-
Pathogenic bacteria remain one of the most persistent threats to global health[1]. These microorganisms are responsible for a wide range of infections resulting not only from clinical exposure, but also from contact with contaminated food, water, and environmental reservoirs. Recent global analyses have shown the devastating consequences of bacterial infections. The escalating threat of antimicrobial resistance (AMR) has emerged as a critical global health crisis, with resistant pathogens undermining the efficacy of current antibiotics and complicating infection control strategies. Bacteria possess inherent mechanisms for acquiring resistance, and the widespread misuse of antibiotics in both clinical and agricultural settings has accelerated this phenomenon. Infections caused by drug-resistant bacteria are associated with higher morbidity, mortality, and economic burden. Consequently, the ability to detect these pathogens rapidly and accurately is vital for guiding targeted therapy, limiting transmission, and informing surveillance efforts[2,3]. In 2019 alone, antimicrobial-resistant pathogenic bacteria were directly responsible for about 1.27 million deaths and contributed to nearly 4.95 million deaths worldwide[1]. Among the six leading bacterial pathogens, Escherichia coli and Staphylococcus aureus accounted for the largest share of infections and fatalities. Crucially, these pathogens are not confined to healthcare settings; they are pervasive in environmental matrices, including soil, water systems, and bioaerosols. These environmental reservoirs act as critical transmission vectors, facilitating the spread of bacteria to humans and triggering outbreaks. Therefore, rapid and reliable detection of pathogens across both clinical and environmental interfaces is essential for protecting human health and preventing epidemics.
These alarming figures underscore the urgent need for accurate and efficient pathogen detection systems. Reliable diagnostic tools not only guide clinical treatment but also safeguard food production, water quality, and environmental health[4]. From preventing hospital-acquired infections[5,6], to monitoring contamination across the food supply chains, developing next-generation biosensing platforms for rapid pathogen detection has become a critical global priority.
Limitations of conventional pathogen detection methods
-
Currently, microbial culture is considered the 'gold standard' for detecting pathogens[7]. This process, however, typically requires several days. This delay renders it impractical for urgent clinical diagnosis or on-site testing. To accelerate detection, researchers have developed molecular diagnostic tools, such as PCR and immunoassays (e.g., ELISA)[8]; however, both approaches face significant limitations. Although PCR is highly sensitive, it cannot distinguish between viable and dead bacteria. It is also susceptible to interference from complex sample matrices[5]. Conversely, immunoassays, which rely on antibodies, encounter a different set of challenges. Their recognition elements are costly to produce, often unstable, and subject to batch-to-batch variation[9]. These drawbacks hinder their widespread application in point-of-care testing (POCT). A comparative overview of conventional pathogen detection methods and emerging phage-based biosensors is shown in Fig. 1.

Figure 1.
Comparative performance of pathogen detection methods. The chart contrasts traditional microbial culture, immunoassays (e.g., ELISA), and PCR with emerging phage-based biosensors. While microbial culture remains the gold standard for detecting viable cells, it is time consuming (2–7 d). Rapid molecular methods (PCR) and immunoassays cannot distinguish live from dead cells, and face challenges due to matrix interference or reagent instability. Phage-based biosensors uniquely combine rapidity (30 min to a few hours) with specific detection of viable bacteria.
Recently, targeted amplicon sequencing has emerged as another approach for pathogen detection. This method involves PCR amplification of conserved regions (e.g., 16S rRNA genes) and subsequent next-generation sequencing to identify microbial taxa based on sequence similarity. Amplicon sequencing offers high taxonomic resolution and can detect mixed or unculturable communities, making it valuable for clinical diagnostics and environmental surveillance. However, it is relatively time-consuming and costly, and it requires bioinformatic analysis which limits its use in point-of-care settings. Moreover, like conventional PCR, it cannot readily differentiate viable from nonviable organisms.
Phages: the emergence of next-generation biorecognition elements
Structure and mechanism of specific recognition
-
Bacteriophages, or simply phages, are viruses that naturally infect bacteria. In recent years, they have attracted significant attention as promising recognition elements in biosensors[5]. A typical phage consists of a protein capsid protecting its genetic material (DNA or RNA), and a tail structure responsible for host recognition. Specifically, tail fibers or tailspike proteins serve as the primary recognition determinants, binding with high specificity to surface receptors such as lipopolysaccharides (LPS), outer membrane proteins, or capsular polysaccharides[6]. This 'lock-and-key' interaction confers exquisite host specificity. Upon attachment, the phage injects its genetic material into the bacterial cell. Subsequently, the infection proceeds via either the lytic or lysogenic cycle[5]. In the lytic cycle, the phage hijacks the host's biosynthetic machinery to replicate, ultimately leading to cell lysis. This lytic activity is classically visualized using a phage plaque assay.
Core advantages of phages as biosensing elements
-
Compared to traditional probes, phages offer several distinct advantages for biosensing. First, they exhibit exquisite host specificity, often distinguishing targets down to the strain level; a trait refined through billions of years of coevolution[5]. This high specificity minimizes cross-reactivity; a common limitation in other detection methods. Crucially, phages naturally possess the ability to distinguish viable from dead bacteria, as they exclusively infect and replicate within metabolically active hosts[6]. This intrinsic capability addresses a major bottleneck in molecular diagnostics (e.g., PCR), which often cannot distinguish live pathogens from residual DNA of dead cells.
From a practical standpoint, phages demonstrate superior stability compared to antibodies[5]. Their robust viral capsids can withstand a wide range of environmental conditions, including fluctuations in pH and temperature. Furthermore, phage production is scalable and cost-effective, requiring only simple propagation in host bacteria.
Finally, the genetic simplicity of phages makes them highly amenable to synthetic biology[5]. Their modular genomes allow for precise reprogramming, such as the insertion of reporter genes or the modulation of host specificity, thereby expanding their versatility for next-generation biosensing applications.
Scope and structure of this review
-
This review adopts an engineering-centric perspective to systematically examine how phage-based biosensors are being redesigned for next-generation pathogen detection. We focus on the modular integration of molecular recognition, signal transduction, and computational design; three foundational pillars that together determine sensor performance.
To provide a clear structural framework, this review is organized around a modular, stepwise engineering workflow illustrated in the redesigned Fig. 2. The discussion begins with phage-based biorecognition strategies and interface design (Steps 1 and 2), followed by synthetic biology tools for programmable phage engineering (Step 1) and signal transduction technologies that convert recognition into measurable outputs (Step 3).
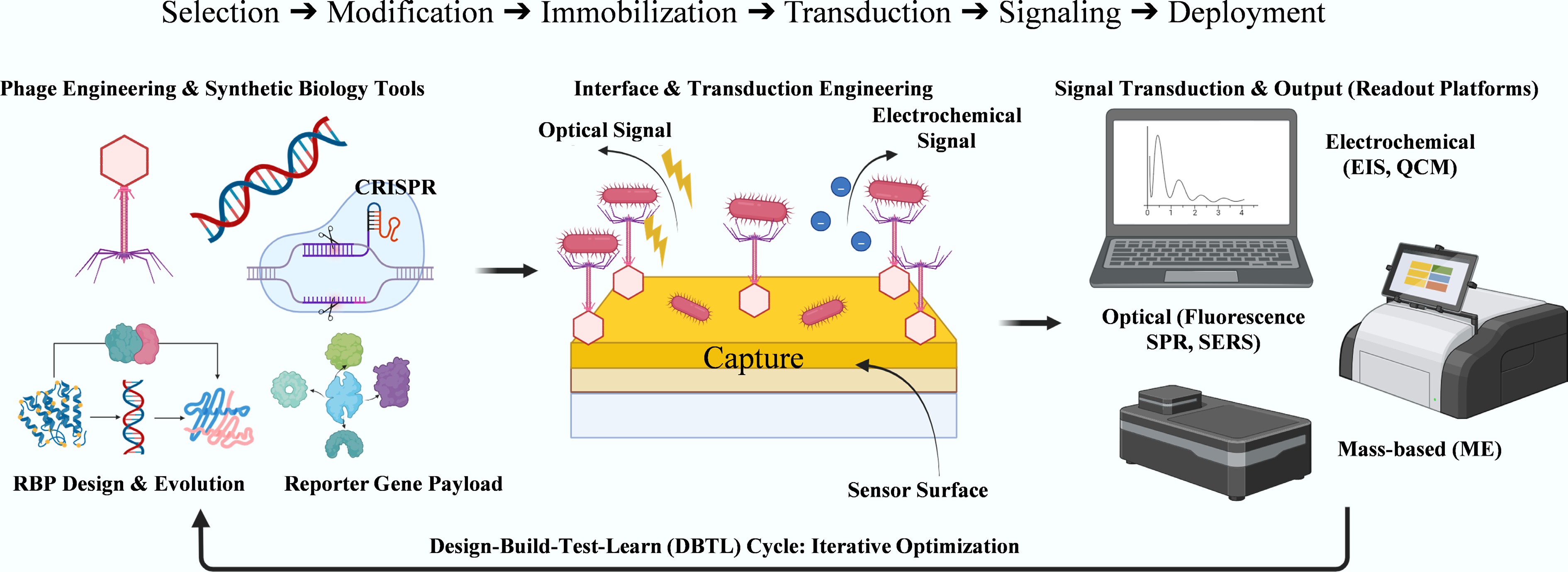
Figure 2.
Modular engineering workflow for phage-based biosensor design. The process comprises three steps: (1) Phage Engineering & Synthetic Biology Tools (e.g., CRISPR, RBP evolution, and reporter gene payload); (2) Interface & Transduction Engineering, including surface functionalization and signal generation; and (3) Signal Transduction & Output, which involves electrochemical, optical, or mass-based readout platforms. This process is embedded within an iterative Design–Build–Test–Learn (DBTL) cycle to optimize performance.
Building upon this foundation, section "Toward intelligent phage biosensors – design, integration, and future directions" examines how computational biology and AI-assisted design are reshaping the biosensor development pipeline, transitioning from empirical discovery to rational design. Finally, we conclude by outlining current challenges and strategic directions toward an intelligent, modular, and application-ready phage biosensing platform.
-
Bacteriophages, firstly discovered over a century ago, have reshaped the understanding of molecular biology[10]. Their remarkable host specificity and resilience under harsh conditions make them highly attractive for bacterial detection[11,12]. The core of this approach lies in bioconjugation; precise chemical or genetic linkage of phages to other materials, creating hybrid structures with enhanced functions[13]. Compared to antibodies, phages are easier to produce, more stable, and uniquely capable of distinguishing live from dead bacteria[14,15]. These features make them ideal biorecognition elements for next-generation biosensors.
Based on advances in phage biology and bioconjugation chemistry[13], researchers have developed integrated recognition strategies that couple biological specificity with controllable surface interfaces. This section outlines how immobilization, amplification, and reporter-based systems lay the foundation for phage biosensing, followed by molecular functionalization and interface engineering that refine performance and precision.
Overview of recognition mechanisms
-
The recognition process in phage-based biosensors utilizes three major strategies: immobilization-based capture, phage amplification, and reporter phage detection (Fig. 3). Each method transforms biological interactions between phages and bacteria into measurable physical or chemical signals.
