-
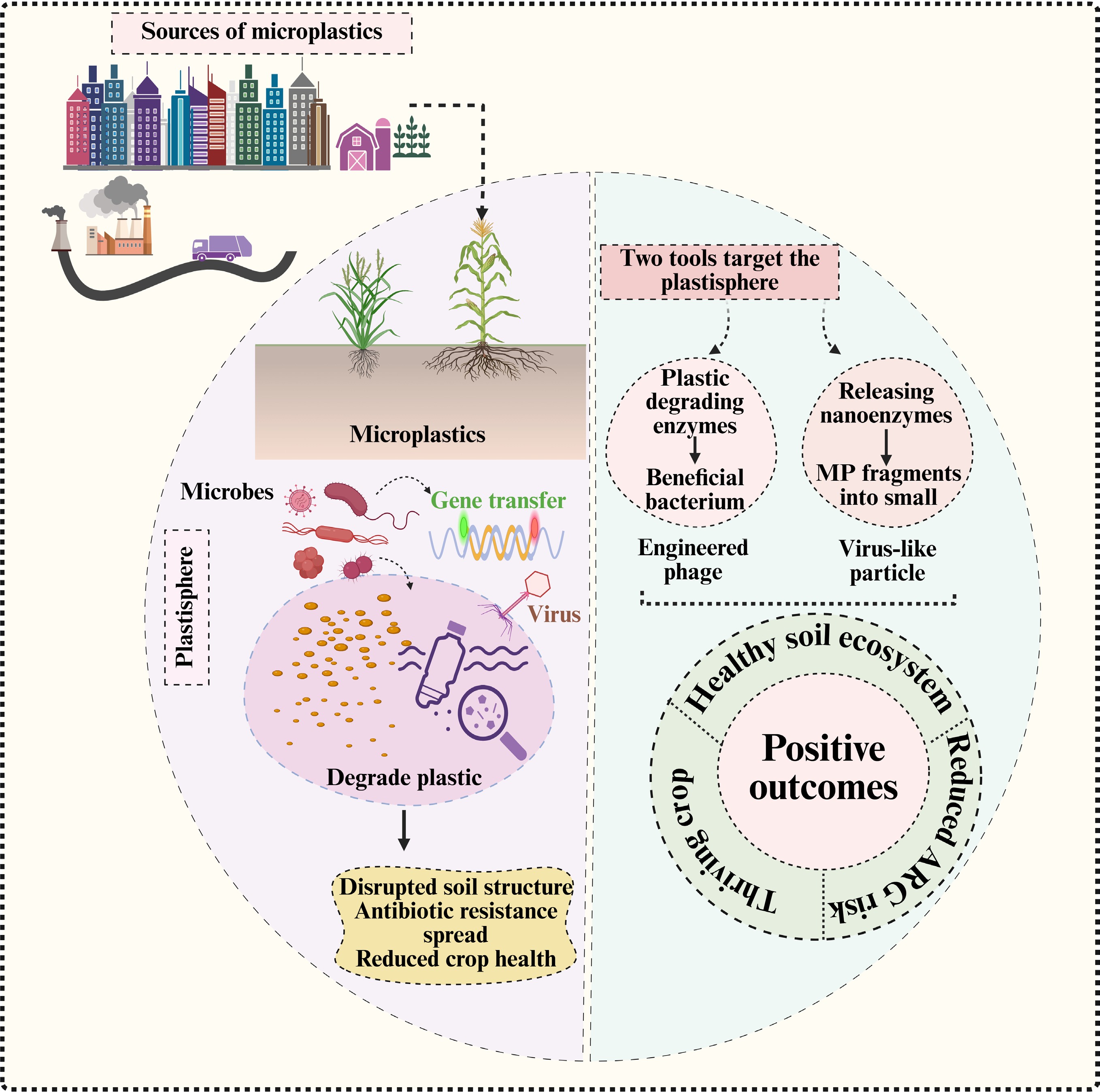
Microplastics (MPs), defined as plastic particles smaller than 5 mm, have become an emerging contaminant of global concern in terrestrial ecosystems, particularly in agricultural ecosystems[1]. Due to human agricultural activities (e.g., plastic film application, sewage irrigation, etc.), MPs in agricultural soils have accumulated to higher levels than those in other ecosystems[2]. The occurrence of MPs in the agricultural soils can disrupt multiple agroecosystem services, by altering soil physical, chemical, and biological processes[3]. For example, by altering aggregate stability and pore architecture, MPs can modify soil water retention, aeration, and microbial habitats that regulate microbial activity and diversity. MPs can also modify microbially mediated nutrient cycling processes and disrupt plant–soil interactions. This may lead to declining soil fertility, and a reduction in crop productivity and quality, ultimately compromising the long-term sustainability of food systems[4]. Moreover, MPs can enter the ecosystem and food web by being taken up by plants or consumed by soil organisms, potentially resulting in bioaccumulation, and posing health risks to humans[5]. The increasing evidence of MPs in the agricultural soil, which acts as both a reservoir, and a secondary source, underscores the urgent need to understand how it influences the ecological processes in the soil, which are vital for food security and environmental sustainability[6].
MPs in the soil have a negative impact on the microbiome in several ways, both directly and indirectly[7]. The microbial environment in the soil can change due to modifications in the porosity, water retention, aeration, and nutrient availability, which in turn influence the gradients and diffusion of oxygen, water, and nutrients within the soil[8]. MPs in the soil release plasticizers, stabilizers, and other detrimental substances that hinder the activity of enzymes and cellular metabolism[9,10]. Similarly, MPs serve as carriers of various pollutants, such as heavy metals, polycyclic aromatic hydrocarbons, or pesticides, leading to severe toxicity that eliminates sensitive microbial communities while promoting the growth of stress-resistant populations[11]. This can lead to a decline in microbial diversity and deteriorate functional processes over time.
MPs generate polymer-associated microhabitats in the soil, which promote the development of surface-associated biofilms, collectively referred to as plastispheres[12]. Microbial cells are densely clustered and enveloped by extracellular polymeric substances in these microhabitats, which facilitates their interactions and forms robust biological connections. These observed conditions are associated with a higher possibility of horizontal gene transfer (HGT), and the accumulation of antibiotic resistance genes (ARGs), indicating the plastisphere as a crucial hub for genetic exchange and microbial adaptation[13]. The formation of MP-associated biofilms modifies soil microbial interaction networks, and influences ecosystem processes mediated by microorganisms[14]. MPs may reduce soil fertility and hinder the ability of plants to absorb nutrients, disrupting the ecological processes driven by microorganisms in the soil, and ultimately limiting agricultural productivity[8]. MPs in the soil serve as both a physical and chemical stressor, indirectly interfering with essential biological processes crucial for sustainable crop production.
On the other hand, soil microbiomes are highly impacted by the presence of MPs in the soil, and closer attention has been directed towards the significance of soil viruses, particularly bacteriophages, which play a crucial role in regulating microbial diversity and function[15]. Viruses influence microbial communities through various processes, including lytic infection, lysogenic conversion, and gene transduction, which alter the movement of nutrients, metabolic processes, and transfer of genetic material within soil ecosystems[16]. In soils, viral processes play a crucial role in carbon and nitrogen cycling by breaking down host cells and releasing labile organic matter. At the same time, lysogeny enables phages to influence host metabolism through auxiliary metabolic genes (AMGs)[17]. MPs create new ecological niches that may alter these virus–host interactions; moreover, the varied microenvironment of the plastisphere enhances the likelihood of interactions between phages and their hosts, potentially resulting in prophage induction, increased gene exchange, and a shift in the equilibrium between lysis and lysogeny[18]. MPs may trigger a cascading effect on soil biogeochemical processes facilitated by microorganisms, influencing the decomposition of soil organic matter and nutrient cycling. These interactions are undoubtedly significant; however, there has been limited research on the relationships among MPs, soil microbes, and viruses. The current understanding regarding how MP pollution alters the functioning of agricultural ecosystems has enhanced our perspective, spanning from the microscopic scale to the broader ecosystem level.
