-
Scrophularia ningpoensis Hemsl., a perennial herb of the genus Scrophularia (Scrophulariaceae), is an important source of traditional Chinese medicinal materials. Its dried roots are rich in bioactive compounds, including iridoid glycosides and phenylpropanoids, which exhibit anti-inflammatory, antioxidant, and neuroprotective effects, making the species valuable for treating neurological and inflammation-related disorders[1,2]. With the expansion of the traditional Chinese medicine industry, demand for high-quality S. ningpoensis seedlings is increasing. The species is mainly propagated vegetatively via bud-bearing rhizomes. Though this preserves genetic uniformity, long-term vegetative propagation can lead to germplasm degradation and pathogen accumulation. Its low propagation coefficient further limits the large-scale, stable production of high-quality medicinal material. In vegetatively propagated medicinal plants, in vitro culture provides a reliable approach to generate healthy, pathogen-free material, mitigating pathogen buildup, maintaining the germplasm's integrity, and enabling rapid multiplication[3,4]. Establishing an efficient, stable, and scalable in vitro regeneration system is therefore essential for germplasm conservation, pathogen-free propagation, and industrial-scale seedling production. Despite extensive pharmacological research on S. ningpoensis, tissue culture studies remain limited. Reports on Scrophularia species are largely restricted to S. nodosa and S. yoshimurae, which have shown that careful selection of the explant type and optimization of plant growth regulator (PGR) treatments can induce callus formation, adventitious rooting, and shoot regeneration[5,6]. Systematic studies on S. ningpoensis are scarce, particularly quantitative comparisons of differentiation responses across explant types under multilevel PGR combinations. Moreover, no studies have applied data-driven approaches to quantitatively analyze and predict its in vitro regeneration.
In vitro regeneration depends on the dedifferentiation and redifferentiation capacity of explants, with explant origin being a major determinant of regeneration efficiency and organ-specific differentiation[7,8]. Explants from different organs differ in their state of cell differentiation, endogenous hormone profiles, and regeneration potential, producing distinct organogenic tendencies[9−11]. For example, stem or petiole explants with well-developed vascular tissues typically favor adventitious shoot formation, whereas leaf explants preferentially form roots, reflecting common organ-specific differentiation patterns[12−14]. Exogenous PGRs, particularly the type, concentration, and ratio of auxins and cytokinins, are key regulators of cell fate and organogenesis, modulating the expression of transcription factors and meristem establishment to determine the direction and efficiency of regeneration[15−17]. Under multifactorial conditions, however, explant responses often exhibit nonlinear and interactive effects, complicating accurate modeling and prediction, increasing experimental costs, and limiting quantitative interpretation and prediction of culture systems.
Tissue culture optimization has traditionally relied on single-factor experiments or linear statistical methods, such as analysis of variance (ANOVA) or regression, which identify significant factors but cannot capture complex nonlinear interactions among multiple variables[18,19]. In practice, explant type, hormone concentration, and culture duration often interact synergistically to determine regeneration outcomes, limiting the predictive power of conventional approaches. Methods that can account for such multivariate interactions are therefore valuable for improving the quantitative analysis and prediction of tissue culture responses. Machine learning (ML) offers a data-driven alternative capable of modeling nonlinear, interactive effects without predefined functional assumptions[18]. ML not only predicts regeneration outcomes accurately but also quantifies the relative contribution of each factor through feature importance analyses, facilitating the identification of key regulators[20,21]. ML has shown promise in medium optimization, callus induction, and micropropagation, improving efficiency while reducing experimental costs[22,23]. Among ML approaches, random forest is well suited to tissue culture datasets because it can capture nonlinear relationships, accommodate complex interactions among predictors, and provide interpretable estimates of variable importance. Recent reviews have also highlighted the growing use of ML approaches, including random forest, in plant tissue culture optimization and prediction studies[21,24]. Despite its potential, ML remains underutilized in nonmodel medicinal plants, and systematic studies integrating organ-specific differentiation responses for predictive modeling of culture responses remain scarce.
Research on S. ningpoensis tissue culture remains limited, particularly regarding organ-specific responses to hormone combinations, and no studies have applied ML for quantitative analysis and predictive modeling of its in vitro regeneration. In this study, leaf and petiole explants of S. ningpoensis were used to systematically evaluate in vitro differentiation under varying combinations of 6-benzylaminopurine (6-BA) and naphthaleneacetic acid (NAA). By integrating conventional statistical analysis with a random forest ML model, we aimed to: (1) establish a stable, high-efficiency regeneration system; (2) characterize organ-specific differentiation patterns under different hormone conditions; (3) develop a predictive model for quantitative interpretation and condition ranking within the tested experimental space; and (4) assess the applicability of ML as a complementary approach for analyzing tissue culture responses.
