-

Figure 1.
Phylogram constructed by maximum likelihood (ML) analyses of the combined tef1-α, rpb2, and cal sequence dataset of Fusarium. Bootstrap support valued for ML and Bayesian posterior probabilities (BYPP) equal to or greater than 70%, and 0.90, are shown above the nodes as ML/BYPP. The tree is rooted to F. concolor (NRRL 13459). The newly generated strain is indicated by black bold. T indicates the ex-type strains.
-
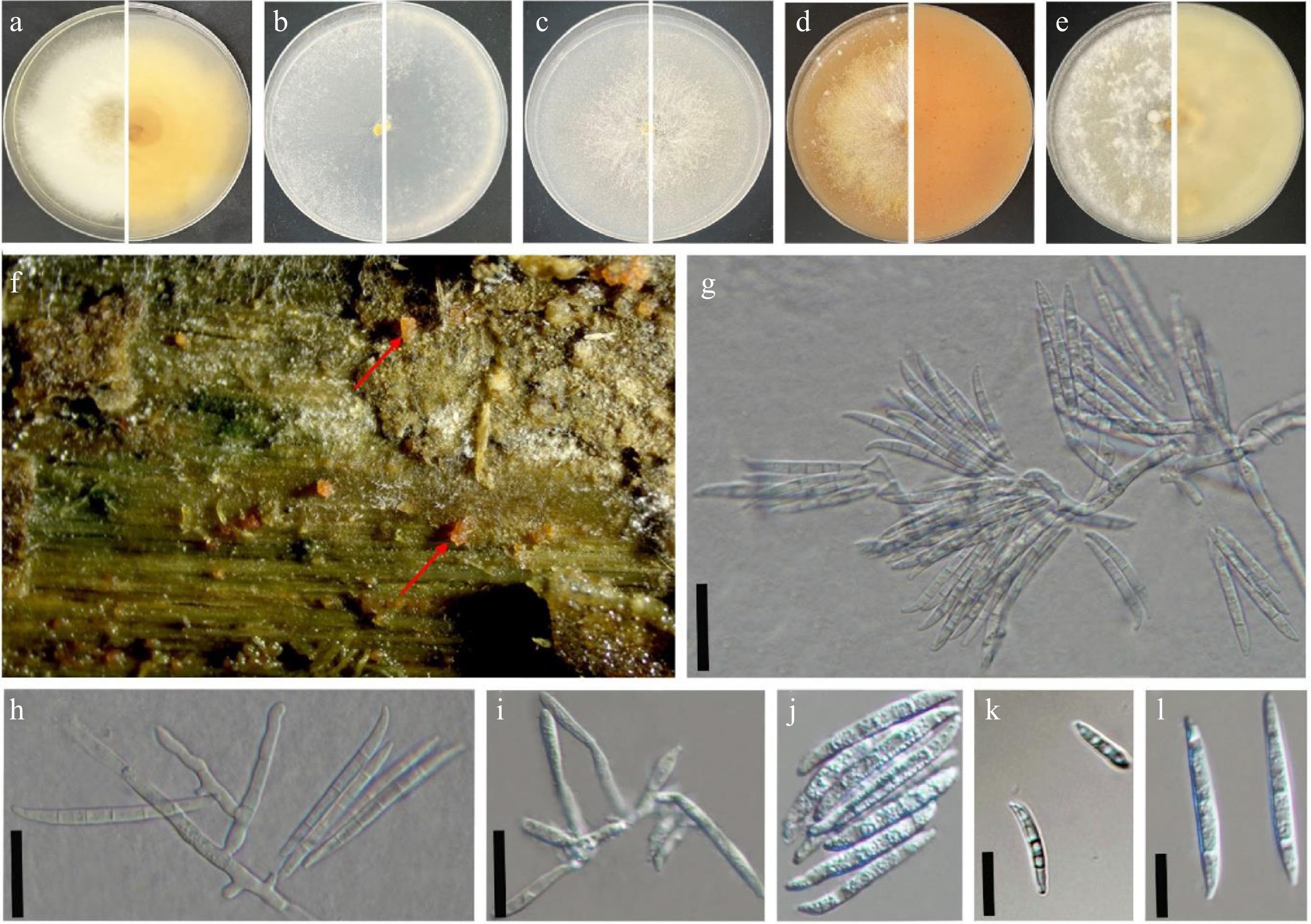
Figure 2.