Therefore, this review aims to provide a comprehensive understanding of the MP–microbiome–virome triad in agricultural soils. The objectives are: (1) to summarize current knowledge regarding the impact of MPs on soil microbial communities, metabolic processes, and ecosystem functions influenced by microbes; (2) to elucidate the potential role of soil viruses in regulating microbial and biogeochemical responses induced by MP pollution in the soil; and (3) to identify key research gaps and propose a framework for future investigations. This review highlights the interconnected roles of microorganisms, viruses, and other biological entities in shaping soil ecosystem resilience, the coupling of carbon and nutrients, and enhancing agricultural productivity, thereby contributing to the broader conversation on soil health and the sustainable management of agroecosystems.
-
MPs in soil function as enduring surfaces that support biofilm formation, allowing prominent microbial genera such as Pseudomonas and Bacillus to thrive among a diverse array of microorganisms[19]. They exhibit remarkable metabolic adaptability, create robust biofilms, demonstrate proficiency in pollutant degradation, produce spores, engage in enzymatic processes, and play a crucial role in nutrient cycling. These microorganisms adapt to non-living environmental stresses associated with MPs, facilitating the interaction of pollutants and the transfer of genes among them, thereby maintaining the ecological and functional dynamics of MPs within soil ecosystems[20].
The expression of functional genes, which play a significant role in the degradation of MPs, signifies a crucial adaptive mechanism in specific microbial communities residing on the surfaces of these materials and thus involves the initiation and control of specific enzymes that degrade MPs. For example, PETase (polyethylene terephthalate hydrolase), and MHETase (mono(2-hydroxyethyl) terephthalate hydrolase) work synergistically to break down polyethylene terephthalate (PET) MPs into environmentally friendly monomers[21]. The activity of these enzymes tends to increase when plastic substrates are present, triggered by environmental factors like the availability of plastic oligomers or associated stressors[22]. The genes responsible for these enzymes are frequently located in bacteria that decompose plastic, such as Ideonella sakaiensis. These genes transfer among various bacterial species, enhancing the ability of microbial communities in the plastisphere to degrade plastic efficiently[23]. Hence, a better understanding of the regulation of these enzymes may lead to the development of bioaugmentation techniques for environmentally friendly plastic remediation.
Toxicity and survival strategies
-
MPs and their associated pollutants exerted physical and chemical pressure on microorganisms. MPs also pose a significant abiotic stress on soil ecosystems, primarily due to their size and the substances they release. MPs exhibit a greater surface area-to-volume ratio, allowing them to navigate soil pores with greater ease and interact with a larger number of microbial cells[24]. Typically, MPs contain synthetic additives, including plasticizers (such as phthalates), flame retardants, stabilizers, and pigments, which gradually leach into the soil structure as time progresses[25]. Additionally, the absence of covalent bonds between these compounds and the polymer matrix increases their potential for degradation when exposed to UV light, temperature fluctuations, and microbial activity[26]. Thus, their release can be harmful, disrupting microbial metabolism and enzyme function, which might lead to the emergence of resistant strains; however, in terrestrial ecosystems, the ecological risk can be exacerbated by the mechanical and/or chemical stresses.
Exposure to MP pollution alters microbial responses through both the physical properties of plastic particles and the chemical toxicity of associated additives and adsorbed contaminants[27]. These selective pressures can drive shifts in microbial community composition and function, leading to the enrichment of ARGs, heavy metal resistance genes, and efflux pump systems[28]. Microorganisms may utilize compounds derived from MP or co-contaminants as alternative sources of carbon and energy, and thus activate genes that assist in the degradation of external substances, manage stress responses, and facilitate the formation of biofilms. HGT, primarily facilitated by plasmids and bacteriophages, can enhance the dissemination of adaptive traits within communities linked to plastispheres[18], which enhance microbial resistance and alter the interactions among microbes within ecosystems, potentially influencing nutrient cycling and the intricate networks of microbial relationships.
Biofilms linked to MP form microhabitats, which are considered to be beyond the adaptation of bacteria, influencing the interactions between viruses and their hosts in the soil environment. The plastisphere influences the dynamics of bacteriophage infections through mechanisms such as lytic, lysogenic, and virus-mediated gene transfer, thereby affecting microbial hosts within the biofilms[29]. As stress induced by MPs affects the microbial communities, virus-mediated processes may aid in later stages of co-adaptation, influencing the composition, stability, and functional capabilities of these communities in contaminated soils[30].
-
Phages are abundant in soil ecosystems and influence the variety and composition of microbial communities through lytic, lysogenic, and virus-mediated gene transfer pathways, subsequently impacting microbial activity. The development of plastisphere biofilms and the presence of MPs in agricultural soils produce distinct microenvironments that may affect the frequency of interactions between phages and hosts, as well as the spread of infections, thus altering the structure of microbial communities and their microscale activities.
Phage-mediated microbial community dynamics
-
By inducing lytic activity and lysogenic conversion, viruses have a significant impact on the function of microorganisms. Lysis can effectively inhibit competing or non-viable bacterial strains, while lysogenic conversion enables phages to survive within host genomes and, in some cases, affect host physiology through regulatory elements or additional metabolic genes. By focusing on and eliminating species that don't aid in the breakdown of plastic, some viruses may promote the growth of helpful bacteria, resulting in less rivalry for resources[31]. Furthermore, when introduced into host genomes, the presence of metabolic genes or regulatory elements in specific temperate phages can enhance stress tolerance, metabolic efficiency, or degrading abilities[18,32]. The selection and gene transfer process facilitated by phages can yield a more specialized and efficient population of bacteria capable of degrading plastic. Bacteriophage regulation may serve as an effective tool for enhancing bioaugmentation techniques in environments contaminated with MP (Fig. 1a, b).
