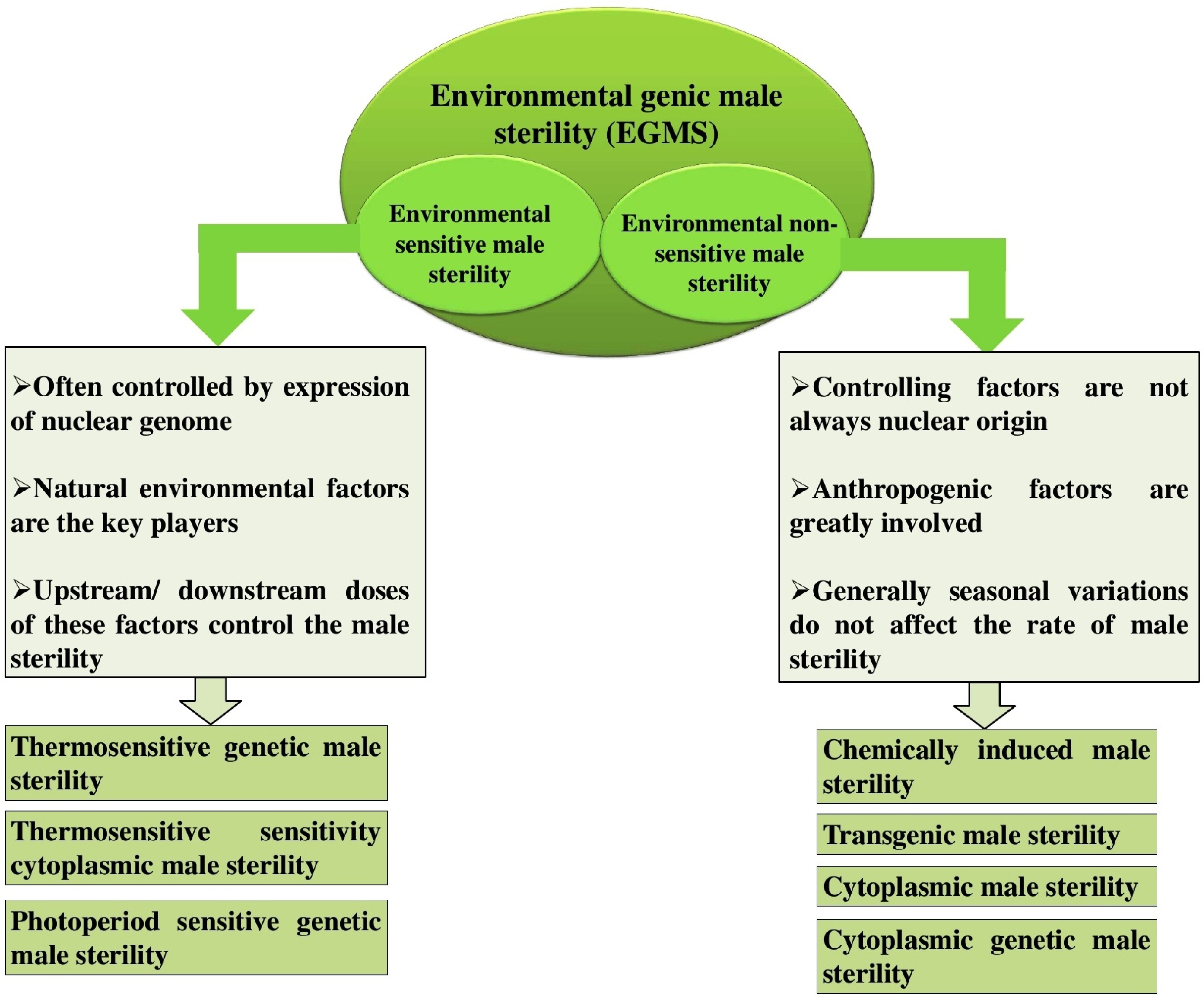
Figure 1.
Types of environmental genic male sterility (EGMS). Environmental sensitive male sterility (ESMS): Male sterility triggered by specific environmental conditions. Environmental non-sensitive male sterility (ENSMS): Male sterility that remains unaffected by environmental variations.