-
Aseptic S. ningpoensis seedlings were obtained in July 2024 from the Dexing Research and Training Center of Chinese Medical Sciences (Dexing, China) and identified by on of the researchers (Wang Sheng). Seedlings were maintained as in vitro stock cultures on Murashige and Skoog (MS) basal medium (30 g/L sucrose, 7.6 g/L agar, pH 5.8) under a 16 h/8 h light/dark cycle at 25 ± 1 °C and 60%−70% relative humidity at the Dexing Research and Training Center of Chinese Medical Sciences, and served as the explant source for all regeneration experiments.
Explant regeneration
-
Leaf and petiole explants were excised from healthy S. ningpoensis seedlings of the third to fourth subculture generation (~8 to 10 cm tall). Petiole segments (~1 cm) and leaf pieces (~5 × 5 mm) were standardized to minimize size-related variability, briefly immersed in sterile water, and inoculated within 1 h. Explants were cultured on MS basal medium (PhytoTechnology Laboratories, USA) supplemented with 6-BA (0−2 mg/L) and NAA (0−1.5 mg/L). Agar (Beijing Dingguo Changsheng Biotechnology Co., Ltd., China) and sucrose (Gu'an County Jingmi Chemical Co., Ltd., China) were added at 7.6 and 30 g/L, respectively. The medium's pH was adjusted to 5.8, and it was autoclaved at 121 °C for 20 min. Explants were maintained in the dark at 25 ± 1 °C and 60%−70% relative humidity.
Callus formation, shoot regeneration, and rooting were evaluated visually. Regeneration rates were calculated as regeneration (%) = (number of explants forming the respective organ / total explants) × 100. Observations were made at 30 and 60 days post-inoculation, and only clearly distinguishable new calli, shoots, or roots were counted. Each hormone combination was applied to both leaf and petiole explants, with three biological replicates of 10 explants per treatment.
Data processing and statistical analysis
-
Data were processed in Python 3.14.2 (Python Software Foundation, USA) with the random seed set to 123[25]. Samples with missing key indicators were excluded, resulting in 162 complete datasets (2 explant types × 28 treatments × 3 replicates). Continuous variables (6-BA, NAA, culture duration, and differentiation rates) were verified and converted to numeric format. Descriptive statistics were performed using the pandas and NumPy tools. Ordinary least squares (OLS) regression and factorial ANOVA were applied to evaluate the effects of explant type, culture duration, hormone concentrations, and their interactions on differentiation.
ML modeling and feature engineering
-
ML modeling was performed to quantitatively analyze shoot and root regeneration under different culture conditions. After exclusion of incomplete observations, 162 complete data points were retained for modeling. The response variables were shooting rate and rooting rate. Callus formation was excluded from the model because it occurred readily in both explant types and generally remained high across treatments, thereby providing limited discriminatory power for regression-based comparison. By contrast, rooting and especially shoot regeneration represented more challenging developmental outcomes and were therefore considered more informative targets for predictive analysis.
The predictor set comprised 13 variables, including explant type, culture duration, 6-BA concentration, NAA concentration, and nine engineered descriptors representing hormone interactions, relative balance, total hormone level, and nonlinear transformations. Specifically, these descriptors included the 6-BA × NAA product term, BA/NAA and NAA/BA ratios, total hormone concentrations (6-BA + NAA), normalized BA and NAA ratios, log-transformed 6-BA and NAA terms, and the difference between log-transformed 6-BA and NAA values. These features were introduced because organogenic responses are often influenced not only by the absolute concentrations of auxin and cytokinin but also by their interaction strength, relative balance, and possible nonlinear effects. After one-hot encoding of explant type and culture duration, the final model input matrix contained 15 features.
Categorical variables, including explant type and culture duration, were encoded using one-hot encoding, whereas hormone concentrations and engineered descriptors were treated as continuous variables. The dataset was randomly divided into a training set (80%) and a test set (20%) using a fixed random seed of 123, and the same split indices were used for both the rooting and shooting models to ensure comparability. The split was performed at the individual observation level rather than by treatment combination, such that replicate-level records from the same treatment could be assigned independently to the training and test sets. The models' stability was evaluated using repeated five-fold cross-validation (three repeats) on the training set, followed by independent assessment on the held-out test set. Performance metrics included the coefficient of determination (R2), root mean square error (RMSE), and mean absolute error (MAE). Random forest regression was used as the primary model for prediction and factor ranking, and grid-based evaluation within the tested hormone range was performed to rank candidate rooting conditions within the tested experimental space.
