-
The global human population is projected to reach approximately 9.8 billion by 2050, placing unprecedented pressure on agricultural systems to meet the rising demand for food—particularly animal-source products[1]. This demand is driven not only by population growth but also by efforts to address hunger and malnutrition, as well as dietary shifts associated with economic development[2,3]. At the same time, agricultural systems face mounting challenges from climate change, limited natural resources, and the persistent burden of infectious and non-infectious diseases affecting livestock, all of which threaten the sustainability of global food production. These pressures are evident in both developed regions, such as the United States (U.S.)[4,5], and developing regions, including Egypt[6,7], underscoring the need for regionally adaptable strategies to safeguard livestock health and productivity.
Reducing production losses is among the most effective and sustainable approaches to improving food security, often yielding greater benefits than increasing animal numbers alone. Strategies such as enhanced biosecurity, improved food safety, strengthened host disease resistance, and the development of environmentally resilient livestock populations are central to this effort[8−10]. Advances in genomics, precision livestock farming, and systems biology have accelerated data-driven approaches to disease prevention, with phenomics—the large-scale characterization of observable traits—emerging as a key framework for integrating genetic, environmental, and management influences on health and productivity.
Cattle play a central role in global food systems by converting non-human-edible plant materials into nutrient-dense meat and milk, particularly in regions unsuitable for crop production[11]. Beyond food production, cattle support soil fertility, provide draft power and leather, and sustain rural livelihoods. However, cattle are affected by a broad spectrum of infectious, metabolic, nutritional, toxic, and multifactorial diseases that compromise productivity, animal welfare, and food safety. In this review, we synthesize phenomic information for 56 major cattle diseases, integrating data on etiology, clinical features, epidemiology, and genomic architecture to demonstrate how phenomics can unify disease understanding, reveal knowledge gaps, and inform precision management and breeding strategies for sustainable cattle production.
-
The names, etiology groups, causative agents or contributing factors, and major clinical manifestations of the 56 cattle diseases analyzed in this review are summarized in Fig. 1 and Supplementary Table S1. Based on primary causation, these diseases were classified into infectious, nutritional and toxic, multifactorial, and other non-infectious categories. This framework provides a structured approach for organizing disease phenotypes while recognizing that many conditions arise from overlapping interactions among host biology, environment, and management.
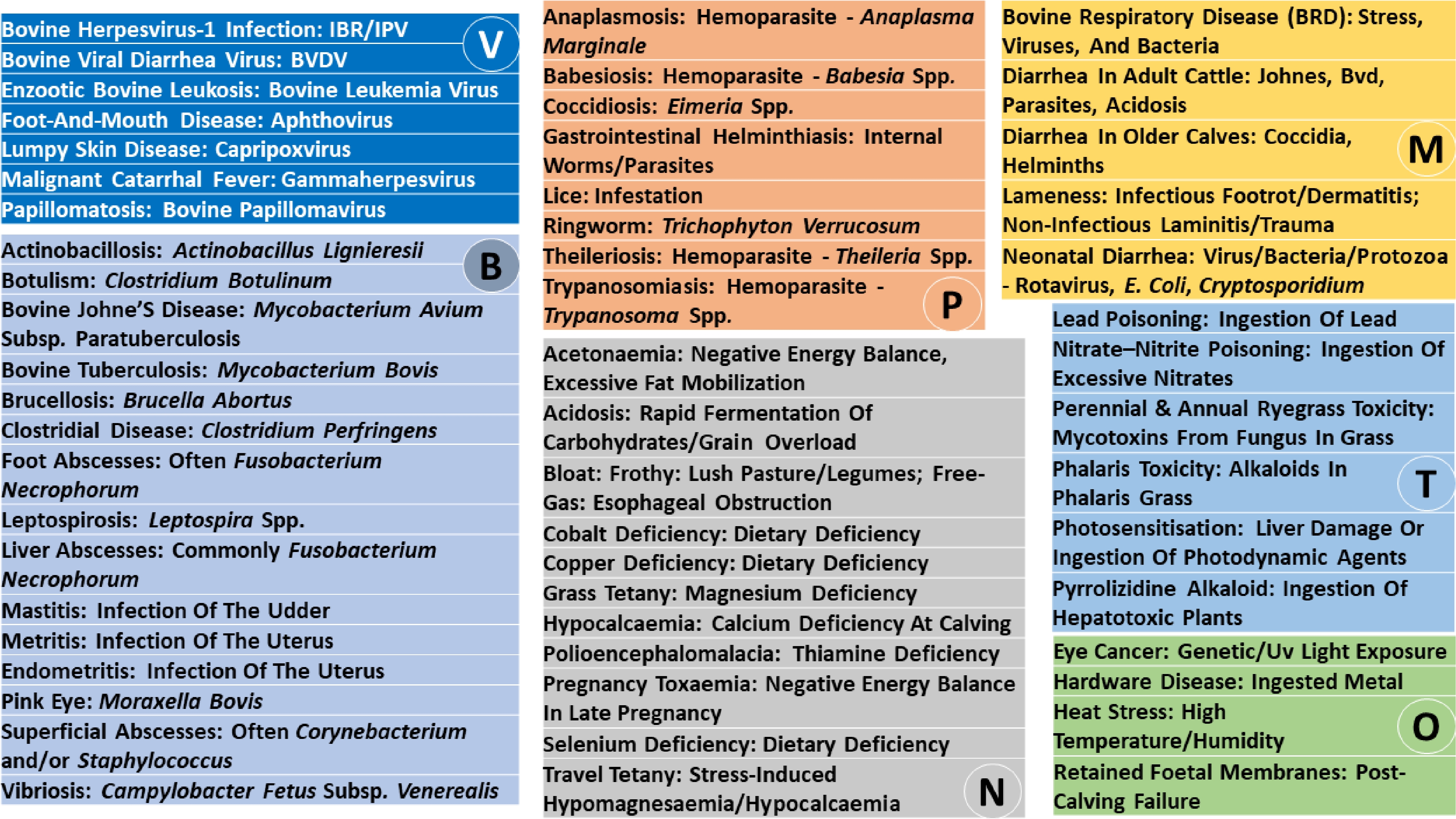
Figure 1.
Cattle diseases and their causative agents or contributing factors. V, viruses; B, bacteria; P, parasites; N, nutrition or metabolic factors; T, toxins; M, multifactorial causes; and O, other factors.
Infectious diseases constitute the largest category and include viral, bacterial, and parasitic conditions with diverse transmission routes, tissue tropisms, and clinical outcomes. Viral diseases range from highly contagious transboundary infections to immunosuppressive and oncogenic agents that predispose cattle to secondary infections and reproductive failure[12,13]. Clinically, they are commonly characterized by systemic febrile illness with respiratory, gastrointestinal, cutaneous, reproductive, or neurologic involvement and may exhibit prolonged subclinical progression in some cases[14−16]. Bacterial diseases span localized, systemic, and zoonotic infections, including chronic enteric and toxin-mediated syndromes often associated with dietary or environmental mismanagement[17−20]. Corresponding clinical manifestations include localized inflammatory lesions, reproductive and enteric disorders, neuromuscular dysfunction, and systemic toxemia, with outcomes ranging from chronic morbidity to acute mortality[14,21]. Parasitic diseases encompass protozoal, helminth, ectoparasitic, and fungal agents, many of which are vector-borne[22−25]. These diseases are dominated clinically by anemia, gastrointestinal dysfunction, dermatologic involvement, impaired growth, and, in severe cases, hemolysis, hypoxia, or death[14,26].
