-
Pear (Pyrus) is an economically important fruit crop worldwide, valued for its rich nutritional profile[1,2]. Fruit quality is critically constrained by stone cell content, as excessive stone cells degrade flesh texture, resulting in undesirable coarseness[3]. Therefore, elucidating the regulatory mechanisms underlying stone cell formation is essential for the genetic improvement of pear fruit quality. Notably, stone cell content varies markedly among different pear cultivars. 'Shannongsu' pear, a novel cultivar derived from a cross between 'Xinli 7' and 'Dangshansu', exhibits desirable traits such as high fruit quality, long storability, very late maturation, and strong antioxidant capacity, making it an excellent model for studying flesh texture improvement.
The formation of stone cells in pear fruit is intrinsically linked to the processes of lignin biosynthesis, transport, and deposition[4,5]. Lignin in pear stone cells is primarily composed of guaiacyl (G) units, with smaller proportions of syringyl (S) and p-hydroxyphenyl (H) units[6]. The biosynthesis of lignin proceeds via the phenylpropanoid pathway, in which cinnamyl alcohol dehydrogenase (CAD) catalyzes the reduction of cinnamyl aldehydes to their corresponding alcohols. This reaction represents the terminal and crucial enzymatic step in monolignol biosynthesis[7]. The expression patterns of lignin biosynthetic genes PbrCAD1, PbrCAD20, PbrCAD27, and PbrCAD31 in pear fruit have been shown to correlate significantly with dynamic changes in stone cell content during development[8]. However, the expression of these structural genes is precisely orchestrated by complex transcriptional programs. Recent research has identified a core transcriptional complex, PbAGL7–PbNAC47–PbMYB73, which functions as a central regulatory module for stone cell formation by directly binding to and activating the promoters of key lignin biosynthesis genes (e.g., PbC3H1 and PbHCT17)[9]. Notably, studies on 'Shannongsu' pear have identified two MYB transcription factors, PbMYB61 and PbMYB308, which interact to form a functionally antagonistic dimer: PbMYB61 directly activates the expression of the lignin polymerization gene PbLAC1 to promote lignification, whereas PbMYB308 negatively regulates lignin synthesis in stone cells by binding to PbMYB61 and inhibiting its transactivation activity[10]. Moreover, phytohormone signaling pathways—such as gibberellin signaling mediated through the PuMYB91–PuERF023 module[11] and reactive oxygen species (ROS) signaling via the PuRBOHF–PuPRX42-like complex—have been demonstrated to cascade-regulate the expression of downstream lignin biosynthetic genes[12], thereby fine-tuning stone cell deposition. Thus, strategically targeting the regulatory hubs governing lignin accumulation represents a core strategy for reducing stone cell content and improving pear fruit texture, offering a coherent framework for molecular breeding and biotechnological intervention.
Calcium, as a structural component of the plant cell wall, plays a vital role in modulating lignin biosynthesis[13]. Exogenous CaCl2 treatment has been demonstrated to effectively suppress lignification and reduce stone cell formation in pear fruit[14]. Conversely, calcium deficiency can induce physiological disorders such as blossom-end hardening, which is associated with abnormal lignin accumulation[15−17]. Furthermore, CaCl2 treatment significantly downregulates the expression of key lignin biosynthesis genes (e.g., CAD1, CAD2, PAL, C4H) and reduces lignin content in pear flesh[18]. As a ubiquitous secondary messenger, calcium ions (Ca2+) are pivotal in plant development and responses to biotic and abiotic stresses[19]. Plants decode intracellular Ca2+ fluctuations via sensor proteins, among which Calmodulin-like proteins (CMLs) are crucial EF-hand Ca2+-binding proteins. Upon Ca2+ binding, CMLs undergo conformational changes and interact with diverse downstream targets to relay signals[19,20]. In Arabidopsis, AtCML24 interacts with transcription factor WRKY46 to regulate aluminum stress resistance[21]. In apple, the interaction between MdCML15 and MaVQ10 promotes MdWRKY75-mediated leaf senescence[17]. Our preliminary research found that in the 'Shannongsu' pear, treatment with 0.01 M calcium reduced lignin and stone cell content, thereby suppressing stone cell formation. Calcium likely regulates lignin biosynthesis and stone cell formation by upregulating the expression of CML genes such as PbCML23 and PbCML46, while downregulating lignin biosynthetic genes, including Pb4CL1b and PbCAD6. These findings suggest that CMLs may participate in the calcium-mediated inhibition of lignin biosynthesis within stone cells of pear fruit.
Recently, the role of transcription factors (TFs) in Ca2+-mediated regulation of lignin biosynthesis in pear fruit has begun to emerge. In 'Nanguo' pear, CaCl2 treatment downregulates PuDof2.5, a TF that directly binds to and represses the promoter activity of the lignin biosynthesis gene PuPRX42-like, thereby inhibiting lignin deposition[14]. Further studies revealed that Ca2+ signaling also suppresses the NAC TF PuNAC21, which forms a transcriptional complex with PuDof2.5 to co-repress the expression of PuPRX42-like and PuCCoAOMT1[22]. Notably, bHLH family TFs are key regulators of plant secondary metabolism and have been implicated in lignin biosynthesis in other species. For instance, Sorghum bicolor SbbHLH1 acts as a negative regulator, repressing genes such as 4CL1 and HCT[23], while rice OsbHLH034 integrates jasmonate signaling to positively regulate lignin deposition for enhanced disease resistance[24]. Nevertheless, the involvement of bHLH TFs in calcium-signal-responsive regulation of lignin biosynthesis in pear fruit has not been reported.
In summary, although exogenous calcium application is known to inhibit lignin biosynthesis and stone cell development, the underlying molecular mechanism involving calcium sensor proteins remains poorly understood. In this study, we identified that the calcium-responsive protein PbCML46 specifically interacts with the bHLH transcription factor PbbHLH96. This interaction enhances the ability of PbbHLH96 to repress the expression of the key lignin biosynthesis gene PbCAD6, thereby reducing lignin accumulation and stone cell formation. This research provides a theoretical basis for using exogenous calcium treatment to improve fruit quality.
-
The 'Shannongsu' pear (Pyrus bretschneideri) plant materials used in this study were provided by the 'Digital Pear Orchard' in Guanxian County, Liaocheng City, Shandong Province, China. 'Shannongsu' pear calli were cultured under sterile, dark conditions at 24 °C on solid MS medium supplemented with 0.5 mg/L 6-BA, 0.5 mg/L IAA, and 1 mg/L 2,4-D.
Nicotiana benthamiana plants were grown in a potting mixture (vermiculite : nutrient soil = 1:1) within a controlled growth chamber maintained at 23 ± 1 °C, 70 ± 5% relative humidity, and a 16-h light/8-h dark photoperiod.
Exogenous calcium treatment of pear fruits
-
At 15, 25, and 35 d after full bloom (DAFB), young fruits of the 'Shannongsu' pear cultivar were sprayed with sugar alcohol calcium (SAC) solutions at concentrations of 0.01, 0.05, and 0.1 mol/L, with water spraying serving as the control (CK). Each treatment was conducted with three biological replicates. The solution was applied until the surface of the young fruits was evenly wetted and the liquid began to drip. Fruit samples were collected at 25, 35, 45, 65, 95, and 180 DAFB. At each time point, ten fruits were randomly selected; the flesh tissue was immediately frozen in liquid nitrogen and stored at −80 °C for subsequent analyses.
