-
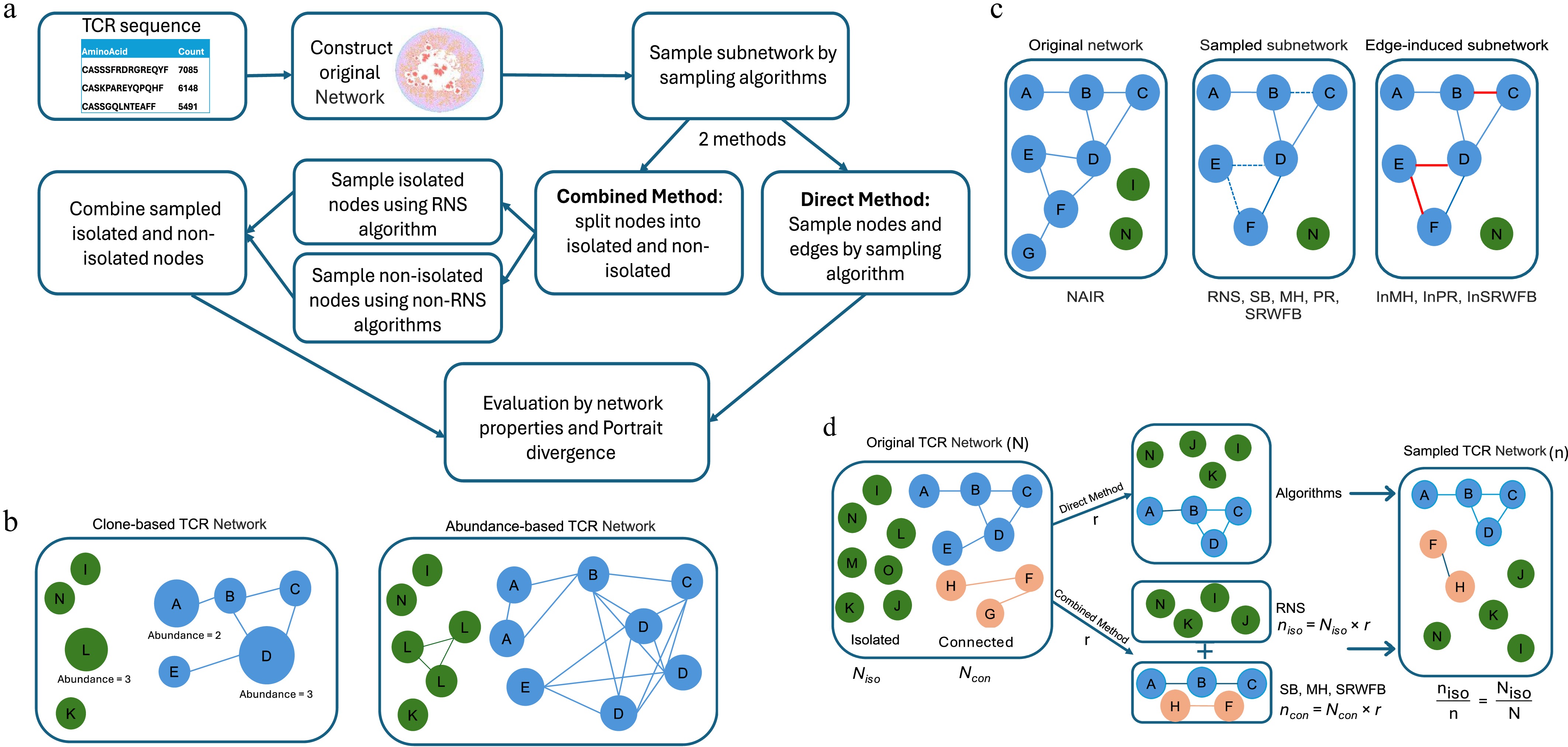
Figure 1.
Schematic chart of subsampling approaches. (a) Analytic flow of network subsampling. (b) Clone-based and abundance-based TCR networks. Both networks are constructed by setting hamming distance between the TCR sequences equals to 1. Nodes in clone-based network correspond individual TCR sequences. In abundance-based network, nodes are expanded based on counts (abundance) of each unique TCR clone. (c) Pseudo examples of original network, and subnetwork by the original algorithm and the induced algorithm. (d) Illustration of the direct and combined strategies. In the direct method, a network is subsampled as a whole using one single algorithm. In the combined method, nodes are partitioned into isolated (Niso) and connected (Ncon) groups. To preserve network sparsity, both groups are subsampled at a consistent rate r, such that niso = Niso × r, and ncon = Ncon × r. This proportional scaling ensures the subnetwork's isolation rate matches the original niso/n = Niso/N, preventing the edge-traversal bias of algorithms from artificially inflating connectivity. The subsampled results are then merged to form the final subnetwork.