-
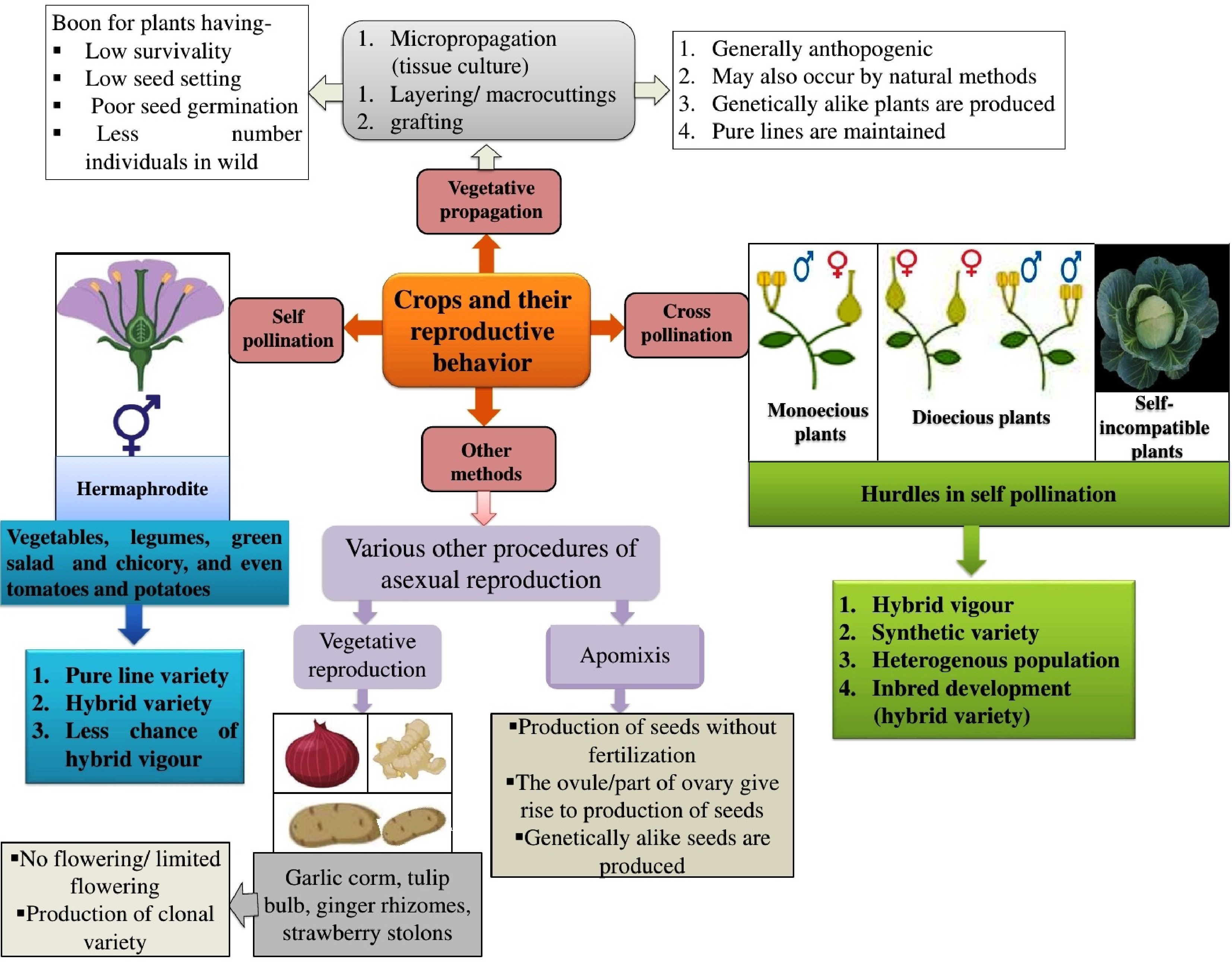
Figure 2.
Different reproductive strategies in plants showcasing the varied reproductive behaviours observed in vegetable crops.
-
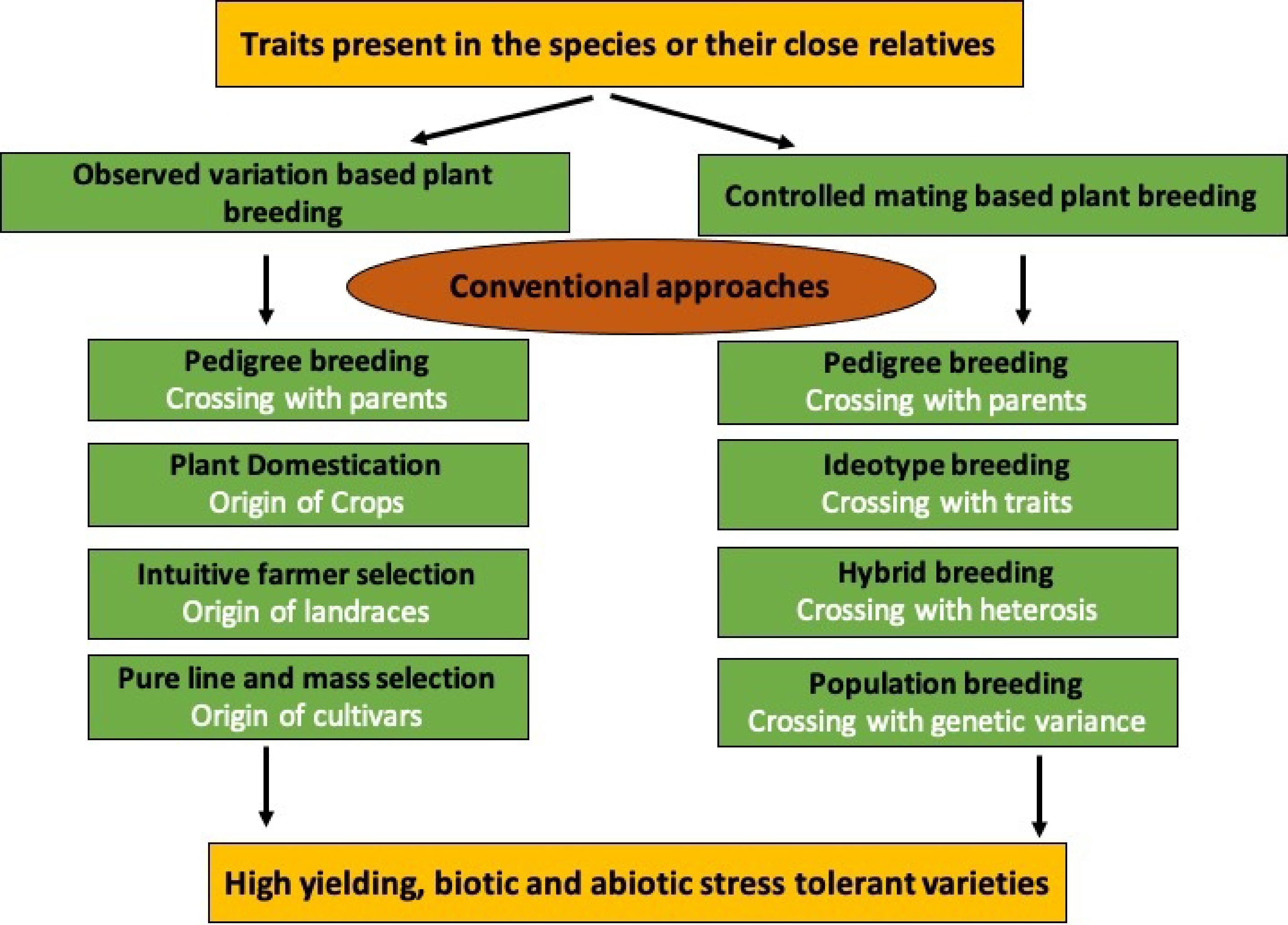
Figure 3.