Transient expression in detached young fruits of 'Shannongsu' pear
-
The full-length coding sequences (CDS) of PbCML46 and PbbHLH96 were cloned into the pCAMBIA2300-35S-GFP overexpression vector. Simultaneously, specific 300-bp fragments of PbCML46 and PbbHLH96 CDS were cloned into the virus-induced gene silencing vector pTRV2 to generate pTRV2-PbCML46 and pTRV2-PbbHLH96, respectively. The recombinant vectors and empty control vectors (pCAMBIA2300-GFP, pTRV2) were individually transformed into Agrobacterium tumefaciens strain GV3101 (pSoup-p19). Positive transformants were selected, and single colonies were inoculated into 5 mL of YEP liquid medium containing kanamycin and rifampicin for activation. The activated culture was diluted 1:20 (v:v) in fresh medium and grown to an OD600 of 0.8. Cells were resuspended in infiltration buffer (10 mM MES, 150 μM acetosyringone, and 10 mM MgCl2) to prepare the final infiltration solution. Pear fruits at 35 DAFB were used for infiltration. Needle holes (1–2 cm deep) were first made in the fruit flesh using a sterile 1-mL syringe. The fruits were then completely submerged in the infiltration solution and subjected to vacuum infiltration at 0.8 kg/cm2. This process was repeated twice, each lasting 7 min. After infiltration, fruits were incubated in darkness for 2 d, followed by a 10-d incubation under continuous light before subsequent analysis. Primer sequences are listed in Supplementary Table S1.
Stable transformation of pear calli
-
The full-length coding sequences (CDS) of PbCML46 and PbbHLH96 were cloned into the overexpression vector pCAMBIA2300-35S-GFP to generate 35S::PbCML46-GFP and 35S::PbbHLH96-GFP recombinant constructs. These were introduced into A. tumefaciens strain LBA4404 via the freeze-thaw method. Two-week-old calli were immersed in an Agrobacterium suspension (OD600 = 0.5) for 20–30 min, blotted dry on sterile filter paper, and then plated onto co-cultivation medium (MS medium + 200 μM acetosyringone) in the dark at 24 °C for 24–48 h. Subsequently, the calli were transferred to selection medium (MS medium + 200 μM acetosyringone + 50 mg/L kanamycin + 500 mg/L carbenicillin) and maintained under dark conditions[25]. Calli were subcultured every three weeks. Transgenic pear calli lines were obtained after six rounds of selection and subculture.
Stone cell distribution analysis
-
Histochemical staining for lignin was performed using the phloroglucinol-HCl method. Pear fruit tissue sections were first treated with HCl for 1 min, followed by incubation in a 12% (w/v) phloroglucinol solution (in 95% ethanol) for 3 min. Sections were rinsed with distilled water, blotted dry, and immediately photographed. Lignified stone cells stained a characteristic purple-red color[26].
Lignin content measurement
-
Lignin content was determined using a commercial Lignin Content Assay Kit (G0708F, Geruisi Biotechnology, Suzhou, China) following the manufacturer's instructions. Briefly, fruit or callus samples were dried at 80 °C to a constant weight and ground into a fine powder. The powder was passed through a 40-mesh sieve. Subsequently, 2 mg of the sample was mixed with 1.5 mL of 80% (v/v) ethanol, vortexed for 10 s, incubated at 50 °C for 20 min, and centrifuged at 12,000 rpm at 25 °C for 10 min. The supernatant was discarded, and the pellet was washed and dried again at 95 °C. The dried residue was incubated with 750 µL of Regent I at 70 °C for 30 min and cooled to room temperature. Next, 300 µL of Regent II and 450 µL of glacial acetic acid were added to the mixture. After centrifugation at 5,000 rpm for 10 min, 200 µL of the supernatant was mixed with 600 µL of glacial acetic acid in a cuvette. Absorbance was measured at 280 nm.
RNA extraction and qRT-PCR analysis
-
Total RNA was extracted from pear flesh tissues under different calcium treatments using an RNA extraction kit (Nanjing Novozan Biotechnology, China). First-strand cDNA was synthesized using the HiScript II Q RT SuperMix for qPCR (+gDNA wiper) kit (Nanjing Novozan Biotechnology, China). qRT-PCR was performed using gene-specific primers (Supplementary Table S1), synthesized by Sangon Biotech (Shanghai, China). PbActin was used as the internal reference gene. Each 20 µL reaction mixture contained 10 µL of 2× SYBR Green PCR Master Mix, 7 µL of ddH2O, 1 µL each of forward and reverse primers, and 1 µL of cDNA template. The PCR protocol was as follows: initial denaturation at 94 °C for 30 s; followed by 40–45 cycles of 94 °C for 5 s, 50–60 °C for 15 s, and 72 °C for 10 s. All reactions were performed with three technical replicates. Relative gene expression levels were calculated using the 2−ΔΔCᴛ method. Primer sequences are listed in Supplementary Table S1.
Y2H assay
-
The CDS of PbCML46 and PbbHLH96 were cloned into the pGBKT7 (bait) and pGADT7 (prey) vectors, respectively, generating pGBKT7-PbCML46 and pGADT7-PbbHLH96. These constructs were co-transformed into yeast strain Y2HGold. Transformants were selected on synthetic dropout (SD) medium lacking leucine and tryptophan (SD/-Leu/-Trp). Protein–protein interactions were assessed on SD medium lacking leucine, tryptophan, histidine, and adenine (SD/-Leu/-Trp/-His/-Ade), supplemented with X-α-Gal. Primer sequences are listed in Supplementary Table S1.
Pull-down assay
-
The CDS of PbCML46 and PbbHLH96 were cloned into the pGEX4T-1 (for GST-tagged protein) and pET32a (for His-tagged protein) vectors, respectively. The recombinant plasmids were transformed into Escherichia coli BL21 (DE3) for fusion protein expression and purification using a commercial protein purification kit (Beyotime Biotechnology, Shanghai, China). For the binding assay, purified His-PbbHLH96 and GST-PbCML46 proteins were incubated together. After extensive washing, bound proteins were eluted and analyzed by western blotting using anti-His and anti-GST antibodies (Abmart, Shanghai, China). Primer sequences are listed in Supplementary Table S1.
LCI assay
-
The CDS of PbbHLH96 and PbCML46 were cloned into the 35S-nLUC and 35S-cLUC vectors, generating PbbHLH96-nLUC and PbCML46-cLUC constructs, respectively. These constructs and empty vector controls were introduced into A. tumefaciens strain GV3101. Bacterial suspensions (OD600 = 0.8) containing the test or control pairs were mixed and infiltrated into the abaxial side of leaves from four- to five-week-old N. benthamiana plants using a needleless syringe. After 48 h of incubation in darkness, leaves were sprayed with 1 mM D-luciferin potassium salt solution, kept in darkness for 5 min, and then imaged using an IVIS Lumina III in vivo imaging system. Primer sequences are listed in Supplementary Table S1.
Y1H assay
-
The CDS of PbbHLH96 was cloned into the pGADT7 vector to generate the pGAD-PbbHLH96 effector construct. A 1,000 bp fragment of the PbCAD6 promoter was cloned into the pHIS2 vector to generate the pHIS2-pPbCAD6 reporter construct. These constructs were co-transformed into the yeast strain Y187. The transformed yeast cells were plated on SD medium lacking tryptophan, leucine, and histidine (SD/-Trp/-Leu/-His) containing an optimal concentration of 3-aminotriazole (3-AT) to suppress background growth. Interactions were assessed based on yeast growth on this selective medium. Primers are listed in Supplementary Table S1.
EMSA assay
-
The CDS of PbbHLH96 was cloned into the His-tagged pET-32a vector. The recombinant plasmid was transformed into E. coli BL21 (DE3), and the His-PbbHLH96 fusion protein was induced and purified. The 1,000 bp PbCAD6 promoter was analyzed using PlantCARE to identify potential cis-elements. Complementary single-stranded oligonucleotides containing the wild-type (WT) or mutated (MUT) putative binding sites were synthesized and labeled with biotin (for the probe) or left unlabeled (for competitors). The binding reactions and detection were performed using the LightShift Chemiluminescent EMSA Kit (Thermo Fisher Scientific, USA) according to the manufacturer's protocol. Probe sequences are listed in Supplementary Table S1.