-
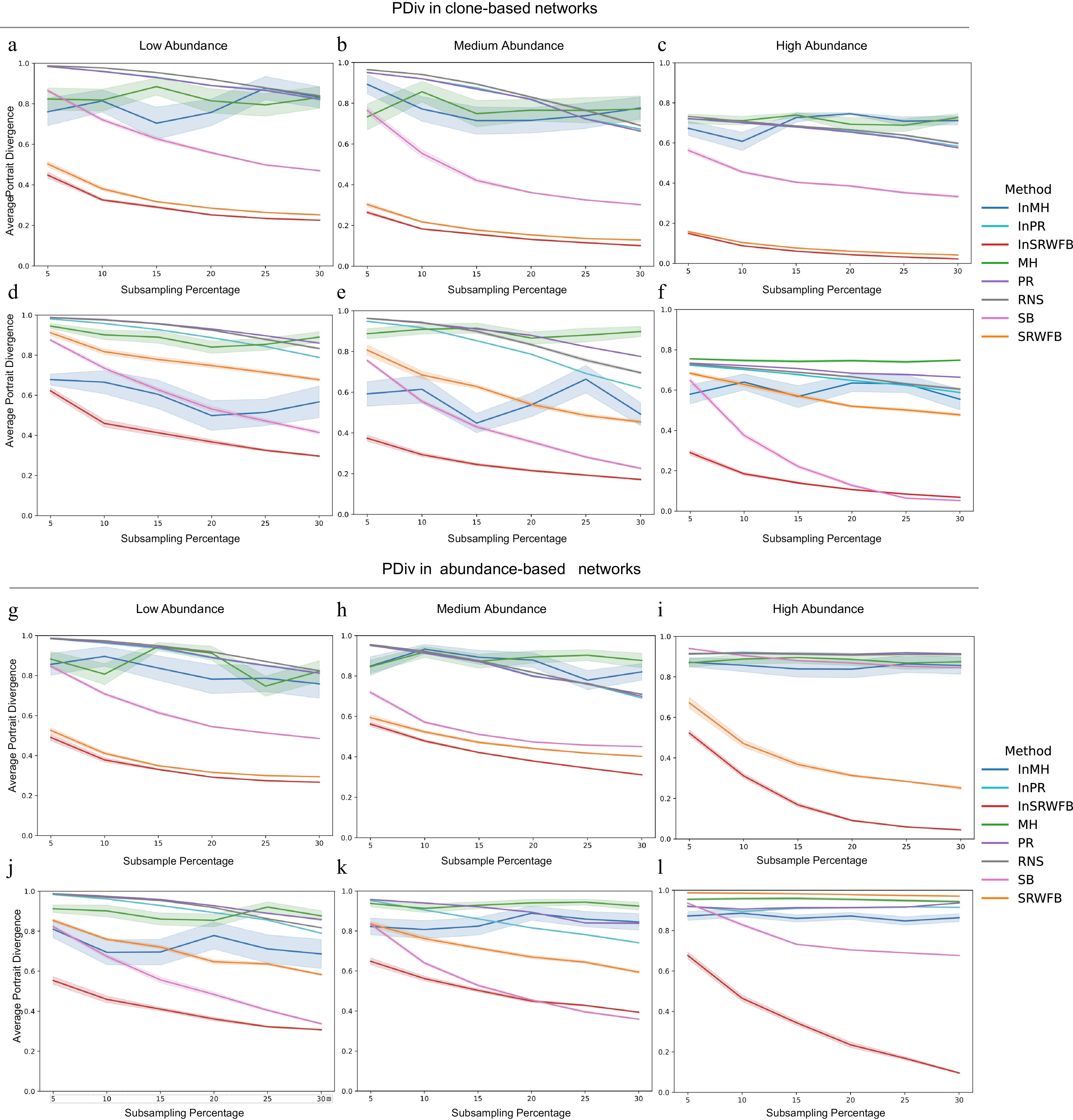
Figure 2.