Non-infectious diseases reflect disruptions in nutrition, metabolism, and environmental exposure, as well as complex interactions among host physiology, management, and stressors. Nutritional and metabolic disorders arise from imbalances between nutrient supply and physiological demand, particularly during growth, lactation, and parturition, and typically manifest as reduced productivity, weakness, recumbency, bloat, or neurologic dysfunction associated with deficiencies in energy, minerals, or vitamins[14,27−30]. Toxic diseases highlight risks associated with contaminated feed, water, and pasture, and are often acute and severe, with clinical presentations determined by the specific agent and commonly involving neurologic, respiratory, hepatic, or dermatologic pathology[31−34]. Multifactorial syndromes, including respiratory disease complexes, enteric disorders, and lameness, emerge from interacting infectious and non-infectious drivers and exhibit overlapping clinical features that are frequently amplified by stress, nutritional status, and co-infection[14,35−40]. Other non-infectious conditions emphasize the roles of environmental stress, mechanical injury, ultraviolet exposure, reproductive physiology, and management practices in shaping disease risk and expression, underscoring the diversity of non-pathogen drivers influencing cattle health outcomes[14,21,41,42].
Diagnostic approaches and phenotypic resolution
-
The diagnostic strategies applied to the 56 cattle diseases analyzed in this review are summarized in Supplementary Table S1. Across disease classes, infectious diseases rely primarily on pathogen detection, serologic evidence of exposure, and lesion-based assessment. Viral diseases are typically diagnosed using molecular assays, antigen or antibody detection, and recognition of characteristic clinical or histopathologic lesions, with approaches selected based on disease stage and transmission risk[14,43,44]. Bacterial diseases are identified through combinations of clinical evaluation, culture-based or serologic testing, and targeted imaging or immunologic assays for systemic or deep-seated infections, particularly those of zoonotic or chronic importance[14,45]. Parasitic diseases are most commonly diagnosed using microscopic or quantitative parasitologic methods, supplemented by molecular or immunologic assays to detect subclinical or carrier states in endemic settings[14,26].
Nutritional and toxic diseases are primarily diagnosed through integration of biochemical testing, dietary or exposure history, and clinical response to intervention. Nutritional disorders are assessed using blood, milk, urine, or tissue biomarkers that reflect energy balance, mineral or vitamin status, and rumen function, often supported by characteristic therapeutic responses or post-mortem findings[14]. Toxic diseases depend on confirmation of exposure through laboratory detection of toxicants or tissue injury patterns, combined with clinical presentation and environmental assessment, highlighting the importance of feed, pasture, and water surveillance in diagnosis and prevention[31,32].
Multifactorial and other non-infectious diseases require integrative diagnostic frameworks that combine clinical scoring, laboratory testing, imaging, and management context. Multifactorial syndromes, including respiratory, enteric, and locomotor disorders, are diagnosed using composite approaches that incorporate pathogen detection, physiologic assessment, and post-mortem evaluation when necessary[14,40]. Other non-infectious conditions rely largely on physical examination and targeted diagnostic tools tailored to mechanical, thermal, reproductive, or neoplastic processes, underscoring the central role of environment and management in disease identification and phenotypic resolution[14,42].
Economic consequences of cattle diseases
-
The economic impacts associated with the 56 cattle diseases reviewed here are summarized in Supplementary Table S1. Across disease classes, infectious diseases impose substantial losses through mortality, reduced productivity, reproductive failure, treatment and control costs, and market or trade restrictions. Viral diseases contribute to production losses through abortion, infertility, reduced growth and milk yield, increased culling, and hide damage, with transboundary infections capable of triggering mass depopulation and severe trade disruptions[14,46,47]. Bacterial diseases further amplify economic burden via chronic morbidity, milk discard, impaired feed efficiency, infertility, premature culling, carcass condemnation, and regulatory constraints, particularly for zoonotic or notifiable diseases[14,48]. Parasitic diseases generate persistent losses due to anemia, mortality, reduced growth and production, reproductive failure, and ongoing costs of treatment and control, especially in endemic regions[14,49].
Nutritional and toxic diseases primarily affect productivity, survival, and reproductive efficiency, often resulting in rapid or irreversible economic losses. Nutritional disorders reduce milk yield, growth, and fertility, increase involuntary culling and mortality, and impose substantial veterinary and management costs[14,50]. Toxic diseases are frequently associated with acute deaths, chronic ill-thrift, neurologic or hepatic damage, hide and carcass losses, food safety concerns, and loss of market access, emphasizing the economic vulnerability of cattle systems to feed, pasture, and water contamination[31,32].
Multifactorial and other non-infectious diseases contribute to cumulative economic impacts through sustained inefficiencies and increased management demands. Multifactorial syndromes, including respiratory disease complexes, enteric disorders, lameness, and calf diarrhea, are associated with high morbidity and mortality, reduced growth and milk production, impaired fertility, and elevated treatment and labor costs[14,40]. Other non-infectious conditions impose losses through premature culling, extended days open, carcass condemnation, chronic productivity decline, and mortality, illustrating how environmental, mechanical, and management-related risks translate directly into economic burden in cattle populations[14,42].
-
Across cattle diseases, the most commonly affected tissue systems include the gastrointestinal tract (rumen, intestines, liver), respiratory system (upper airways and lungs), integumentary system (skin, hooves, hair), reproductive tract (uterus, placenta, fetus, udder), nervous system, hematopoietic/blood system, and lymphoid tissues (Fig. 2, Supplementary Table S2). Infectious agents frequently target epithelial surfaces and immune tissues as portals of entry and replication, whereas nutritional and toxic disorders disproportionately affect metabolically active organs such as the liver, brain, muscle, and rumen, reflecting systemic metabolic disruption[14,15].