Graphical overview of conventional breeding methods illustrating the traditional approaches used in vegetable crops.
-
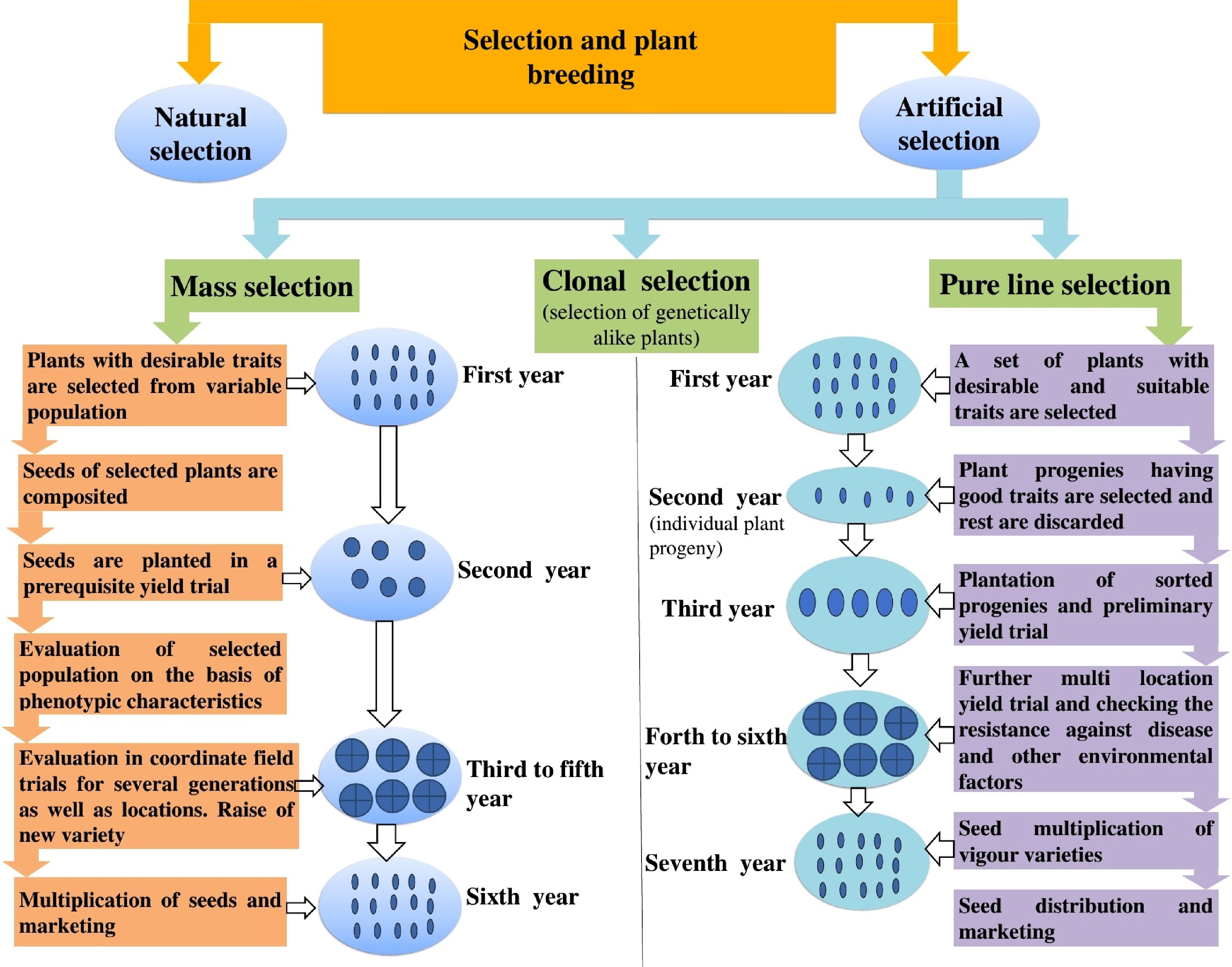
Figure 4.
Role of various selection strategies in plant breeding. This figure demonstrates how different selection methods contribute to the improvement of vegetable crop plants.