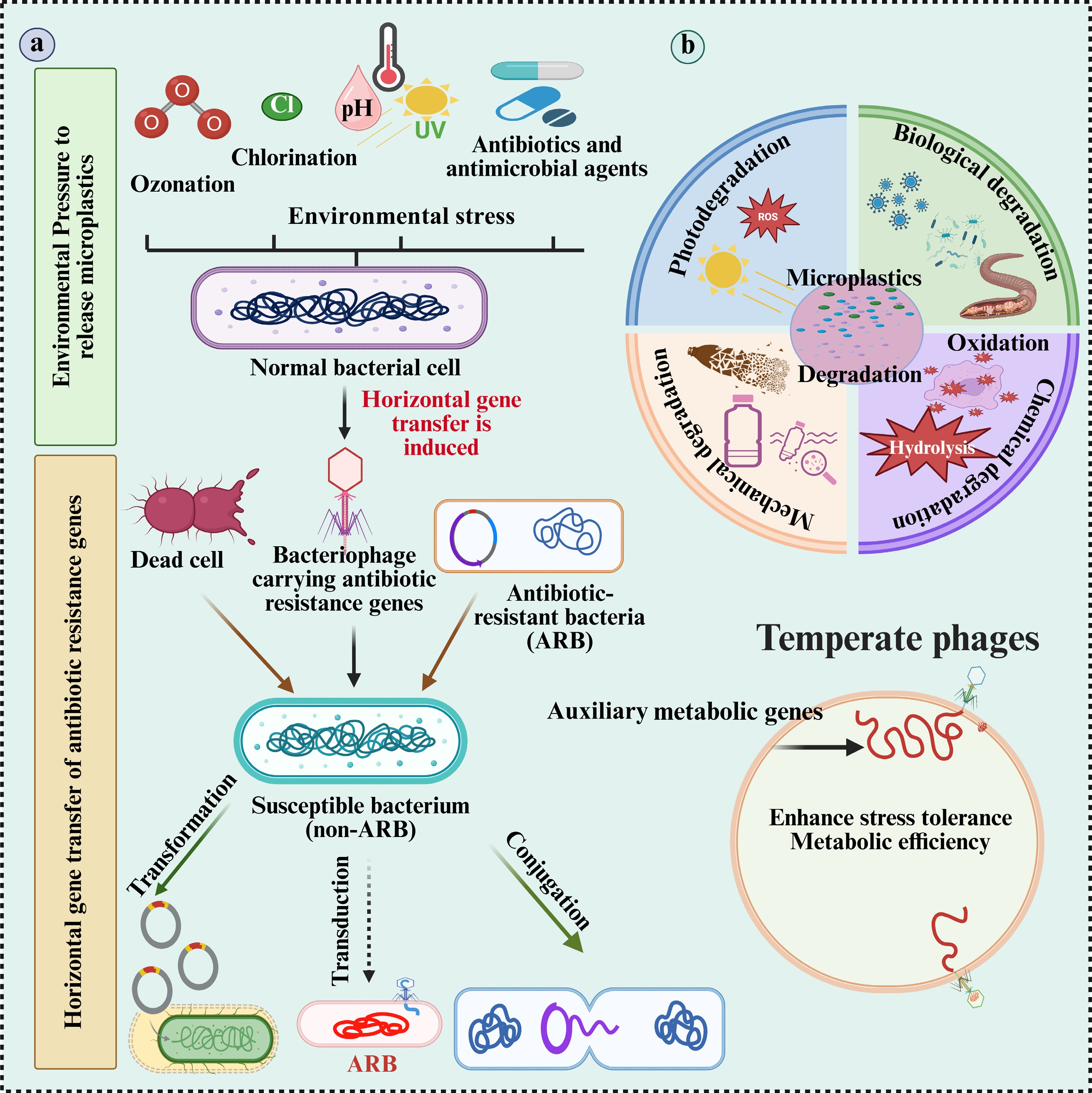
Figure 1.
Schematic representation of the proposed role of microplastics (MPs) in microbial adaptation under environmental stress. (a) Environmental stressors, such as ozonation, chlorination, UV radiation, pH changes, and antibiotics which accelerate the oxidation, hydrolysis, and ionization of MPs. These processes promote the degradation of MPs and the generation of reactive oxygen species (ROS). (b) The same stressors also induce horizontal gene transfer (HGT), particularly transduction, wherein bacteriophages carrying antibiotic resistance genes (ARGs) transfer these genes to susceptible bacteria, converting them into antibiotic-resistant bacteria (ARB). Additionally, dead cells may release ARGs that are acquired via transformation. MP degradation enhances microbial stress tolerance and metabolic efficiency, creating a feedback loop that supports the survival and proliferation of ARB under persistent environmental stress. ARB: antibiotic-resistant bacteria; ARGs: antibiotic resistance genes.
In MP-associated environments, the intricate relationships between bacterial defense mechanisms, such as CRISPR-Cas, facilitate the coevolution of phages and their bacterial hosts, which are shielded from invasive phages that target specific sequences by CRISPR-Cas systems. Nevertheless, HGT, which entails the acquisition of beneficial traits, may also be inhibited by these systems[33]. Accordingly, CRISPR-Cas may prevent genetic exchange in biofilms associated with MP while also contributing to the stability of specific bacterial populations[34]. The total impact on the ecosystem will likely depend on the prevalence of infection, the types of flora and fauna present in the area, and the climatic conditions. The lifespan and susceptibility of microorganisms to phage infection can be altered using CRISPR-Cas systems; however, applying these findings to soils requires caution[35]. The characteristics of the soil microenvironment considerably restrict the interactions between phages and their hosts in terrestrial ecosystems. Therefore, phages are essential to the microbial dynamics of soils contaminated by MP. However, their contributions are frequently underappreciated, and they are best described as context-dependent regulators rather than straightforward catalysts for the decomposition of plastic.
Viral shuttle of genetic information
-
Through the dissemination of functional genes, virus-mediated HGT, particularly through bacteriophages, can change the structure of microbial communities. Nonetheless, the extent to which phages disseminate genes that degrade plastic among soil ecosystems remains ambiguous. Transduction is the mechanism by which bacteriophages transfer genetic material between bacterial hosts. This can occur primarily through two mechanisms: specialized transduction, in which particular genes located near prophage integration sites are transferred during phage excision, and generalized transduction, in which random bacterial DNA is inadvertently incorporated into phage particles[36]. Phage-mediated processes may facilitate the transfer of ARGs among species in MP-associated biofilms, which are characterized by dense cell aggregation. This could increase the predominance of multidrug-resistant populations in terrestrial and aquatic ecosystems[37]. At the same time, plastic-degrading genes could spread through phages, accelerating the formation of microbial communities with enhanced biodegradation capacities[34]. The importance of viruses as gene carriers in contaminated environments has been confirmed by recent metagenomic studies of plastisphere communities, which have revealed phage-related sequences linked to both resistance determinants and catabolic processes. This gene transfer may aid in the breakdown of plastic, but it may also have negative ecosystem impacts by increasing the resistance of natural microbiomes to environmental changes. This highlights the importance of enhancing our understanding of phage-mediated horizontal gene transfer to develop effective and sustainable bioremediation techniques (Fig. 1a, b).
The ecosystem and public health may suffer greatly if phages transfer resistant traits. Through runoff, erosion, aerosols, or plant growth-related processes, resistant bacteria or their genes can spread from soils to nearby ecosystems, disrupting food production systems[38]. A recent study indicates that phages can transport genes that enhance the virulence of their hosts and improve their stress resilience, thereby increasing their likelihood of survival and pathogenicity[32]. Thus, the widespread presence of MPs in aquatic and terrestrial environments, along with the discovery of virulence factors and ARGs in microbial communities associated with plastic waste and their viral counterparts, suggests a complex and poorly understood mechanism of gene transfer that could unintentionally promote the survival of highly resilient microbial populations.
Virus-microbe synergy in plastic degradation
-
Bacteriophages may be used as vectors to introduce or mobilize genes associated with plastic breakdown throughout bacterial communities, according to the emerging concept of utilizing viruses to facilitate the delivery of enzymes or genes. There is still little proof, nevertheless, to justify its use in soil environments. Phages inject their genetic material into the cells of vulnerable hosts. Depending on the life cycle of the phages, this genetic material may include catabolic genes that are either active temporarily during lytic infection or preserved through lysogenic integration in both engineered designs and rare natural events[39]. In the lytic pathway, phage infection triggers the rapid expression of enzyme genes in tandem with phage replication. Consequently, new phage particles and active enzymes are released into the surrounding environment as a result of this process, potentially causing bacterial cells to rupture[40]. In some niches, particularly within the biofilms prevalent in MPs, this may lead to the breakdown of plastic. The lysogenic cycle, on the other hand, utilizes a temperate phage that incorporates its DNA into the host's genetic material, converting the host bacteria into a reliable and persistent source of plastic-degrading enzymes[41]. Under ideal conditions, this gene delivery could expand the number of bacterial community members capable of degradative activities. However, transforming non-degraders into effective degraders necessitates verified expression, suitable secretion/localization, and quantifiable increases in degradation rates, all of which have not been fully evaluated in complex soil communities (Fig. 1a).