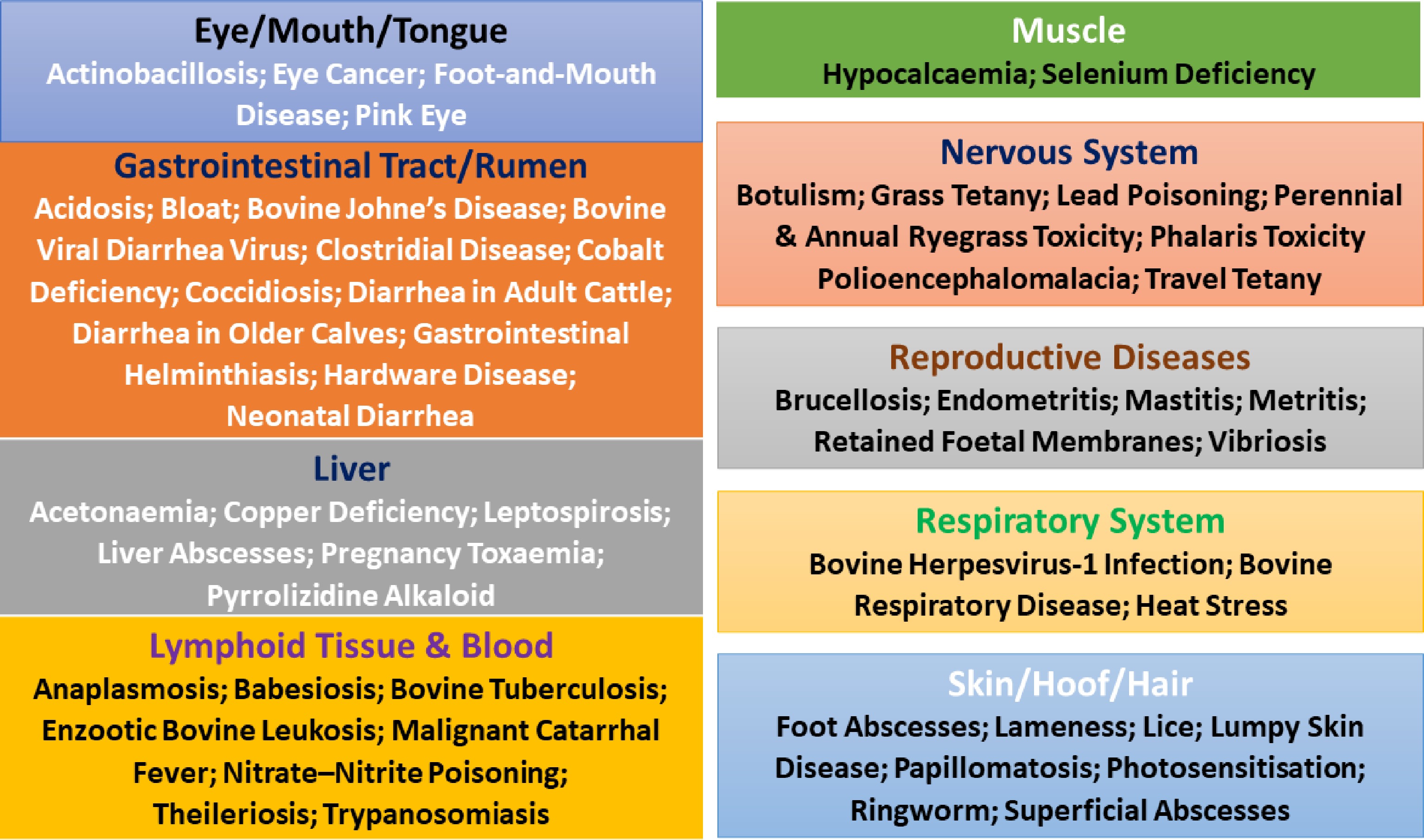
Figure 2.
Cattle diseases and primary target tissues. Cattle diseases often exhibit strong tissue specificity, with distinct disorders predominantly affecting particular organs or biological systems.
Diseases may involve single dominant tissues or multiple organ systems. Single-tissue diseases include papillomatosis and ringworm (skin), mastitis (mammary gland), pink eye (cornea and conjunctiva), and ocular squamous cell carcinoma (ocular epithelium). In contrast, multisystem diseases are common in cattle, such as bovine viral diarrhea virus (intestinal epithelium and lymphoid tissue), enzootic bovine leukosis (lymph nodes, heart, abomasum, uterus), malignant catarrhal fever (blood vessels and mucous membranes), tuberculosis (lungs, lymph nodes, liver, spleen), and leptospirosis (kidneys and liver), where systemic spread or immune-mediated injury drives widespread tissue damage[14,45].
Several key tissues are repeatedly targeted by different diseases, regardless of etiology. The intestinal mucosa is affected by viral (BVDV), bacterial (Johne's disease), parasitic (coccidiosis, helminths), nutritional (acidosis), and multifactorial diarrheal syndromes, making it a central site of disease convergence[14,15]. The liver is a shared target in nutritional (acetonaemia, pregnancy toxaemia, copper deficiency), toxic (pyrrolizidine alkaloids, photosensitisation), bacterial (liver abscesses, leptospirosis), and metabolic disorders, reflecting its role in detoxification and metabolism[14,31]. Similarly, the nervous system is affected by nutritional (grass tetany, polioencephalomalacia), toxic (lead, ryegrass, phalaris), and metabolic stress–related conditions, often manifesting as tremors, blindness, or paralysis[14,30].
When grouped by disease category, clear tissue patterns emerge. Infectious diseases (viral, bacterial, parasitic) primarily affect epithelial surfaces, lymphoid tissues, blood cells, lungs, intestines, skin, reproductive organs, and eyes, often with systemic dissemination[14,15,45]. Nutritional and toxic diseases preferentially involve metabolically sensitive tissues, especially the liver, brain, muscles, rumen, and blood, with lesions reflecting deficiencies, imbalances, or toxic injury rather than direct pathogen damage[14,30,31]. Multifactorial diseases concentrate on functional systems under stress, such as the lungs (bovine respiratory disease), intestines (diarrhea complexes), and locomotor apparatus (lameness), where environment, nutrition, and infection interact[14,40].
Finally, other non-infectious conditions highlight mechanical, environmental, and physiological vulnerabilities in cattle. Hardware disease affects the reticulum, diaphragm, and pericardium, heat stress disrupts the hypothalamus, cardiovascular, and respiratory systems, and retained fetal membranes involve persistent attachment of the uterus and placenta, emphasizing how non-pathogen factors can still cause severe tissue-specific pathology with systemic consequences[14,42].
Age and physiological stage dependence
-
Cattle diseases show clear clustering by physiological age (Fig. 3, Supplementary Table S3). Newborn calves (0–30 d) are predominantly affected by neonatal diarrhea, early-life viral respiratory infections (e.g., BHV-1 in maternally naïve calves), and nutritional deficiencies such as selenium deficiency (white muscle disease), reflecting immature immunity and colostrum dependence[51,52]. Young calves (1–6 months) are highly susceptible to enteric and skin diseases, including Coccidiosis, Ringworm, Pink Eye, Lead Poisoning, and early parasitic infections, as well as nutritional disorders such as cobalt deficiency; this period coincides with rumen development and management transitions[37,51,52].