Average Portrait Divergence (PDiv) between the original network and the subnetwork. PDiv under the direct strategy for clone-based networks with (a) low, (b) medium, and (c) high abundance level. PDiv under the combined strategy for clone-based networks with (d) low, (e) medium, and (f) high (f) abundance level. PDiv under the direct strategy for abundance-based networks with (g) low, (h) medium, and (i) high abundance level. PDiv under the combined strategy for abundance-based networks with (j) low, (k) medium, and (l) high abundance level. Each curve represents PDiv change across different subsampling percentages (5% to 30%) for one of the subsampling algorithms, including Metropolis-Hastings (MH), PageRank (PR), Random Node Sampling (RNS), Snowball Sampling (SB), and SRWFB, and Induced Metropolis-Hastings (InMH), Induced PageRank (InPR), Induced Simple Random Walk with Fly Back (InSRWFB). For each subsampling percentage, 20 replicates were performed per method, and the lines represent the mean PD across replicates. Shaded areas indicate mean+/- standard error. Lower PD values indicate greater structural similarity between the subnetwork and original networks.
-
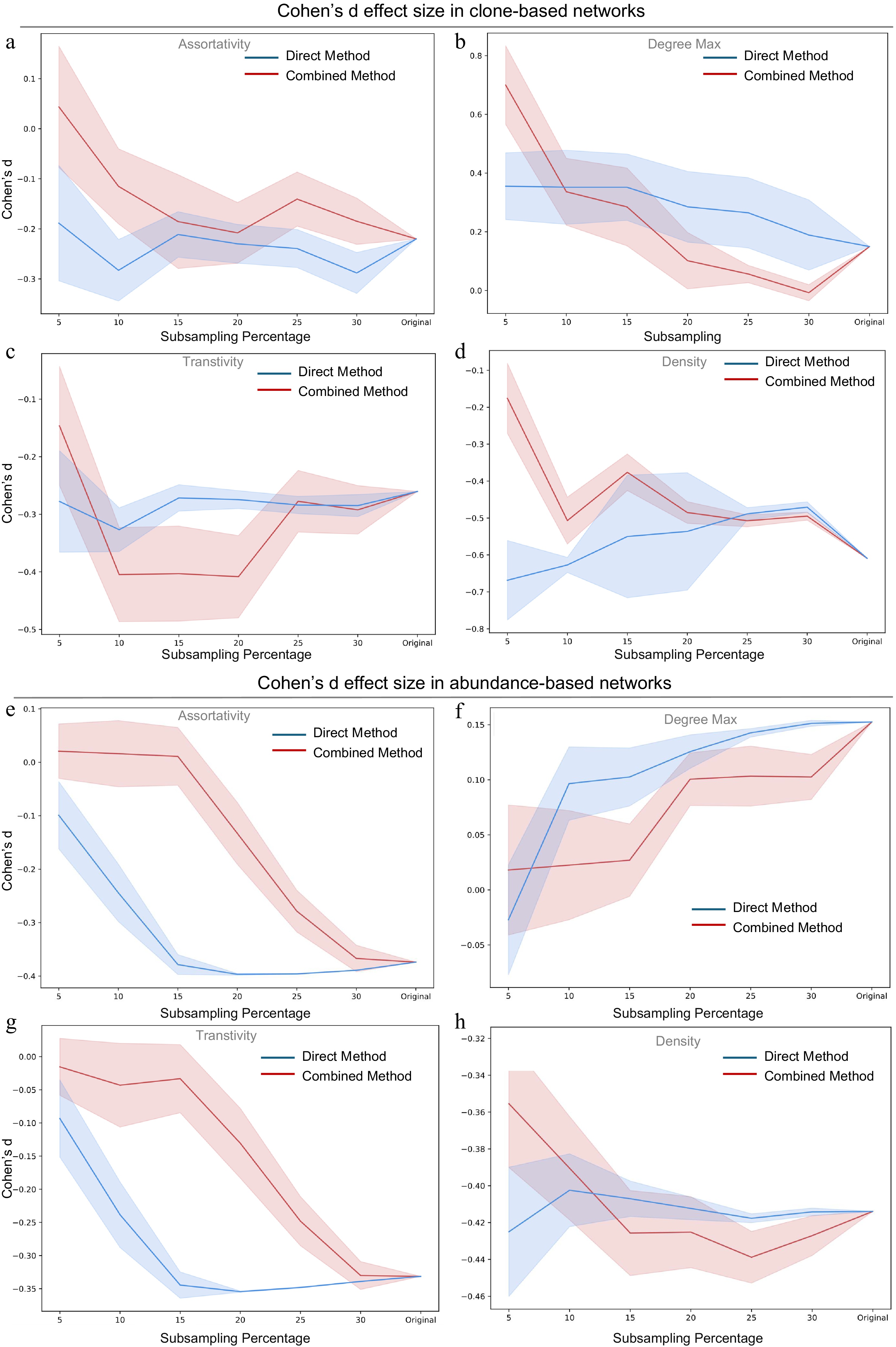
Figure 3.