Virus-assisted enzyme delivery systems have been proposed as a viable bioremediation strategy, particularly in environments where native microorganisms lack sufficient enzymatic activity to degrade plastics effectively. To ensure accurate gene delivery and reduce any unexpected environmental effects, phages can be designed to target particular bacterial strains. This strategy, however, could also raise biosafety issues, as gene transfer through phages may unintentionally spread other functional genes, such as those linked to virulence factors or antibiotic resistance[42]. Therefore, even though virus-assisted enzyme delivery presents a potentially valuable strategy to improve plastic biodegradation in contaminated environments, its application necessitates careful risk assessment and regulatory control.
The application of synthetic biology for environmental remediation has revealed promising prospects, particularly through the development of modified bacteriophages that enhance plastic degradation[43]. Furthermore, phage genomes can be modified using synthetic biology to improve enzyme expression levels, stabilize enzymes in a variety of conditions, and control phage life cycles to optimize degradation and reduce effects on microbial populations[44]. In addition to providing enzymes, it is suggested that synthetic phages have regulatory or detecting gene circuits. However, the viability of triggering environmentally responsive mechanisms in complex soil matrices remains primarily theoretical and has not yet been confirmed by actual data[45]. By improving the natural breakdown processes without introducing additional microbial species, this strategy could help preserve the environmental equilibrium. Phage cocktails, made possible by synthetic biology, comprise modified phages intended to target different kinds of polymers or bacteria[46], making them valuable due to their wide variety of applications against various contaminants and microbial populations. Despite the great potential of modified phages, their use presents serious ethical and biosafety concerns, including the potential hazards of HGT and their environmental impacts. Consequently, any investigation into synthetic phage-based methods should be limited to carefully monitored experimental environments. To be successful, these types of projects require thorough testing and efficient containment strategies. It is crucial to put preventative regulatory measures in place before conducting any ecological assessments.
A proposed containment technique involves modifying phages to rely on specific host-derived tRNAs or metabolites, thereby facilitating efficient replication predominantly in genetically modified host bacteria within regulated settings. To function as a replication protein or a crucial capsid component, the phage might be genetically altered to require an amino acid that is not naturally present[47]. The phage is unable to complete its life cycle or reproduce within natural, unaltered bacterial communities because this particular metabolite or amino acid is exclusively present in regulated remediation settings, such as specialized bioreactors or enhanced soils. Genetic circuits designed to induce self-inactivation in phages or transition them to a non-replicative state after a predetermined number of replication cycles can be integrated into the genome to enhance safety[48]. For instance, a phage's integrated CRISPR-Cas system may be programmed to activate and eliminate crucial phage genomic sequences after a few generations. A 'dead-man's switch' would be inserted as a result, preventing the phage from surviving for very long. Furthermore, inducible promoters that regulate genes responsible for cell death may be present in specially engineered phages[49]. These promoters can be activated by an artificial, external stimulus that is uncommon in nature, such as a particular chemical inducer or a particular wavelength of light. At the conclusion of the remediation procedure, a toxin or nuclease would be released in response to this stimulus, eliminating the remaining altered phages.
Net ecological effect of phages
-
The intricate processes by which bacteriophages impact the plastisphere have been explained in the sections above, which also portray them as beneficial bioremediation agents and potential sources of ecological risk. A conceptual conundrum arises from the apparent contradiction regarding the circumstances under which the balance shifts in favor of a net positive or net negative result. A conceptual framework is proposed that integrates phage traits, environmental constraints, and microplastic–interface properties to support more predictive interpretations of phage effects (Fig. 2). In this framework, net outcomes emerge from dynamic interactions among phage traits, environmental filters, and microplastic-interface properties, rather than being intrinsic to phages alone.
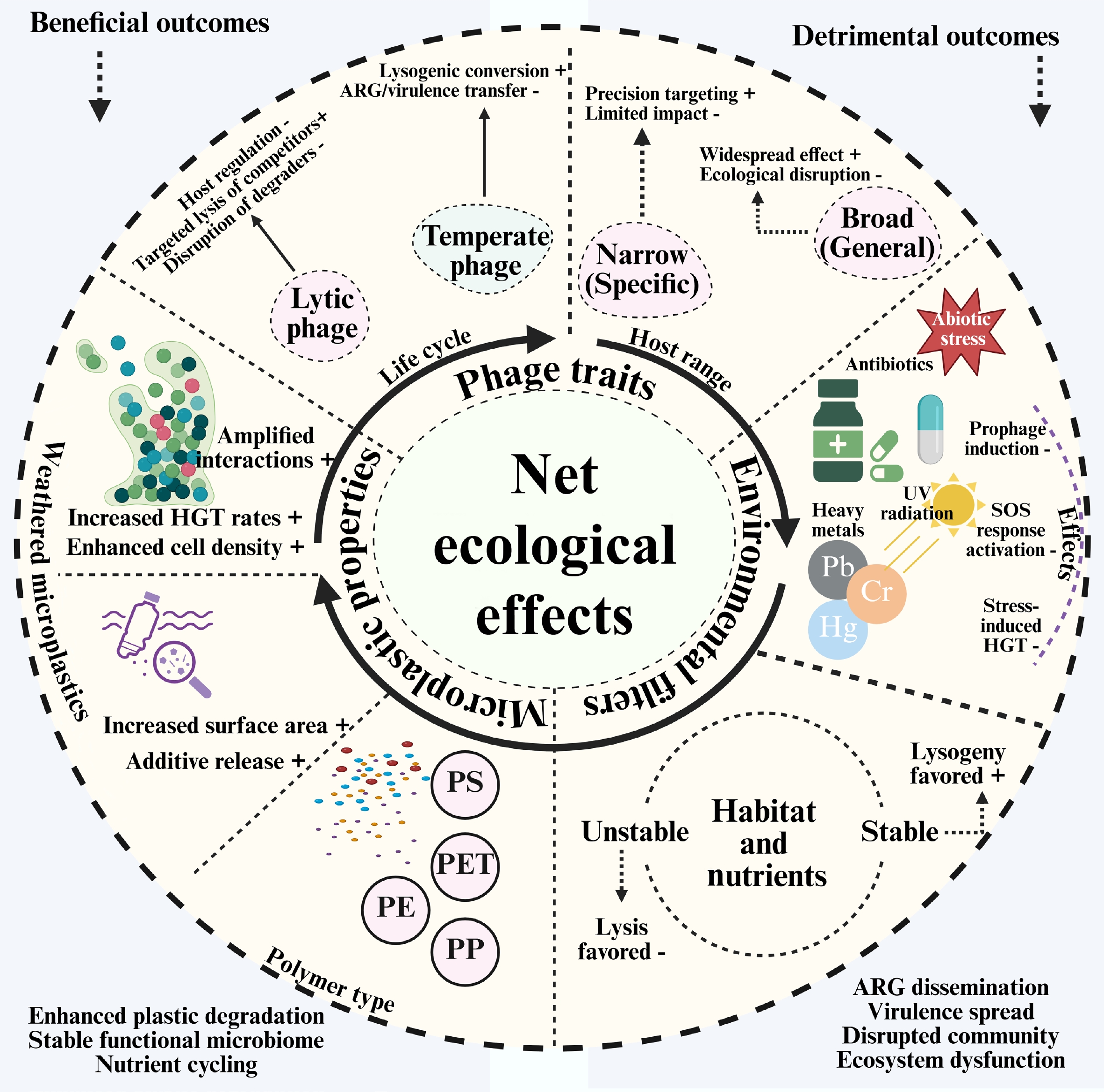
Figure 2.
Conceptual framework determining the net ecological effect of bacteriophages in the plastisphere.