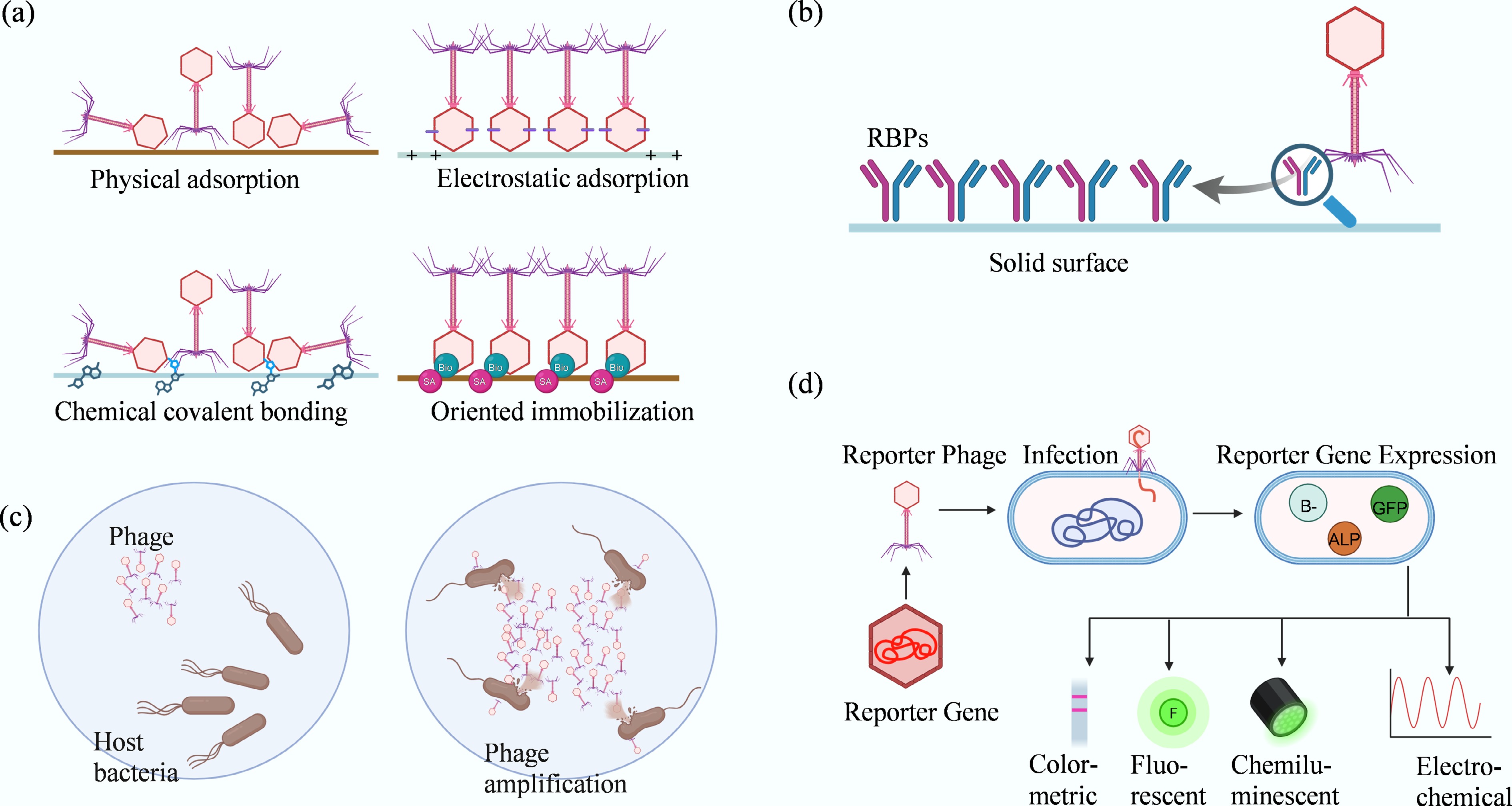
Figure 3.
Major phage-based recognition strategies for pathogen detection. This figure illustrates the primary mechanisms utilized in phage biosensing. (a) Phage immobilization strategies on sensor surfaces: physical adsorption, relying on hydrophobic and van der Waals forces, often results in random orientation; electrostatic adsorption, utilizing head–tail charge differences for alignment; covalent bonding, providing strong attachment but limited orientation control; and affinity or bioorthogonal chemistry, achieving high-precision, site-specific, and oriented immobilization. (b) Comparison of whole phages vs recombinant receptor-binding proteins (RBPs). (c) Detection based on phage amplification in viable hosts. (d) Reporter phage detection, illustrating how engineered phages drive the expression of signal-generating genes (e.g., GFP, ALP) upon infection, to produce colorimetric, fluorescent, chemiluminescent, or electrochemical signals.
Immobilization-based capture
-
Immobilization remains the cornerstone of most phage biosensors[13,14]. The efficacy of bacterial capture, and consequently the sensitivity, stability, and response time, is governed by the immobilization strategy employed. Early efforts relied on physical adsorption via hydrophobic and van der Waals forces, an approach that, while simple, suffers from poor reproducibility and random orientation[11].
To achieve more reliable orientation, electrostatic attraction has been leveraged to guide phage alignment. At physiological pH, the inherent dipole moment of phages (typically with negatively charged capsids and positively charged tails) facilitates 'head-down' attachment to positively charged substrates[16,17]. Coatings such as polyethyleneimine (PEI) and APTES have been shown to significantly improve monolayer stability and uniformity[18]. Further refinements, such as PEI pre-orientation combined with π–π stacking or electrical biasing of CNT electrodes, have demonstrated enhanced detection sensitivity[13,17].
Covalent methods utilize chemical cross-linking to provide stronger and more reproducible attachment. Surfaces functionalized with carboxyl, amino, or aldehyde groups form stable bonds with phage capsid residues, typically via EDC/NHS coupling[13−20]. Although robust, these reactions often result in random orientation. To overcome this, directional control has been achieved through affinity and genetic strategies. For instance, biotin-streptavidin interactions yield high specificity under mild conditions[21], while phage display technology enables the insertion of affinity tags directly into capsid proteins for autonomous self-assembly[22,23]. More advanced techniques employ bioorthogonal 'click' reactions between azide- or alkyne-modified residues and complementary surface groups, allowing for atomic-level control over phage orientation and density[13,24].
Phage amplification-based detection
-
Beyond static binding, phage biosensors can leverage the phage life cycle itself for signal amplification. A single infective phage replicates within a viable host, releasing hundreds of progeny virions upon lysis[25]. This biological multiplication acts as an intrinsic signal enhancer. To visualize the amplified phages, lateral flow assays (LFAs) are commonly employed. For instance, Cox et al.[26] demonstrated that γ phages incubated with Bacillus anthracis produced progeny detectable on gold nanoparticle-labeled strips, forming visible test lines within 2 h. This workflow, which couples biological amplification with immunochromatographic detection, merges high sensitivity with the portability of low-cost diagnostics. By tuning infection time and labeling chemistry, detection limits can be optimized for various clinical or environmental matrices, offering a sensitive, yet field-deployable biosensing approach.
Reporter phage-based detection
-
Reporter systems integrate signal generation directly into the infection process. Engineered phages are designed to carry reporter genes, such as those encoding beta-galactosidase[14], luciferases (e.g., LuxAB or NanoLuc)[27,28], alkaline phosphatase (ALP), or GFP[29], which are expressed exclusively in metabolically active cells. This mechanism provides a direct link between infection events and signal output. Upon infection, the host machinery expresses the encoded enzyme, which subsequently catalyzes a reaction producing colorimetric, fluorescent, or luminescent signals.
For instance, Pires et al.[30] constructed reporter phages by strategically replacing nonessential genomic regions with foreign reporter cassettes. Similarly, Zhang et al.[31] developed a NanoLuc-based reporter phage capable of detecting E. coli O157:H7 in ground beef at concentrations as low as 10 CFU within 9 h. A distinct advantage of this approach is the high signal-to-noise ratio, as dead or inactive cells fail to support transgene expression and thus yield no background signal. Furthermore, certain enzymes (e.g., ALP) can generate electroactive products that can be quantified by amperometry, enabling sensitive electrochemical readouts[32].
Phage-derived receptor-binding proteins (RBPs)
-
Receptor-binding proteins (RBPs) play a vital role in defining recognition specificity[11,14]. These tail fibers, or tailspike proteins, act as molecular 'keys' that bind to unique receptors on bacterial surfaces, such as lipopolysaccharides, outer membrane proteins, or capsular polysaccharides. Each RBP–receptor interaction dictates the host range and binding strength of the phage, allowing strain-level discrimination. While intact phages provide multivalent binding that enhances avidity and robustness under variable conditions, isolated or recombinant RBPs offer greater design flexibility[15]. They can be genetically fused to functional tags, immobilized on solid supports, or integrated into hybrid biosensing platforms as modular biorecognition elements. However, RBPs alone lack the structural protection of the phage capsid and are generally less stable, making surface engineering and fusion design essential to preserve their activity. These complementary features, including robust whole-phage recognition and tunable RBP-based specificity, form a versatile foundation form a versatile foundation for developing both selective and customizable biosensors.
Together, these recognition strategies, immobilization, amplification, and reporter-based detection bridge biological interaction with measurable signals. They form the initial design layer for phage-based biosensors, establishing selectivity and sensitivity that subsequent molecular and interface engineering can further refine.
A comparative analysis of these immobilization techniques is provided in Supplementary Table S1.
Molecular functionalization and phage display
-
Beyond natural recognition, molecular functionalization allows phages to serve as programmable nanoplatforms. This process primarily targets the capsid surface to introduce novel recognition, signaling, or assembly motifs.
Phage display links genotype to phenotype by fusing foreign peptides or proteins to capsid proteins[33]. This technique enables the creation of vast combinatorial libraries, which can be screened through affinity selection to identify peptides with high specificity for bacterial receptors or abiotic surfaces. While classical M13 systems remain popular, lytic T4 and T7 phages have emerged as robust alternatives. The T7 system, in particular, offers distinct advantages for displaying large or toxic proteins at high density, thereby underpinning the design of biosensors with tailored pathogen recognition capabilities[34,35]. The iterative biopanning process used in phage display to enrich high-affinity binders is illustrated in Fig. 4.
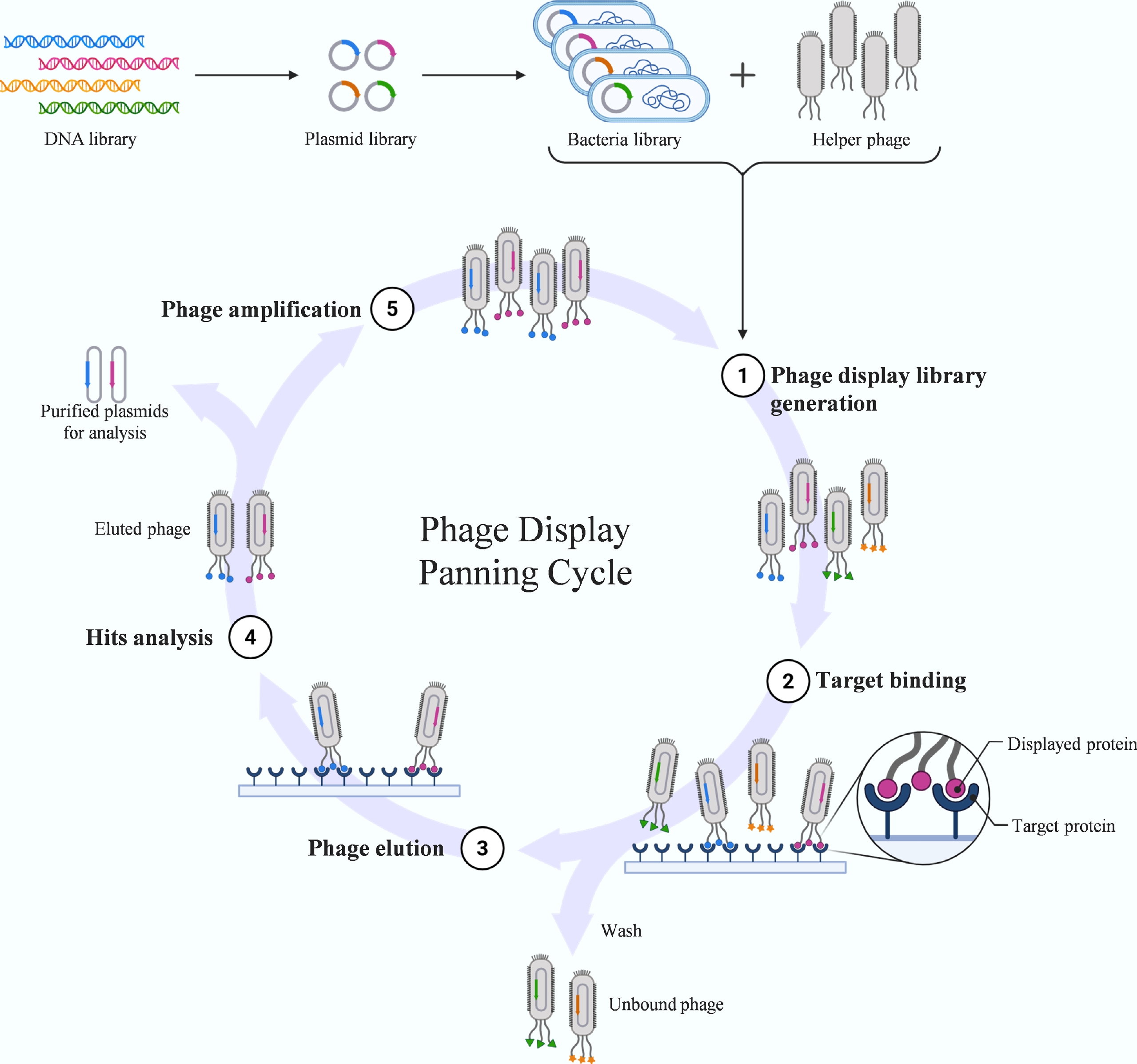
Figure 4.