-
To characterize the organ-specific regeneration potential of S. ningpoensis, leaf and petiole explants were evaluated for callus formation, shoot regeneration, and rooting under varying hormone combinations over 60 d. Leaf and petiole explants exhibited distinct organogenic biases under cytokinin–auxin treatments (Supplementary Tables S1 and S2). Leaf explants showed strong rhizogenic competence, with rooting frequencies increasing between 30 and 60 d under most hormone combinations and reaching a maximum of 96.7% ± 3.33% at 1.0 mg L−1 6-BA + 1.0 mg L−1 NAA after 60 d (Fig. 1a). In contrast, shoot regeneration was limited in leaves but robust in petiole explants, which achieved maximal shoot formation at 0.5 mg L−1 6-BA + 0.2 mg L−1 NAA (40.0% ± 10.0% at 30 d; 50.0% ± 11.55% at 60 d) (Fig. 1b). Both explant types readily formed calli across a broad range of hormone conditions (Fig. 1c), with callus proliferation predominating at 30 d and organ differentiation becoming more apparent by 60 d. Representative phenotypes under optimal regeneration conditions are shown in Fig. 1d. Together, these results reveal pronounced organ-specific regeneration competence in S. ningpoensis, with leaves preferentially adopting a rhizogenic fate and petioles favoring shoot organogenesis.
Effects of explant type and hormone concentrations on regeneration
-
Mean regeneration rates differed significantly between explant types (Fig. 2a; Supplementary Table S3). Petioles produced higher shoot regeneration at both 30 d (6.90% ± 3.42%) and 60 d (12.05% ± 3.28%) compared with leaves (0.51% ± 0.41% and 1.28% ± 0.81%, respectively). In contrast, leaves exhibited higher rooting rates at 30 d (32.31% ± 6.89%) and 60 d (43.33% ± 7.62%) than petioles (4.52% ± 1.18% and 19.76% ± 3.85%, respectively). Callus induction exceeded 85% in most treatments and was slightly higher in petioles.
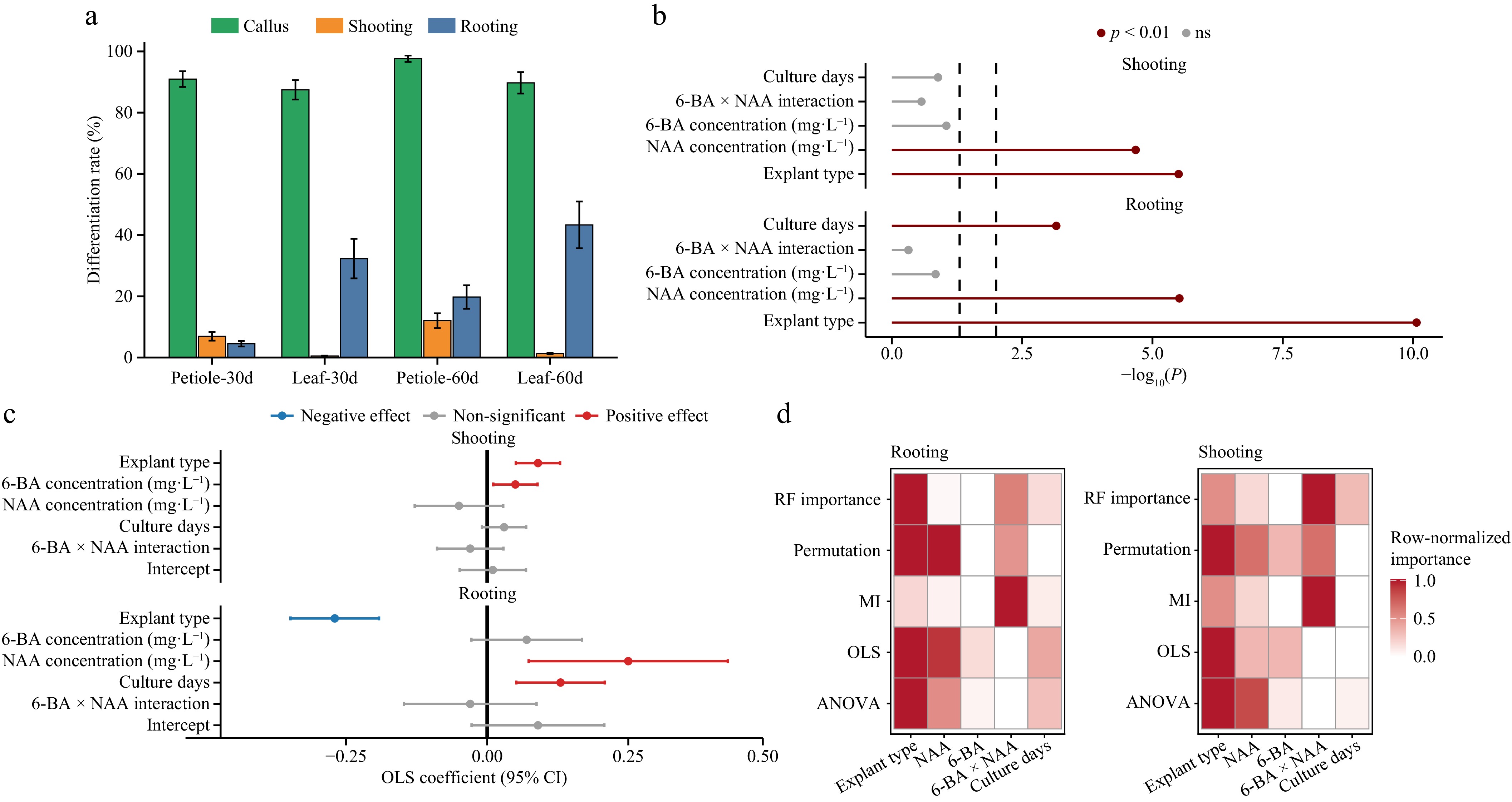
Figure 2.
Statistical effects of explant type and hormone combinations on differentiation rates. (a) Mean (± SE) differentiation rates of leaf and petiole explants cultured for 30 and 60 d. Callus induction (green), shoot formation (orange), and rooting (blue) are shown. (b) Forest plot of two-way ANOVA results. The x-axis represents −log10(P), and dashed vertical lines indicate significance thresholds. Colors correspond to p < 0.01 and nonsignificant results (ns). (c) Ordinary least squares (OLS) regression coefficients with 95% confidence intervals. Positive effects are shown in red, negative effects in blue, and nonsignificant effects in gray. The solid vertical line represents zero effect. (d) Consensus