-
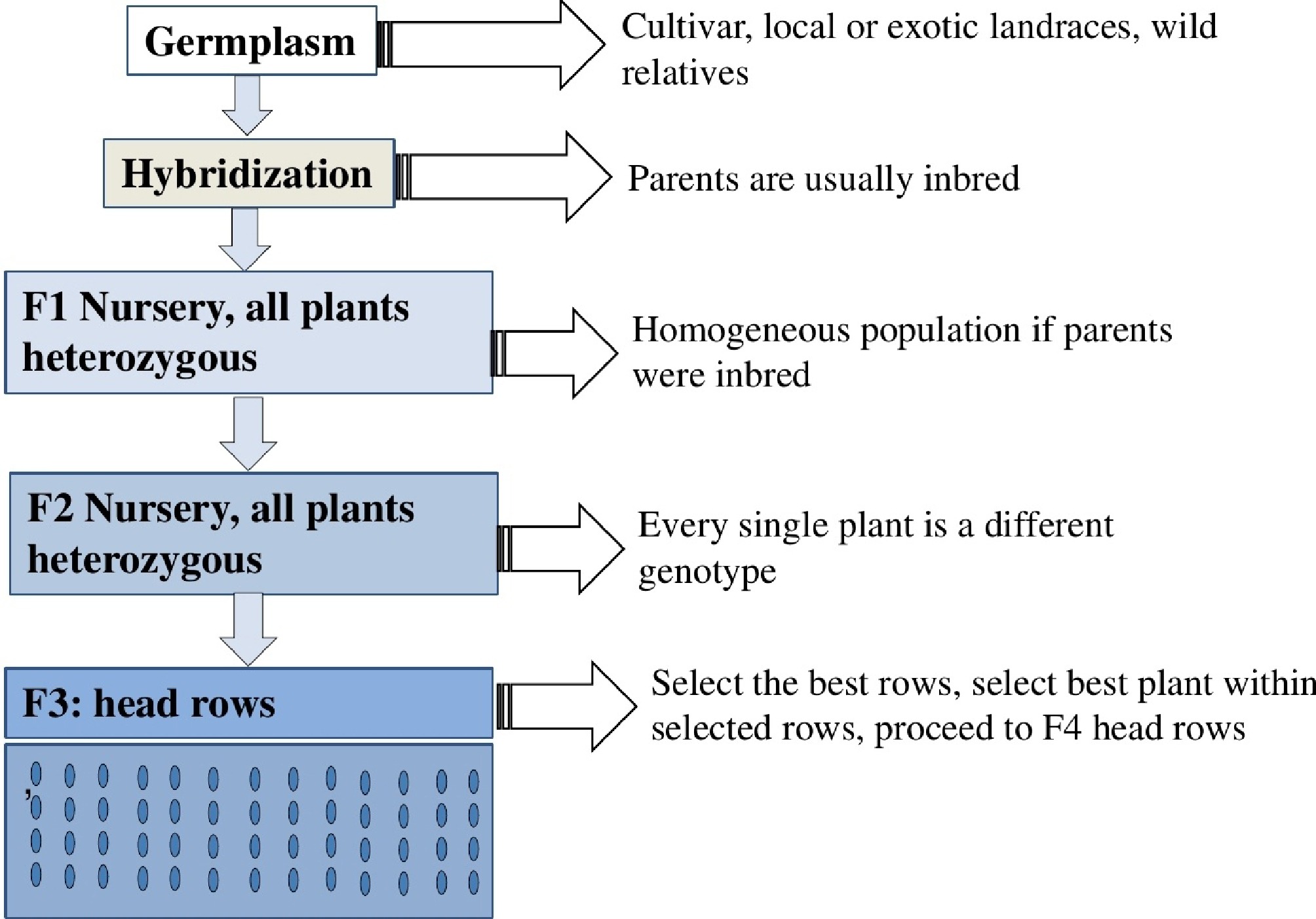
Figure 5.
Graphical presentation of cultivar development in self-pollinated species using the pedigree method, illustrating the typical steps involved in developing new cultivars for self-pollinated species.
-
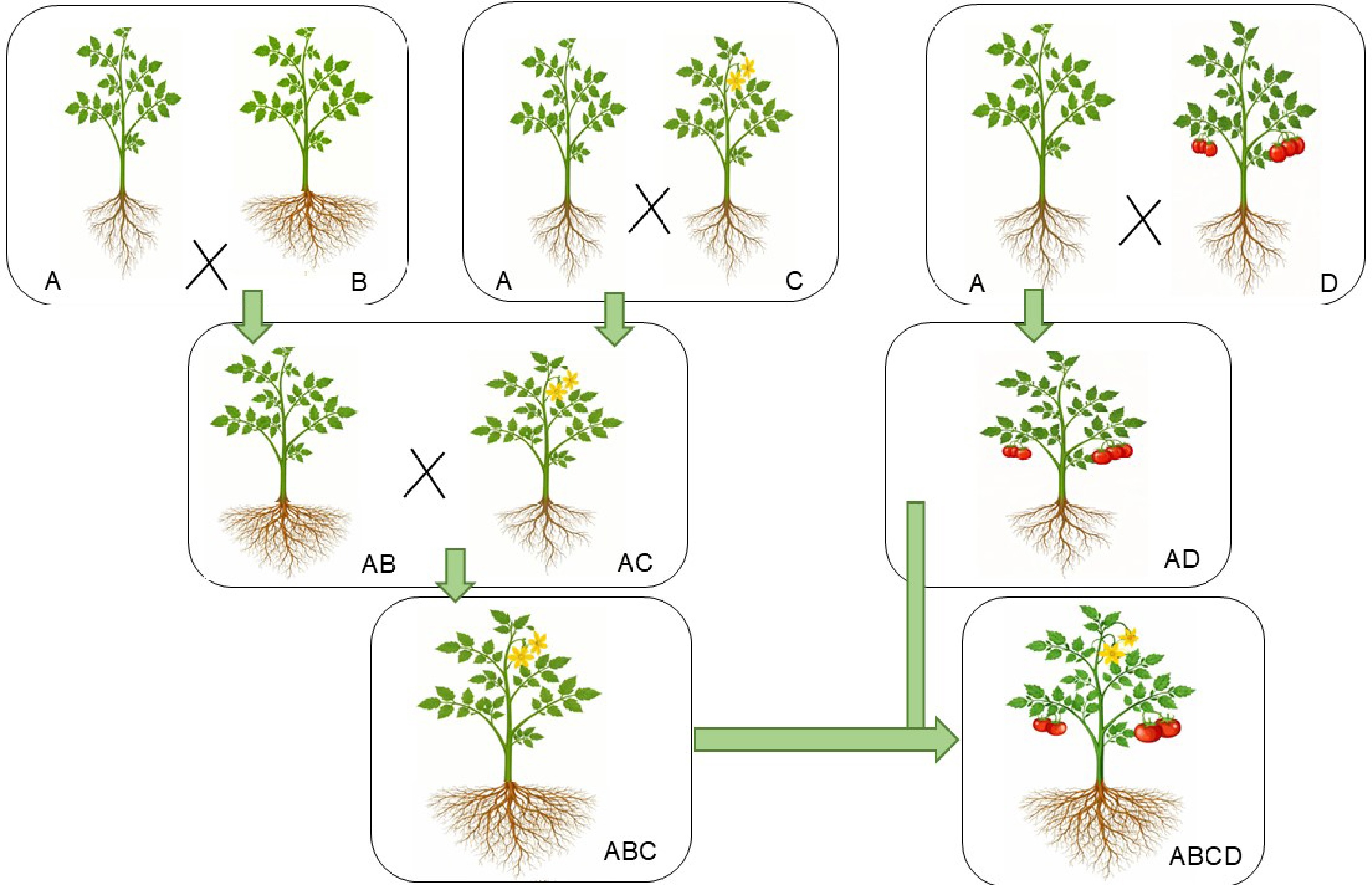
Figure 6.