Biopanning workflow in phage display. This diagram illustrates the iterative affinity selection cycle used to isolate high-affinity peptides. Phage display links genotype to phenotype by fusing foreign peptides to capsid proteins. The panning process involves: (1) binding the phage library to an immobilized target protein; (2) washing to remove unbound phages; (3) elution of specifically bound phages; and (4) amplification of the eluted pool in host bacteria for the next round of selection. Through multiple iterations, peptides with high specificity and affinity are enriched.
Phage capsids can be engineered through precise genomic editing or chemical conjugation[23]. Modern synthetic biology tools, such as BRED and CRISPR–Cas9, enable the targeted insertion or replacement of genes to introduce reporter or binding domains[30]. Complementary to genetic methods, chemical modification utilizes native amino or carboxyl residues for covalent conjugation. However, traditional chemical approaches often lack positional specificity, potentially yielding heterogeneous products[15].
To achieve site-specific control, researchers have introduced non-canonical amino acids bearing azide or alkyne groups into specific capsid positions[36]. These residues facilitate bioorthogonal reactions, such as copper-catalyzed azide–alkyne cycloaddition (CuAAC) or enzyme-mediated Sortase A ligation[37,38]. These techniques allow for the controlled attachment of fluorescent labels, probes, or affinity ligands with atomic-level precision, producing multifunctional, homogeneous phage scaffolds for advanced biosensing interfaces.
Interface control strategies
-
Beyond molecular recognition, the efficacy of a biosensor relies heavily on the physicochemical interaction between engineered phages and the sensor surface. Precise interface design is essential to ensure the correct orientation, structural stability, and biological activity of immobilized phages.
Passive physical adsorption often leads to random orientation, which can occlude tail fibers and significantly reduce bacterial capture efficiency. In contrast, controlled orientation preserves the functional exposure of recognition domains. Strategies to achieve this include electrostatic pre-alignment, affinity bridging, and bioorthogonal coupling[16,39]. Furthermore, proper tuning of immobilization density and linker flexibility is critical for improving sensor reproducibility (Fig. 5).
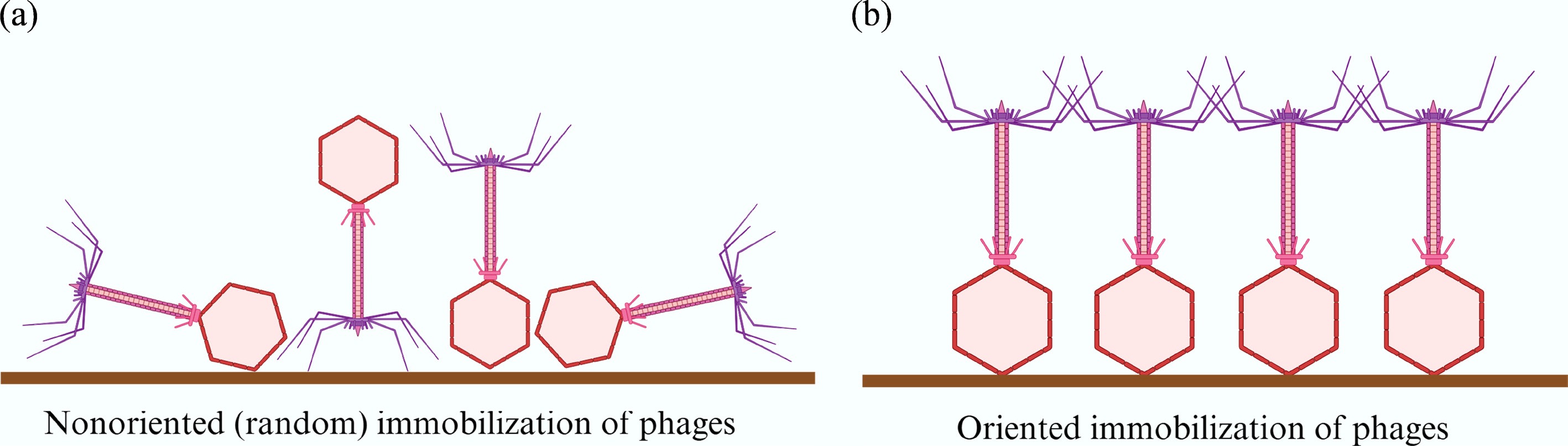
Figure 5.
Schematic comparison of random vs oriented phage immobilization strategies. (a) Non-oriented (random) immobilization often results in disordered alignment, which can occlude receptor-binding tail fibers and reduce bacterial capture efficiency. (b) Oriented immobilization aligns phages vertically on the sensor surface, preserving the functional exposure of tail fibers and thereby enhancing bacterial binding capacity.
The choice of immobilization strategy strongly depends on the nature of the biorecognition element. Whole phages, with their robust capsids and inherent dipole moments, are typically immobilized via physical adsorption, electrostatic interaction, or covalent cross-linking. These methods are simple but often result in random orientation, which may occlude tail fibers and reduce binding efficiency.
In contrast, recombinant receptor-binding proteins (RBPs) are structurally less stable and more sensitive to denaturation. They usually require scaffolded surfaces or affinity tags (e.g., His-tags, biotin, or SpyTag) to achieve oriented and functional immobilization. Since RBPs are monovalent and lack the multivalent interactions provided by intact phages, careful surface design, such as the use of flexible linkers or antifouling layers is critical to preserving their activity[14,22].
Engineered hybrid constructs, such as reporter phages or enzyme-tagged capsids often integrate genetically encoded affinity domains (e.g., gold-binding peptides, cellulose-binding modules) to enable self-directed immobilization. This approach has been successfully demonstrated in biosensors using ALP–GBP fusion enzymes expressed by reporter phages, which directly anchor to gold electrodes while simultaneously generating an electrochemical signal[32].
By tailoring immobilization strategies to the biochemical nature of the recognition element, researchers can optimize biosensor performance for different detection platforms. A summary comparison of these strategies is provided in Supplementary Table S1.
Genetically encoded affinity tags, such as silica-binding peptides (Si-Tags) or gold-binding sequences, facilitate substrate-specific attachment without the need for complex chemical cross-linking[40,41]. When combined with flexible linkers, typically glycine–serine repeats ([GGGGS]n), these tags enhance the mobility of the immobilized phage. This design prevents steric hindrance and maintains correct protein folding, ensuring optimal accessibility to target bacteria[39,42].
The integration of bioorthogonal chemistry with nanomaterials has enabled the construction of robust, electrically conductive interfaces. For instance, phages covalently immobilized on gold nanoparticle–carbon nanotube electrodes have achieved detection limits for E. coli O157:H7 as low as 3 CFU/mL[43]. These hybrid designs highlight how molecular engineering and surface chemistry can work synergistically to enhance real-world sensor performance.
Looking forward, emerging trends in interface engineering include automated target enrichment via magnetic separation[23], integration with on-chip microfluidics[44], and the use of computational modeling to virtually optimize immobilization geometry. These developments aim to unify molecular design and interface control, paving the way for automated, high-throughput biosensor fabrication.
Integration of recognition and signal transduction
-
Recognition and interface control form the biological and physical foundations of phage-based biosensors. To generate a diagnostic output, the captured biological interaction must be converted into measurable signals using signal transduction technologies, such as optical, electrochemical, or mass-sensitive systems, as discussed below.
Immobilization-based sensors integrate naturally with label-free platforms such as Surface Plasmon Resonance (SPR) and Quartz Crystal Microbalance (QCM), where bacterial binding induces shifts in refractive index or resonance frequency[45,46]. In contrast, amplification and reporter-based systems employ enzymatic or luminescent reporters that produce optical or electrochemical outputs[26,31]. Together, these hybrid approaches allow phage biosensors to achieve high specificity, real-time detection, and operational stability, even in complex environments[32].
The continued convergence of phage engineering, surface chemistry, and intelligent computation will enable modular, programmable biosensors that can detect, analyze, and adapt to microbial threats in real time.
Comparative evaluation and limitations of recognition strategies
-
While phage-based biosensors encompass multiple recognition strategies, each presents distinct strengths and context-dependent limitations that warrant careful consideration for real-world deployment. Immobilization-based capture enables rapid detection and is well-suited for integration with label-free transduction platforms such as SPR or QCM[47]. However, this strategy often suffers from random phage orientation, reduced capture efficiency in viscous or complex matrices, and vulnerability to non-specific adsorption[39]. In contrast, phage amplification strategies leverage the biological replication of infective phages for signal enhancement[26], but typically require extended incubation times (often exceeding 2 h) and are sensitive to environmental factors such as temperature or sample toxicity that may inhibit phage activity[25]. Reporter phages provide a high degree of specificity and the unique advantage of distinguishing viable cells by linking infection with signal generation[27,31]. Yet, they require stable genetic engineering, may display reduced robustness across diverse bacterial strains, and face regulatory barriers that limit their applicability in field-based or commercial diagnostic settings[48]. These trade-offs highlight the importance of aligning recognition strategy selection with practical performance requirements, including detection speed, sensitivity, and operational context.
The selection of an optimal recognition strategy depends not only on technical performance, but also on specific deployment contexts. For example, immobilization-based systems are ideal for rapid screening in laboratory settings where sample composition is controlled. In contrast, phage amplification approaches, though slower, are well-suited for field-deployable diagnostics in resource-limited environments[25,26] due to their simplicity and portability. Reporter phages offer superior specificity and viability discrimination[27,48], making them attractive for regulated clinical diagnostics despite higher technical complexity. Table 1 summarizes these trade-offs to support application-driven decision-making.
Table 1. Comparative evaluation of phage-based recognition strategies
Strategy Advantages Limitations/trade-offs Time to result Suitable applications Immobilization Rapid detection; real-time compatible Orientation effects; non-specific binding; surface fouling 30 min−1 h Real-time sensors (SPR, QCM) Phage amplification High sensitivity; biological signal amplification Longer time-to-result; viability-dependent; less robust 2−4 h Low-cost lateral flow strips Reporter phages High specificity; detects viable cells only Requires genetic engineering; regulatory concerns 4−9 h (typical) Clinical diagnostics; water testing These immobilization approaches not only govern bacterial capture and signal specificity, but also directly influence compatibility with downstream signal transduction platforms. For example, oriented phage alignment enhances sensitivity in QCM and SPR, while robust surface anchoring is essential for electrochemical impedance-based platforms. A more detailed analysis of transduction methods is presented in section "Signal transduction technologies".
To mitigate cross-host infection and improve diagnostic accuracy, several optimization strategies are commonly adopted. At the biorecognition level, selecting or engineering RBPs with higher receptor selectivity and validating host range against representative strain panels can reduce off-target binding. At the infection step, using strictly lytic phages and tuning multiplicity of infection (MOI) and incubation windows can minimize unintended amplification from weakly permissive hosts. At the signal-capture level, oriented immobilization and antifouling surface chemistries help suppress non-specific adsorption. At the same time, orthogonal confirmation (e.g., dual-phage targeting, or a secondary molecular readout) further reduces false positives in complex matrices.