DLR assay
-
The 1,000 bp promoter fragment of PbCAD6 was cloned into the pGreenII 0800-LUC vector to generate the reporter construct pPbCAD6::LUC. The CDS of PbCML46 and PbbHLH96 were cloned into the pGreenII 62-SK vector to generate effector constructs (35S::PbCML46, 35S::PbbHLH96), with the empty pGreenII 62-SK vector serving as a negative control. All constructs were transformed into A. tumefaciens strain GV3101 (pSoup-p19). Bacterial suspensions (OD600 = 0.8) containing the reporter and effector constructs were mixed at a 1:1 (v/v) ratio and co-infiltrated into the leaves of four- to five-week-old N. benthamiana plants. After 48 h, luciferase activity was measured using a dual-luciferase reporter assay system. Firefly luciferase (LUC) activity was normalized to Renilla luciferase (REN) activity for each sample. Supplementary Table S1 contains a list of the primers.
Statistical analysis
-
IBM SPSS Statistics 22 (SPSS) Statistics 22 (SPSS, Inc., Chicago, IL, USA) and GraphPad Prism 8.0 were used for statistical analysis. The data were expressed as mean ± standard error (SE). A p-value less than 0.05 was considered statistically significant.
-
Building on previous findings that low-concentration exogenous calcium treatment significantly induced PbCML46 expression in pear fruits at different developmental stages, concurrent with reduced stone cell content and lignin accumulation[27], we functionally characterized PbCML46 as a candidate regulatory gene. Both overexpression (PbCML46-OE) and virus-induced gene silencing (PbCML46-VIGS) constructs were generated and introduced into 'Shannongsu' pear fruits at 35 d after full bloom (DAFB) via Agrobacterium tumefaciens-mediated vacuum infiltration. Phenotypic analysis 12 d post-infiltration revealed that, relative to the empty vector control, PbCML46-OE fruit sectors exhibited markedly reduced lignified staining areas, whereas PbCML46-VIGS fruit sectors displayed intensified lignin staining (Fig. 1a). PbCML46 overexpression significantly decreased lignin content and stone cell content in the infiltrated region of the pear fruit; in contrast, suppression of PbCML46 expression significantly increased lignin and stone cell contents in the area surrounding the infiltrated region (Fig. 1c, e). qRT-PCR analysis confirmed significant upregulation of PbCML46 in overexpression lines, accompanied by suppression of the key lignin biosynthetic gene PbCAD6. Consistent with this transcriptional profile, lignin content was significantly lower in PbCML46-OE fruits. Conversely, in PbCML46-VIGS fruits, PbCML46 expression was downregulated, while PbCAD6 expression was elevated (Fig. 1b, d).
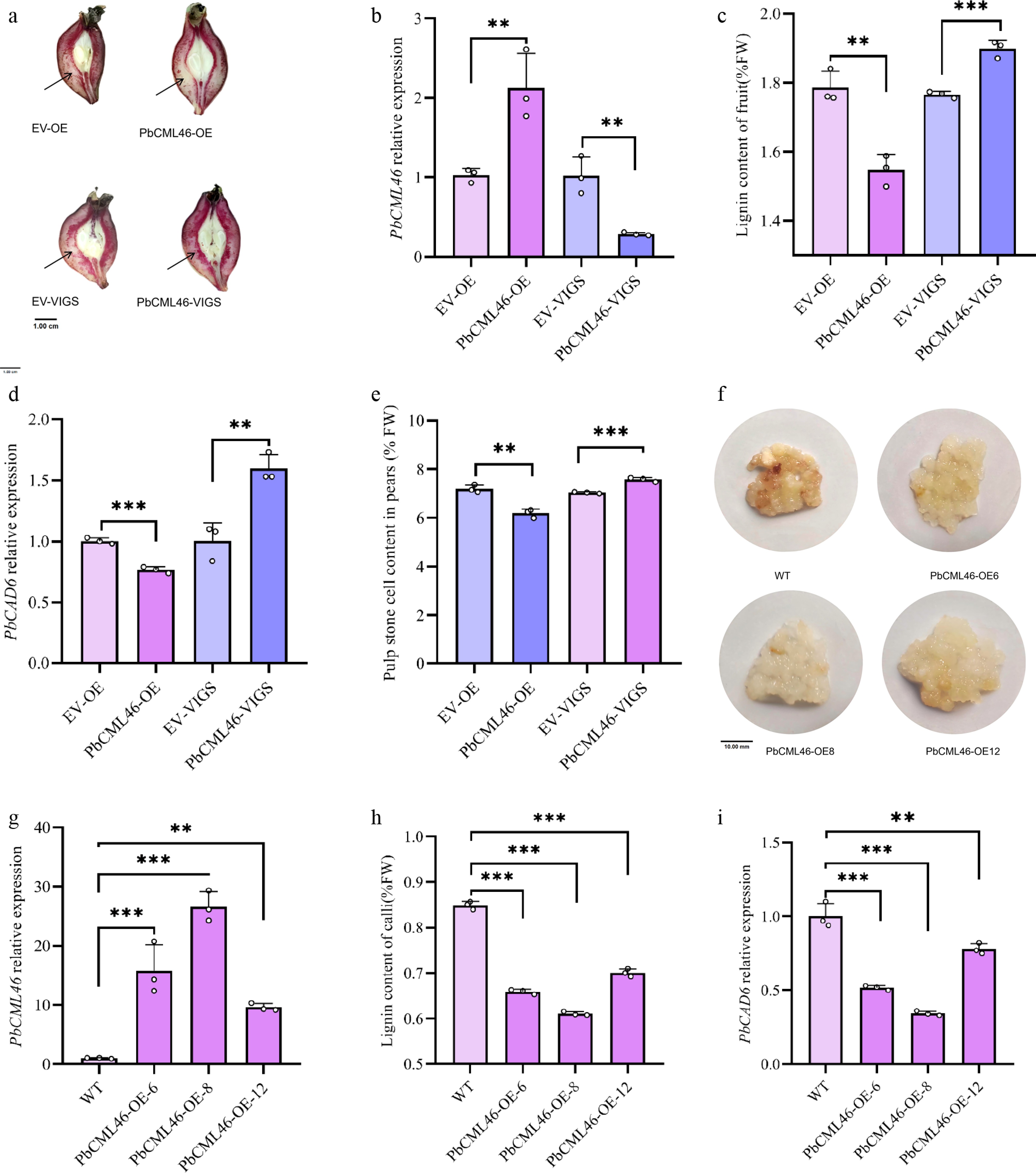
Figure 1.
PbCML46 functions in lignin biosynthesis of stone cells in pear fruit. (a) Histochemical localization of stone cell tissue in pear fruits with empty pCAMBIA, empty pTRV, PbCML46 overexpression, and PbCML46 silencing (pear young fruits 35 d after full bloom). (b) qRT-PCR analysis of the relative expression levels of PbCML46 in the infiltration zones of pear fruits with empty pCAMBIA, empty pTRV, PbCML46 overexpression, and PbCML46 silencing. (c) Measurement of lignin content in the infiltration zones of pear fruits with empty pCAMBIA, empty pTRV, PbCML46 overexpression, and PbCML46 silencing. (d) qRT-PCR analysis of the relative expression levels of PbCAD6 in the infiltration zones of pear fruits with empty pCAMBIA, empty pTRV, PbCML46 overexpression, and PbCML46 silencing. (e) Measurement of stone cell content in the osmotic sites of pear fruits with empty pCAMBIA, empty pTRV, PbCML46 overexpression, and PbCML46 silencing. (f) Lignin staining phenotype of wild-type and PbCML46-overexpressing callus. (g) qRT-PCR validation of the relative expression level of PbCML46 in stably overexpressing callus. (h) Measurement of lignin content in stably overexpressing callus. (i) qRT-PCR analysis of the relative expression level of PbCAD6 in stably overexpressing callus. Values represent the mean ± SD of three independent biological replicates. ANOVA analysis and Tukey's test were used (** p < 0.01; *** p < 0.001).