The primary function of lytic phages is top-down regulation. Host specificity has a significant influence on their overall impact. Selective lysis can, in some cases, shift community composition by suppressing particular host populations; however, phages do not 'target' organisms based on ecological function, and whether lysis favors degraders or non-degraders is context dependent. Furthermore, regardless of their significance, non-specific or broad-host-range lytic activity can kill crucial degraders, such as Bacillus or Pseudomonas, halting biodegradation, and destroying the functional community[50]. Through lysogeny, temperate phages act as genetic modulators. By altering the host bacterium's DNA to confer beneficial characteristics, such as enzymes that degrade plastic (like PETase), genes that enhance its stress tolerance, or elements that facilitate biofilm formation, integrating a prophage can be beneficial[51]. The primary risk occurs when the prophage is activated. In addition to killing the host, environmental stressors can trigger a synchronized lytic cycle that disperses any encoded genes, such as virulence factors or ARGs, across the biofilm[52]. For focused actions that don't significantly alter the environment, a host range that incorporates a phage with a limited host range is also beneficial. Conversely, a phage with a broad host range is more likely to have both beneficial and detrimental effects. For instance, it might widely enhance a catabolic function or disperse ARGs, which would make its ecological effects less predictable and perhaps more extensive[53].
Phage behavior is altered by its surroundings, which act as powerful filters and switches. In addition to exerting selection pressure on bacteria, antibiotics, heavy metals, nutrient deficiencies, and UV radiation are potent regulators of phage dynamics. These stressors can activate bacterial DNA damage responses (e.g., the SOS response), which can trigger prophage excision and the transition from lysogeny to lytic replication[54]. As a result, the system may transition from a helpful, stable lysogeny to one of unchecked gene transduction and destructive lytic growth. The same pressures can select for resistant microorganisms; phages may then influence the distribution of resistance determinants indirectly through transduction and by reshaping host population dynamics. The frequency of interactions between phages and hosts is also influenced by the physical composition of the soil and the availability of nutrients[16]. While nutrient-rich, stable settings, such as established biofilms, may support more temperate interactions and lysogenic stability, oligotrophic (nutrient-poor) conditions often encourage lytic cycles and 'kill-the-winner' dynamics (Fig. 2).
The MP particle actively shapes the interaction rather than acting as a passive substrate. Different populations of bacteria are attracted to varying types of plastics, and these microbes, in turn, host diverse groups of phages[55]. Thus, the surface area of MPs increases and becomes rougher with weathering, which promotes the formation of biofilms and facilitates cell-to-cell contact. These conditions may increase the potential for genetic exchange (e.g., conjugation, transformation, and transduction), although in situ rates and dominant pathways remain poorly quantified in soils. Phage-related responses may also result from environmental stress brought on by the polymer's chemical additives leaching. The plastisphere creates an extraordinarily dense and interconnected microenvironment by collecting bacteria and viruses on a limited surface area[56]. This may increase local phage–host encounter probabilities relative to adjacent bulk soil, although the magnitude of this contrast depends on soil moisture connectivity and adsorption processes. This implies that each MP may develop into a hotspot for both the propagation of harmful ARGs and advantageous gene transfer[57]. The overall ecological effect is depicted as a scale that balances things out in this context (Fig. 2). The position of the scale is determined by the total weight of factors that are pushing for a net positive effect (such as stable acquisition of catabolic genes and targeted lysis of competitors), vs those that are resulting in a net adverse effect (such as induction-driven ARG bloom and disruption of essential degraders). By understanding this intricacy, we may utilize phages as a sophisticated tool for managing the plastisphere's environment rather than a crude one.
-
The interplay between microorganisms and viruses represents a promising, nature-inspired strategy for mitigating microplastic pollution and restoring soil functions impaired by MP contamination[58]. Phages can be regarded as possible modulators of microbial community dynamics via infection and gene transfer; however, the phage host range limits host targeting, and HGT may encompass both advantageous and detrimental traits[59]. This collaboration enhances the adaptability of microorganisms and their functional capabilities, resulting in a more effective and sustainable process for degrading contaminants, such as MPs. This technology may offer a strategy based on biological principles to attain more adaptive and sustainable remediation by combining microbial metabolism with virus-mediated gene delivery and regulation[60]. Pathogenic bacteria or rapidly proliferating non-degraders may overshadow or hinder beneficial degraders, such as Pseudomonas or Bacillus species, thereby reducing the overall effectiveness of plastic biodegradation. Phage predation can affect microbial competition; however, it does not directly inhibit pathogens or resistance genes, and in certain circumstances, it may even facilitate their spread[61]. Moreover, alterations in population dynamics induced by phages may affect the longevity of specific degradative groups; however, increased degradation is more likely for hydrolysable polymers (such as PET) than for polyolefins like polyethylene (PE), which necessitate different oxidative or abiotic preconditioning. Some phages undergo lysogenic conversion, integrating into the host genome and potentially imparting advantageous traits that enhance bacterial metabolism, increase stress resistance, or promote biofilm formation, thereby further assisting degraders. Phage-mediated mechanisms can serve as ecological filters, modifying the composition of microbial populations linked to MP. The results may differ, shaped by the environmental factors and community dynamics[18]. This selection pressure promotes and maintains a population of bacteria proficient in plastic degradation, which is essential for advancing microbial bioremediation efforts to mitigate plastic pollution in terrestrial and aquatic ecosystems.
It has been suggested that Virus-Like Particles (VLPs) containing nanoenzymes could provide a theoretical framework for examining catalytic systems on MP surfaces, potentially facilitating polymer transformation in a controlled environment. VLPs are nanoscale protein assemblies that mimic natural viruses but are devoid of any viral genetic material, therefore rendering them incapable of inducing disease. However, the incapacity to spread diseases does not suggest that they are devoid of potential harm to the ecosystem[62]. These particles can be designed to encapsulate or display catalytic enzymes or enzyme-mimetic substances. Nonetheless, the most efficacious approach to activate enzymes is to use polymers that contain esters, such as PET. The direct enzymatic cleavage of polyolefins, including PE and polypropylene (PP), is significantly limited and generally necessitates prior activation by oxidation or abiotic techniques. Nanoenzyme-loaded VLPs exhibit a notable ability to target and attach to MP surfaces, utilizing the structural benefits of VLPs, such as their uniform size, extensive surface area, and adeptness in traversing complex biofilms or soil matrices. When the nanoenzymes are optimally positioned, they promote the degradation of polymer chains by hydrolysis, oxidation, or other enzymatic processes. This technique deconstructs long-chain molecules into smaller, more environmentally friendly fragments[63]. Encapsulation or immobilization within VLPs may enhance enzyme stability under specific conditions; however, it is uncertain if these configurations effectively address concerns regarding substrate accessibility, sorption, and inactivation in soils. VLPs can be modified with targeting ligands to enhance their selectivity for specific plastics or environmental niches, thereby enabling a more precise bioremediation strategy[64] (Fig. 3). Their nanoscale dimensions facilitate their diffusion within soil ecosystems, thereby enhancing interaction with more widely dispersed MP particles. The modularity of VLP design is essential, as it reduces the simultaneous administration of many enzymes or synergistic catalysts, which could improve degradation rates and expand the range of substrates[65]. This method leverages the most effective aspects of virology, nanotechnology, and enzymology to develop resilient and efficient catalytic systems for MP remediation (Fig. 3). Nevertheless, concerns persist regarding large-scale production, ecological stability, and potential impacts on natural systems, necessitating further evaluation and improvement. VLPs with nanoenzymes may offer a novel strategy to alleviate plastic pollution by degrading polymer chains at the nanoscale[66].