-
The preceding section described how bacteriophages recognize bacteria through immobilization, amplification, and reporter strategies. These biological mechanisms form the foundation of phage-based biosensing. However, natural phages often lack the precision or tunability required for practical diagnostics. To meet complex sensing needs, their molecular components must be redesigned and optimized.
Synthetic biology provides a comprehensive toolkit to reprogram phages beyond their native biology[6]. While these methods can serve diverse purposes, from expanding host range to constructing programmable biological circuits, one of their most direct and impactful applications lies in improving pathogen recognition and detection. By combining genetic design, controlled evolution, and modular assembly, researchers can now tailor phages with enhanced binding specificity, signal generation, and regulatory logic.
This section summarizes four key technologies that enable this transformation: genome synthesis and rebooting, genome editing, directed evolution, and genetic circuit design. Together, they establish an integrated 'design–build–test–learn' (DBTL) framework to support the development of engineered phages for next-generation biosensing platforms. The major synthetic biology tools and their functional roles in phage engineering are summarized in Fig. 6.
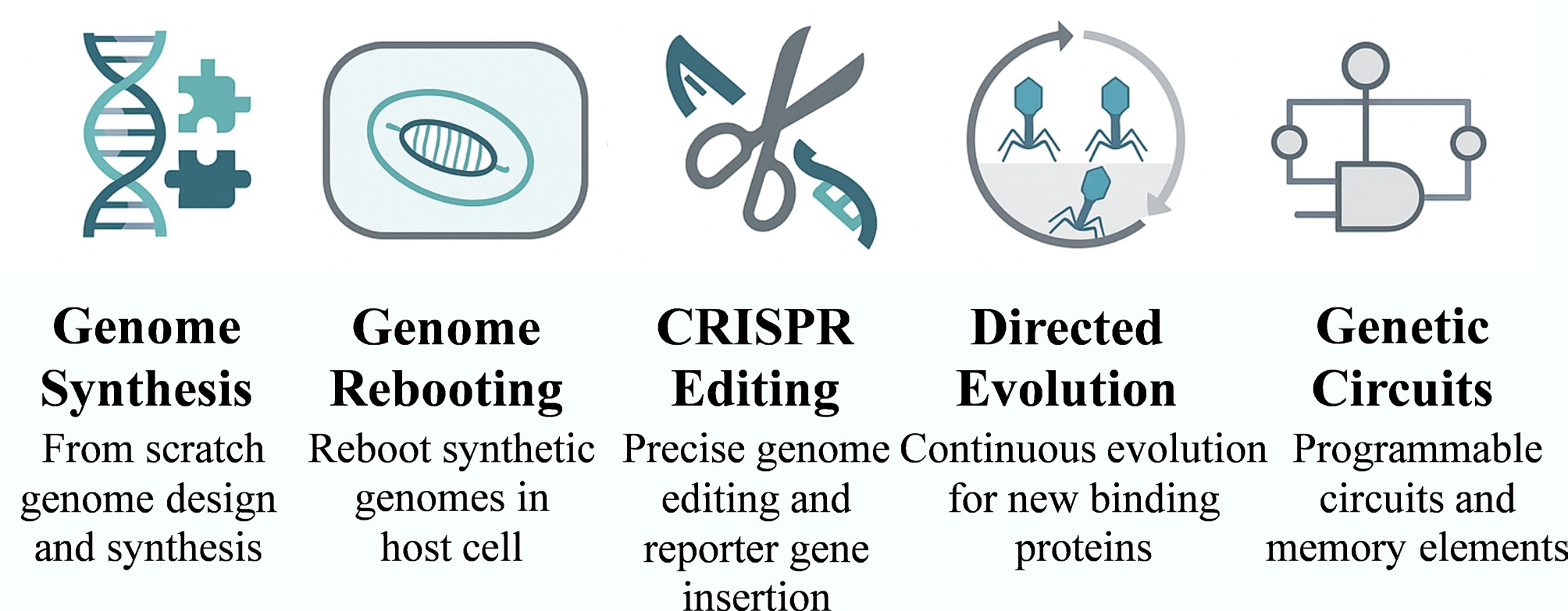
Figure 6.
Synthetic biology toolkit for engineering bacteriophages. The schematic illustrates core technologies for phage design, including genome synthesis and rebooting, CRISPR-based genome editing, directed evolution, and genetic circuit construction. These technologies collectively allow precise control over phage structure and function for pathogen-oriented biosensing applications.
Genome synthesis, assembly, and rebooting
-
Advances in synthetic biology now allow not only sequencing and analysis of phage genomes, but also their de novo construction[49]. A major goal of this capability is to rebuild or redesign phages for controlled infection, host-range modification, and incorporation of sensing modules that enhance pathogen detection. Yet, transforming synthetic DNA into functional viral particles remains technically challenging, especially during cloning and the so-called 'rebooting' phase, in which synthetic genomes must be activated within a compatible host.
Early progress depended on developing efficient DNA assembly systems. Gibson et al. introduced the Gibson Assembly, an in vitro technique that joins multiple DNA fragments in a single reaction, enabling rapid construction of large viral genomes[50]. However, when scientists attempted to assemble complete phage genomes inside their natural bacterial hosts, toxic phage genes often disrupted cell viability and hindered cloning.
To overcome this obstacle, Jaschke et al. established a yeast-based platform for genome assembly using Saccharomyces cerevisiae as an intermediate cloning host[51]. This strategy allowed researchers to assemble and modify lytic phage genomes without lethality to the host, opening the door to engineering phages that target Gram-negative bacteria. Building on this foundation, Ando et al. demonstrated a modular yeast-based system that accommodated multiple phages and even allowed tail-fiber swapping to retarget host specificity[52]. Structural studies of tail proteins, such as those from the T7 phage, further revealed the critical domains responsible for recognition of the bacterial receptor[25,53].
Engineering phages infecting Gram-positive bacteria required alternative rebooting systems. The Loessner lab achieved a breakthrough by employing L-form bacteria, cell-wall-deficient variants that can directly take up synthetic phage DNA[54]. Remarkably, these Listeria L-forms could reboot phages active against both Listeria and Staphylococcus aureus, demonstrating cross-species compatibility. This discovery liberated phage engineering from the constraints of natural hosts and made it possible to reconstruct functional phages solely from sequence information.
These synthetic and cross-host assembly platforms have transformed genome-level manipulation from an experimental challenge into a designable process. For biosensor development, they provide the foundation for building phages with integrated reporter modules, broadened host range, or custom-tuned infectivity, all crucial features for reliable and selective pathogen detection.
Beyond in vivo rebooting systems, recent efforts have explored cell-free phage synthesis using transcription–translation (TX–TL) systems. These platforms enable the in vitro expression and assembly of functional bacteriophages from synthetic DNA without the need for live bacterial hosts. For example, Shin et al. demonstrated the cell-free production of the T7 phage using a PURE system, resulting in infectious virions assembled entirely in vitro. Such approaches offer advantages in speed, biosafety, and modularity. They also enable tight control over reaction conditions and facilitate automated, high-throughput genome screening. Although current yields remain lower than in vivo systems, ongoing optimization of energy supply, DNA packaging, and co-factor balance is rapidly closing this gap. As cell-free platforms mature, they may become central to next-generation biosensor workflows by allowing host-independent phage prototyping and distributed manufacturing[55].
Genome editing
-
While full genome synthesis allows bottom-up phage construction, most engineering efforts still rely on precise modification of existing phage genomes to introduce functional elements such as reporter genes or recognition modules. Early editing approaches depended on homologous recombination, but success rates were extremely low because only a small fraction of phages incorporated the desired mutations, making selection laborious and inefficient.
The introduction of CRISPR–Cas systems revolutionized phage genome editing. In 2014, pioneering researchers transformed the bacterial CRISPR immune mechanism into a powerful tool for targeted editing of lytic phages[56,57]. Their approach used engineered host cells expressing CRISPR–Cas complexes that selectively cleaved unmodified phage DNA while providing repair templates carrying the desired sequence changes. Only phages that successfully integrated the new DNA survived, achieving nearly 100% editing efficiency.
This method proved widely applicable across diverse phage species. For instance, Tao et al. edited the chemically modified genome of phage T4, which contains hydroxymethylated and glycosylated cytosines that are normally resistant to standard molecular tools[58]. Through CRISPR-mediated repair, targeted gene insertions and deletions became routine. As a result, CRISPR-based editing now allows insertion, replacement, or deletion of genes almost anywhere within a phage genome, enabling rapid generation of specialized constructs such as reporter phages. The workflow from design to selection has been shortened from weeks to days, drastically improving throughput and reproducibility.
Beyond diagnostics, this strategy also facilitates functional exploration of phage biology, such as the study of host-recognition domains and packaging signals. Yet for biosensing, its main contribution lies in the controlled integration of reporter or affinity modules, which directly enhance pathogen detection sensitivity and specificity.
From a translational perspective, the cost of phage engineering is driven less by genome editing itself than by downstream screening, validation (host-range, stability, and safety), and quality control during manufacturing. While CRISPR-enabled editing can accelerate prototyping in research settings, large-scale deployment typically favors standardized chassis, modular receptor-binding protein (RBP) swapping, and batch-propagation workflows that reduce per-assay reagent costs. Therefore, application-driven design often balances 'tailored specificity' with 'manufacturable modularity', using phage panels or cocktails when broad coverage is needed.
Directed evolution platforms
-
Phage-based biosensors rely heavily on specific recognition elements, including tail fibers and binding peptides discovered through phage display[59]. Natural diversity, however, cannot always meet new detection demands. When novel pathogens or emerging bacterial strains appear, these interactions must be re-engineered. Directed evolution provides a powerful route to achieve this.
Traditional in vitro evolution methods separate mutation, selection, and amplification into discrete steps, limiting throughput. To accelerate this process, the laboratory of David R. Liu developed phage-assisted continuous evolution (PACE) in 2011[60]. PACE integrates all three steps within a continuous bioreactor that exploits the life cycle of filamentous M13 phages. Mutation, expression, selection, and amplification occur simultaneously, allowing proteins to evolve over hundreds of generations in only a few days (Fig. 7).