To further validate these observations, stable PbCML46-overexpressing transgenic calli were established from 'Shannongsu' pear using Agrobacterium-mediated transformation. Compared to wild-type calli, PbCML46-OE calli exhibited reduced lignin staining (Fig. 1f). qRT-PCR analysis verified significant upregulation of PbCML46 and concomitant downregulation of PbCAD6 in overexpression lines, along with a significant reduction in lignin content (Fig. 1g–i). Collectively, these results indicate that PbCML46 is a key regulator of lignin biosynthesis in stone cells of pear fruit.
PbCML46 interacts with PbbHLH96 to regulate lignin biosynthesis in stone cells of pear fruit
-
To further elucidate the molecular regulatory mechanism of PbCML46, a yeast two-hybrid (Y2H) screen was performed using PbCML46-BD as bait to identify its interacting proteins, which led to the identification of the bHLH family transcription factor PbbHLH96 (Supplementary Fig. S1). Yeast cells co-transformed with PbCML46-BD and PbbHLH96-AD grew on stringent selective medium (-Leu/-Trp/-Ade/-His) and exhibited a distinct blue color in the presence of X-α-gal, confirming a specific protein–protein interaction (Fig. 2a).
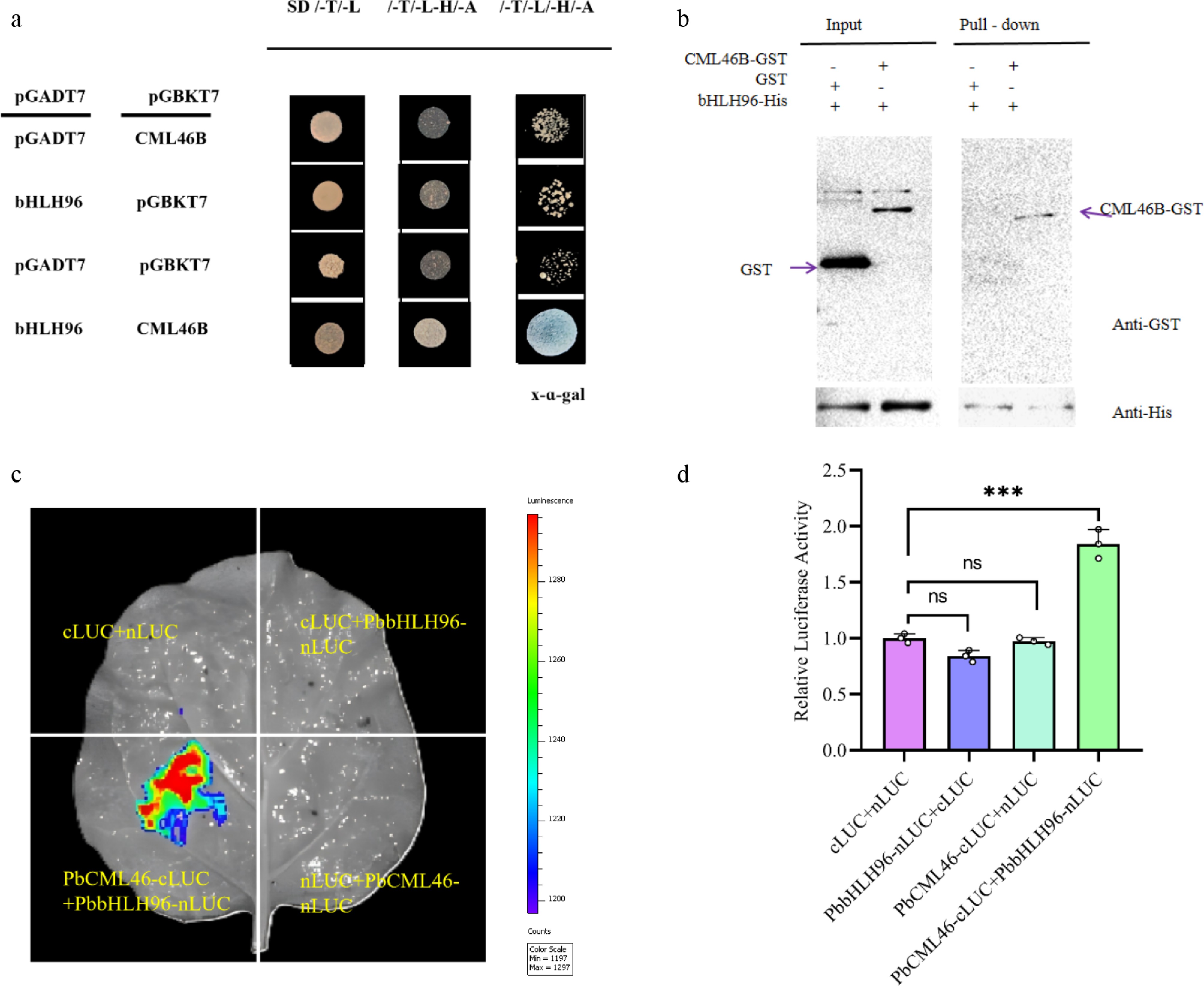
Figure 2.
PbCML46 interacts with PbbHLH96 to regulate lignin biosynthesis in stone cells of pear fruit. (a) Yeast two-hybrid assay showing the interaction between PbCML46 and PbbHLH96. (b) In vitro pull-down assay validating the interaction between PbCML46 and PbbHLH96. '+' and '−' indicate the presence and absence of the specified protein, respectively. (c), (d) Luciferase complementation imaging assays demonstrating the interaction between PbCML46 and PbbHLH96 in planta. Values represent the mean ± SD of three independent biological replicates. One-way ANOVA was performed (*** p < 0.001).
To validate this interaction, an in vitro GST pull-down assay was conducted. The results showed that PbCML46-GST immobilized on glutathione beads specifically pulled down PbbHLH96-His, whereas the GST control protein did not (Fig. 2b), confirming direct binding in vitro. Additionally, an in vivo luciferase complementation imaging (LCI) assay in tobacco leaves demonstrated that co-expression of PbCML46-cLUC and PbbHLH96-nLUC generated a strong luciferase signal, significantly higher than that observed in all negative control combinations (Fig. 2c, d). Collectively, these results confirm that PbCML46 specifically interacts with PbbHLH96, both in vitro and in vivo.
PbbHLH96 binds to the PbCAD6 promoter and represses its activity
-
The bHLH family of transcription factors plays a pivotal role in regulating lignin biosynthesis in plants. To investigate the function of PbbHLH96 in pear fruit, we first examined its expression pattern in fruit flesh, where lignin content decreased following exogenous sugar alcohol calcium (SAC) treatment. qRT-PCR results showed that SAC treatment significantly induced PbbHLH96 expression at different fruit developmental stages (Supplementary Fig. S2), leading us to hypothesize that PbbHLH96 might act as a negative regulator of lignin biosynthesis.