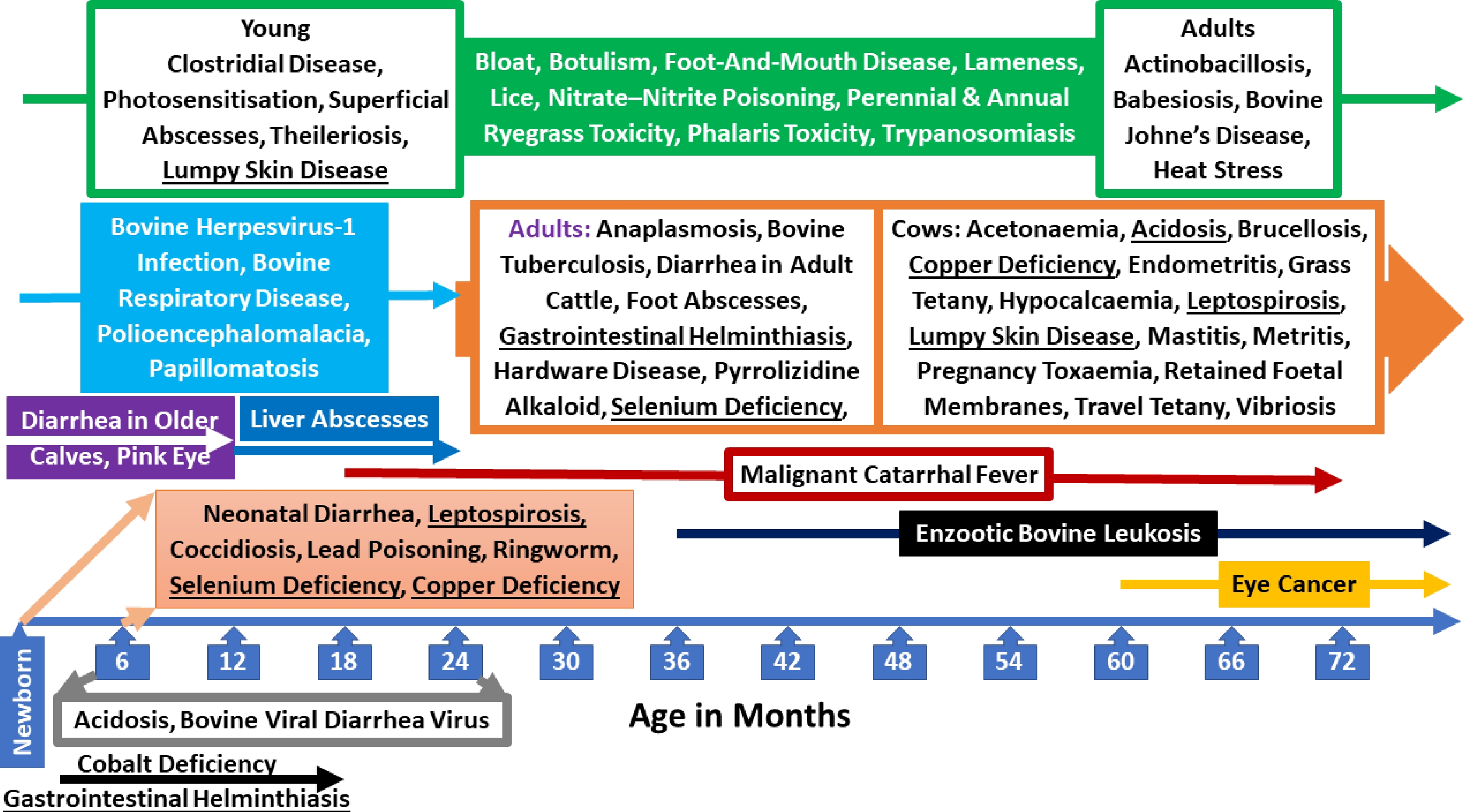
Figure 3.
Cattle diseases and age dependence. Cattle diseases show distinct age-related patterns. Diseases affecting all age groups are shown in green, while those occurring at different physiological stages or in different animal types are underlined.
Older calves and juveniles (6–18 months), particularly post-weaning and feedlot cattle experience a high incidence of bovine respiratory disease, Papillomatosis, gastrointestinal helminthiasis, Polioencephalomalacia, liver abscesses, and acidosis, driven by stress, dietary change, and commingling[52,53]. Mature adults (> 2–3 years) are more commonly affected by chronic and cumulative-exposure diseases such as enzootic bovine leukosis, Johne's disease (clinical signs typically ≥ 3 years), bovine tuberculosis, anaplasmosis, eye cancer, hardware disease, and pyrrolizidine alkaloid toxicity, reflecting long incubation periods and bioaccumulation[37,54].
Cows, especially during late gestation and early lactation, exhibit a distinct disease profile. Periparturient cows (−3 weeks to +6 weeks relative to calving) are at highest risk for hypocalcaemia (0–3 d post-calving), acetonaemia (2–6 weeks postpartum), metritis, endometritis, mastitis, retained fetal membranes, and pregnancy toxaemia, due to negative energy balance, immune suppression, and reproductive tract exposure[52,55]. Grass tetany typically affects older lactating cows, while travel tetany and heat stress disproportionately impact adult, high-producing cows under environmental or transport stress[37,55].
Across categories, infectious diseases show strong age dependence: acute viral, parasitic, and enteric diseases dominate in calves and juveniles, whereas chronic bacterial and vector-borne infections predominate in adults[51,53,54]. Nutritional and toxic diseases cluster around growth (calves), diet transitions (feedlot cattle), and metabolic stress (periparturient cows), while many toxicities affect all ages, but cause more severe outcomes in young or stressed animals[37]. Multifactorial diseases such as BRD, diarrheal syndromes, and lameness span all ages but peak during management stress points, underscoring the value of age-stratified phenomics for prevention and precision livestock management[52,53,55].
Seasonal variation across production environments
-
We used information for the U.S. and Egypt to compare the seasonality patterns (Fig. 4, Supplementary Table 4). Interestingly, many cattle diseases show broadly similar seasonal tendencies, particularly those driven by temperature, moisture, and management cycles. Respiratory diseases (e.g., Bovine Herpesvirus-1, Bovine Respiratory Disease) peak in winter or winter–spring in both countries, reflecting cold stress, crowding, and reduced ventilation during housing or confinement periods[14,51]. Likewise, vector-borne diseases (e.g., anaplasmosis, babesiosis, pink eye, lumpy skin disease) consistently peak in summer or summer–autumn, corresponding to heightened activity of flies and ticks in warm conditions in both regions[56,57]. Chronic infections such as enzootic bovine leukosis, Johne's disease, tuberculosis, and venereal diseases show year-round occurrence in both countries, reflecting persistent transmission, independent of season[58].