heatmap of variable importance across analytical methods (ANOVA, OLS, mutual information, permutation importance, and random forest). Columns represent the explanatory variables (explant type, NAA, 6-BA, 6-BA × NAA interaction, and culture duration). Color intensity indicates the row-normalized relative importance (0–1), with darker shades representing higher importance. All analyses were performed on three biological replicates (n = 3). Detailed statistical results are provided in Supplementary Tables S3−S7, and the multimethod consensus ranking is summarized in Supplementary Table S8.
Two-way ANOVA identified explant type and NAA concentration as significant factors affecting shoot regeneration (p < 0.01), whereas 6-BA showed a weaker effect (p = 0.09) (Fig. 2b; Supplementary Table S4). Rooting was significantly influenced by explant type, NAA concentration, and culture duration (p < 0.01), wherea 6-BA and the interaction terms were not significant (Fig. 2b; Supplementary Table S5). OLS regression analysis confirmed the strong contribution of explant type to both shoot and root formation (p < 0.001). A positive association between 6-BA concentration and shoot regeneration was detected (p = 0.048) (Fig. 2c; Supplementary Tables S6 and S7). A multimethod consensus analysis integrating ANOVA, OLS, mutual information, permutation importance, and random forest confirmed the dominant regulatory role of explant type (Fig. 2d; Supplementary Table S8).
ML modeling of regeneration outcomes
-
ML models were developed to quantitatively predict shooting and rooting responses from explant type, culture duration, 6-BA concentration, NAA concentration, and the engineered descriptors representing hormones' interaction and balance. The final predictor set comprised 13 variables and expanded to 15 encoded input features after one-hot encoding of explant type and culture duration. For rooting prediction, the random forest model achieved a cross-validation R2 of 0.59 and a test R2 of 0.78, with RMSE = 0.15 and MAE = 0.11. Extra Trees and Gradient Boosting models showed comparable performance (test R2 = 0.78 and 0.77, respectively). Predictive performance for shoot regeneration was lower, with the best random forest model reaching a test R2 of 0.57, indicating moderate predictive ability. This relatively lower performance may be related to the sparse and uneven distribution of shoot regeneration responses across treatments, with many observations concentrated at low values. Nevertheless, the ML models consistently outperformed the dummy baseline predictor for both targets (Fig. 3a, b; Supplementary Tables S9 and S10).