Gene pyramiding for vegetable crop enhancement and high-yield varieties. This figure highlights the role of gene pyramiding in developing vegetable crops with multiple desirable traits. The key traits include: Trait A, plants with simple shoot and root systems; Trait B, plants with high-yielding roots and simple shoots; Trait C, plants with potent flowering alongside simple root and shoot systems; Trait D, plants with a high rate of fruit setting and simple shoot and root systems, respectively.
-
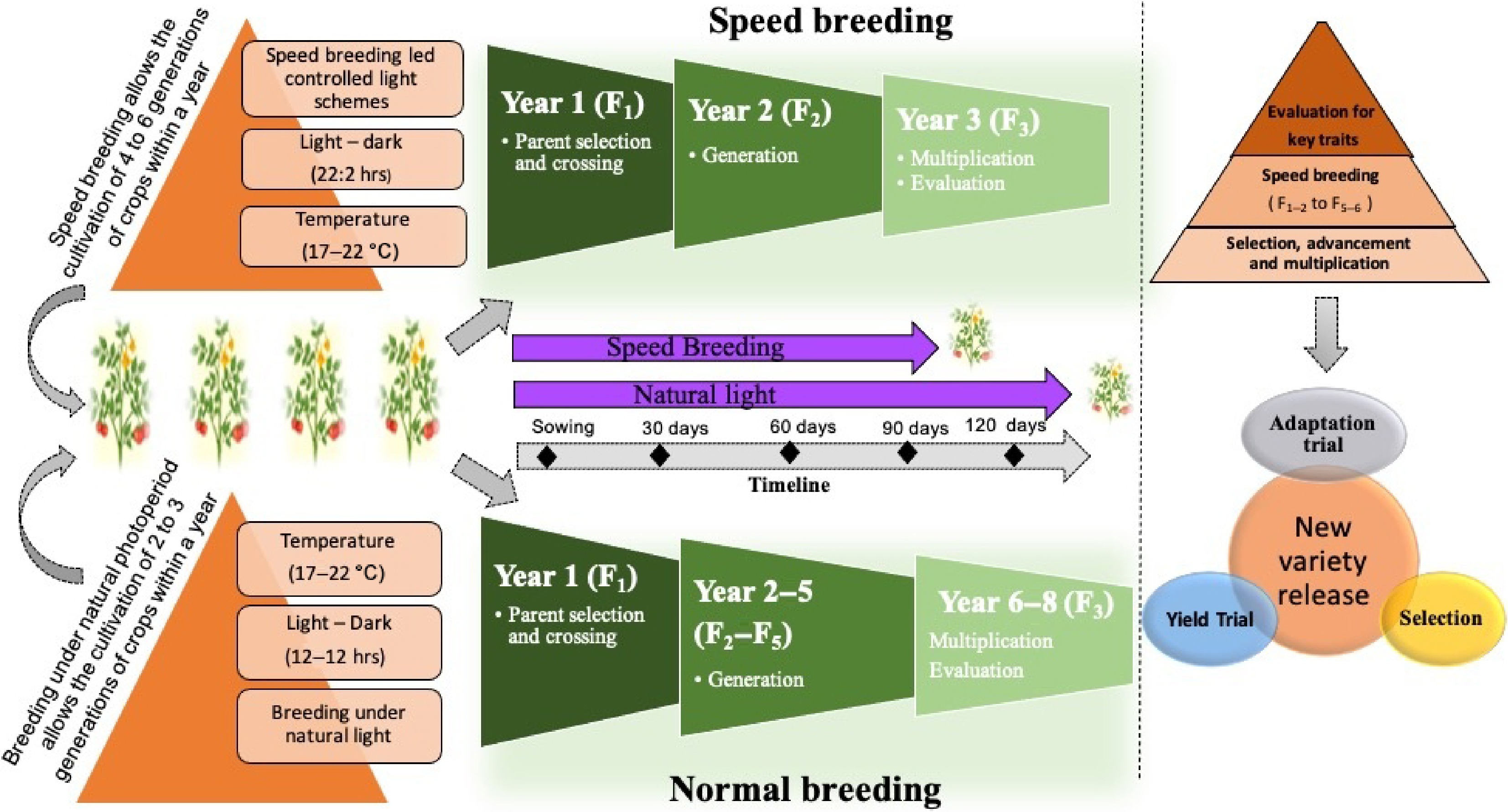
Figure 7.