Cohen's d effect size of original network and subnetworks using Induced Simple Random Walk with Fly Back (InSRWFB) at different subsampling percentages (5% to 30%). Cohen's d effect size of (a) assortativity, (b) maximum degree, (c) transitivity, and (d) density by InSRWFB for clone-based networks. Cohen's d effect size of (e) assortativity, (f) maximum degree, (g) transitivity, and (h) density by InSRWFB for abundance-based networks. Cohen's d values were computed based on 11 patients at two time points to assess the magnitude and direction of change in the four network properties. For each patient and time point, 20 independent subsampling replicates were generated, and the resulting d values were averaged across replicates. Blue and red lines represent the direct and combined strategies, respectively. Shaded areas indicate mean +/− standard error.
-
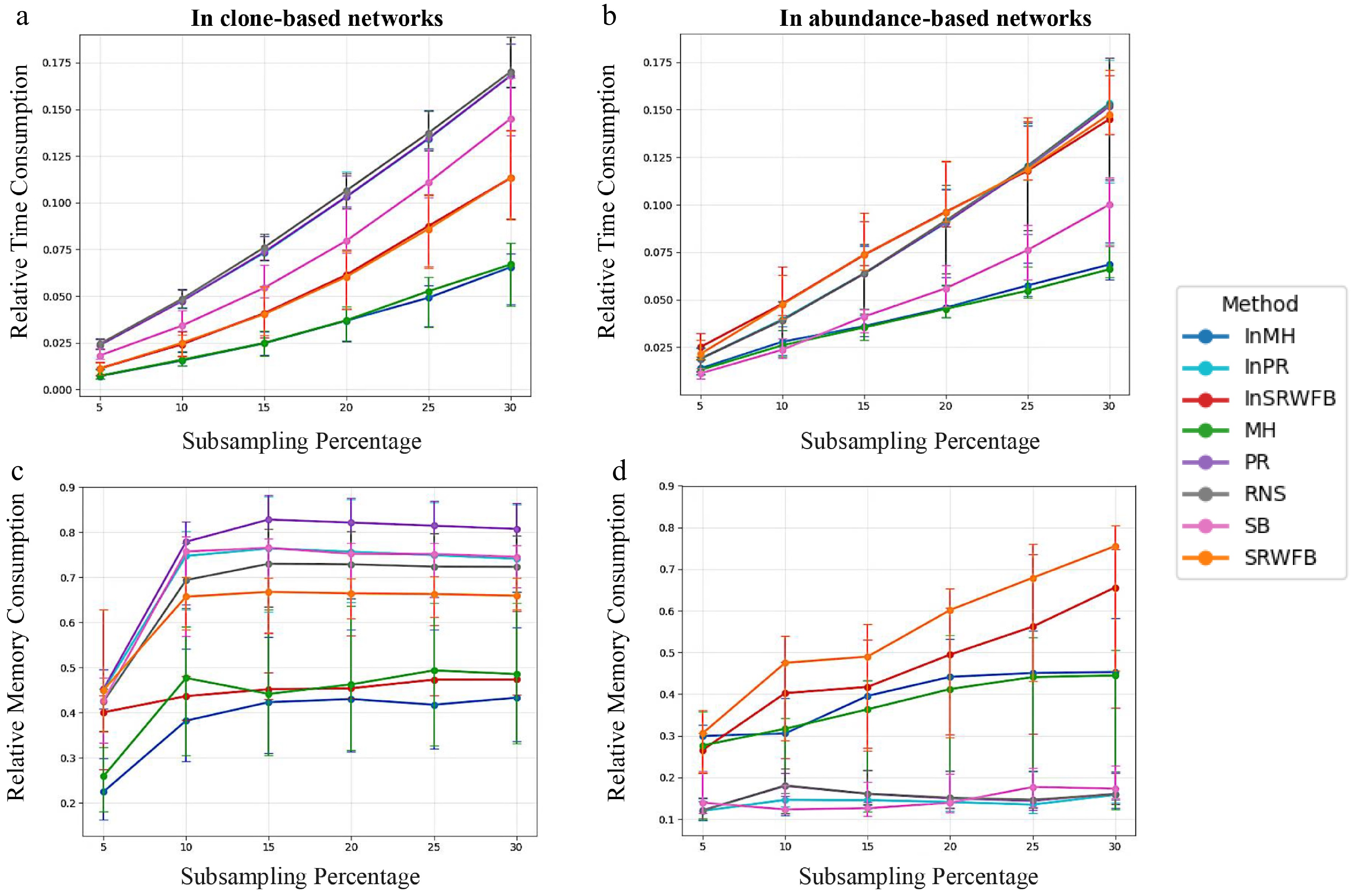
Figure 4.