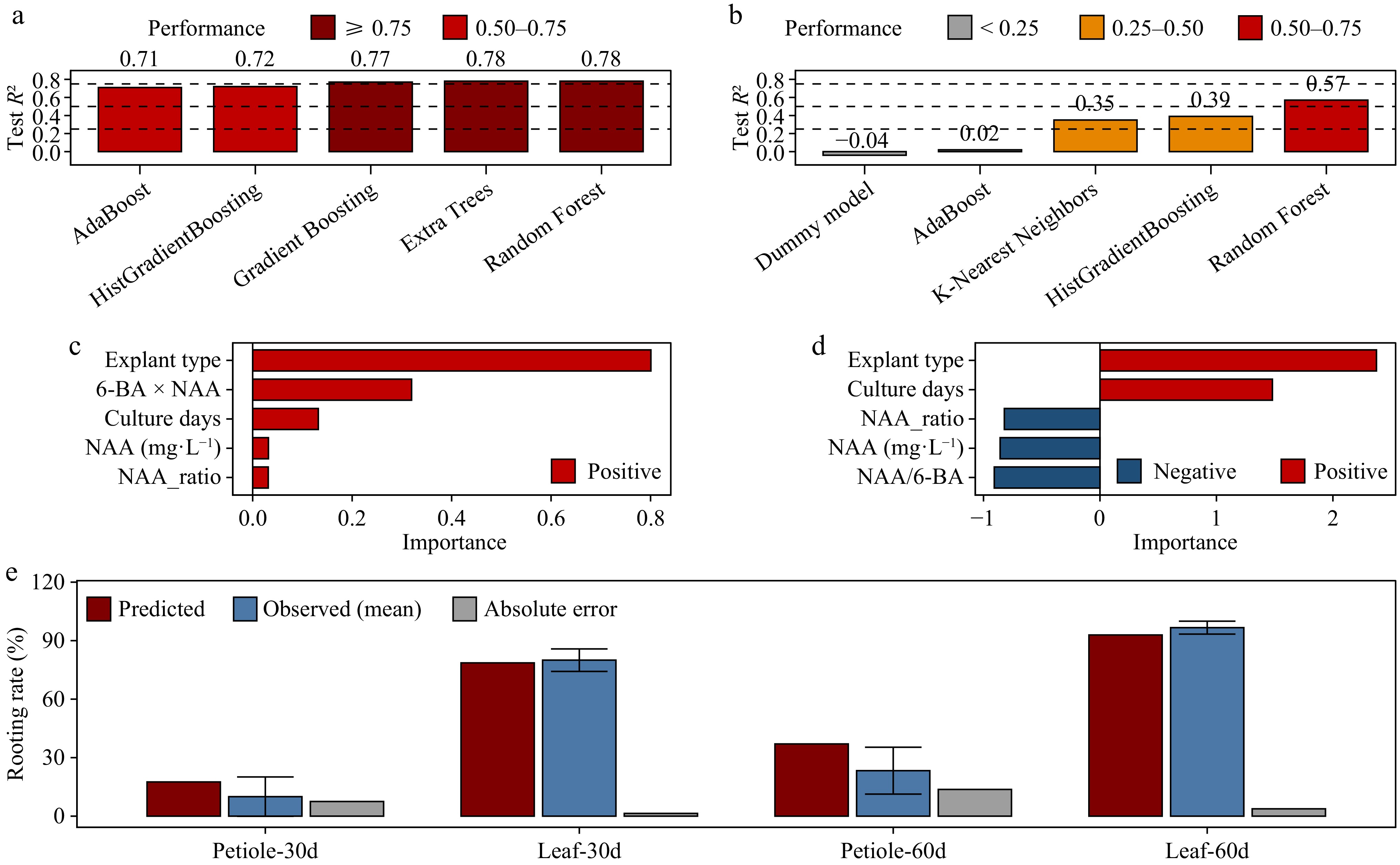
Figure 3.