Fusarium flavens (SDBR-CMU869 = KUNCC 25-20161, ex-type). (a)–(e) Seven-day-old culture on PDA, SNA, YEA, V8 juice agar, and OA incubated at 30 °C (top and reverse). (f) Natural symptoms of stem disease on Cannabis sativa (red arrow indicates the symptoms). (g), (h) Sporodochial conidiophores. (i) Phialides. (j)–(l) Macroconidia found on PDA media incubated at 30 °C. Scale bars: (g)–(i) = 100 μm; (k), (l) = 10 μm.
-
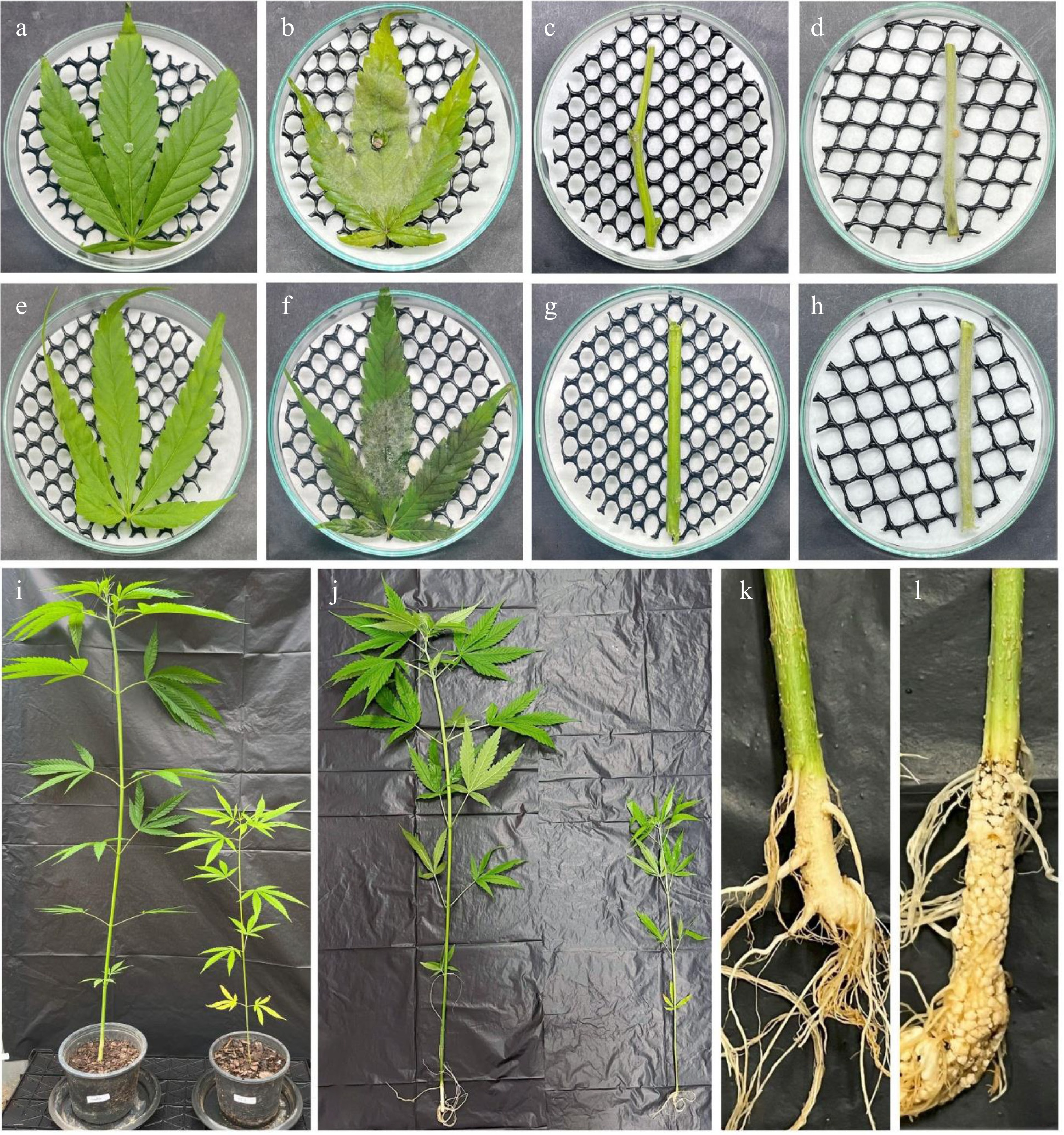

Figure 3.