To test this hypothesis, we performed Agrobacterium-mediated transient overexpression and gene silencing of PbbHLH96 in pear fruit. The results demonstrated that PbbHLH96 overexpression significantly suppressed lignin biosynthesis in stone cells at the infiltration site of pear fruits, as evidenced by reduced histochemical staining intensity in fruit sections, along with decreased lignin content and stone cell content. Conversely, silencing PbbHLH96 promoted lignin synthesis, resulting in increased lignin content and intensified staining, as well as significantly increased stone cell content in the infiltrated region of pear fruits (Fig. 3a–c, e). Stable overexpression of PbbHLH96 in pear calli similarly confirmed its inhibitory effect on lignin deposition (Supplementary Fig. S3a–S3c).
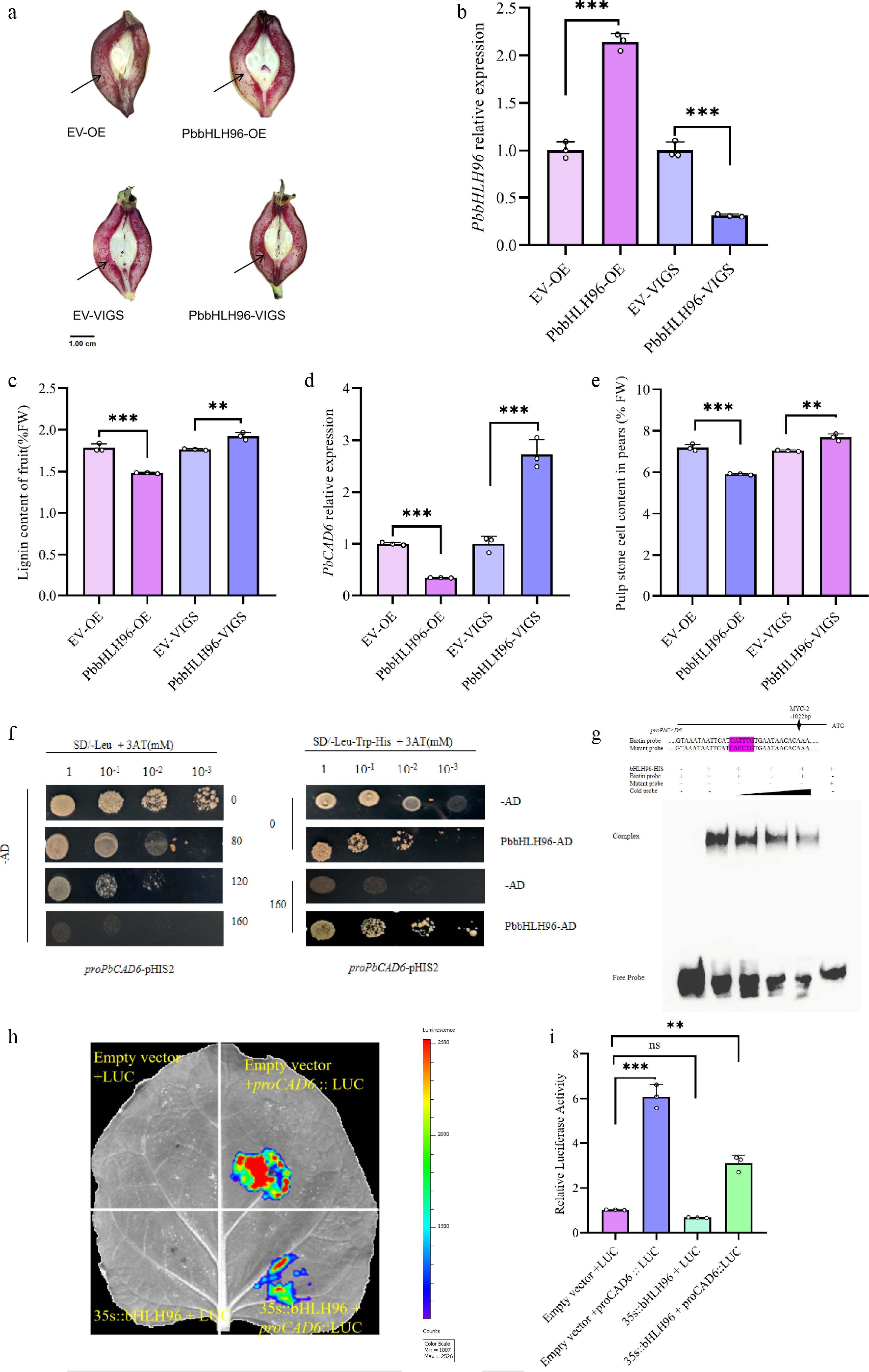
Figure 3.
PbbHLH96 regulates lignin biosynthesis in pear stone cells by binding to and repressing the PbCAD6 promoter. (a) Transient transformation of 'Shannongsu' pear fruits at 35 d after full bloom (DAFB) with PbbHLH96 overexpression and silencing constructs. Longitudinal fruit sections were stained with phloroglucinol-HCl and photographed 12 d after infiltration. qRT-PCR analysis of (b) PbbHLH96 and (c) PbCAD6 expression in transiently transformed fruits. (d) Lignin content measurement at the infiltration sites. (e) Determination of stone cell content in the infiltration site regions of pear fruits with empty pCAMBIA, empty pTRV, PbbHLH96 overexpression, and PbbHLH946 silencing. (f) Yeast one-hybrid assay showing the interaction between PbbHLH96 and the PbCAD6 promoter. (g) Electrophoretic mobility shift assay (EMSA) confirming that PbbHLH96 binds in vitro to the MYC cis-element within the PbCAD6 promoter. (h), (i) Luciferase reporter assay demonstrating that PbbHLH96 represses PbCAD6 promoter activity. Values represent the mean ± SD of three independent biological replicates. Statistical analysis was performed using ANOVA and Tukey's test (** p < 0.01; *** p < 0.001).
Cinnamyl alcohol dehydrogenase (CAD) is the key terminal enzyme in the lignin monomer biosynthesis pathway. Our previous study indicated that exogenous calcium treatment significantly suppresses PbCAD6 expression. We therefore investigated whether PbbHLH96 is involved in regulating PbCAD6. qRT-PCR analysis revealed that, compared to controls, PbbHLH96 overexpression in both transiently transformed fruits and stable overexpression calli significantly inhibited PbCAD6 expression, whereas PbbHLH96 silencing markedly upregulated PbCAD6 (Fig. 3d; Supplementary Fig. S2d).
To determine whether PbbHLH96 directly binds to the PbCAD6 promoter and regulates its expression, we conducted a yeast one-hybrid (Y1H) assay. Under optimized screening conditions (medium containing 180 mM 3-AT), PbbHLH96 was confirmed to bind directly to the PbCAD6 promoter (Fig. 3f). Furthermore, an electrophoretic mobility shift assay (EMSA) showed that PbbHLH96 binds to the MYC cis-element within the PbCAD6 promoter. This binding was abolished by the addition of unlabeled cold probes or by mutation of the PbCAD6 promoter sequence (Fig. 3g). Finally, a luciferase reporter assay (LUC) confirmed that PbbHLH96 significantly represses the luciferase activity driven by the PbCAD6 promoter (Fig. 3h, i). These results demonstrate that PbbHLH96 directly binds to the MYC element in the PbCAD6 promoter region, repressing its transcription and thus negatively regulating lignin biosynthesis in pear fruit.
PbCML46 interacts with PbbHLH96 to enhance the repression of PbCAD6 expression
-
To analyze the functional consequence of the PbCML46–PbbHLH96 complex on the promoter of downstream PbCAD6, we performed a dual-luciferase reporter (DLR) assay. The results confirmed that PbbHLH96 represses the promoter activity of the key lignin biosynthetic gene PbCAD6. Notably, co-expression of PbCML46 with PbbHLH96 significantly enhanced this repressive effect (Fig. 4a, b), indicating that PbCML46 cooperatively augments the transcriptional repression exerted by PbbHLH96 on PbCAD6.