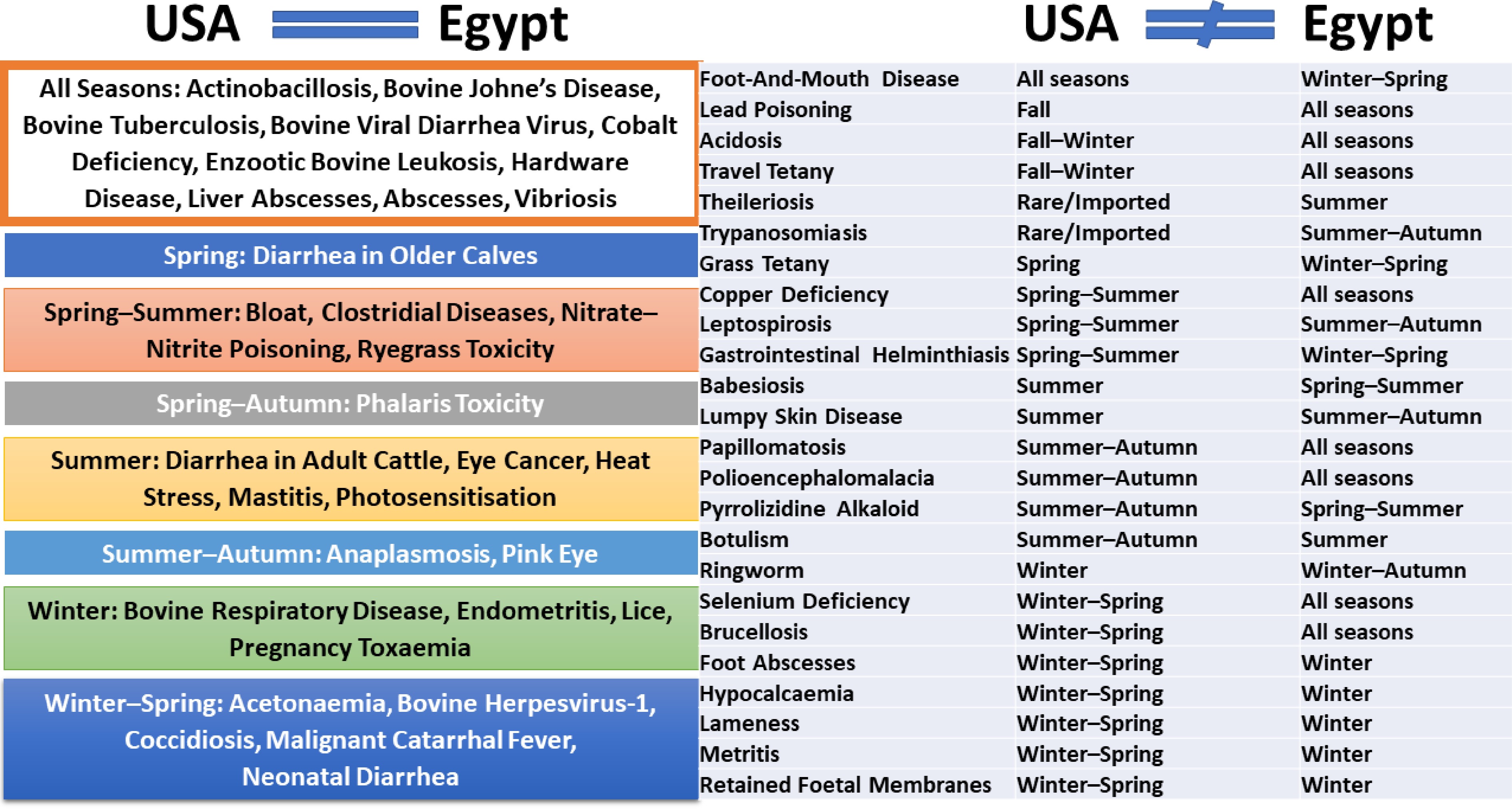
Figure 4.
Cattle diseases and seasonality dependence. Cattle diseases exhibit seasonal variation that may be consistent or differ across geographic regions. Disease seasonality is compared between the U.S. and Egypt.
Despite these similarities, climatic contrasts between temperate U.S. regions and Egypt's arid to semi-arid environment generate important differences. In Egypt, diseases linked to heat, solar radiation, and vectors (e.g., mastitis, photosensitisation, eye cancer, heat stress) show stronger and more prolonged summer peaks than typically observed in the U.S., where summer disease pressure is often regionally limited[56,59]. Conversely, winter-associated diseases (e.g., lice, ringworm, foot abscesses, lameness) are generally more pronounced in the U.S., where prolonged cold, snow, and housing increase moisture and animal contact, while Egypt's milder winters reduce the duration and intensity of these risks[14,57].
Differences in production systems further shape disease seasonality. In the U.S., intensive feedlot systems amplify fall–winter peaks of acidosis, liver abscesses, and BRD due to diet transitions, transport, and commingling stress[15,51]. In Egypt, cattle are more commonly raised in smallholder or mixed crop–livestock systems, where year-round exposure to variable feed quality contributes to more uniform seasonal patterns for nutritional deficiencies (e.g., cobalt, copper, selenium) and metabolic diseases, with sharper peaks around winter calving and early lactation. Pasture-associated toxicities and mineral imbalances also differ, reflecting variation in forage species, irrigation, and fertilization practices[60].
Social and husbandry practices also contribute to seasonal differences. In Egypt, synchronized calving during cooler months, communal grazing, and limited biosecurity increase winter–spring peaks for neonatal diarrhea, metritis, retained fetal membranes, and brucellosis[59,60]. In contrast, U.S. systems often distribute calving across seasons, depending on beef vs. dairy operations, leading to more regionally variable seasonality for reproductive and neonatal diseases[14,15]. Vector control intensity, housing design, and access to veterinary services further modulate disease expression between the two countries[56,57].
In summary, common seasonal drivers—temperature, moisture, vectors, and physiological stress—produce broadly similar disease patterns in U.S. and Egyptian cattle, while differences in climate severity, production intensity, and social management explain divergent seasonal peaks. These contrasts underscore the importance of region-specific, age- and season-stratified phenomics approaches, as disease risk cannot be extrapolated directly across production systems or climates without accounting for environmental and socio-management context[15,51,59].
Prevalence patterns and geographic contrasts
-
The reported prevalence rates of cattle diseases were used to evaluate similarities and contrasts between the U.S. and Egypt (Fig. 5, Supplementary Table S5). Across both countries, infectious diseases dominate overall prevalence, particularly viral and bacterial infections characterized by efficient transmission or prolonged subclinical phases. Viral diseases such as bovine viral diarrhea virus (BVDV), bovine herpesvirus-1 (BHV-1), and enzootic bovine leukosis consistently show high prevalence (often > 40%) in both countries, reflecting persistent infection, imperfect biosecurity, and incomplete vaccination coverage, with subclinical carriers playing a central epidemiological role[14,15]. In contrast, a substantial group of conditions—largely toxic, plant-associated, or rare multifactorial disorders—remain sporadic in both countries, including malignant catarrhal fever, botulism, lead poisoning, nitrate–nitrite poisoning, ryegrass and phalaris toxicities, photosensitization, and pyrrolizidine alkaloid toxicity. Their sporadic occurrence supports the interpretation that these diseases arise primarily from point-source exposure or unusual management failures rather than sustained transmission cycles[14].