Pathogenicity tests on detached hemp leaves, stems, and seedlings. (a) Control leaves, and (c) stems, inoculated with a PDAC plug. (b), (d) Symptoms after being inoculated with Fusarium flavens (SDBR-CMU869 = KUNCC 25-20161) mycelium plug for 7 d. (e) Control leaves, and (g) stems, inoculated with distilled water (DW). (f), (h) Symptoms after being inoculated with F. flavens (SDBR-CMU869 = KUNCC 25-20161) spore suspension for 7 d. Inoculation of hemp seedling control with (i) DW, compared with (j) that inoculated with F. flavens (SDBR-CMU869 = KUNCC 25-20161) spore suspension. (k) The roots of a plant inoculated with DW (control) shows no symptoms. (l) The roots of a plant inoculated with F. flavens (SDBR-CMU869 = KUNCC 25-20161) spore suspension shows root rot symptoms (30 d).
-
Locus Primer name PCR procedures Ref. tef1-α EF1/EF2 94 °C for 3 min, followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 55 °C for 50 s, elongation at 72 °C for 1 min, and final extension at 72 °C for 10 min [24,25] rpb2 RPB2-5f/
RPB2-7cR95 °C for 5 min, followed by 35 cycles of denaturation at 95 °C for 1 min, annealing at 52 °C for 2 min, elongation at 72 °C for 1.5 min, and final extension at 72 °C for 10 min [26] cal Cal-228F/
Cal-2Rd94 °C for 5 min, followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 59 °C for 30s, extension at 72 °C for
1 min and final step at 72 °C for 10 min[25] Table 1.
The PCR thermal cycle programs for gene amplification used in this study.
-
Taxon Strain GenBank assession number tef1-α rpb2 cal Fusarium aberrans CBS 131385T MN170445 MN170378 MN170311 F. aberrans CBS 131387 MN170446 MN170379 MN170312 F. arcuatisporum LC12147 MK289584 MK289739 MK289697 F. brevicaudatum NRRL 43638 GQ505665 GQ505843 GQ505576 F. bryceae BRIP 74865c PP209369 PP209368 — F. caatingaense URM 6779 LS398466 LS398495 — F. camptoceras CBS 193.65 MN170450 MN170383 MN170316 F. cateniforme CBS 150.25T = ATCC 11853 MN170451 MN170384 MN170317 F. caulendophyticum GUCC 191050.1 = CGMCC3.25474T OR043881 OR043826 OR043731 F. caulendophyticum GUCC 191050.2 OR043882 OR043827 OR043732 F. caulicola GUCC 191051.1 OR043883 OR043828 OR043733 F. citri CGMCC 3.19467T MK289617 MK289771 MK289668 F. citrullicola SDBR-CMU422 OP020920 OP020928 OP020924 F. clavus CBS 394.93 = BBA 64265 = NRRL 25795 GQ505597 GQ505775 GQ505509 F. clavus CBS 126202T MN170456 MN170389 MN170322 F. clavus CBS 130395 = NRRL 34032 = UTHSC 98- 2172 GQ505635 GQ505813 GQ505547 F. clavus CBS 119881 = MRC 8412 MN170457 MN170390 MN170323 F. clavus NRRL 32871 = FRC R-9561 GQ505619 GQ505797 GQ505531 F. coffeatum CBS 635.76T = BBA 62053 = NRRL 208 MN120755 MN120736 MN120696 F. coffeatum CBS 430.81 = NRRL 28577 MN120756 MN120737 MN120697 F. compactum NRRL 36323 GQ505648 GQ505826 GQ505560 F. concolor NRRL 13459 GQ505674 GQ505852 — F. croceum CBS 131777 MN170463 