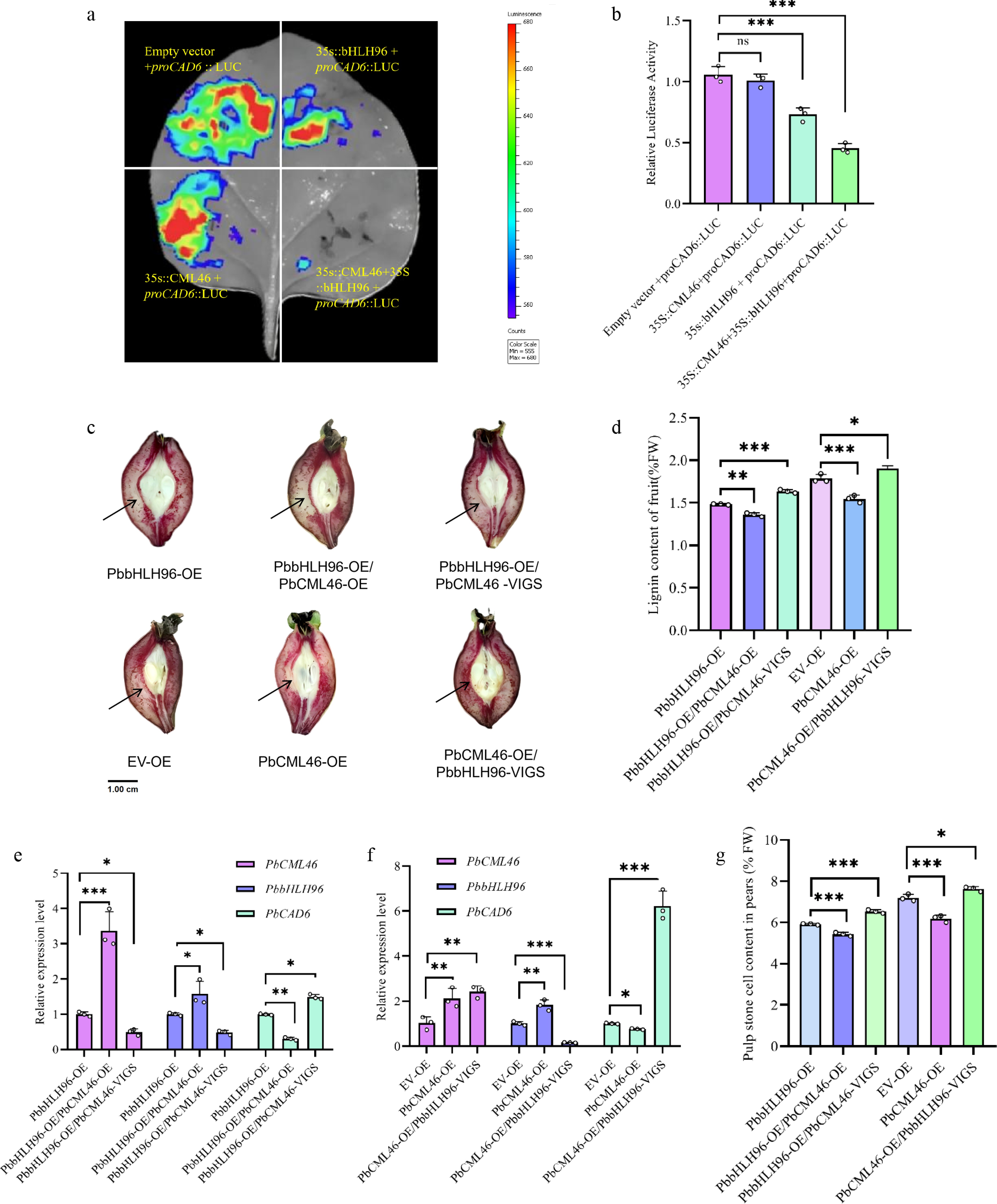
Figure 4.
PbCML46 interacts with PbbHLH96 to enhance its regulatory effect on PbCAD6 and inhibit lignin biosynthesis in pear stone cells. (a) Dual-luciferase reporter (DLR) assay showing that the interaction between PbCML46, and PbbHLH96 enhances the repression of PbCAD6 promoter activity by PbbHLH96. (b) Quantitative analysis of the DLR assay shown in (a). (c) Transient co-transformation in pear fruit via co-injection of overexpression and silencing constructs. Longitudinal fruit sections were stained with phloroglucinol-HCl and photographed 12 d after infiltration. (d) Lignin content measurement (e), (f) qRT-PCR analysis of PbCAD6, PbCML46 and PbbHLH96 expression in the infiltrated areas. (g) Determination of stone cell content in the infiltration regions of pear fruits after instantaneous injection. Values represent the mean ± SD of three independent biological replicates. Statistical analysis was performed using ANOVA (* p < 0.05; ** p < 0.01; *** p < 0.001).
To further validate their functional interplay in planta, we employed a transient transformation system in pear fruit. Compared to PbbHLH96 overexpression alone, co-overexpression of PbCML46 and PbbHLH96 significantly suppressed lignin accumulation in stone cells at the infiltration site of pear fruits, as evidenced by reduced histochemical staining intensity of stone cells surrounding the infiltration site in fruit sections, as well as significantly decreased lignin content and stone cell content in the infiltrated region. Co-expression of PbbHLH96 overexpression with PbCML4 silencing resulted in significantly increased lignin and stone cell contents compared with PbbHLH96 overexpression alone (Fig. 4c, d, g), qRT-PCR analysis revealed that, compared with overexpression of PbbHLH96 alone, co-overexpression of PbCML46 and PbbHLH96 significantly reduced the transcript level of PbCAD6, whereas co-expression of PbbHLH96 overexpression with PbCML46 silencing led to a significant upregulation in the relative expression level of PbCAD6 (Fig. 4e).
Notably, compared with the empty vector control, overexpression of PbCML4 alone reduced both lignin and stone cell contents in pear fruits (Fig. 4c, d, g) and significantly downregulated the relative expression level of PbCAD6 in the infiltrated region of pear fruits (Fig. 4f) . However, co-expression of PbCML46 overexpression with PbbHLH96 silencing abolished this suppression, leading to significantly elevated lignin and stone cells deposition. qRT-PCR analysis showed that the relative expression level of PbCAD6 in the infiltrated region of pear fruits co-expressing PbCML46 overexpression with PbbHLH96 silencing was significantly higher than that in fruits overexpressing PbCML46 alone (Fig. 4c, d, f, g). These results demonstrate that PbCML46 significantly enhances the repression of PbCAD6 specifically through its interaction with PbbHLH96, while PbCML46 itself does not directly regulate PbCAD6 transcription. Furthermore, the PbCML46-PbbHLH96 complex further reduces stone cell formation and lignin accumulation in pear fruits. This finding provides novel mechanistic insight into the molecular regulatory network controlling lignin synthesis.
-
The formation of stone cells in pear fruit is a key factor limiting its eating and commercial quality, the essence of which is the aberrant deposition and polymerization of lignin in parenchyma cells[6,28]. Although extensive literature documents that exogenous calcium treatment can effectively inhibit lignin biosynthesis in pear fruit stone cells[14,16,12,29], the molecular mechanism underlying calcium signal regulation of lignin biosynthesis in pear stone cells remains unclear. This study deciphers the molecular pathway through which exogenous calcium treatment regulates lignin synthesis in pear fruit stone cells via the PbCML46-PbbHLH96 module, providing new insights into the molecular basis of calcium signaling-mediated regulation of lignin deposition in pear fruit.