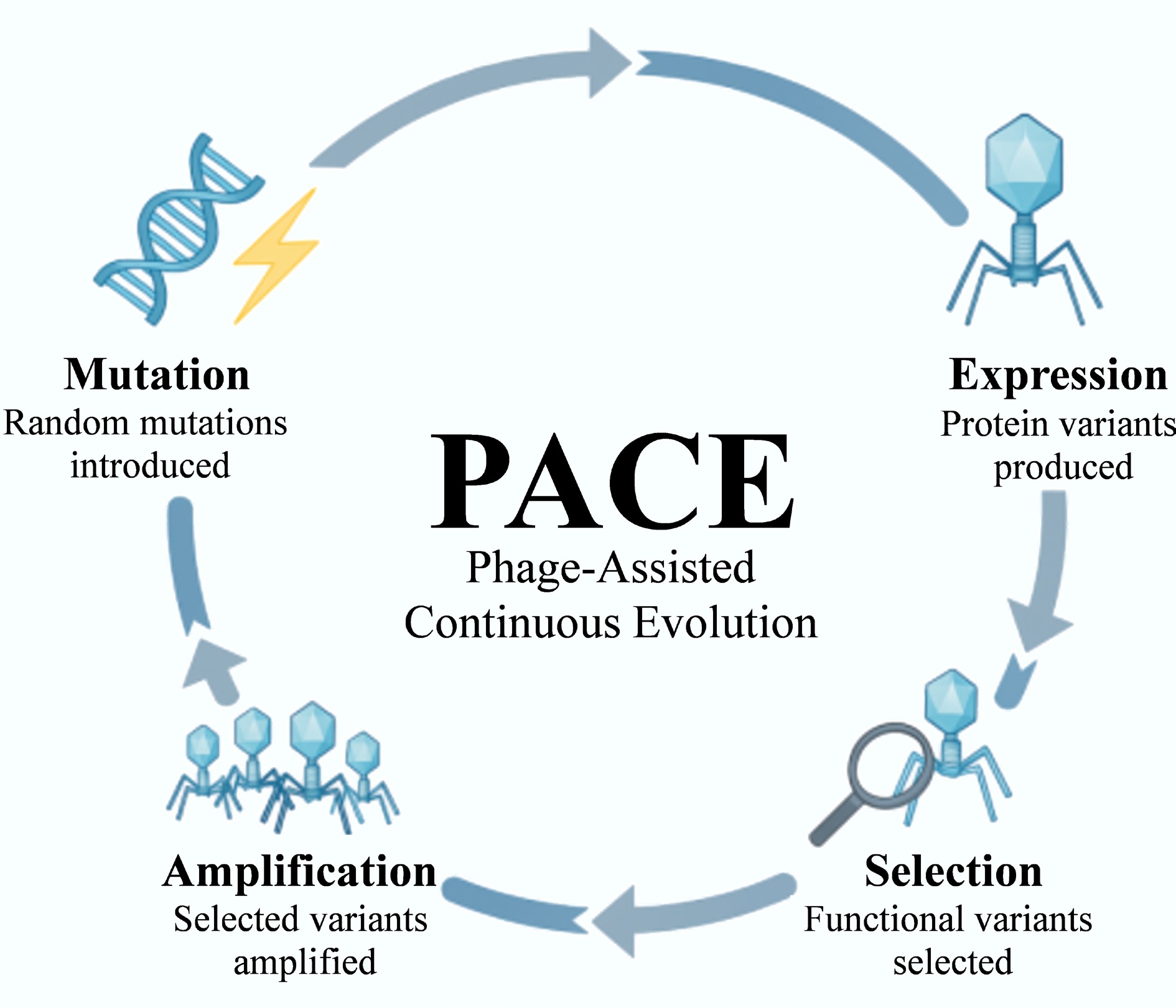
Figure 7.
Principle of phage-assisted continuous evolution (PACE). The diagram illustrates the continuous cycle of mutation, expression, selection, and amplification within an M13 phage system. This integrated bioreactor approach enables the rapid evolution of functional proteins or receptor-binding domains (RBPs) for biosensing applications, achieving hundreds of evolutionary generations in just a few days.
Badran et al. used PACE to evolve a Bacillus thuringiensis toxin that recognizes a new insect gut receptor, thereby overcoming natural resistance[61]. Although this example falls outside pathogen sensing, it demonstrates the method's capacity to rapidly generate new protein–protein interactions. For biosensor design, such speed and scalability are invaluable for evolving receptor-binding proteins (RBPs) with improved affinity or altered host specificity.
By applying PACE or related continuous-evolution systems, researchers can now evolve phage-derived proteins or enzymes directly toward desired biosensing characteristics, higher binding strength, broader host range, or optimized enzyme kinetics for reporter activity. These advances transform phage engineering from a static design problem into an adaptive optimization process, bringing engineered biosensors closer to real-world pathogen detection challenges.
Genetic circuits and memory elements
-
Recent advances in synthetic biology have expanded phage-based biosensing from simple detection toward programmable information processing. By integrating genetic circuits, engineered phages can regulate outputs, perform logic operations, and even record biological events.
To reduce signal interference among transcriptional modules, Voigt's lab developed orthogonal T7 RNA polymerase variants that independently control different promoters[62]. This modular design enables multi-input and multi-output sensing in a single system. Later studies divided T7 RNA polymerase into subunits to fine-tune transcriptional resource allocation[63].
Phage-derived recombinases, such as integrases, were also adapted to construct genetic logic and memory devices. Using these enzymes, researchers built all 16 two-input Boolean gates in living cells[64,65] and circuits capable of recording event order[66,67]. Unlike transcription factor-based systems, integrase-mediated memory rewrites DNA sequences, producing stable and heritable records[67].
These concepts can be incorporated into phage biosensors to create intelligent detection systems that activate only under defined conditions or record infection events. Such designs bridge molecular recognition with computation and mark a step toward programmable biological devices.
Applications and extensions
-
Synthetic biology tools, including genome synthesis, CRISPR editing, directed evolution, and circuit design have enabled a wide range of engineered phage applications. Among them, biosensing and pathogen detection remain central themes.
In diagnostics, synthetic biology simplifies the construction of reporter phages. Jacobs et al. first used luciferase phages for rapid drug-susceptibility testing in Mycobacterium tuberculosis[68]. Later, Kim et al. inserted the luxCDABE operon into phages for self-luminescent Salmonella detection[27]. Smaller phage-derived proteins such as tailspike RBPs can also function independently, as shown in the ELITA assay for Salmonella serotyping[69].
Phages have also been used as biotemplates and delivery vehicles. Engineered M13 phages formed manganese dioxide nanowires for electrochemical glucose sensing[70]. In therapeutics, engineered phages are being tested as new weapons against antibiotic resistance[71]. Meanwhile, therapeutic phages expressing dispersin B or CRISPR payloads have been developed to target resistant bacteria[72−74]. Endolysin engineering has also evolved: Lavigne's group created Artilysins by fusing antimicrobial peptides to lysins to attack Gram-negative bacteria[75]. Concurrently, Buchanan's laboratory expanded lysin host range via domain swapping[76]. These examples share the same principle: programmable phage design for selective recognition and controlled response.
Virus-like particles (VLPs) further illustrate this modularity. MS2-based VLPs with customizable inner and outer surfaces can load functional cargos or display targeting ligands[77,78]. Similarly, engineered phage libraries have been proposed as rapid-response platforms for detecting emerging pathogens[79].
Collectively, these advances shift phage engineering from empirical modification toward rational, modular design. CRISPR-based editing supports precise reporter insertion, directed evolution tunes host specificity, and genetic circuits enable logic control. Together, they form the molecular foundation for next-generation programmable phage biosensors that couple pathogen recognition with intelligent signal processing.
Critical outlook on synthetic biology tools
-
While synthetic biology offers a transformative toolkit for engineering phage-based biosensors, its true impact lies in translating these tools into measurable performance improvements. For example, CRISPR–Cas–mediated editing has dramatically increased the efficiency and precision of genetic modifications in lytic phages, achieving near 100% editing success compared to traditional homologous recombination methods, which typically yield below 1%[56,57]. Directed evolution platforms such as phage-assisted continuous evolution (PACE) allows rapid affinity maturation of receptor-binding proteins (RBPs), completing hundreds of evolutionary generations within just 48–72 h[60]. This significantly reduces development timelines and improves binding specificity for emerging or low-affinity targets. Genome rebooting strategies, especially those leveraging L-form bacteria, have expanded host range engineering beyond natural constraints, allowing synthetic phages to target clinically or environmentally relevant strains previously inaccessible to standard systems[54]. Collectively, these advances enhance the robustness, reproducibility, and design agility of phage-based biosensors, shifting development from empirical optimization to rational, modular design. Their continued integration into sensor pipelines represents a crucial step toward scalable, application-ready biosensing platforms.
-
Section "Recognition and interface engineering in phage-based biosensors" introduced several strategies for using phages as recognition elements. In this section, the signal transduction technologies, mechanisms that convert specific interactions between phages and bacteria into measurable and analyzable signals, are emphasized.
The construction of an effective biosensor requires the precise pairing of a recognition module with an appropriate transduction method. The compatibility between these components directly dictates the system's overall performance, governing critical parameters such as sensitivity and response time. Accordingly, this section is organized around two central themes. First, we elucidate the physical principles underlying commonly used transduction methods, with an examination of how these principles influence detection capabilities, including the potential for label-free analysis. Second, we analyze the integration of these strategies with previously discussed recognition techniques. This approach demonstrates how specific biological and physical combinations can be leveraged to enhance detection outcomes.
Broadly, phage-based biosensors are categorized into three primary classes based on their signal transduction mechanism: optical, electrochemical, and mass-sensitive. Each modality offers distinct advantages and is suited to specific application scenarios. Collectively, they provide a versatile toolkit for the design of robust phage-based pathogen detection systems. Representative label-free and real-time signal transduction mechanisms used in phage-based biosensors are illustrated in Fig. 8.
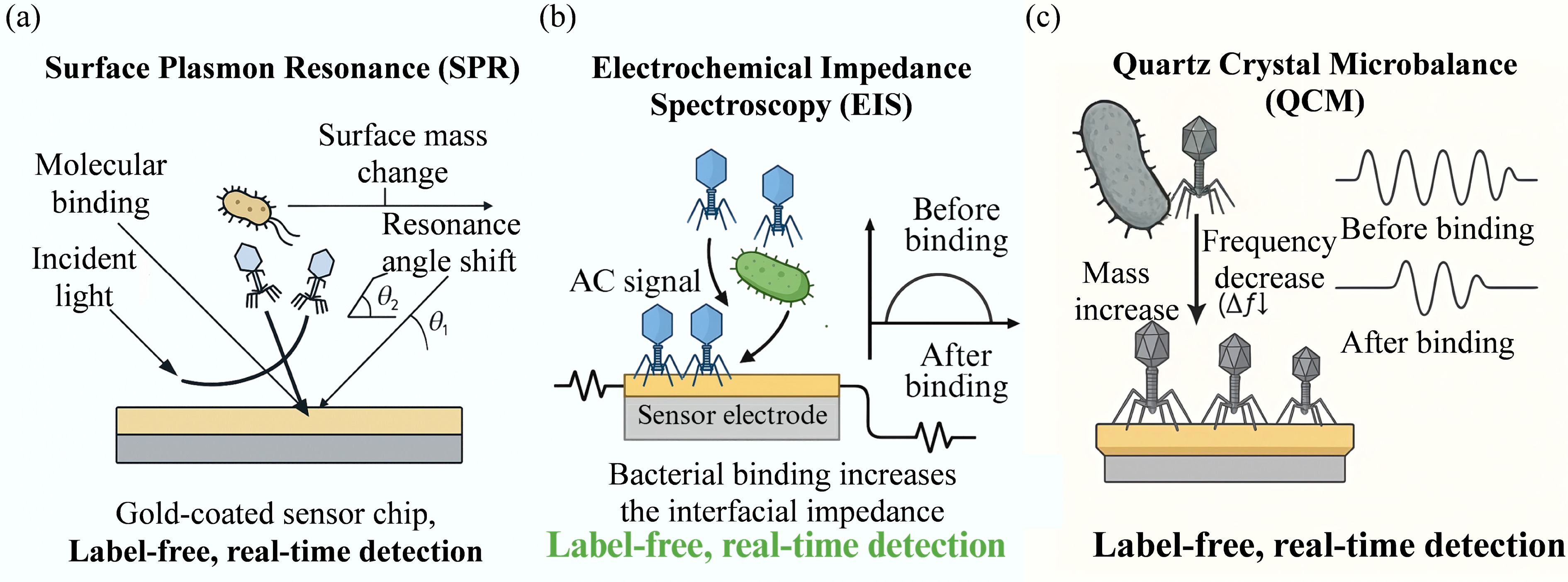
Figure 8.
Schematic illustration of representative label-free, real-time signal transduction mechanisms in phage-based biosensors. (a) Surface Plasmon Resonance (SPR) detects bacterial binding events by monitoring refractive index changes near a gold-coated sensor surface. The interaction between phage-coated surfaces and bacterial targets induces a measurable resonance angle shift (Δθ). (b) Electrochemical Impedance Spectroscopy (EIS) measures changes in interfacial impedance at a phage-functionalized electrode. When bacteria bind to immobilized phages, electron transfer is hindered, resulting in an increase in the impedance signal. (c) Quartz Crystal Microbalance (QCM) detects bacterial capture by monitoring frequency shifts (Δf). The increase in mass on the sensor surface causes a corresponding decrease in resonance frequency, allowing for direct mass quantification.
Detailed case studies integrating these sensing methods with transduction technologies are summarized in Table 2 and Supplementary Table S2.