Machine learning-based analysis of regeneration predictions and factor importance. (a) Test performance (R2) of optimized models for predicting rooting rates. Bars indicate the predictive performance on independent test sets, with colors representing performance categories. Dashed horizontal lines indicate model classification thresholds. (b) Test performance (R2) of optimized models for predicting shooting rates. Model types are shown on the x-axis, and the test R2 on the y-axis; colors indicate performance categories. (c) Top five features contributing to rooting rate predictions in the optimized random forest model. Bar length indicates relative importance; red bars denote positive contributions. (d) Top five features contributing to shooting rate predictions in the optimized random forest model. Red and blue bars indicate positive and negative contributions, respectively; bar length represents the relative effect size. (e) Comparison of predicted and observed rooting rates for model-ranked conditions within the tested experimental space. Red bars indicate predicted values, blue bars indicate observed means, and gray bars indicate absolute prediction errors. Error bars denote the SE of observed values. Treatment groups correspond to explant type and culture duration. For leaf explants, the highest-ranked rooting condition identified by the model corresponded to the best-performing treatment observed experimentally. For petiole explants, the displayed comparisons are based on the experimentally tested hormone combinations closest to the model-ranked conditions and therefore do not represent a prospective validation of untested predictions. All models were trained and evaluated on three biological replicates (n = 3). Detailed performance metrics, feature importance analyses, and predicted vs observed comparisons are provided in Supplementary Tables S8−S14.
Feature-importance analyses also indicated that explant identity was the dominant determinant of regeneration outcome, in agreement with the conventional statistical analyses. Mutual information analysis ranked the 6-BA × NAA interaction as the strongest contributor to rooting predictions, followed by explant type and log-transformed NAA concentration, whereas the type of explant showed the highest importance for shoot regeneration, with hormone interaction features contributing secondarily (Supplementary Tables S11−S13). Permutation importance analysis of the optimized random forest model yielded a similar ranking pattern (Fig. 3c, d), supporting the robustness of the main factor hierarchy identified across analytical approaches. The difference in predictive performance between the rooting and shooting models also suggests that prediction errors were more broadly distributed for shoot regeneration than for rooting, consistent with the lower overall model fit for shoot responses.
Model-based ranking of candidate conditions within the tested experimental space
-
To further evaluate the models' behavior within the tested experimental space, the best-performing regression model (random forest) was used to rank candidate hormone conditions for rooting (Fig. 3e). The highest-ranked rooting condition was leaf explants cultured for 60 d under 1.0 mg L−1 6-BA + 1.0 mg L−1 NAA (Supplementary Table S14), which was the same as the best-performing treatment identified in the factorial experiment. Under this condition, the predicted rooting rate was 92.90%, compared with an observed value of 96.67% ± 3.33%, indicating close agreement between the model's prediction and experimental observations. At 30 d under the same hormone combination, the predicted rooting rate (78.60%) was consistent with the observed value (80.00% ± 5.77%). For petiole explants, the model-ranked rooting responses were substantially lower, consistent with the overall experimental trend, although the corresponding model-ranked conditions were not prospectively validated. Overall, these results indicate that the random forest model effectively captured the relative response pattern of rooting within the tested parameter space and provided a useful basis for condition ranking and quantitative interpretation.
-
This study characterizes the organ-dependent regenerative behavior of S. ningpoensis, establishes a reproducible in vitro regeneration platform, and demonstrates that ML can serve as a useful complementary framework for quantitative interpretation and predictive analysis of regeneration responses within the tested experimental space. By integrating conventional statistical inference with random forest modeling, this work provides both practical value for regeneration protocol development and a quantitative analytical framework that may support future optimization studies in other nonmodel medicinal plants.
Explant origin emerged as the dominant determinant of regenerative trajectory. Consistent with prior reports[26−28], tissues derived from distinct organs exhibit divergent developmental outcomes during in vitro regeneration, reflecting intrinsic differences in cellular composition, developmental status, and hormonal responsiveness. In the present study, petiole explants showed a stronger capacity for shoot regeneration, whereas leaf explants exhibited a marked tendency toward rooting, indicating clear organ-specific differences in regenerative competence[29,30].
Auxin–cytokinin balance is widely recognized as a major regulator of regeneration polarity[31,32], but its functional impact depends on local signaling architecture. In model species, auxin transport, local auxin accumulation, and the associated signaling pathways have been implicated in root initiation and regeneration-related developmental reprogramming[33,34]. These observations may provide a possible framework for interpreting the stronger rooting response of leaf explants in S. ningpoensis; however, no direct molecular or physiological evidence is currently available in this species.