Impact of photoperiod on standard and speed breeding methods in plant breeding, depicting the differences in photoperiod effects between conventional breeding and speed breeding methods.
-
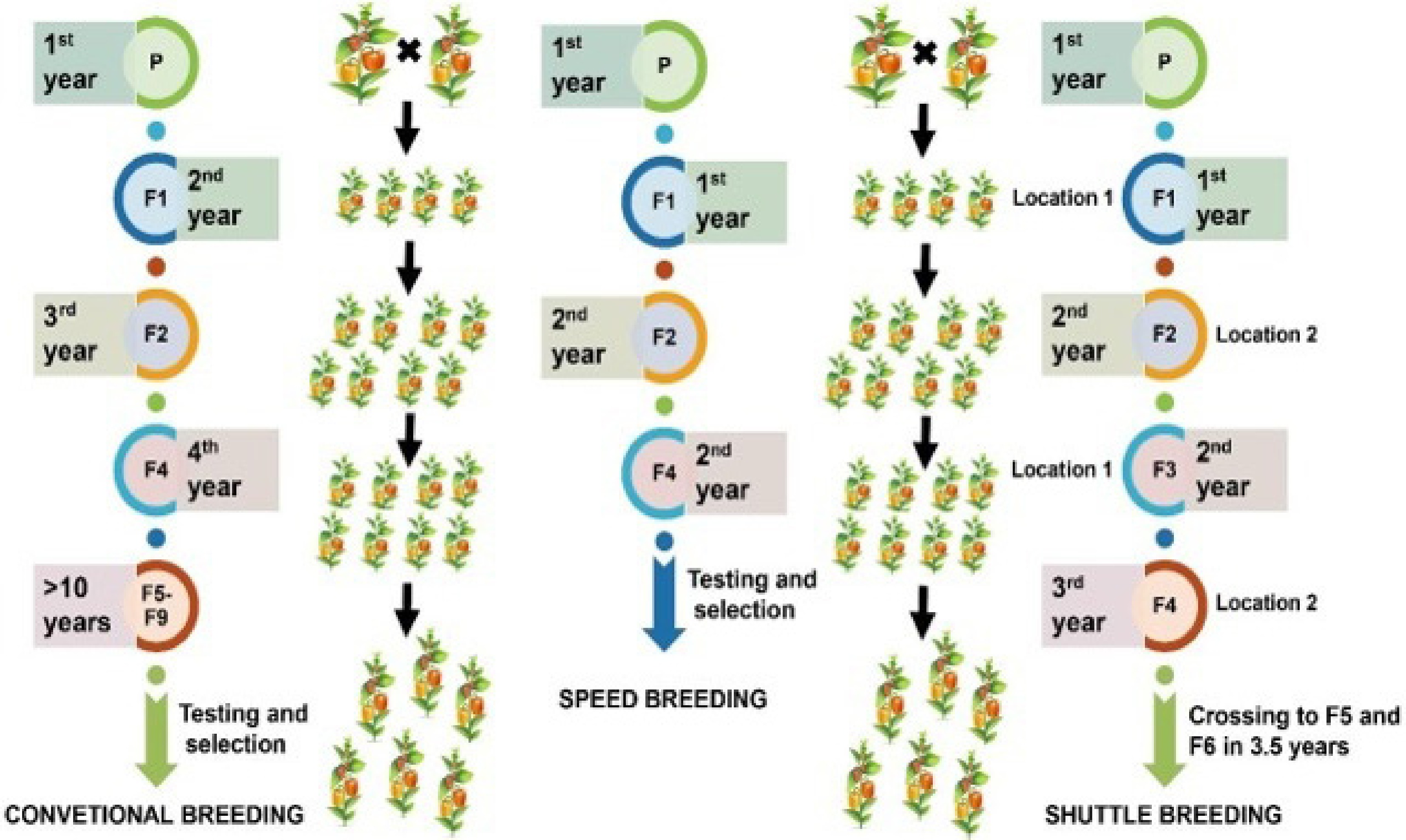
Figure 8.
Comparative representation of conventional, speed, and shuttle breeding techniques in vegetable crop improvement, showcasing the efficiency and timeframes.
-
Types of stress tolerance Breeding approaches Ref. Heat stress tolerance Selection and evaluation of breeding lines under high temperature conditions to identify heat-tolerant genotypes.
Use of physiological trait-based selection, such as canopy temperature and membrane stability.
Implementation of molecular and genomic-assisted breeding to accelerate identification of stress-resilient lines.[6,7] Cold tolerance Conventional breeding to select genotypes adapted to low-temperature environments.
Development of genetically modified plants carrying cold-tolerance genes.
Genome editing technologies are applied to introduce or modify genes regulating cold stress response.
[8]Salt stress tolerance Breeding programs aimed at developing cultivars with stable yield under saline conditions.
Introgression of salt-tolerance alleles into elite, high-yielding backgrounds.
Application of quantitative trait loci (QTL) mapping and marker-assisted selection (MAS) to identify and transfer salt-tolerant traits.
Marker-assisted backcrossing and genome-editing techniques for precise genetic improvement.[9,10] Drought stress tolerance Selection for superior yield stability under both drought and non-stress conditions.
Incorporation of drought adaptation traits, such as root architecture and water use efficiency.
Use of molecular markers, genomic selection, and genome-editing tools like CRISPR/Cas9 to enhance drought resilience.[11] Waterlogging tolerance Developing hybrids through crossing tolerant and susceptible genotypes.