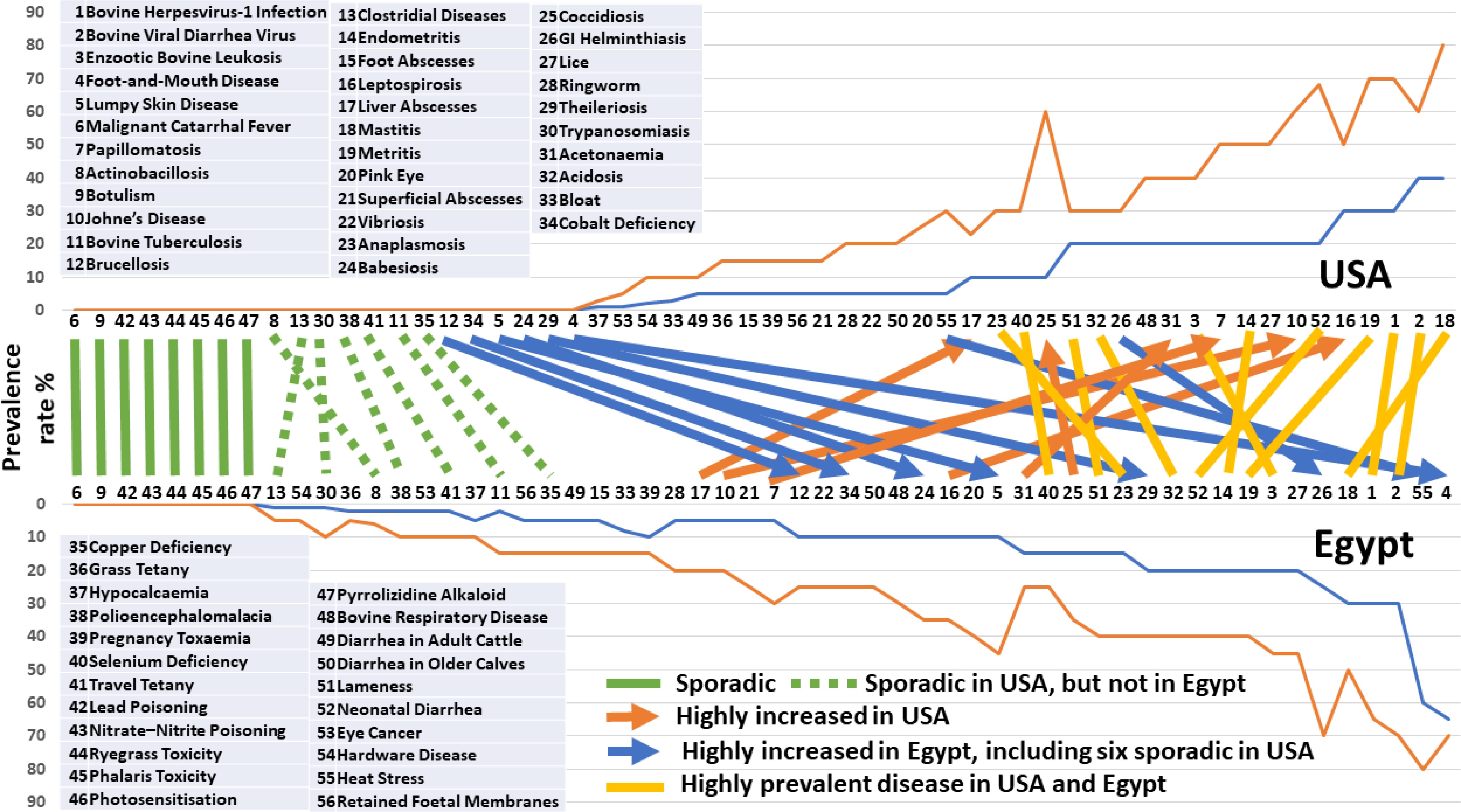
Figure 5.
Cattle diseases and prevalence rate. Disease prevalence rates were compared between the U.S. and Egypt.
Parasitic and vector-borne diseases show a strong geographic gradient, with consistently higher prevalence in Egypt. Gastrointestinal helminthiasis, anaplasmosis, babesiosis, theileriosis, lumpy skin disease, and foot-and-mouth disease frequently reach moderate-to-high prevalence or outbreak levels in Egypt, whereas many remain sporadic in the U.S. This pattern reflects climatic conditions favorable to arthropod vectors, year-round environmental exposure, limited vector control, and challenges in vaccination coverage and animal movement regulation in Egyptian production systems[26,60]. These contrasts underscore the dominant role of ecology and vector pressure rather than host susceptibility alone.
Nutritional and metabolic diseases exhibit moderate-to-high subclinical prevalence in both countries, but are strongly shaped by production intensity, diet formulation, and physiological stress. Conditions such as acidosis, ketosis, selenium deficiency, lameness, mastitis, endometritis, metritis, and neonatal diarrhea are common across systems, reflecting universal biological vulnerabilities associated with lactation, parturition, housing, early-life immunity, and metabolic demand[14,61]. However, certain mineral deficiencies (notably cobalt and copper) appear sporadic in the U.S. yet reach low-to-moderate prevalence in Egypt, consistent with differences in forage composition, ration balancing, mineral supplementation, and water sanitation in smallholder systems[60−62].
Several diseases occur at substantially higher prevalence in the U.S., notably Johne's disease (20%–68% vs. 5%–20%), leptospirosis (30%–70% vs. 10%–35%), papillomatosis (20%–50% vs. 5%–30%), liver abscesses (10%–30% vs. 5%–20%), and subclinical metabolic disorders such as ketosis and acidosis. These patterns are closely associated with high-density dairy and feedlot systems, longer productive lifespans, intensive feeding strategies based on high-energy diets, and enhanced diagnostic surveillance that increases detection of subclinical disease[15,51,61]. For example, liver abscesses and ruminal acidosis are tightly linked to grain-based finishing systems, while Johne's disease accumulates over time in large, relatively closed herds, making prevalence strongly age- and management-dependent rather than environmentally driven[15].
Overall, the contrast between the U.S. and Egypt illustrates how production intensity amplifies chronic and subclinical disease, whereas climate and vector ecology amplify infectious and parasitic burdens. Diseases that remain sporadic in both countries highlight the limited role of endemic transmission, while shared high-prevalence conditions underscore common physiological constraints of cattle production. Together, these complementary patterns support the value of comparative phenomics to disentangle host genetics, environment, and management effects on disease prevalence, and to guide context-specific interventions rather than one-size-fits-all control strategies[15,26,51].
-
In this review, we used the CattleQTLdb as the primary source of information[63] to summarize QTL mapping and association studies for 56 major cattle diseases. Although substantial evidence indicates that host genetics plays an important role in disease susceptibility in cattle, the majority of reported diseases still lack disease-specific QTL annotations in current databases. Among the 56 diseases examined, only 16 have documented genomic studies to date (Supplementary Table S6), highlighting a pronounced imbalance in genetic research coverage across disease traits.
When stratified by disease prevalence in the U.S., a clear trend emerges. Among diseases with sporadic occurrence, only 2 of 21 (10%)—foot-and-mouth disease and bovine tuberculosis—have been subjected to QTL mapping. In contrast, diseases with higher prevalence show progressively greater representation in genomic studies: 1 of 5 (20%) for prevalence rates of 1%–10%, 3 of 10 (30%) for 5%–30%, 1 of 4 (25%) for 10%–60%, 6 of 11 (55%) for 20%–70%, and 3 of 5 (60%) for 30%–80% (Fig. 6). These patterns suggest that diseases with higher prevalence are more likely to attract sustained scientific and public attention, thereby motivating investment in genomic research. Conversely, the low and unpredictable prevalence of many diseases limits the availability of large, well-phenotyped populations required for statistically robust QTL mapping[56,64].