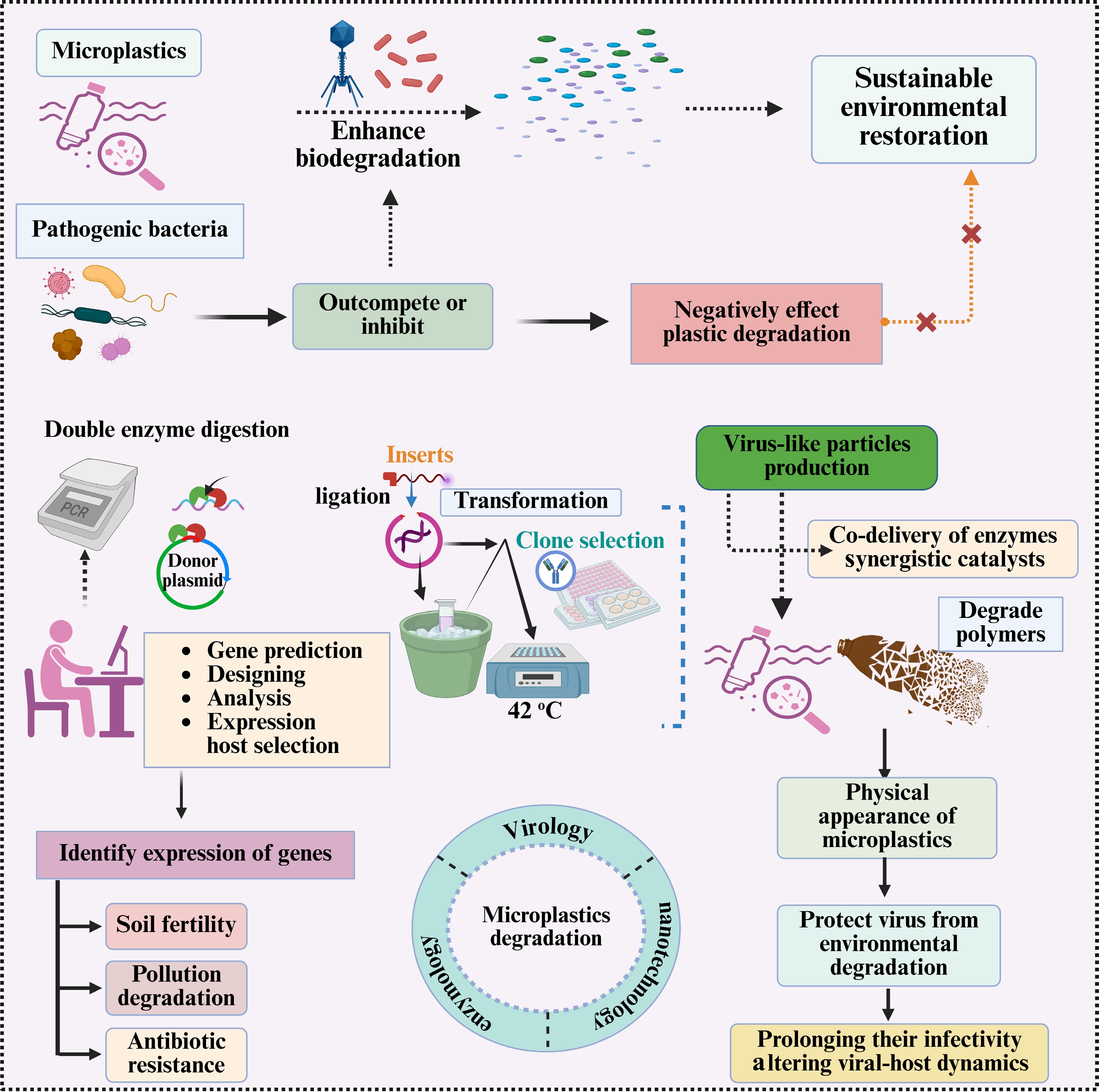
Figure 3.
A multi-step approach combining genetic engineering and viral-like particles to improve microplastic degradation and soil bioremediation. It includes: (1) double enzyme digestion for plasmid preparation, gene prediction, and expression host selection; (2) ligation and transformation for clone selection (42 °C); and (3) virus-like particle production to co-deliver synergistic enzymes while protecting viral infectivity. It also highlights the challenges posed by pathogenic bacteria that may inhibit plastic degradation, as well as strategies to monitor the impacts of gene expression on soil fertility, pollution degradation, and antibiotic resistance.
-
Phage-microbe systems have recently been shown to be capable of enhancing MP pollutant degradation and modulating community structure; however, significant challenges remain before these discoveries can be applied in field-scale, environmentally sustainable applications. Future studies must be cross-disciplinary, moving beyond transient lab studies to investigate the complex, long-term dynamics of soil ecosystems impacted by MPs.
Knowledge gaps
-
There is still a significant knowledge gap regarding long-term field data on the interactions between MPs and viruses, despite growing awareness of the environmental impacts of MP pollution and the significance of viruses, particularly bacteriophages, to the dynamics of microbial communities. Due to this gap, it is challenging to understand how these interactions evolve in the natural world and to develop effective and efficient strategies to mitigate their consequences. The lack of long-term research examining how the MP–virus interaction varies across various ecosystems (soil, freshwater, and marine) over extended periods, spanning multiple seasons and years, is a primary concern.
● The majority of current research relies on laboratory experiments or brief field surveys that capture static snapshots of the phenomenon under study. Thus, detailed information is lacking on how viral populations colonize MP surfaces over time, how viral infection cycles affect the succession and functionality of microbial communities involved in plastic degradation, and how these relationships respond to environmental factors such as temperature, moisture content, and pollution levels.
● In-situ processes and rates of viral-mediated HGT on mobile elements are still poorly understood. Genes that make bacteria resistant to antibiotics, and that degrade plastic are among the genes that viruses are known to facilitate. The effectiveness of MPs as hotspots for this type of gene flow in nature, however, is not well understood. This knowledge is crucial when assessing ecological concerns, such as the spread of resistance genes or virulence factors linked to viral activity on MPs[67,68].
● The impact of viral lysis and lysogeny on microbial metabolism and ecosystem functioning in MP-contaminated environments represents another significant knowledge gap. We must understand the long-term effects of virus-induced cell death or integration on microbial community resilience, nutrient cycling, and pollutant degradation to forecast ecosystem responses and the duration of bioremediation efforts.
● Insufficient research has been carried out on the interactions between MP-associated viruses and chemical pollutants or plastic additives, including their roles as stressors or vectors[69]. This interaction may alter the propagation of viruses, the susceptibility of microorganisms to infection, and the general toxicity of contaminated environments (Table 1).