Collection and evaluation of germplasm for waterlogging-tolerant traits.
Investigation of genetic mechanisms involved in anaerobic stress adaptation.
Utilization of QTL mapping to locate genes associated with tolerance.[12,13] Table 1.
Different breeding approaches for various kinds of stress tolerance.
-
Vegetable crops Breeding method Trait improved Ref. Tomato (Solanum lycopersicum) Backcross Enhanced tolerance to late blight, improved fruit texture [15] Brinjal/eggplant (Solanum melongena) Backcross Resistance to shoot and fruit borer, and yield stability [16] Chilli/pepper (Capsicum annuum) Pedigree Better fruit quality, improved disease tolerance, consistent pungency [17] Okra (Abelmoschus esculentus) Pure-line Increased yield, early maturity, uniform fruit traits [18] French bean (Phaseolus vulgaris) Pure-line High pod uniformity, disease, and pest resistance [18] Cowpea (Vigna unguiculata) Mass selection Larger fruit size, stress tolerance, and vine vigor [19] Cucumber (Cucumis sativus) Mass selection Bulb size uniformity, improved shelf life, and early maturity [20] Bottle gourd (Lagenaria siceraria) Recurrent selection Compact head formation, tolerance to pests [20] Onion (Allium cepa) Pedigree High curd compactness, adaptability to temperature stress [21] Cabbage (Brassica oleracea var. capitata) Mass selection Improved root color, sugar content, and shape regularity [14] Cauliflower (Brassica oleracea var. botrytis) Recurrent selection Resistance to powdery mildew, enhanced cold tolerance [22] Carrot (Daucus carota subsp. sativus) Mass selection Increased leaf yield, resistance to downy mildew [23] Pea (Pisum sativum) Backcross Higher fruit size, drought tolerance, and yield stability [24] Spinach (Spinacia oleracea) Pedigree Enhanced leaf yield, heat, and drought tolerance [25] Bitter gourd (Momordica charantia Mass selection High fruit set, color intensity, and disease tolerance [20] Table 2.
Overview of conventional breeding approaches in vegetable crops and their associated trait improvements.
-
Marker type Use in vegetable crops Advantages Ref. RFLP (Restriction Fragment Length Polymorphism) Tomato, cabbage Reliable, co-dominant marker; useful for gene tagging and varietal fingerprinting; not affected by environmental conditions [28,30] AFLP (Amplified Fragment Length Polymorphism) Broccoli, pepper, carrot High polymorphism detection; suitable for genome-wide screening; useful for linkage map construction [30] SSR (Simple Sequence Repeats/microsatellites) Cucumber, tomato, brinjal Highly polymorphic, reproducible, co-dominant; ideal for genetic diversity and parentage studies [28] SNP (Single Nucleotide Polymorphism) Pepper, cucumber, tomato Abundant across the genome; co-dominant; suitable for high-throughput and genomic selection studies [30] SCAR (Sequence Characterized Amplified Region) Tomato, chili Highly specific and reproducible; derived from RAPD markers; efficient for detecting specific genes or traits [31] InDel (Insertion–Deletion) Tomato, pepper Stable marker type; valuable for fine mapping and polymorphism studies in breeding populations [31] CAPS (Cleaved Amplified Polymorphic Sequence) Chili, tomato High specificity and co-dominant; useful for mapping and marker-assisted selection of quality traits [31] SRAP (Sequence-Related Amplified Polymorphism) Cabbage, lettuce, cucurbits Targets open reading frames; detects functional polymorphisms related to agronomic traits [29] CRISPR/Cas9 (Genome Editing System) Tomato, cucumber, lettuce Enables targeted gene modification for stress tolerance, disease resistance, and yield improvement [29,32] Table 3.
List of molecular markers used in vegetable crops and their specific advantages.
-
S. No. Vegetable crop Key factor Influence/observation Ref. 1 Pea (Pisum sativum) Photoperiod length
(22 h light/17 °C day-night cycle)Accelerated flowering and generation turnover; up to six generations per year achieved under controlled light regimes [39] 2 Hot pepper (Capsicum spp.) Light spectrum and far-red proportion Far-red supplementation (30–90 µmol·m−2 s−1) enhanced early bud initiation and shortened the reproductive phase to achieve four generations annually. [40] 3 Cabbage (Brassica oleracea var. capitata) Supplemental lighting
(blue/red ratio 2:1)An extended photoperiod increased budding rates up to threefold compared to field conditions. [24,41,42] 4 Chinese kale (Brassica alboglabra) Far-red (3–6 W·m−2) illumination Budding and flowering rates improved by 20%–35% relative to control treatments. [41,42] 5 Tomato (Solanum lycopersicum 'Micro Tom') Blue light exposure (~100 µmol·m−2 s−1) Stimulated earlier flowering by 3–4 d and faster fruit ripening under indoor photoperiod systems [43] 6 Tomato (Solanum lycopersicum 'Mini Chal') Supplementary UV-A radiation
(0.4 W·m−2)Mild UV-A stress induced floral initiation and reduced the vegetative period [11] 7 Cowpea (Vigna unguiculata) Planting density modulation Closer spacing (60 × 20 cm) advanced flowering time and improved seed set efficiency [44] 8 Spotted bean (Phaseolus vulgaris) Temperature control (20/18 °C) Moderate thermal regimes enhanced flowering and reduced cycle duration to under 60 d [26] 9 Soybean (Glycine max) Light intensity (480 µmol·m−2 s−1) Optimal light intensity reduced the generation cycle to roughly 75 d [45] 10 Faba bean (Vicia faba) Photoperiod variation (14–22 h) Long-day exposure (18 h) produced the most effective balance between rapid cycling and yield consistency. [46] 11 Amaranth (Amaranthus spp.) Temperature (30–35 °C, 16 h light) Induced early floral transition and enabled up to six generations per year under controlled temperature. [47] 12 Chinese cabbage (Brassica rapa) CO2 enrichment (380 vs 100 ppm) Elevated CO2 promoted photosynthetic efficiency and faster seed maturation [41,42] 13 Pea (Pisum sativum) Temperature + light combination Warmer regimes with 22-h photoperiod shortened the flowering phase by 2.5 × compared to field setups. [48] 14 Bean
(Phaseolus vulgaris L.)Planting at different densities
(10 × 30 cm, 20 × 30 cm, 30 × 30 cm,
40 × 30 cm, 50 × 30 cm, 60 × 30 cm)Planting density of 20 × 30 cm resulted in the highest yield [26] Table 4.