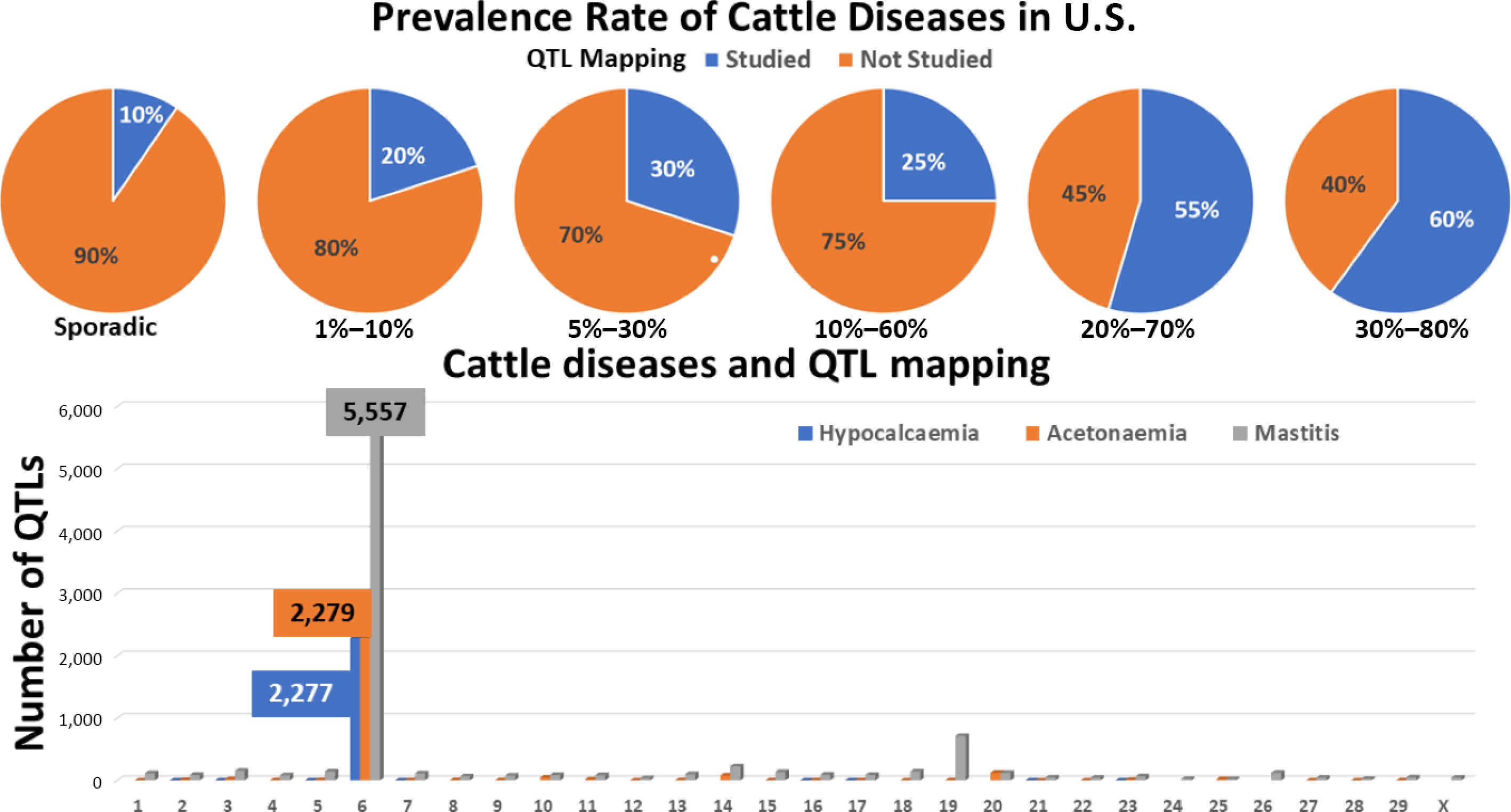
Figure 6.
Cattle diseases and QTL mapping. QTL mapping activity is enriched for diseases with high prevalence rates. For three diseases, QTL loci show pronounced enrichment on chromosome 6.
Beyond prevalence, several practical and conceptual challenges further constrain QTL studies of cattle diseases[63,65]. Foremost among these is phenotype definition. Many diseases are recorded as binary clinical outcomes or broad syndromic diagnoses, such as respiratory disease complex or enteric disorders, which obscure underlying biological heterogeneity. Such coarse phenotypes reduce estimated heritability and dilute genetic signals, making QTL detection inefficient or unreliable[66,67]. In addition, environmental and management factors—including housing, nutrition, hygiene, climate, and veterinary interventions—often dominate disease expression. These influences can overwhelm host genetic effects, particularly for infectious and metabolic diseases, and substantially reduce statistical power in the absence of careful experimental or analytical control[65,68].
Ethical and logistical constraints further limit progress. Unlike laboratory species, controlled pathogen challenge experiments in cattle are highly restricted, costly, and often ethically unacceptable, greatly reducing opportunities for precise dissection of genetic resistance mechanisms[65]. Finally, it is important to note that the absence of disease-specific QTL annotations does not necessarily imply a lack of genetic investigation. For several diseases, genetic associations have been reported for correlated or proxy traits—such as immune response, antibody titers, somatic cell count, or survival—rather than for disease incidence itself. Because these traits are not always curated as disease phenotypes, relevant QTLs may fall outside the scope of disease-centered annotations in current QTL databases[63,69].
Diseases with intensive QTL mapping
-
In contrast to diseases lacking genomic evidence, a subset of cattle diseases has been subjected to intensive QTL mapping, in some cases yielding hundreds or even thousands of loci. Among the 16 cattle diseases for which QTLs have been identified, nine are associated with fewer than 100 loci, whereas the remaining traits show substantially larger numbers, ranging from 216 loci for retained fetal membranes to 8,735 loci for mastitis (Supplementary Table S6). This wide variation reflects both biological complexity and differences in research intensity across traits.
For several extensively studied diseases—including bovine tuberculosis, bovine respiratory disease, Johne's disease, and mastitis—QTLs have been identified on all autosomes plus cases on the X chromosome. To date, no QTLs have been reported on the Y chromosome for any cattle disease, consistent with its limited gene content and low representation in genomic studies. Among autosomes, chromosome 6 stands out as a major hotspot, harboring 2,277 QTLs for hypocalcaemia (milk fever), 2,279 for acetonaemia (ketosis), and 5,557 for mastitis. As illustrated in Fig. 6, QTLs for hypocalcaemia are concentrated in two major regions, those for acetonaemia are distributed across a few additional loci, and mastitis-associated QTLs are widely dispersed across the chromosome, indicating differing genetic architectures among these traits.
Diseases that have attracted intensive genomic investigation share several defining characteristics that make them particularly amenable to genetic analysis[63,70]. Foremost among these is economic importance. Conditions such as mastitis, reproductive disorders, and metabolic diseases impose substantial financial losses through reduced productivity, increased veterinary costs, and premature culling. Their direct economic impact has driven sustained investment in large-scale genomic studies[71].
In addition, heavily studied diseases often have well-defined and routinely recorded proxy phenotypes. Somatic cell count, for example, serves as a quantitative and heritable indicator of mastitis susceptibility, and has enabled high-powered GWAS across multiple breeds and production systems[72]. Similarly, fertility, survival, and longevity traits are commonly used as indirect measures of disease resilience, further expanding the scope of genetic analyses.
These diseases also tend to exhibit moderate heritability, allowing detectable genetic variation despite strong environmental influences. Although individual loci generally have small effects, the availability of large sample sizes and repeated phenotypic records facilitates the identification of numerous QTLs with cumulative relevance[70,73]. Moreover, diseases with extensive QTL mapping frequently display strong pleiotropic relationships with production traits. Many loci influence both health and performance, contributing to inflated QTL counts while simultaneously complicating biological interpretation[74].