Table 1. An overview of various viral databases and their potential use in soil microplastic studies
Database Primary focus areas Main features Limitations specific to general virology Soil MP research limitations Ref. NCBI RefSeq Viral
(viral genomes; proteins; transcripts)High-quality reference sequences; Taxonomic breadth; Data standardization; Comprehensive metadata Standardized annotation framework; Integration with NCBI resources; Facilitation of viral research; Cross-disciplinary utility Representation bias; Homology-dependent curation; Limited environmental virome data; Genomic diversity challenges Incomplete coverage of environmental viruses; Difficulty in novel viral discovery; Annotation limitation; Host-assignment complexity [70,71] IMG/VR (Integrated Microbial Genomes/Virus)
(Studying viral genomes and their integration with microbial genomes in various environments)Viral genetic diversity; Host-virus interactions; Functional annotation; Ecosystem-level implications Scalability and dataset size; Integrated analysis tools; Ecological context mapping; Host prediction frameworks; Downloadable and visualizable data; Continuous updates ensure that researchers have access to the latest insights Sampling biases; Error-prone predictions (inaccuracies stemming from incomplete genome assemblies); Data resolution gaps (insufficient in cases where a detailed functional role, life cycle, or ecological impacts of specific virus need further exploration) Underrepresentation of soil viral sequences (lag behind other environments like marine and human gut ecosystems); Challenges in linking the virus to microbial hosts (dense and diverse with intertwined tropic network); Depth and assembly gaps (low-abundance and challenging to resolve due to sequencing depth limitation) [72,73] GVD (Giant virus database) Access to genomics, proteomics, and phylogenetic data of Nucleocytoviricota, which are notable for their genome sizes reaching megabases. (Supporting taxonomic classification and evolutionary studies; Enabling exploration of interactions between giant viruses and their host organisms; Investigating the role in terrestrial and aquatic ecosystems) Advanced data integration (functional dynamics of giant viruses); Search capabilities (access tailored datasets); Visualization tools (improve accessibility and interpretation of data); Global coverage (aquatic systems, forest soils, wastewater, and human-associated habitats); Interoperability with other databases Narrow scope (limits studying RNA viruses, bacteriophages, or other small genome DNA viruses); Limited clinical relevance (i.e., studying human pathogenic viruses or animal-associated viromes); Bias in sampling (difficult to detect viruses in environmental samples); Data completeness Lack of direct linkages (do not explicitly focus on giant virus activity in relation to MP particles); Limited environmental metadata (lack of information specific to soil ecosystems impacted by MPs); Neglect of abiotic factors (integration in hydrophobicity, chemical sorption, and mechanical effects of MPs on microbial communities); Host specificity constraints (giant viruses infect protists and algae while MP-associated microbiomes may involve bacteria, archaea, and other small organisms) [74] ● Methodological limitations also exist, especially with regard to the creation of high-resolution, affordable instruments for simultaneous metagenomic and viromic monitoring in complex field settings, as well as the lack of standardized procedures for data collection, analysis, and interpretation. Comparing studies across ecosystems becomes challenging as a result.
● The lack of standardized, comprehensive, and well-curated viral databases is a primary technical issue in the functional annotation of viral groups. Because of this, it is challenging to precisely identify and describe the functions of viral sequences present in environmental samples (Table 1). The genomes of bacteria and archaea differ greatly from those of viruses. They lack globally conserved marker genes that facilitate organism classification, such as the 16S rRNA gene in bacteria, and frequently have novel genes with uncertain functions. This makes it challenging to infer the function of viral sequences or to attribute them to particular groups with confidence. Although current viral databases, such as GVD (giant virus database), IMG/VR (Integrated Microbial Genomes/Virus), and NCBI RefSeq Viral, are helpful, they do not fully represent the diversity of viruses, particularly those environmental viruses like bacteriophages that are connected to MPs or soil ecosystems.
● There are many viral genes that are either unannotated or insufficiently annotated since many viral sequences obtained from metagenomic and viromic investigations are categorized as 'viral dark matter', meaning they lack homologs in reference databases.
● Furthermore, compared to bacteria, viruses have less sophisticated functional annotation systems. The short and extremely varied sequences of viral genes often make homology-based annotation techniques more challenging. Additionally, the functions of viral proteins have not been experimentally verified, which makes it more difficult to use computational methods to generate precise predictions. Because viruses are known to exhibit high amounts of genomic recombination and mosaicism, tools for assembling and classifying viral genomes struggle. This complicates downstream annotation.
● Additionally, it is challenging to incorporate multi-omics data (such as metagenomics, metatranscriptomics, and proteomics) due to the need for sophisticated bioinformatics methods and computing resources that are currently being created to determine the roles of viruses in the environment.
● To overcome these challenges, we must collaborate to establish community standards for viral functional ontologies, build and manage viral genome databases with environmental isolates, and develop viral-specific annotation methods using structural modeling and machine learning. These kinds of developments are critical to understanding the impact of viruses on ecosystems in MP-contaminated areas and applying them to environmental cleanup and ecosystem restoration.
Emerging tools
-
The way we study and address the connections between viruses, microbes, and MPs is set to alter due to new techniques. We can now analyze these complex systems and develop targeted bioremediation strategies with an unprecedented degree of detail thanks to new technologies like single-cell viromics, AI-driven phage-host prediction, and synthetic virus-like particles (Fig. 4). These cutting-edge technologies must be integrated in order to transform fundamental ecological information into solutions that can be used widely. A few of them are discussed below:
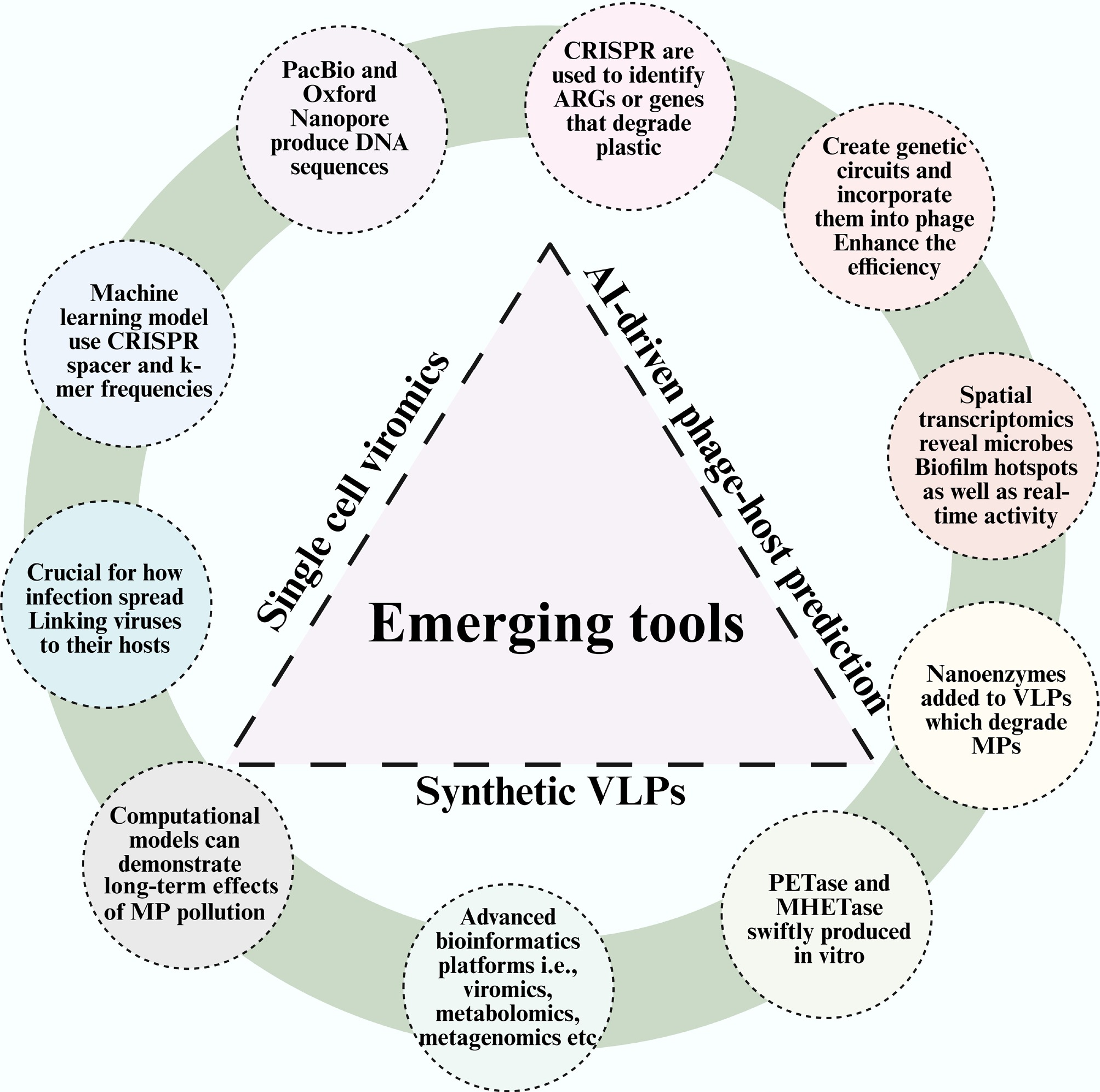
Figure 4.