Likewise, regeneration-related regulatory pathways identified in model plants, including LATERAL ORGAN BOUNDARIES DOMAIN (LBD)- and WUSCHEL-related homeobox (WOX)-associated networks, have been shown to participate in root organogenesis and meristem establishment[35−37]. Although these pathways may be relevant to organ-specific regeneration in S. ningpoensis, their involvement remains hypothetical and will require future experimental validation. By contrast, the stronger shoot regeneration observed in petiole explants may reflect differences in tissue organization and developmental competence that favor shoot organogenesis under cytokinin-containing conditions. Overall, the organ-specific differentiation patterns observed in S. ningpoensis are likely associated with intrinsic differences in tissue identity and hormone responsiveness, although the underlying molecular basis remains to be determined.
From an analytical perspective, classical statistical models (ANOVA and OLS regression) identified explant type and NAA concentration as significant determinants of regeneration outcomes, with explant type exerting the strongest influence. These findings (Fig. 2) were fully consistent with the variable importance rankings generated by the random forest model (Fig. 3c, d), reinforcing the biological validity of the conclusions. However, linear statistical frameworks are inherently limited in capturing higher-order interactions among explant identity, hormone concentration, and hormonal ratios, factors that likely regulate regeneration in a nonlinear and synergistic manner[18,19]. ML approaches, including random forest, provide flexible frameworks for modeling complex relationships among tissue culture variables and for evaluating the relative importance of predictors, and have increasingly been applied in plant tissue culture optimization and regeneration studies[21,38].
The random model showed stronger predictive performance for rooting (R2 = 0.78), whereas the prediction of shoot regeneration was more moderate (R2 = 0.57) and should be interpreted more cautiously. In addition, the lower cross-validation performance relative to the held-out test result for rooting suggests that some degree of performance variability cannot be excluded, likely owing to the limited sample size and variation introduced by data partitioning. The broad agreement between ML-derived feature importance and conventional statistical analyses suggests that the model captured biologically relevant patterns in the dataset while providing an additional quantitative perspective on factor ranking. Within the tested experimental space, model-based ranking identified the same rooting condition as the best-performing treatment observed experimentally, supporting the practical value of the approach for quantitative interpretation rather than prospective optimization. Under 1.0 mg/L 6-BA and 1.0 mg/L NAA for 60 d, the predicted rooting rate for leaf explants (92.90%) closely approximated the observed value (96.7%), indicating that the model captured key regulatory features of the regeneration landscape. Such agreement between the predicted and observed values suggests that ML may be useful for supporting condition prioritization and quantitative interpretation in future protocol refinement[23]. The comparatively lower cross-validation performance for shoot induction likely reflects the limited sample size and skewed distribution of shooting rates, which may restrict the learning efficiency for extreme-value observations. Additionally, the current model was built exclusively on phenotypic and exogenous culture parameters and did not incorporate endogenous hormonal or molecular regulatory variables, potentially constraining predictive depth. Future model development may benefit from expanded datasets, incorporation of biologically informed composite features (e.g., hormone ratios), and advanced model tuning strategies. Prediction uncertainty may also have been influenced by the relatively narrow hormone concentration ranges tested and by unavoidable differences in the physiological status of explants at the time of inoculation, both of which may contribute additional biological variation. For shoot regeneration in particular, predictive performance may be improved in future studies by incorporating additional descriptors related to explants' physiology, endogenous hormone status, or image-based phenotypic traits, as well as by evaluating alternative ensemble or hybrid modeling strategies. Integration of ML with molecular or multiomics datasets may also improve biological interpretability and predictive robustness.
In summary, this study established an efficient and reproducible in vitro regeneration system for S. ningpoensis and revealed clear organ-specific differences in rooting and shoot regeneration between leaf and petiole explants. Within the tested experimental space, conventional statistical analyses and random forest modeling consistently identified explant type as the dominant factor influencing regeneration outcomes, whereas the highest-ranked rooting condition predicted by the model corresponded to the best-performing treatment observed experimentally. Although the present study focused on S. ningpoensis, the analytical framework used here may also provide a useful reference for quantitative optimization of tissue culture systems in other nonmodel medicinal plants with similarly complex organ-specific regeneration responses. These findings indicate that ML can serve as a useful complementary framework for quantitative interpretation and predictive analysis of tissue culture responses, although its extrapolative value beyond the tested hormone combinations will require future prospective validation.