Finally, intensive investigation across breeds, environments, and analytical frameworks has led to repeated detection of partially overlapping loci. Consequently, exceptionally high QTL counts reflect not only the underlying biological complexity of these diseases, but also the cumulative effects of research intensity and data aggregation across studies and databases[63,75].
Future directions: from phenomics to precision health
-
Future progress in understanding genetic control of cattle diseases requires rethinking trait definitions, study designs, and data integration strategies, rather than relying solely on traditional disease phenotypes[64,67].
First, emphasis should shift toward intermediate and biologically informative phenotypes, such as immune response traits, pathogen load, inflammatory markers, or time-to-infection measures. These traits are often more heritable and genetically tractable than binary disease outcomes[66,76].
Second, greater use of longitudinal and precision phenotyping is needed. Repeated measurements across life stages can capture disease resilience, recovery, and tolerance, providing richer genetic signals than single clinical diagnoses[68,77].
Third, integrating multi-omics data—including genomics, transcriptomics, epigenomics, and microbiome profiles—can help disentangle host–pathogen–environment interactions and identify regulatory mechanisms underlying disease susceptibility[78].
Fourth, improved trait harmonization and ontology mapping across studies would facilitate more accurate annotation of disease-related QTLs in public databases. This would help bridge the gap between published genetic evidence and curated resources[63,69].
Finally, genomic selection frameworks that incorporate disease resistance as part of overall resilience indices may offer a practical path forward, even when individual disease QTLs are small or context-dependent[73,79].
-
Cattle diseases represent a complex and interconnected phenotypic landscape shaped by host biology, environment, management, and genetics. By synthesizing phenomic information across 56 major diseases, this review highlights recurring principles that transcend individual etiologies. Diseases cluster around a limited number of vulnerable tissues—particularly the gastrointestinal tract, liver, respiratory system, skin, reproductive organs, and blood—reflecting shared physiological bottlenecks in metabolism, immunity, and barrier function. Likewise, disease burden is strongly structured by age and physiological state, with early life, dietary transitions, and the periparturient period emerging as critical windows of susceptibility.
Comparative analyses between the U.S. and Egypt reveal that while many diseases follow broadly similar seasonal patterns driven by temperature, moisture, vector activity, and management stress, the magnitude and timing of disease peaks are strongly modulated by climate severity, production intensity, and husbandry practices. These findings emphasize that disease risk cannot be extrapolated across regions without accounting for environmental and socio-management context, reinforcing the need for region-specific phenomic frameworks.
From a genomic perspective, our analysis demonstrates a pronounced imbalance in research coverage. Diseases with high prevalence and clear economic impact have attracted intensive QTL mapping, often revealing thousands of loci and shared genomic hotspots, particularly on chromosome 6. In contrast, the majority of cattle diseases remain genomically undercharacterized, largely due to challenges in phenotype definition, environmental confounding, ethical constraints on experimental infection, and limited availability of large, well-phenotyped populations. Importantly, the absence of disease-specific QTLs does not imply a lack of genetic control but rather reflects the limitations of traditional binary disease phenotypes.
Looking forward, progress in cattle disease genomics will depend on a shift toward more informative phenotypes, including intermediate, longitudinal, and resilience-related traits, combined with precision phenotyping and multi-omics integration. Harmonization of disease definitions and improved annotation of health-related traits in public databases will be essential to translate existing genetic evidence into actionable knowledge. Ultimately, embedding phenomics within genomic selection and precision livestock management offers a practical pathway to reduce disease burden, enhance resilience, and improve the sustainability of cattle production systems under growing global demand and environmental change.
This work is supported by: (1) the Organic Agriculture Research and Extension Initiative, project award no. 2022-51300-38058, and the Foundational and Applied Science Program/Animal Breeding, Genetics and Genomics, project award nos 2023-67015-39566 and 2023-67015-40080 from the U.S. Department of Agriculture's National Institute of Food and Agriculture; and (2) the Fulbright Commission Egypt to ZJ. Search engines and AI-based tools (including ChatGPT) were used solely to assist with data searching, collection, and enrichment under the author's instruction. The author takes full responsibility for the content of this work. Any opinions, findings, conclusions, or recommendations expressed in this publication are those of the author(s) and should not be construed to represent any official USDA or U.S. Government determination or policy.
-
Not applicable.
-
The authors confirm their contributions to the paper as follows: study conception and design, writing − original draft: Jiang Z; data collection: Jiang Z, Ibrahim A, Sallam A, AbdelMaksoud A, Abouelhag H, Hakim A; analysis and interpretation of results: Jiang Z, Michal JJ, Sharma A, Kasimanickam RK, Wang Y, Gao Y, Fragomeni B, Miller LC; writing − review: Jiang Z, Effat M, Hassan M, Sayme S. All authors reviewed the results and approved the final version of the manuscript.
-
This is a review and all information here are from published literature.
-
The authors declare that they have no conflict of interest.
- Supplementary Table S1 Cattle disease data summary.
- Supplementary Table S2 Tissue variation.
- Supplementary Table S3 Age variation.
- Supplementary Table S4 Seasonality variation.
- Supplementary Table S5 Prevalence variation.
- Supplementary Table S6 QTL mapping.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of Nanjing Agricultural University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Jiang Z, Michal JJ, Sharma A, Kasimanickam RK, Wang Y, et al. 2026. The phenomics landscape of major diseases in cattle – a comprehensive review. Animal Advances 3: e015 doi: 10.48130/animadv-0026-0008
The phenomics landscape of major diseases in cattle – a comprehensive review
- Received: 14 February 2026
- Revised: 14 February 2026
- Accepted: 10 March 2026
- Published online: 06 May 2026
Abstract: Cattle convert non-human-edible plant materials into nutrient-dense meat and milk, support soil fertility through manure, provide leather and draft power, and sustain rural livelihoods, making them central to global food systems and well positioned to meet the rising demand for animal-derived protein as the human population approaches 9.8 billion by 2050. However, cattle diseases frequently compromise productivity, animal welfare, and food security, causing substantial economic losses. Here, we synthesize and characterize phenomic information for 56 major cattle diseases to provide an integrative framework linking disease expression with host biology, management practices, and environmental factors. To establish foundational phenomic structure, diseases were classified into three categories: infectious; nutritional and toxic; and multifactorial and other non-infectious diseases, with causative factors, clinical signs, diagnostic approaches, and economic impacts reviewed accordingly. We further examined phenomic variation, focusing on tissue specificity, age dependence, seasonality, and disease prevalence. Using data from the U.S. and Egypt, we demonstrate that seasonality and prevalence are often region-specific and shaped by production systems and environmental conditions. Integration of disease phenomics with genomic data from CattleQTLdb reveals a pronounced imbalance in research coverage: QTL mapping is heavily enriched for high-prevalence and economically important diseases, such as mastitis and metabolic disorders, with chromosome 6 emerging as a major genomic hotspot. In contrast, most cattle diseases lack disease-specific genomic annotations, largely due to challenges in phenotype definition, environmental confounding, and limited data availability. Overall, this review highlights how phenomics can bridge epidemiology, physiology, and genomics to uncover shared disease principles, expose knowledge gaps, and guide precision livestock management and breeding for disease resilience.