Table 2. Performance comparison of representative phage-based biosensor architectures
Sensing method Transduction Target pathogen LOD Time Matrix Key innovation Ref. Reporter phages Fluorescence (NanoLuc) E. coli O157:H7 < 10 CFU 9 h Ground beef Integration of ultrabright NanoLuc luciferase for extreme sensitivity [31] Phage amplification Colorimetric (LFA) Bacillus anthracis 2.5 × 104 CFU/mL 2 h Buffer Coupling biological amplification with portable lateral flow immunochromatography [26] Immobilization-based capture Mass-sensitive (QCM-D) MRSA Not specified Real-time Buffer Dual-step recognition (phage capture + antibody binding) enabling strain differentiation [46] Reporter phages Amperometry (ALP) E. coli 1 CFU/100 mL 8 h Drinking water Magnetic bead pre-enrichment combined with self-immobilizing fusion enzymes (ALP–GBPs) [80] Optical methods
-
Optical biosensors characterize biomolecular interactions by monitoring variations in optical properties, such as intensity, wavelength, phase, or polarization[80]. These methodologies are generally classified into two categories: label-free, real-time detection methods, such as Surface Plasmon Resonance (SPR), and signal-generating techniques, such as fluorescence or colorimetry[45].
The selection of a transduction strategy depends on the specific analytical requirements. Label-free techniques are particularly advantageous for determining binding kinetics and affinity, making them ideal for studying surface-level processes and optimizing sensor interfaces. Conversely, signal-based methods often exhibit superior performance when analyzing complex matrices or when high sensitivity is required. By leveraging enzymatic amplification, these approaches facilitate the detection of ultra-low concentrations of target molecules, even in heterogeneous biological samples.
Surface plasmon resonance (SPR)
-
Surface plasmon resonance (SPR) detects molecular interactions by monitoring changes in the refractive index near a sensor surface, which correlate directly with mass accumulation. When molecules bind to or dissociate from the sensor chip, the resonance angle shifts, enabling real-time tracking of these events.
A primary advantage of SPR is its label-free nature, which allows for the monitoring of binding events in real time, and the calculation of kinetic parameters such as association and dissociation rates. Consequently, this technique is frequently employed to investigate the interaction dynamics between phages and bacteria. However, precise measurement of resonance angle shifts requires sophisticated, often costly, optical instrumentation. This requirement poses a challenge for implementing SPR in portable or point-of-care testing (POCT) formats.
Researchers also utilize SPR to evaluate and optimize immobilization-based capture strategies. For example, Arya et al. immobilized T4 phages on gold chips using chemical cross-linkers to construct a sensor capable of specifically detecting Escherichia coli K12[45]. The system achieved a detection limit of 5 × 105 CFU/mL and demonstrated strong selectivity against non-host strains. This study underscores the efficacy of SPR for the dynamic, label-free monitoring of phage–bacteria interactions.
Fluorescence and colorimetric methods
-
Unlike SPR, which passively tracks surface changes, both fluorescence and colorimetric methods actively generate signals following a recognition event. This mechanism provides intrinsic signal amplification, often resulting in superior sensitivity for detecting low-abundance pathogens.
Fluorescence detection is characterized by high sensitivity and can be achieved through surface capture or reporter gene expression. For instance, Oda et al. developed a fluorescent probe by fusing the green fluorescent protein (GFP) gene to the capsid protein SOC in the T-even phage PP01[29]. This construct is specifically bound to Escherichia coli O157:H7, allowing cells to be visualized and counted under a fluorescence microscope within 10 min. Notably, this method could distinguish between live, dead, and viable but non-culturable (VBNC) bacteria.
Reporter phage systems further enhance sensitivity by utilizing enzymatic reactions to convert a single recognition event into a cascade of fluorescent molecules. Hinkley et al. engineered T7 phages to express an ultrabright NanoLuc luciferase fused to a cellulose-binding module (CBM)[48]. Upon infection and lysis, the released enzyme anchored to cellulose membranes, triggering a strong fluorescent signal that enabled the visualization of single colony-forming units (CFUs) in drinking water. Alternatively, isolated phage tailspike proteins were used in combination with fluorescent antimicrobial peptides to construct a paper-based platform[48]. This tool successfully detected Pseudomonas aeruginosa and simultaneously assessed antibiotic susceptibility.
Colorimetric assays offer cost-effectiveness and visual interpretability, making them ideal for field-deployable diagnostics. Like fluorescence, these methods often rely on enzymatic turnover, but produce visible colored precipitates or solution changes. Hoang & Dien engineered phage PP01 to express cytochrome c peroxidase (CCP)[81]. Following the infection with E. coli O157:H7, the expressed CCP enzyme catalyzed a reaction that produced an orange-red signal, achieving a detection limit of 1 CFU/mL in apple juice. Signal amplification is also pivotal in lateral flow assays (LFAs). Alcaine et al. demonstrated that phages engineered to express alkaline phosphatase (ALP) significantly improved LFA sensitivity through substrate precipitation, lowering the detection limit for E. coli in water by 100-fold to 100 CFU per 100 mL[82].
To address the limitation of narrow host ranges in single-phage systems, researchers have developed cocktail and universal capture strategies. Yang et al. formulated a phage cocktail targeting Klebsiella pneumoniae, which increased detection coverage from 42%–62% to 91.6%[83]. To further optimize LFA performance, Ilhan et al. replaced traditional antibodies with phages, finding that phage-based LFAs were approximately ten times more sensitive[84]. Similarly, using phage tailspike proteins alone could yield high-performance LFAs[48]. In a broader approach, Wu et al. introduced gold nanoparticles coated with thiol-phenylboronic acid, which covalently bind to cis-diol groups on bacterial surfaces[85]. Acting as 'universal capture agents', these nanoparticles simplify probe preparation and support the development of broad-spectrum detection platforms.
Surface-enhanced raman scattering (SERS)
-
Surface-enhanced raman scattering (SERS) is an ultrasensitive spectroscopic technique that provides molecular-level 'fingerprint' information. Under optimal conditions, SERS can achieve signal amplification factors of 1014–1015, enabling the detection of single molecules[86]. This enhancement arises primarily from localized electromagnetic fields known as 'hot spots', which typically form in the narrow gaps between noble-metal nanostructures.
However, reliance on randomly aggregated nanoparticles to generate these hot spots often results in poor reproducibility and a lack of control. To address this limitation, Fabris proposed a controlled 'bottom-up' synthesis strategy in which gold nanoparticles are fixed at subnanometer distances using bifunctional linker molecules[87]. This approach offers precise control over hot spot formation, improving both signal intensity and reproducibility while minimizing background fluorescence.
In phage-based biosensing, SERS is frequently used as a signal-amplification mechanism, coupled with immobilization-based capture. For instance, Ilhan et al. functionalized gold nanoparticles with Raman reporter molecules to enhance the sensitivity of phage-based lateral flow assays (LFAs)[84]. This integration improved the detection limit by approximately 1,000-fold, achieving a sensitivity of 102 CFU/mL and demonstrating the potential of SERS for rapid pathogen detection.
Fiber-optic sensors
-
Fiber-optic biosensors enable real-time monitoring, both remotely and directly at the source. The fundamental principle involves attaching biorecognition elements to the tip of an optical fiber, which serves as a waveguide to deliver excitation light and to collect the emitted return signal[88]. Due to their small footprint and capability for remote operation capabilities, these sensors are particularly well-suited for in vivo diagnostics and monitoring within complex or hazardous environments.
In the context of phage-based sensing, bacteriophages are typically immobilized on the fiber tip via surface functionalization techniques. This assembly is then integrated with signal-generating modalities, such as fluorescence, to construct a compact, all-in-one platform. This integrated architecture supports real-time pathogen detection and significantly enhances the practicality of biosensors for deployment in real-world settings.
Electrochemical biosensors
-
Electrochemical biosensors function by transducing biological recognition events into measurable electrical signals, such as current, potential, or impedance. These platforms are characterized by high sensitivity, cost-effectiveness, and the potential for miniaturization, while maintaining robust performance even in complex sample matrices. These attributes make them particularly well-suited for rapid, on-site pathogen detection.
Electrochemical impedance spectroscopy (EIS)
-
Electrochemical impedance spectroscopy (EIS) is a label-free technique widely used to monitor interfacial changes at sensor surfaces. It operates by characterizing the variation in electrode impedance during molecular binding events. In phage-based biosensing, EIS is typically coupled with immobilization strategies. Fundamentally, since captured bacterial cells act as insulating layers, they disrupt the conductivity at the electrode surface.
Methodologically, EIS applies a small-amplitude sinusoidal voltage perturbation to the system and monitors the resulting current response to determine the impedance at the electrode–solution interface. When bacteria bind to immobilized phages, they hinder electron transfer, leading to a measurable increase in interfacial electron-transfer resistance (Rct). This configuration offers dual advantages: high sensitivity to surface perturbations, and the elimination of secondary labeling steps.
However, EIS is susceptible to nonspecific binding (NSB), which can generate false signals; therefore, precise surface engineering is critical. Cheng & Brajter-Toth addressed this by fabricating stable, negatively charged self-assembled monolayers (SAMs) of thioctic acid (lipoic acid) on gold electrodes[89]. These SAMs generated strong electrostatic repulsion against similarly charged interferents, increasing interfacial resistance by over three orders of magnitude and thereby enabling selective charge-based screening.
Carbon nanotubes (CNTs), while promising for their conductivity, pose challenges due to the hydrophobicity of their pristine surfaces, which promotes significant NSB. To mitigate this, Chen et al. proposed a noncovalent functionalization strategy[90]. They utilized polyethylene oxide (PEO) chains to form a neutral, antifouling layer, followed by the attachment of specific probes to the polymer termini. This approach significantly enhanced the selectivity of CNT-based biosensors by suppressing background noise.
Amperometric methods
-
Amperometry is a widely utilized electrochemical technique for quantifying target analytes. It functions by measuring the redox current generated by electroactive species at a constant applied potential. The magnitude of the resulting current is directly proportional to the analyte concentration. Consequently, signal amplification strategies that increase the concentration of electroactive products are essential for enhancing sensitivity in these systems.
In the context of phage-based biosensing, amperometry is often integrated with reporter phage systems. This integration converts a single bacterial recognition event into an amplified electrochemical output via enzymatic catalysis. For instance, El-Moghazy et al. developed a system utilizing engineered T7 phages to detect Escherichia coli on fresh produce[32]. Upon infection, these phages induce the release of alkaline phosphatase (ALP), which catalyzes the conversion of a substrate into 1-naphthol, an electroactive species. The generated 1-naphthol is subsequently oxidized on screen-printed electrodes modified with single-walled carbon nanotubes (SWCNTs). The SWCNT coating significantly enhances electrode conductivity and electron transfer rates, enabling sensitive pathogen detection in complex food matrices.
Similarly, Wang et al. constructed an integrated detection platform for trace levels of E. coli in drinking water[80]. The protocol involved the initial enrichment of bacteria from 100 mL water samples using phage-coated magnetic beads. Captured cells were subsequently infected with engineered phages expressing a bifunctional fusion enzyme, ALP-gold-binding peptides (ALP-GBPs). This enzyme simultaneously anchors to the gold electrode via the GBP domain and catalyzes a reaction that induces the deposition of silver particles on the electrode surface. The resulting silver oxidation current peaks correlated with bacterial concentration, enabling the detection of viable E. coli at the single-cell level.
Potentiometric methods
-
Applications of potentiometric methods in phage-based sensing remain relatively limited compared to other electrochemical modalities. However, recent advances in materials science and theoretical modeling have significantly enhanced the efficacy of these methods for target detection. Potentiometry measures the potential difference between a working electrode and a reference electrode under zero-current conditions. Variations in this potential arise from shifts in the local charge density at the interface, typically induced by specific analyte–surface interactions[91].
Similar to EIS, potentiometry is inherently sensitive to interfacial surface modifications. Consequently, this technique is frequently coupled with immobilization strategies in which phages serve as the recognition elements on the electrode surface. Stable immobilization is a prerequisite for generating a detectable potential shift upon bacterial binding. Therefore, precise control over the design of the immobilization layer is essential to ensure reliable sensor performance.