Calcium ions serve as ubiquitous secondary messengers involved in regulating various physiological processes during plant growth, development, and biotic stress responses[19,30]. The transmission and decoding of calcium signals depend on calcium-binding proteins[31,32]. CMLs, as a crucial class of calcium signal sensors, play a central role in decoding calcium signals and relaying them to downstream targets[20,33]. In this study, to elucidate the mechanism by which PbCML46 responds to calcium to inhibit stone cell lignin synthesis, we performed a Y2H screen to identify PbCML46-interacting proteins. Through Y2H, pull-down, and LCI assays, the bHLH family transcription factor PbbHLH96 was identified as an interaction partner of PbCML46 (Fig. 2). Notably, previous studies have shown that CMLs can influence lignin biosynthesis through diverse mechanisms. In pear fruit, CML38 interacts with WRKY46, modulating lignin-related physiological disorders via the downstream NAC187-CCR module[34]. In moso bamboo, the calmodulin-like protein PeCML4 interacts with the calcium-dependent protein kinase PeCDPK32 to cooperatively activate the activity of the key lignin polymerization peroxidase PePRX2, thereby promoting lignin monomer polymerization[35]. Furthermore, overexpression of PbCML3 increases stone cell content in pear fruit[29]. Whereas PbCML46 described herein exerts an inhibitory effect. Such functional diversity likely reflects differences in downstream interacting partners and the signaling pathways involved.
The bHLH family represents an important component of the plant lignin biosynthesis regulatory network, modulating lignin synthesis through interactions with other proteins and in response to hormonal and environmental signals. In rice, OsbHLH034 interacts with the jasmonate signaling repressor OsJAZ9 to subsequently activate downstream lignin synthesis-related genes[24]. Conversely, heterologous expression of sorghum SbbHLH1 in Arabidopsis inhibits lignin production[23]. In pear, numerous studies have demonstrated the involvement of MYB transcription factors, such as PbMYB24, PbMYB169, PbMYB6, and PbMYB308, in the regulation of lignin biosynthesis[10,26,36,37]. The NAC transcription factor PpNAC187 also regulates lignin synthesis[38]. However, bHLH transcription factors have not been reported in this context. In this study, we found that the bHLH transcription factor gene PbbHLH96 is a negative regulator of lignin biosynthesis in pear fruit stone cells. Following exogenous calcium treatment, PbbHLH96 expression was upregulated, concomitant with reduced stone cell and lignin contents (Supplementary Fig. S2), suggesting its involvement in calcium-mediated inhibitory regulation. Transient transformation assays showed that overexpression of PbbHLH96 significantly decreased the transcript level of PbCAD6, a key lignin biosynthesis gene, and reduced lignin accumulation, stone cell content, and stone cell distribution in fruit sections, whereas silencing PbbHLH96 produced opposite effects. Yeast one-hybrid (Y1H), electrophoretic mobility shift assay (EMSA), and dual-luciferase reporter (LUC) assays demonstrated that PbbHLH96 specifically binds to the MYC element (CATTTG) in the PbCAD6 promoter and represses its transcriptional activity, establishing its function as a transcriptional repressor (Fig. 3 and Supplementary Fig. S2). This preliminarily defines PbbHLH96 as a transcriptional repressor. However, whether PbbHLH96 regulates other lignin biosynthesis genes and whether its repressive activity is subject to post-translational modification requires further investigation.
Previous studies have shown that WRKY-CaM/CML interactions can influence the transcriptional activity of WRKY factors within calcium signaling pathways[21,39,40]. Furthermore, the PpCML38-PpWRKY46 interaction inhibits the transcriptional activity of PpWRKY46 in the calcium signaling pathway[34]. In this study, we found that PbCML46 interacts with PbbHLH96 which does not involve co-binding to downstream DNA. Instead, PbCML46 significantly enhances the ability of PbbHLH96 to transcriptionally repress PbCAD6, thereby negatively regulating lignin biosynthesis in pear fruit stone cells. This conclusion was confirmed through dual-luciferase reporter (LUC) assays and co-expression validation in pear fruit (Fig. 4). This indicates that the PbCML46-PbbHLH96 complex synergistically regulates PbCAD6 to inhibit lignin biosynthesis in pear stone cells. PbCML46 itself does not possess direct transcriptional regulatory activity, and its negative regulatory function in lignin biosynthesis in pear fruit stone cells is strictly dependent on the presence of PbbHLH96. Based on these findings, we propose a working model for the regulation of lignin biosynthesis in pear fruit stone cells by exogenous calcium through the PbCML46–PbbHLH96 module (Fig. 5). Upon entry of exogenous calcium into the cell, cytosolic Ca2+ concentration increases. On one hand, this induces PbbHLH96 expression, and PbbHLH96 directly binds to the MYC cis-element in the PbCAD6 promoter to repress its expression, thereby inhibiting lignin and stone cell synthesis. On the other hand, PbCML46 is activated upon Ca2+ binding, undergoes conformational changes, and interacts with PbbHLH96 to further enhance the repression of PbCAD6 expression, thereby inhibiting lignin and stone cell accumulation. Additionally, PbCML46 may upregulate PbbHLH96 expression through a feedback mechanism, creating a cascade amplification effect that ultimately leads to robust suppression of lignin biosynthesis and stone cell deposition. This model elucidates the molecular mechanism by which calcium signaling fine-tunes lignin metabolism in pear fruit stone cells through the PbCML46–PbbHLH96 module, providing a new theoretical foundation for the improvement of pear fruit quality. Despite the key role of the PbCML46–PbbHLH96 module in negatively regulating lignin biosynthesis in pear fruit stone cells, several limitations remain to be addressed. First, the interaction domains and critical amino acid residues mediating the PbCML46–PbbHLH96 interaction have not been identified, and the impact of varying Ca2+ concentrations on their binding affinity requires quantitative analysis. Second, the precise mechanism by which PbCML46 enhances the transcriptional repressive activity of PbbHLH96 remains unclear, whether through stabilization of the PbbHLH96 protein, promotion of its nuclear localization, recruitment of co-repressor complexes, or induction of conformational changes that enhance DNA-binding affinity, warrants further investigation using biochemical and cell biological approaches. Third, while this study focused primarily on PbCAD6, whether the complex regulates additional lignin biosynthesis genes (e.g., PbCCR, PbLAC) remains to be explored; genome-wide approaches such as ChIP-seq or transcriptomic analysis will be valuable for defining the full downstream regulatory network. Fourth, the conclusions of this study are largely based on transient transformation in pear fruits and stable transformation in callus; whether this regulatory module is conserved in other horticultural crops and whether it exhibits dose-dependent responses to different calcium signal intensities remains to be validated.