Computation Time and Memory Consumption Median relative runtime across 22 TCR samples for each subsampling method and percentage (5%–30%) in (a) clone-based, and (b) abundance-based networks. Median relative peak memory across the same samples in (c) clone-based, and (d) abundance-based networks. Error bars indicate the interquartile range (IQR; 25th–75th percentiles).
-
Patient Proportion of nodes Abundance level ≥ 100 ≥ 200 ≥ 500 P1 6.50% 2.90% 1.40% Medium P2 17.70% 10.50% 6.20% High P3 3.40% 1.10% 0.50% Low P4 8.20% 4.10% 1.80% Medium P5 0.90% 0.30% 0.00% Low P6 8.60% 6.10% 3.00% High* P7 4.70% 2.80% 1.50% Medium P8 5.60% 3.30% 1.60% Medium* P9 2.80% 0.90% 0.50% Low P10 6.70% 5.10% 2.40% High P11 0.20% 0.10% 0.00% Low * * Indicates representative patients selected from each group for evaluation of sampling methods. Table 1.
Distribution of TCR node abundance.
-
Algorithm type Description Key parameters Random Node Sampling (RNS) Selects nodes uniformly at random from the network. − SnowBall (SB) Starts from a set of seed nodes and expands by connecting edges. k − Max number of neighbors added per cycle Page Rank (PR) Nodes are sampled based on their PageRank score in an iterative process. α (damping factor) − 0.85 Metropolis-Hastings (MH) Relies on edge connections and follows a Markov Chain Monte Carlo (MCMC) process. Acceptance depends on node's degree Simple Random Walk with Fly Back (SRWFB) Starts with a random node and performs a random walk with a predefined probability of returning to the starting node. p (fly-back probability) − 0.15,
iteration time − 100Induced-Page Rank (InPR) Retains all original edges between selected nodes. Inherits from PR Induced-Metropolis-Hastings (InMH) Retains all original edges between selected nodes. Inherits from MH Induced-Simple Random Walk with Fly Back (InSRWFB) Retains all original edges between selected nodes. Inherits from SRWFB Table 2.
Summary of sampling algorithms.
-
Metric Description Network Portrait Divergence (PDiv) Assesses similarity of 2 networks by analyzing 'Network Portrait'. Values range from 0 to 1. Network properties Max degree The maximum number of edges connected to a single node. Density The ratio of the number of actual edges to the possible number of edges. Assortativity Measures how strongly nodes with similar properties preferentially connect. Transitivity Measures the tendency of similar nodes to connect to each other. Table 3.
Evaluation metrics.
-
Subsampling percentage Relative time: Median (Min, Max) Relative memory: Median (Min, Max) Clone-based network Abundance-based network Clone-based network Abundance-based network 5 0.96% (0.6%, 3.1%) 2.4% (1.0%, 7.2%) 3.1% (0.1%, 7.2%) 10.8% (0.8%, 29.9%) 10 2.01% (1.2%, 4.6%) 4.5% (2.5%, 14.6%) 2.8% (1.6%, 10.6%) 13.3% (1.1%, 38.4%) 15 3.62% (2.1%, 8.5%) 7.3% (4.1%, 15.2%) 4.4% (1.5%, 19.2%) 17.2% (1.1%, 37.1%) 20 5.10% (3.4%, 9.8%) 9.2% (5.7%, 18.1%) 6.8% (1.2%, 19.7%) 16.8% (1.2%, 30.7%) 25 7.43% (4.8%, 12.9%) 11.8% (7.9%, 20.9%) 10.9% (1.6%, 38.7%) 20.3% (3.9%, 31.3%) 30 10.17% (6.9%, 16.3%) 13.8% (10.5%, 23.2%) 13.7% (1.2%, 48.0%) 20.7% (4.4%, 27.7%) Table 4.
Relative time and memory consumption of InSRWFB across subsampling percentages.
Figures
(4)
Tables
(4)