Foundational work by Chan & Anderson demonstrated that diazonium compounds react specifically with sulfhydryl groups (–SH) in proteins[92]. This discovery established a chemical basis for attaching biomolecules, such as enzymes and antibodies, to modified electrode surfaces via cysteine residues. This chemistry has since become a cornerstone for the fabrication of potentiometric biosensors.
Mass-sensitive methods
-
This category of techniques allows the monitoring of physical and chemical changes occurring on two-dimensional sensor surfaces. Frequently integrated with immobilization-based capture strategies, these methods enable real-time, label-free detection by measuring shifts in resonant frequency or mechanical parameters. Upon the binding of bacteria to phages, the sensor interface experiences mass accumulation, increased layer thickness, or variations in viscoelastic properties. The sensing system directly transduces these interfacial perturbations into quantifiable signals.
Quartz crystal microbalance (QCM)
-
Quartz crystal microbalance (QCM) and surface plasmon resonance (SPR) are widely used label-free sensing methodologies. While both techniques detect molecular binding by monitoring mass changes at the sensor surface, they differ in their transduction mechanisms: SPR relies on optical signals, whereas QCM detects shifts in resonant frequency.
The fundamental component of a QCM system is a piezoelectric quartz crystal that oscillates when a voltage is applied. Mass accumulation on the crystal surface is quantified using the Sauerbrey equation, which establishes a direct linear relationship between the decrease in oscillation frequency and the increase in adsorbed mass. When immobilized phages capture target bacteria, the resulting increase in mass induces a detectable decrease in frequency. This mechanism enables the real-time, label-free monitoring of binding events with sensitivity down to the nanogram or picogram level. Compared to SPR, QCM instrumentation is generally more cost-effective, structurally simpler, and more amenable to miniaturization.
However, the high sensitivity of QCM also presents analytical challenges. The sensor response can be influenced by factors other than mass, such as surface roughness, solution viscosity, and the viscoelastic properties of the adsorbed layer. These non-Sauerbrey effects can potentially distort data interpretation. Consequently, rigorous experimental design and data analysis are required to decouple these variables and ensure measurement reliability.
QCM is particularly effective for characterizing surface-immobilized phage systems, facilitating the study of binding kinetics and the development of rapid diagnostic tools. For instance, Guntupalli et al. employed QCM with dissipation monitoring (QCM-D) in a dual-recognition strategy to detect methicillin-resistant Staphylococcus aureus (MRSA)[46]. First, broad-spectrum phages were used to capture all S. aureus strains. Subsequently, antibodies targeting an MRSA-specific protein were introduced. A sample was confirmed as MRSA-positive only if both recognition steps induced distinct frequency shifts, thereby ensuring high specificity.
Furthermore, Olsson et al. investigated the impact of phage surface density on capture efficiency[93]. Their study on tailed (E79) and tailless (PRD1) phages revealed an optimal density window for bacterial capture. It was observed that capture efficiency declined when the phage layer was either too sparse or overly crowded (steric hindrance). This insight provides critical guidance for the rational design of future phage-based sensor interfaces.
Magnetoelastic (ME) sensors
-
Magnetoelastic (ME) sensors represent a distinct class of mass-sensitive devices capable of wireless, non-contact target detection. Fundamentally, these sensors utilize magnetostrictive materials that oscillate at a natural resonance frequency when subjected to an alternating magnetic field. Similar to QCM, ME sensors respond to mass loading; the specific binding of bacteria to surface-immobilized phages induces a quantifiable decrease in resonance frequency.
A critical advantage of ME technology is its ability to conduct remote interrogation. The sensor can be excited and its signal monitored via magnetic fields without physical connection, making it ideal for real-time monitoring within sealed containers or opaque media. Furthermore, sensitivity is inversely proportional to sensor mass, meaning that miniaturization significantly enhances detection performance.
Lakshmanan et al. pioneered the application of this technology for pathogen detection by immobilizing filamentous phages onto ME sensors to detect Salmonella Typhimurium[94]. Their findings demonstrated the feasibility of the method and highlighted that reducing sensor dimensions markedly improved mass sensitivity. Subsequently, Wang et al. advanced this approach for food safety applications[14,95]. They developed a standardized protocol for washing, filtering, and concentrating bacteria from spinach leaves prior to detection. The resulting system achieved a detection limit of 8 × 102 CFU/mL in complex matrices while maintaining high specificity.
-
The preceding sections described the development of phage-based biosensors, outlining the transition from molecular recognition to signal transduction. We examined strategies to enhance phage–host interactions, including genetic modification, surface engineering, and the integration of diverse signaling modalities. Each stage in this conventional workflow depends on precise biological and physical optimization.
Historically, this 'build-then-test' approach has driven significant progress in biosensor research. However, as synthetic biology and computational tools advance, the field is undergoing a paradigm shift. Rather than merely fine-tuning existing isolates, researchers are now asking a fundamental question: can biosensors be designed rationally rather than empirically?
This perspective shifts the focus to the upstream design phase, presenting distinct challenges such as defining precise detection goals, selecting or engineering specific phage components, and predicting their behavior before wet-lab validation. In the past, phage discovery relied heavily on environmental sampling and trial-and-error screening. This process is constrained by serendipity and captures only a fraction of natural viral diversity, a limitation often referred to as the 'viral dark matter' problem. For instance, a comprehensive study analyzing over 3,000 metagenomic samples expanded the known viral gene pool 16-fold; yet, more than half of these sequences lacked homology to known genes[96].
To bridge this gap, computational biology has emerged as a critical enabler. Emerging tools now facilitate the identification of novel phages, the prediction of bacterial hosts, and the mapping of key structural proteins, such as receptor-binding proteins (RBPs). These capabilities mark a transition from discovery-based exploration to design-based workflows, in which components are generated from predictive models rather than through random screening. Nevertheless, every digital model requires empirical validation. Consequently, each computational design must still be tested and refined through the molecular engineering and interface control strategies described previously. This transition from discovery-driven screening to design-driven, AI-assisted biosensor development is summarized in Fig. 9.
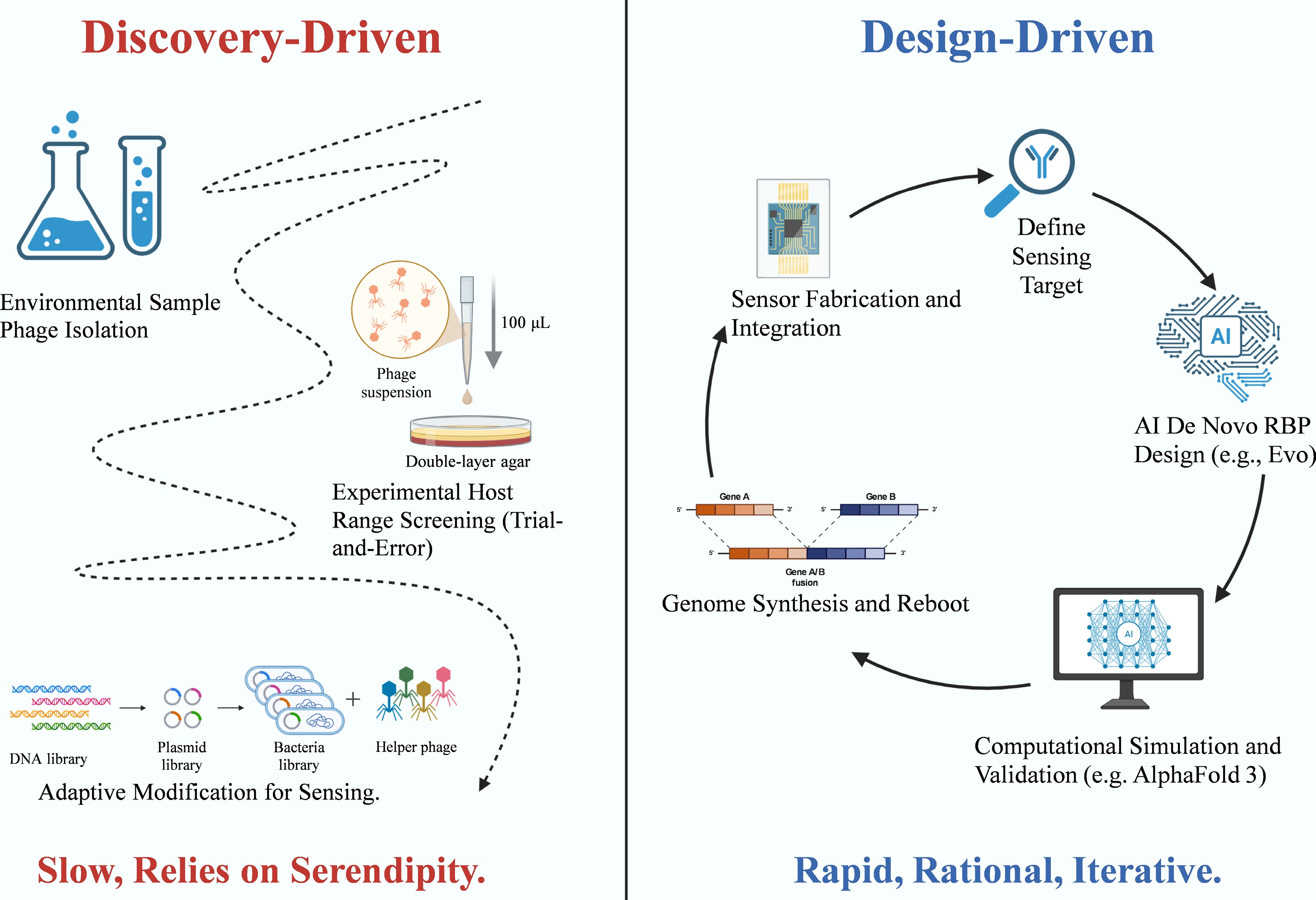
Figure 9.
Paradigm shift in phage sensor development: from discovery-driven to design-driven. The diagram illustrates the transition from empirical isolation to rational, AI-assisted design workflows. The Discovery-Driven approach (left) relies on environmental sampling and trial-and-error host range screening, a process often limited by serendipity. In contrast, the Design-Driven approach (right) employs a rapid, iterative cycle utilizing AI for de novo receptor-binding protein (RBP) design and computational simulation (e.g., AlphaFold 3), followed by genome synthesis, rebooting, and integration with sensor platforms.
Computational tools for phage design
-
The convergence of metagenomic sequencing and synthetic biology has fundamentally transformed the paradigm of phage-based biosensor design. Reliance on environmental screening and empirical trial-and-error assembly has been superseded by AI-assisted computational workflows, which enable the rational design of phage components with predefined functional goals. This design-centric approach is underpinned by four strategic pillars: host prediction, receptor-binding protein (RBP) annotation, protein structure modeling, and genome rewriting.
Host prediction remains a critical step in identifying bacterial targets for newly discovered or engineered phages. Early computational tools, such as VirHostMatcher and CRISPR-based matching systems, relied on co-evolutionary signals, including oligonucleotide frequency and spacer sequence homology[97,98]. However, these methods often exhibited limited efficacy when applied to short DNA fragments, bacteria lacking CRISPR-Cas systems, or complex virus–host networks[99,100]. Subsequent advancements, such as WIsH, utilized probabilistic models to improve prediction accuracy on short contigs[101]. More recently, VirHostMatcher-Net advanced this field by integrating sequence similarity with attention-based learning algorithms to further enhance predictive performance[102].
Parallel advancements have been observed in RBP annotation. Traditional sequence alignment tools often struggled to identify fast-evolving tail proteins because of high sequence divergence[103−106]. Machine learning models, including artificial neural networks, random forests, and support vector machines, overcame this by identifying RBPs based on amino acid composition patterns[95,107]. A significant leap occurred with the introduction of deep learning models such as PhANNs and DeepVP, which substantially increased both prediction accuracy and processing speed[108,109]. These tools are now capable of classifying structural proteins across massive phage datasets, while tools like PPR-Meta facilitate the discrimination between phage and plasmid sequences[110]. The reliability of these analyses is further supported by well-curated databases such as UniProt[111].