Schematic representation of various emerging tools that elucidate the interaction of viruses, bacteria, and MPs.
● Viral genomes and their interactions with particular microbial hosts can be sequenced at the single-cell level using environmental samples, primarily due to single-cell viromics. It is crucial for understanding how infections spread without the need to cultivate the viruses and for linking viruses to their hosts in the complex plastisphere.
● CRISPR spacers and k-mer frequencies are examples of genomic information that machine learning models can use to make exact predictions about the interactions between phages and hosts. This bypasses the laborious, antiquated methods of cultivating phages and rapidly identifies the finest ones for targeted biocontrol or enzyme delivery.
● Long, continuous DNA sequences produced by platforms such as PacBio are revolutionary for assembling entire viral genomes from complex metagenomes. This helps fill the gaps in viral databases, and addresses the genetic mosaicism of phages.
● In the field, specific viral sequences, ARGs, or genes that degrade plastic can be identified using engineered CRISPR systems. This is an efficient and affordable method to monitor bioremediation progress and environmental risks.
● It is possible to design genetic circuits and incorporate them into phage genomes to regulate their life cycle, enhance the efficiency of plastic-degrading enzyme delivery, or create environmental sensors that initiate degradation pathways only upon encountering MPs.
● On the surface of an MP particle, spatial transcriptomics and other techniques can reveal the locations of microbial and viral gene expression. This identifies the locations of the biofilm's functional 'hotspots' as well as the real-time activity of the degradation pathways.
● Custom enzyme cocktails, or nanoenzymes, can be loaded into engineered VLPs and directed towards specific kinds of plastic polymers. These stable, non-replicating nanoreactors enable the precise degradation of MPs without the environmental hazards associated with live viruses.
● New plastic-degrading enzymes (like PETase and MHETase) that are more stable, active, and capable of breaking down a greater variety of substrates can be produced rapidly in vitro, and then, these enhanced genes can be transferred using phages or VLPs.
● It is essential to have sophisticated bioinformatics systems that incorporate information from viromics, metabolomics, metagenomics, and metatranscriptomics. With an emphasis on significant regulatory nodes and novel functions, they present the MP-microbe-virus network from a systems-level perspective.
● Prior to application in the field, computer models based on real-world data on viral predation, HGT rates, and microbial metabolism can simulate the long-term environmental impacts of MP contamination and the effectiveness of various virus-assisted remediation techniques.
Call for interdisciplinary collaboration
-
Experts from all disciplines must collaborate to provide comprehensive, practical, and long-lasting solutions to the expanding issue of MP pollution and its complex interactions with viral ecology. Ecological scientists, virologists, and policymakers must collaborate to address the myriad biological, environmental, and sociopolitical facets of virus-MP interactions. To aid in the restoration of ecosystems and the mitigation of pollution, each group contributes its unique perspective and skill set.
These domains must collaborate to generate new ideas to overcome issues, including societal challenges, ecological uncertainty, and technical limitations. For instance, collaboration can help establish monitoring systems that utilize metagenomic, viromic, and environmental health data; standardize procedures for implementing virus-based remediation; and enhance the consistency of risk assessment techniques. Cross-disciplinary research consortia and international networks can exchange information, combine resources, and expedite the process of converting laboratory findings into field applications with broad applicability. Additionally, interdisciplinary cooperation facilitates education and capacity development. It enables policymakers and scientists to collaborate on training programs that equip communities and practitioners with the skills and information necessary to implement virus-assisted remediation safely and successfully. Additionally, this collaboration encourages the integration of local experiences and indigenous knowledge, thereby enhancing scientific methods and highlighting their cultural significance.
-
A better understanding of virus–microorganism interactions is essential for developing cleaner, more adaptive, and environmentally sustainable strategies for managing MP pollution. Phages can alter the composition of microbial communities by lysing specific bacterial species, which regulates the quantity of microorganisms that degrade plastic, as well as those that are harmful or competitive. By acknowledging ecological complexity, emphasizing practical applications, ensuring safety, and promoting cooperative innovation, the scientific community can maximize the potential of microbial-virus networks for comprehensive, effective, and sustainable MP remediation. We must comprehend the 'duality' that these phages produce if we are to maximize their potential in the future. Future research and implementation must be grounded in context-specific risk assessments and strict regulations to strike a balance between efficacy and environmental safety. By maximizing remediation efficiency while maintaining ecological safety, this enables the development of workable, secure, and scalable solutions to the worldwide MP problem.
-
Not applicable.
-
The authors confirm their contributions to the paper as follows: Babar Iqbal: supervision, visualization, and writing – the original draft. Amir Abdullah Khan: software analysis. Jian Hu: investigation, software analysis. Qiang Liu: writing, review & editing. Chen Wang: writing, review & editing. Guanlin Li: conceptualization, investigation, project administration, resources, writing, review & editing. Mao Ye: visualization, writing, review & editing. All authors reviewed the results and approved the final version of the manuscript.
-
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
-
This work was supported by the National Natural Science Foundation of China (Grant No. 32350410400), and the Zhenjiang Municipal Policy Guidance Project of International Science and Technology Cooperation (Grant No. GJ2023005).
-
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
-
Microplastics as abiotic stressors alter the functioning of soil ecosystems.
Bacteriophages are strong but overlooked agents in microplastic-contaminated soils.
VLPs equipped with nanoenzymes enable focused, direct attack on polymer chains and phage-assisted microbial augmentation.
These concepts require a single framework to link lab science with long-term ecosystem recovery.
Viral-mediated HGT can enhance the capacity to degrade plastic and raise the risk of transmitting virulence genes and antibiotic resistance.
-
Full list of author information is available at the end of the article.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Iqbal B, Khan AA, Hu J, Liu Q, Wang C, et al. 2026. Soil microplastics hidden web: interaction of microbes and viruses as a frontier for sustainable ecosystem recovery. Agricultural Ecology and Environment 2: e006 doi: 10.48130/aee-0026-0003
Soil microplastics hidden web: interaction of microbes and viruses as a frontier for sustainable ecosystem recovery
- Received: 12 November 2025
- Revised: 26 December 2025
- Accepted: 15 January 2026
- Published online: 28 February 2026
Abstract: Microplastics (MPs) act as an emerging contaminant in agricultural ecosystems, affecting the health and function of the soil. Their presence can alter soil physicochemical properties, disrupt nutrient cycling processes, and ultimately influence crop productivity and quality. Meanwhile, MPs can also modify the soil microbiome by forming polymer-associated biofilms known as plastispheres, and altering the interactions between various microbial taxa. With a focus on the impact of bacteriophages on microbial communities and traits through processes like lysis, lysogeny, and horizontal gene transfer, this review summarizes the current understanding of soil-specific interactions between microorganisms and viruses. While examining the dual ecological effects of viral horizontal gene transfer on both plastic-degrading capacities, and the spread of antibiotic resistance, possible relationships between phage activity and soil biogeochemical processes are evaluated. Together with carefully planned biotechnological approaches, such as phage-assisted microbial augmentation and the application of virus-like particles, innovative analytical techniques, including metagenomics, viromics, and single-cell methodologies, enable the investigation of new diagnostic methods and potential approaches to support soil recovery under MP stress. In conclusion, a practical framework is proposed that connects diagnostics to management, focusing on continuous monitoring, standardized indicators, and a comprehensive risk assessment. This review demonstrates that MPs serve not only as physical pollutants but also as an abiotic stressor that alter the interactions among microbes, viruses, and soils. This finding indicates that the soil virome plays a significant yet underexplored role in the sustainability of agriculture.