MN170396 MN170329 F. curculicola PPRI 20458T = PREM 61345 MF787266 MN605062 — F. duofalcatisporum CBS 384.92T = NRRL 36448 GQ505652 GQ505830 GQ505564 F. duofalcatisporum CBS 264.50 = NRRL 36401 GQ505651 GQ505829 GQ505563 F. equiseti CBS 245.61 = NRRL 20697 — JX171595 — F. equiseti NRRL 26419 = CBS 307.94 = BBA 68556T GQ505599 GQ505777 GQ505511 F. equiseti NRRL 36136 = CBS107.07, IMI 091982 GQ505644 GQ505822 GQ505556 F. equiseti NRRL 36466 = CBS 414.86 GQ505653 GQ505831 GQ505565 F. extenuatum LLC1501 OP487158 OP486728 OP486039 F. fasciculatum CBS 131382T MN170473 MN170406 MN170339 F. fasciculatum CBS 131383 MN170474 MN170407 MN170340 F. fasciculatum CBS 131384 MN170475 MN170408 MN170341 F. fecundum LC15875 OQ125250 OQ125544 OQ125281 F. flagelliforme NRRL 36269 GQ505645 GQ505823 GQ505557 F. flavens SDBR-CMU869 = KUNCC 25-20161T PX380329 PX380327 PX380330 F. goeppertmayerae BRIP 64547d OQ626868 — — F. gracilipes NRRL 43635T GQ505662 GQ505840 GQ505573 F. guilinense CBS 161.25 MN170448 MN170381 MN170314 F. guilinense MFLUCC 24-0626 PV394837 PV394827 PV297811 F. hainanense LC11638 MK289581 MK289735 MK289657 F. hainanense MFLUCC 24-0630 PV394834 PV394828 PV297812 F. heslopiae BRIP 74746a PP209371 PP209370 — F. humuli CGMCC 3.19374 = CQ1039T MK289570 MK289724 MK289712 F. humuli CQ1048 MK289571 MK289725 MK289713 F. incarnatum CBS 132.73 MN170476 MN170409 MN170342 F. ipomoeae CGMCC 3.19496 = LC12165 = M0111T MK289599 MK289752 MK289704 F. ipomoeae LC12166 MK289600 MK289753 MK289706 F. irregulare LC7188 MK289629 MK289783 MK289680 F. jinanense LC15878 OQ125131 OQ125521 OQ125271 F. khuzestanicum IRAN 4863C PP858510 PP858506 PP858502 F. khuzestanicum SCUA-Af-108–2 PP858512 PP858508 PP858504 F. kotabaruense Indo172 LS479445 LS479859 LS479429 F. kotabaruense MFLUCC 24-0633 PV394836 PV394831 PV297813 F. lacertarum NRRL 20423 = ATCC 42771 = CBS 130185T = IMI 300797 GQ505593 JX171581 GQ505505 F. lacertarum NRRL 36123 = CBS102300, BBA 70843 GQ505643 GQ505821 GQ505555 F. longicaudatum CBS 123.73T = ATCC 24370 = IMI 160825 = NRRL 25477 MN170481 MN170414 MN170347 F. longifundum NRRL 36372 GQ505649 GQ505827 GQ505561 F. luffae LC12167 MK289601 MK289754 MK289698 F. makinsoniae BRIP 64547a OQ626867 — — F. melonis SDBR-CMU424 OP020922 OP020930 OP020926 F. mianyangense LC15879 OQ125232 OQ125510 OQ125335 F. mianyangense MFLUCC 24-0625 PV394834 PV394829 PV297814 F. monophialidicum NRRL 54973 MN170483 KC808362 MN170349 F. mucidum CBS 102395 MN170485 MN170418 MN170351 F. multiceps NRRL 43639 GQ505666 GQ505844 GQ505577 F. nanum CGMCC 3.19498 = LC12168 = GXGL14-2T MK289602 MK289755 MK289651 F. nanum LC1384 MK289611 MK289764 MK289661 F. nanum LC1385 MK289612 MK289765 MK289662 F. nanum LC1516 MK289613 MK289766 MK289663 F. neoscirpi NRRL 26922 GQ505601 GQ505779 GQ505513 F. neosemitectum CBS 189.60 MN170489 MN170422 MN170355 