Beyond classification, artificial intelligence now drives de novo biological design. Early implementations, such as DeepBind, demonstrated that convolutional neural networks could learn DNA- and RNA-binding specificities[112]. Recent generative language models, such as ProGen and Evo, can generate novel functional enzymes, while AlphaFold 3 predicts biomolecular interactions between proteins, nucleic acids, and small molecules with atomic precision[113,114]. These models not only predict molecular behavior but also guide the construction of synthetic phage components. Experimental validation of these computational predictions has been achieved through genome rebooting in L-form bacteria and classical genome synthesis, exemplified by the de novo construction of the phiX174 phage[49,54,115].
Collectively, these computational tools accelerate the biosensor development pipeline. They significantly shorten the design-build-test cycle, conserve experimental resources, and enhance confidence in design parameters. By synergizing AI, synthetic biology, and virology, the field is transitioning toward a smarter, more deliberate methodology for biosensor engineering.
Bridging in silico design and wet-lab validation
-
Despite advances in computational design and the development of robust experimental biosensors[39,42], a primary bottleneck remains the scarcity of high-quality training datasets. Current AI models often rely on small, biased, or unbalanced data derived predominantly from culturable phages or clinically relevant strains, which constrains their generalizability to novel or rare targets.
Experimental implementation faces significant technical challenges, particularly regarding genome rebooting. Synthetic constructs can exhibit toxicity, and cross-species activation is not always viable. While L-form bacteria offer a broader host range for genome rebooting, technical complexities persist[54]. Furthermore, the construction and screening of synthetic genomes are resource-intensive processes that limit the speed of design iteration.
A critical discrepancy exists between in silico predictions and physical reality. A predicted protein interaction may appear optimal in theory, but fail in practice due to factors such as pH fluctuations, ionic strength, or complex sample matrices. As discussed in section "Recognition and interface engineering in phage-based biosensors", physicochemical factors, including surface orientation, linker length, and immobilization chemistry, play a decisive role in sensor performance. Current computational models rarely account for these interfacial nuances. Moreover, as highlighted in section "Signal transduction technologies", signal transduction is highly context-dependent; not every predicted binding event translates into a measurable diagnostic output.
To bridge this gap, model outputs must be tightly integrated with sensor design constraints, incorporating hardware specifications, material properties, and biochemical environments. Ultimately, the design cycle must be grounded in empirical validation. By closing the loop between design, construction, testing, and learning, the field can advance toward biosensors that are not only intelligent but also practically robust in practice.
From biosensors to theranostics: expanding the application space
-
Phage-based biosensors possess the potential to transcend simple pathogen detection, paving the way for 'theranostics', a paradigm that integrates diagnostics with therapeutic intervention. These systems are designed not only to identify harmful bacteria, but also to execute actionable responses, such as delivering antimicrobial agents or modulating the local microbial environment in situ.
Recent studies have demonstrated the viability of this approach beyond theoretical models. For instance, engineered phages have been equipped with dispersin B to degrade bacterial biofilms[72], while others utilize CRISPR-Cas tools to selectively eliminate antibiotic resistance genes[74]. By coupling these therapeutic modules with detection circuits, phages can be engineered to respond conditionally, releasing antimicrobial payloads only upon the detection of specific resistance markers.
Artificial intelligence facilitates the precision design of such systems. Large language models (LLMs) trained on vast viral, bacterial, and resistance gene datasets can propose personalized phage architectures tailored to the genetic profile of a patient's infection or to complex environmental samples[116]. When combined with microfluidic chips and portable sensing hardware, these designs enable the construction of systems capable of real-time detection, analysis, and actuation. Furthermore, virus-like particles (VLPs) offer a modular platform for simultaneous detection and treatment[77,78]. These particles can be engineered to target specific bacteria, report their presence via fluorescence or electrochemical signals, and deliver functional cargos such as enzymes, drugs, or CRISPR machineries.
Looking ahead, phage biosensors are poised to evolve into fully programmable autonomous agents. Beyond passive detection, they will possess the capacity for decision-making and actuation, heralding a new class of intelligent synthetic biological devices.
Perspectives
-
Although most engineered phage biosensors are currently at the laboratory validation stage, they have shown promising conceptual feasibility and increasing potential for real-world application, particularly in point-of-care diagnostics and environmental monitoring. The first challenge lies in modular design. Current biosensor architectures are predominantly tailored for a single bacterial target and are difficult to adapt to other targets. To create scalable and flexible systems, standardized modular components are required, allowing receptor-binding proteins (RBPs), signaling elements, and transduction platforms to be assembled like interchangeable building blocks. Achieving this necessitates well-defined biological–physical interfaces and standardized assembly frameworks. Synthetic biology provides the necessary tools to realize such modularity through precise genetic and molecular engineering[72,74].
The second challenge concerns robustness in complex sample matrices. While phage-based sensors often perform well in controlled laboratory environments, they can be affected by signal interference in real-world samples, such as food, blood, or wastewater. Emerging solutions, including advanced immobilization chemistries, antifouling coatings, and magnetic bead–based target enrichment show promise in mitigating nonspecific adsorption and improving selectivity[32]. However, these methods have yet to be fully integrated into standardized design pipelines.
The third key issue is computational modeling and data integration. Intelligent biosensor systems must balance multiple interacting layers, protein structure, binding kinetics, genetic control, and transduction hardware. Recent advances in foundation models for protein-small molecule interactions[117] and large-scale biological language models[116] suggest a pathway toward unified, multi-domain predictive design. Nevertheless, progress remains limited by the scarcity of high-quality, cross-domain datasets and the lack of standardized evaluation benchmarks across biology and engineering disciplines.
A fourth priority is the establishment of shared, open-source databases. A curated repository of engineered phage components, covering synthetic RBPs, modular lysins, and tailored tail fibers, would accelerate innovation and reproducibility. Although databases such as UniProt[111] and PVP-SVM[107] provide valuable resources, none yet focus specifically on biosensor-oriented design. A dedicated, annotated repository of phage-derived sensing elements would significantly enhance interoperability and data-driven engineering.
Finally, biosafety and ethical governance must be embedded within the development process. Engineered phages capable of replication or therapeutic gene delivery pose potential risks of dual use and environmental release. The convergence of sensing and therapeutic applications further amplifies these concerns. To ensure safe implementation, future systems should integrate genetic kill switches, controls on horizontal gene transfer controls, and robust containment strategies, while regulatory frameworks must evolve in parallel with advances in synthetic biology.
Looking ahead, phage biosensor research is shifting from discovery-driven to design-driven paradigms. With the integration of computational modeling, deep learning, and genome synthesis, researchers can now rationally design phage components, predict their behavior, and iteratively refine them within closed design-build-test-learn loops. This approach accelerates innovation and enhances predictive accuracy, enabling programmable biosensing architectures that bridge biology and computation.
Ultimately, realizing the full potential of phage-based biosensors will require a unified framework that connects biology, engineering, and information science. Each subsystem, from molecular recognition to signal transduction, must interface seamlessly with both experimental and digital design layers. Such integration marks a broader shift in synthetic biology: from building isolated biological tools to constructing intelligent, adaptive systems.
Another important consideration for practical deployment is the intrinsic host range limitation of phages. While high-strain-level specificity is advantageous for selectivity, it also poses challenges for designing broadly applicable or multiplexed biosensors. Future strategies may involve using phage cocktails, modular RBP libraries, or AI-guided host prediction to ensure effective recognition of diverse clinical or environmental isolates.
Moreover, phage-based sensors could be further enhanced by coupling detection with antibiotic resistance profiling. This may include integrating CRISPR-based systems that detect resistance genes, reporter phages engineered to respond to resistance-associated promoters, or combining phage capture with downstream nucleic acid amplification for resistance markers. Such hybrid approaches would support more actionable diagnostics, especially in clinical settings where rapid detection and resistance identification are both critical for informed treatment decisions.
Compared with qPCR-based fluorescence detection systems, phage-based biosensors offer several distinctive advantages. While qPCR provides high sensitivity and specificity, it requires thermal cycling equipment, trained personnel, and often 1–2 h of reaction time. In contrast, phage biosensors can be integrated into portable, label-free platforms and directly detect viable pathogens without nucleic acid extraction. Additionally, engineered phages enable infection-coupled signal amplification and can be tailored for multiplex or viability-selective detection, making them suitable for low-resource and point-of-care environments.
Phage biosensors are thus emerging as versatile platforms not only for pathogen detection, but also for theranostic and systems-level applications. Beyond diagnostics, they offer a foundation for programmable biological devices capable of sensing, decision-making, and actuation. This transformation redefines phage biosensors from simple analytical instruments into dynamic, designable components of synthetic biological ecosystems, bringing us closer to the era of truly intelligent living sensors. The practical implementation of phage-based diagnostics will likely proceed in stages, with high-value use cases such as foodborne pathogen detection and water quality testing serving as early deployment targets.
Reagent stability and shelf life remain practical determinants of deployability. Although many phages tolerate broader pH and temperature ranges than protein antibodies, their activity can still decay during long-term storage or repeated freeze–thaw cycles. In practice, stability can be improved through formulation (e.g., optimized buffers and stabilizing excipients), dry-format preservation (e.g., lyophilization or paper-based immobilization), and packaging strategies that reduce humidity and thermal stress. Importantly, the effective 'validity period' should be evaluated within the intended matrix and workflow, as complex samples and disinfectants may accelerate the loss of infectivity. Establishing standardized QC metrics (titer, infectivity, and binding performance after storage) will be essential for commercialization and field deployment.
-
It accompanies this paper at: https://doi.org/10.48130/biocontam-0026-0004.
-
During the preparation of this work, the authors used ChatGPT to improve readability. After using this tool, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
-
The authors confirm their contributions to the paper as follows: Wei Wang: conceptualization, visualization, writing − original draft; Yuanyuan Xue: writing − review & editing; Zihan Xu: writing − review & editing; Mao Lin: formal analysis; Qingshun Guo: formal analysis; Li Cui: supervision, funding acquisition, writing − review & editing. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article and its supplementary information files.
-
This work was supported by the National Key Research and Development Program of China (Grant No. 2022YFF0713100), the Natural Science Foundation of China (Grant No. 22241603), and Fujian Province Fujian Science & Technology Innovation Leading Talent Program (Grant No. 2024J011016).
-
The authors declare that they have no conflict of interest.
-
Full list of author information is available at the end of the article.
- The supplementary files can be downloaded from here.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Wang W, Xue Y, Xu Z, Lin M, Guo Q, et al. 2026. Phage-based biosensors for pathogen detection. Biocontaminant 2: e007 doi: 10.48130/biocontam-0026-0004
Phage-based biosensors for pathogen detection
- Received: 19 December 2025
- Revised: 23 March 2026
- Accepted: 04 April 2026
- Published online: 21 April 2026
Abstract: Pathogenic bacteria continue to pose a severe and persistent threat to global public health, necessitating the development of rapid, sensitive, and reliable detection systems. Bacteriophages (phages), owing to their exquisite host specificity and biological robustness, have emerged as powerful biorecognition elements in biosensor development. This review provides a systematic analysis of recent advances in phage-based biosensing, with a particular emphasis on their modular design. We first discuss three fundamental phage-based recognition strategies for sensor construction: immobilization-based capture, phage amplification, and reporter phage systems. The application of synthetic biology tools, such as CRISPR genome editing and directed evolution, alongside interface engineering strategies, such as site-specific immobilization is discussed to precisely regulate the phage-sensor interface and enhance biosensor performance. Various signal transduction platforms, ranging from label-free techniques to reporter-based amplification are summarized, and their roles in translating biological recognition into quantifiable outputs are described. The convergence of phage engineering and computational tools for next-generation biosensor design is highlighted, particularly within a design-build-test-learn framework. Finally, we explore future directions, including AI-assisted phage design, theranostic applications, and the transition toward programmable, adaptive biosensing systems, aiming to establish intelligent platforms for precise clinical diagnosis and on-site environmental monitoring.