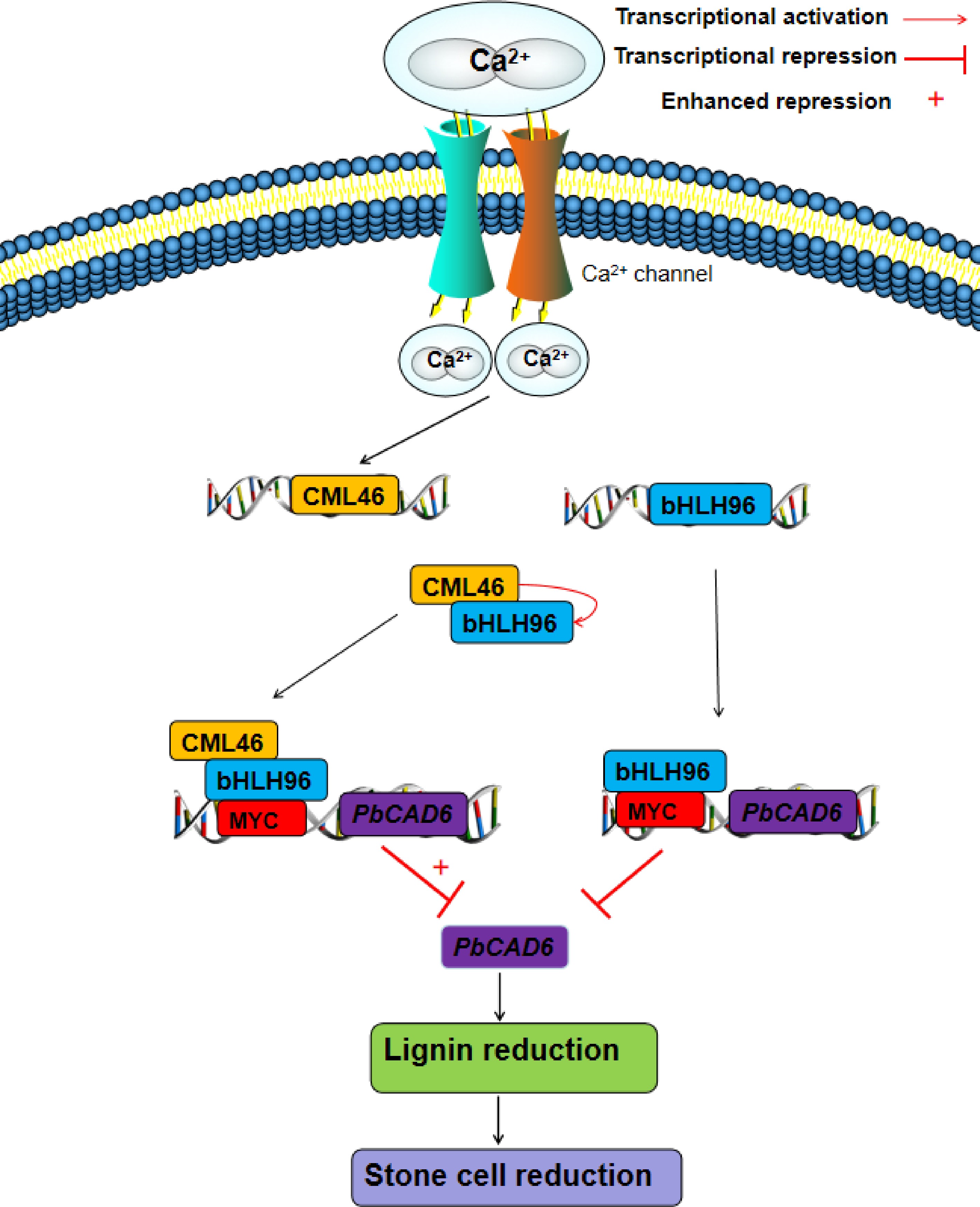
Figure 5.
A proposed regulatory model for the roles of PbCML46 and PbbHLH96 in modulating lignin biosynthesis within stone cells in response to Ca2+ signals in 'Shannongsu' pear. Upon the entry of exogenous calcium ions into the cell, the cytosolic Ca2+ concentration increases. On one hand, this directly induces the expression of PbbHLH96, which in turn binds to the MYC cis-element in the PbCAD6 promoter and represses its expression, thereby inhibiting lignin biosynthesis and stone cell formation. On the other hand, PbCML46 is activated upon Ca2+ binding, undergoing a conformational change that enables its interaction with PbbHLH96, further potentiating the inhibitory effect of PbbHLH96 on downstream PbCAD6 expression, and consequently suppressing lignin and stone cell accumulation. Furthermore, PbCML46 may upregulate PbbHLH96 expression through a feedback mechanism, forming a cascade amplification effect that ultimately leads to a pronounced inhibition of lignin biosynthesis and stone cell formation. This model elucidates the molecular mechanism by which calcium signaling finely regulates lignin metabolism in pear fruit stone cells through the PbCML46-PbbHLH96 module, providing a new theoretical foundation for the improvement of pear fruit quality.
In summary, this study elucidates the molecular mechanism by which calcium inhibits lignin biosynthesis in pear stone cells through the PbCML46-PbbHLH96 regulatory module. Our findings provide a theoretical basis for the application of exogenous calcium in improving fruit quality.
This work was supported by the Key R&D Program of Shandong Province, China (Grant No. 2023LZGC015), the National Natural Science Foundation of China (Grant No. 32572980), and the Taishan Scholar Climbing Project (Grant No. tspd20240811).
-
The authors confirm contributions to the paper as follows: study conception and design, supervision, funding acquisition: Zhang Z, Chen X; experimental guidance: Qi S, Zhu Y, Yu H; material processing: Wang N, Chen X; data analysis: Wang N, Chen X, Li A; experimental work: Li A, Cao R; material management: Cao R; draft manuscript preparation: Li A. All authors reviewed the results and approved the final version of the manuscript.
-
Data supporting the findings of this study are available from the corresponding author upon reasonable request. All relevant data are included within the manuscript and its supplementary materials.
-
The authors declare that there is no conflict of interest.
-
accompanies this paper online at: https://doi.org/10.48130/frures-0026-0011.
- Supplementary Table S1 Sequences for the gene-specific primers used in this work.
- Supplementary Fig. S1 Yeast two-hybrid interaction assay.
- Supplementary Fig. S2 Relative expression levels of PbbHLH96 in pear flesh at different developmental stages following calcium polyol (SAC) treatment.
- Supplementary Fig. S3 Analysis of PbbHLH96-overexpressing transgenic callus of 'Shannongsu' pear.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Li A, Qi S, Zhu Y, Yu H, Wang N, et al. 2026. Ca2+ mediated PbCML46-PbbHLH96 module negatively regulates lignin biosynthesis in pear fruit. Fruit Research 6: e015 doi: 10.48130/frures-0026-0011
Ca2+ mediated PbCML46-PbbHLH96 module negatively regulates lignin biosynthesis in pear fruit
- Received: 16 March 2026
- Revised: 25 March 2026
- Accepted: 01 April 2026
- Published online: 15 April 2026
Abstract: Stone cell formation, resulting from aberrant lignin deposition in parenchyma cells, is a key determinant of pear fruit quality. Although exogenous calcium application is known to inhibit lignin biosynthesis and stone cell development, the underlying molecular mechanism involving calcium sensor proteins remains poorly understood. This study aimed to elucidate the molecular pathway by which calcium signaling modulates lignin biosynthesis. Our findings demonstrate that the calmodulin-like protein PbCML46 significantly suppresses lignin accumulation. This function was validated through complementary approaches, including transient injection in pear fruit and stable overexpression in pear calli. A yeast two-hybrid screen revealed that PbCML46 specifically interacts with the bHLH transcription factor PbbHLH96, an interaction further confirmed by pull-down and luciferase complementation imaging (LCI) assays. Further yeast one-hybrid (Y1H), electrophoretic mobility shift assay (EMSA), and luciferase (LUC) reporter assays analysis showed that PbbHLH96 directly binds to the MYC cis-element in the PbCAD6 promoter and functions as a transcriptional repressor. Dual-luciferase reporter (DLR) and in vivo co-expression assays indicated that PbCML46 markedly enhances the repressive activity of PbbHLH96 on the PbCAD6 promoter, an effect strictly dependent on PbbHLH96. In conclusion, this study reveals a Ca2+-initiated regulatory cascade in which calcium signaling promotes the interaction between the sensor PbCML46 and the transcriptional repressor PbbHLH96, leading to cooperative suppression of the key lignin biosynthetic gene PbCAD6 and thereby negatively regulating lignin deposition in pear stone cells. These findings provide novel mechanistic insights into how calcium signaling improves fruit quality.
-
Key words:
- Pear /
- PbCML46 /
- PbbHLH96 /
- Lignin biosynthesis /
- Transcriptional repression /
- Calcium signaling