F. nothincarnatum LC18436 OQ125147 OQ125509 OQ125290 F. oryzicola IRAN 4864C PP858509 PP858505 PP858501 F. oryzicola SCUA-Af-125–2 PP858511 PP858507 PP858503 F. pernambucanum MUM 1862T = URM 7559 LS398489 LS398519 — F. pernambucanum URM 6810 LS398485 LS398515 — F. pernambucanum MUM 1864 = URM 7554 LS398486 LS398516 — F. pernambucanum SDBR-CMU744 PX233235 — PX233226 F. pernambucanum SDBR-CMU745 PX233236 — PX233227 F. persicinum CBS 479.83 MN170495 MN170428 MN170361 F. radicigenum GUCC 197371.1 OR043907 OR043851 OR043752 F. radicigenum GUCC 197425.1 OR043908 OR043852 OR043753 F. radicigenum GUCC 197221.1 = CGMCC3.25478T OR043909 — OR043754 F. rhinolophicola KUMCC 21–0449 OR026001 OR025917 OR022061 F. ruthhalliae BRIP 72406h OP627085 OP627084 — F. scirpi NRRL 36478 GQ505654 GQ505832 GQ505566 F. serpentinum CBS 119880 MN170499 MN170432 MN170365 F. sulawesiense Indo138 LS479443 LS479855 LS479422 F. sulaweaiense MFLUCC 24-0629 PV394835 PV394830 PV297816 F. sylviaearleae BRIP 72816b OR269444 OR269438 — F. tanahbum-buense Indo176 LS479448 LS479863 LS479432 F. tangerinum JZG-2022b OP487189 OP486758 OP486067 F. toxicum CBS 406.86 MN170508 MN170441 MN170374 F. weifangense LC18333 OQ125107 OQ125515 OQ125276 F. wereldwij-sianum NL19-94.009 MZ921850 MZ921718 MZ921538 The new taxon is indicated in bold. The ex-type strains are indicated by T. Table 2.
GenBank accession numbers of the Fusarium incarnatum-equiseti species complex (FIESC) used in phylogenetic analysis.
-
Genes Percentage similarity across similar species tef1-α F. citri
NRRL 52765 (98.24%)
JF740839Fusarium sp.
FS5 (98.10%)
JQ244848Fusarium sp.
CMW55738 (97.94%)
MZ966178rpb2 F. humuli
hnxryc2 (98.82%)
OR257587F. humuli
JXRC 11 (98.71%)
MZ824673F. humuli
GL 17-3 (98.71%)
OQ512810cal F. humuli
LC18763 (99.27%)
OQ125277F. humuli
LC18553 (99.27%)
OQ125278F. humuli
CMML20-13 (98.99%)
OL331025Table 3.
Nucleotide BLAST similarity results among several similar strains of Fusarium flavens.
-
Compared strains Gene region/locus tef1-α rpb2 cal F. humuli
(CQ 1039, ex-type)2.8%
(18/629 bp)
MK2895701.1%
(10/874 bp)
MK2897240.5%
(3/531 bp)
MK289712F. weifangense
(LC18333, ex-type)3.4%
(19/551 bp)
OQ1251072.1%
(15/751 bp)
OQ1255151.4%
(8/548 bp)
OQ125276F. caulendophyticum
(CGMCC3.25474, ex-type)2.9%
(10/650 bp)
OR0438812.0%
(19/932 bp)
OR0438261.6%
(10/595 bp)
OR043731Table 4.
Nucleotide differences among closely related strains of Fusarium flavens.
-
Treatment Number of plants (n) Plant height
($\overline{\rm x} $ Disease incidence (%) Control plant 15 83.82 ± 5.67 no symptoms Inoculated plant 15 56.13 ± 9.03 73% Disease severity was evaluated based on plant growth performance. Growth reduction (%) relative to the control was calculated and reported in the Results section. Table 5.
Comparison of plant height between control and inoculated plants.
Figures
(3)
Tables
(5)