Various speed breeding factors and their influence on vegetable crops.
-
S.No. Conventional breeding Molecular breeding Speed breeding 1. Uses conventional techniques for plant crossbreeding to merge desired characteristics from two parent plants. Breeding is focused on phenotype, making it difficult to determine a pure line of genes. Breeding using a genotype-based approach is more accurate in identifying a pure gene line. It brings novel characteristics from allied species or cultivars. Uses methods for controlled environments to speed up the breeding cycle. Entails adjusting growth factors (temperature, light, etc.,) to encourage quicker plant development and more generations in a shorter time. 2. Dominant genes are typically the only ones chosen. It depends on recombination and natural genetic variation. Recessive alleles can be chosen using markers and the identification of certain gene locations, which is a quicker process than the traditional method.
Identifies plants with desired characteristics using molecular markers/DNA sequences.Speed breeding is a viable and effective solution for vegetable crops with huge populations. It can be used on a variety of vegetable crops and requires less work than conventional breeding techniques. 3. Less technological expertise and genetic understanding is required. It may be done using local methods and equipment, thus being less expensive. Molecular plant breeding is more reliable and effective. Due to the need for sophisticated equipment and techniques, it is costly. Needs specific equipment and facilities. Requires a large upfront setup cost for facilities in controlled environments. 4. Usually, takes 10–15 years of generation time to develop a new variety. A new variety can be released in a relatively shorter amount of time. By eliminating some of the trial-and-error procedures, breeding time can be significantly reduced. Significantly reduces the time needed to develop new varieties from years to months. 5. This technique is less accurate as unwanted characteristics could be added. Frequently used in a variety of vegetable crops. More accurate than traditional breeding, focusing on particular features or genes. Utilized for vegetable crops that have particular qualities that are desired, like increased nutritional value, drought tolerance, or disease resistance. With four to six generations every year, it has been successfully used on a variety of vegetable crops and is helpful in research settings seeking quick genetic improvement, or for vegetable crops with lengthy breeding cycles. Table 5.
Comparative overview of conventional and molecular breeding approaches, and the complementary role of speed breeding as a generation advancement tool.
-
Approach Vegetable crop Advantage Limitation Conventional Tomato, potato, chilli, brinjal, broccoli, bottle gourd, etc. Established resistance traits Slow, labor-intensive Molecular Tomato, cucumber, lettuce, etc. Precise trait introgression Costly, infrastructure-heavy Speed Lettuce, tomato, pepper, cabbage, Chinese kale Reduced cycle time Controlled environment required Integrated Tomato, pepper, soyabean, pea Rapid and precise breeding Requires expertise and funding Table 6.
List of conventional, molecular, and integrated speed breeding approaches in vegetable crops.
-
Vegetable crop Target trait/gene Reported outcome/success Approx. success rate Limitations/knowledge gaps Ref. Tomato Seedlessness (SlAGAMOUS-LIKE) Stable seedless fruit 60%–70% Moderate editing efficiency; few multi-trait studies; limited field trials [65] Tomato Ripening (SlEXP1) Delayed ripening, extended shelf life 65% Mostly greenhouse-tested; off-target effects possible [62] Tomato GABA enrichment Nutritionally enhanced fruit 50%–60% Lab-scale validation; field performance not fully evaluated [65] Cucumber Salt tolerance (ion transport genes) Improved salinity tolerance 55%–60% Limited field validation; genotype-dependent efficiency [66] Broccoli Flowering time (BoFLC2) Delayed flowering observed 50%–55% Transformation efficiency varies; limited multi-trait editing [61] Lettuce Heat tolerance Increased tolerance to heat stress 50%–55% Greenhouse data only; off-target mutations need assessment [67] Garlic Storage and shelf life Improved post-harvest traits 30%–40% Polyploid genome; low transformation efficiency [68] Pepper Fruit quality and disease resistance Enhanced fruit traits, disease resistance 50%–55% Mostly greenhouse trials; multi-trait editing is rarely explored [69] Spinach Downy mildew resistance Resistant lines developed 45%–50% Limited field data; editing efficiency varies by cultivar [10] Eggplant Anthocyanin content Increased antioxidant levels 50% Few studies; field validation lacking [32] Carrot Root shape and carotenoid content Improved root quality and nutritional traits 40%–50% Transformation efficiency is low; few multi-trait edits [70] Cauliflower Flowering time and curd development Delayed flowering and uniform curd size 45%–50% Genotype-dependent; field-level validation limited [71] Potato Late blight resistance Enhanced disease tolerance 40%–50% Tetraploid genome complicates editing; off-targets possible [9,16] Rapeseed Abiotic and biotic stress tolerance Positive results in field trials 60% Protocol optimization is needed for consistent efficiency [4] Table 7.
Applications and challenges of CRISPR/Cas9 in vegetable crop improvement.
Figures
(8)
Tables
(7)