This research was funded by the National Key Research and Development Program of China (2023YFC3503801); the Fundamental Research Funds for the China Academy of Chinese Medical Sciences (JBKY2025A02); the China Agriculture Research System (CARS-21), funded by the Ministry of Finance and the Ministry of Agriculture and Rural Affairs; and the Beijing Nova Program for Science and Technology Innovation.
-
The authors confirm their contributions to this study as follows: conceptualization, writing – review and editing: Liu Y, Wang S; methodology, formal analysis: Liu Y, Song J; software and ML modeling, writing – original draft preparation: Liu Y; investigation: Liu Y, Song J, Shi K, Zhu H, Dai X; supervision: Wang S, Zhang C; funding acquisition: Wang S. All authors reviewed the results and approved the final version of the manuscript.
-
The data supporting the findings of this study are contained within the article and its supplementary materials.
-
The authors declare that they have no conflict of interest.
-
# Authors contributed equally: Yunyun Liu, Jun Song
- Supplementary Table S1 Organ-specific in vitro differentiation responses of Scrophularia ningpoensis under different hormone treatments for 30 days.
- Supplementary Table S2 Organ-specific in vitro differentiation responses of Scrophularia ningpoensis under different hormone treatments for 60 days.
- Supplementary Table S3 Differentiation rates of leaf and petiole explants of Scrophularia ningpoensis at 30 and 60 days post-inoculation.
- Supplementary Table S4 Two-way ANOVA for shooting rate of Scrophularia ningpoensis.
- Supplementary Table S5 Two-way ANOVA for rooting rate of Scrophularia ningpoensis.
- Supplementary Table S6 OLS regression analysis for shooting rate of Scrophularia ningpoensis.
- Supplementary Table S7 OLS regression analysis for rooting rate of Scrophularia ningpoensis.
- Supplementary Table S8 Identification of key regulatory factors using multiple analytical approaches.
- Supplementary Table S9 Baseline model performance for differentiation rate prediction.
- Supplementary Table S10 Performance of optimized machine learning models for differentiation rate prediction.
- Supplementary Table S11 Mutual information–based importance analysis of factors affecting differentiation rate.
- Supplementary Table S12 Permutation importance of predictors in the optimized model.
- Supplementary Table S13 Comparative mutual information ranking of regulatory factors.
- Supplementary Table S14 Comparison of model-predicted and experimentally observed rooting rates for model-ranked conditions within the tested experimental space.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Liu Y, Song J, Shi K, Dai X, Zhu H, et al. 2026. Development of an in vitro organ-specific regeneration system for Scrophularia ningpoensis and predictive modeling of culture conditions using machine learning. Medicinal Plant Biology 5: e011 doi: 10.48130/mpb-0026-0008
Development of an in vitro organ-specific regeneration system for Scrophularia ningpoensis and predictive modeling of culture conditions using machine learning
- Received: 28 February 2026
- Revised: 01 April 2026
- Accepted: 10 April 2026
- Published online: 29 April 2026
Abstract: Scrophularia ningpoensis relies mainly on vegetative propagation, a practice that can accelerate germplasm degradation and pathogen accumulation, thereby limiting the large-scale production of healthy seedlings. Here, we established an in vitro regeneration system using leaf and petiole explants cultured on Murashige and Skoog (MS) medium supplemented with different combinations of 6-benzylaminopurine (6-BA) and naphthaleneacetic acid (NAA), and applied machine learning (ML) to model the regeneration responses within the tested experimental space. Regeneration showed clear organ-specific patterns: Leaf explants exhibited strong rooting capacity, reaching 96.7% ± 3.3% at 1.0 mg/L 6-BA + 1.0 mg/L NAA, whereas petiole explants showed higher shoot regeneration, with a maximum of 50.0% ± 11.6% at 0.5 mg/L 6-BA + 0.2 mg/L NAA. Factorial analysis of variance and ordinary least squares regression identified explant type and NAA concentration as the major determinants of regeneration. Random forest models showed moderate predictive performance for shoot regeneration (R2 = 0.57) and stronger predictive performance for rooting (R2 = 0.78), with explant type consistently ranked as the most influential variable. Notably, the highest-ranked rooting condition predicted by the model was the same as the best-performing treatment identified experimentally. Together, these findings establish an efficient regeneration system for S. ningpoensis, demonstrate explant-specific responses to phytohormones, and support ML as a complementary tool for quantitative interpretation and predictive analysis of tissue culture responses in nonmodel medicinal plants.













