-
Global warming, characterized by escalating frequency of extreme heat events, poses a significant threat to global food security by potentially perturbing the spatial distribution and diversity of major food crops. According to the Food and Agriculture Organization of the United Nations (FAO), the average temperature in many countries has risen by more than 1.5 °C from 1961 to 2021[1]. Projections based on authoritative emission scenarios indicate that global temperatures will continue to rise by more than 1.5 °C over the next 10 years[2], posing a direct threat to staple food production. A 1 °C temperature increase in the environment is associated with an average yield reduction of 6.0% for wheat, 3.2% for rice, 7.4% for maize, and 3.1% for soybean, respectively[3]. The impact is particularly acute in low-latitude regions, where 10%–31% of current food crop-producing areas will be affected even under a 2 °C global warming scenario. This proportion is projected to rise to 20%–48% under a 3 °C warming scenario, alongside a marked decline in potential food crop diversity[4]. Compounding this challenge, the continuous growth of the global population is expected to double food demand, thereby posing huge challenges to future agriculture sustainability.
Rice (Oryza sativa L.) is one of the most important food crops globally, feeding more than half of the world's population. Rice is sensitive to heat stress (> 35 °C) throughout its life cycle, spanning both vegetative and reproductive stages. Although high temperature during the vegetative stage can inhibit growth, significant negative impacts typically require sustained high temperature (e.g., > 35 °C for several days). If the heat stress is mild, these impacts can be largely reversed after returning to normal temperatures[5]. In contrast, the reproductive stage (including floral organ formation, male and female gamete development, flowering, fertilization, and grain filling) is particularly sensitive to heat stress[6,7]. During the booting and heading/flowering stages, sustained high temperature can impair rice productivity by reducing spikelet number, triggering floral organ malformation and pollen abortion, disrupting pollination and fertilization, and ultimately leading to a severe decline in seed-setting rate[8]. High temperature during the grain filling stage is detrimental to the development of endosperm structure, leading to increased chalkiness and a consequent deterioration in grain quality, taste, size, and overall yield[9]. These impacts are evidenced by significant production losses across major rice-producing areas. For instance, China experienced substantial loss in many main producing areas by extreme high-temperature events. In the Yangtze River Basin, it was conservatively estimated that 3 × 107 hectares of rice were affected by high temperature in 2003, with a loss of 5.2 × 107 tons of rice grain[10]. In Anhui Province, 333,000 hectares of rice suffered heat stress during the flowering-stage in 2003, resulting in a seed setting rate that is lower than 60%[11]. In 2013, the city of Ma'an also suffered heat stress again, with a direct economic loss of more than 150 million yuan[12]. In 2006, Southwest China suffered an unprecedented extremely high-temperature with maximum temperatures exceeding 35 °C ranged from 40 to 60 d, resulting in a rice yield reduction of more than 25%, and even over 50% in some local areas[13]. In South China, extreme high-temperature events tend to occur during the filling stage of early rice and the vegetative growth stage of late rice, with every 1.0 °C increase in temperature leading to an 8.1% reduction in the yield of early rice[14]. Similarly, during El Niño-induced high-temperature events in Southeast Asia's Mekong Delta, the yield per unit area of rice decreased by 15%–20%[4]. Therefore, deciphering the molecular mechanisms underlying heat stress is critical, as it enables more accurate predictions of climate change impact on rice production. This knowledge provides a vital scientific basis for devising adaptive agricultural policies, stabilizing global food supplies, and ensuring farmers' income.
Historically, research on plant thermotolerance primarily focused on the vegetative stage, likely due to its experimental tractability and the visibility of stress phenotypes. However, given that reproductive development is more vulnerable to heat stress and holds greater relevance for agricultural productivity, research efforts have increasingly pivoted towards the reproductive phase over the past decade. This shift reflects the growing recognition that reproductive failure is the primary cause of yield loss under heat stress, driving a surge in studies targeting flowering, pollination, and grain development. This review synthesizes advances in understanding the molecular regulatory networks governing heat tolerance in the rice reproductive stage, and proposes potential strategies for developing rice cultivars with stable and high yield under high-temperature conditions.
-
Heat stress severely impairs spikelet development, leading to floral organ malformation and various abnormal phenotypes, such as floret abortion, smaller and incompletely closed glumes, abnormal filaments, pollen sterility, and defective female gametophyte development[15]. This process involves a molecular regulatory network in which transcription factors such as members of the MADS (MCM1, AGAMOUS, DEFICIENS, SRF)-box and FRS (FAR1-RELATED SEQUENCE) families, and plant hormones like jasmonic acid (JA) and auxin interact to maintain floral organ formation under high temperature stress (Fig. 1).
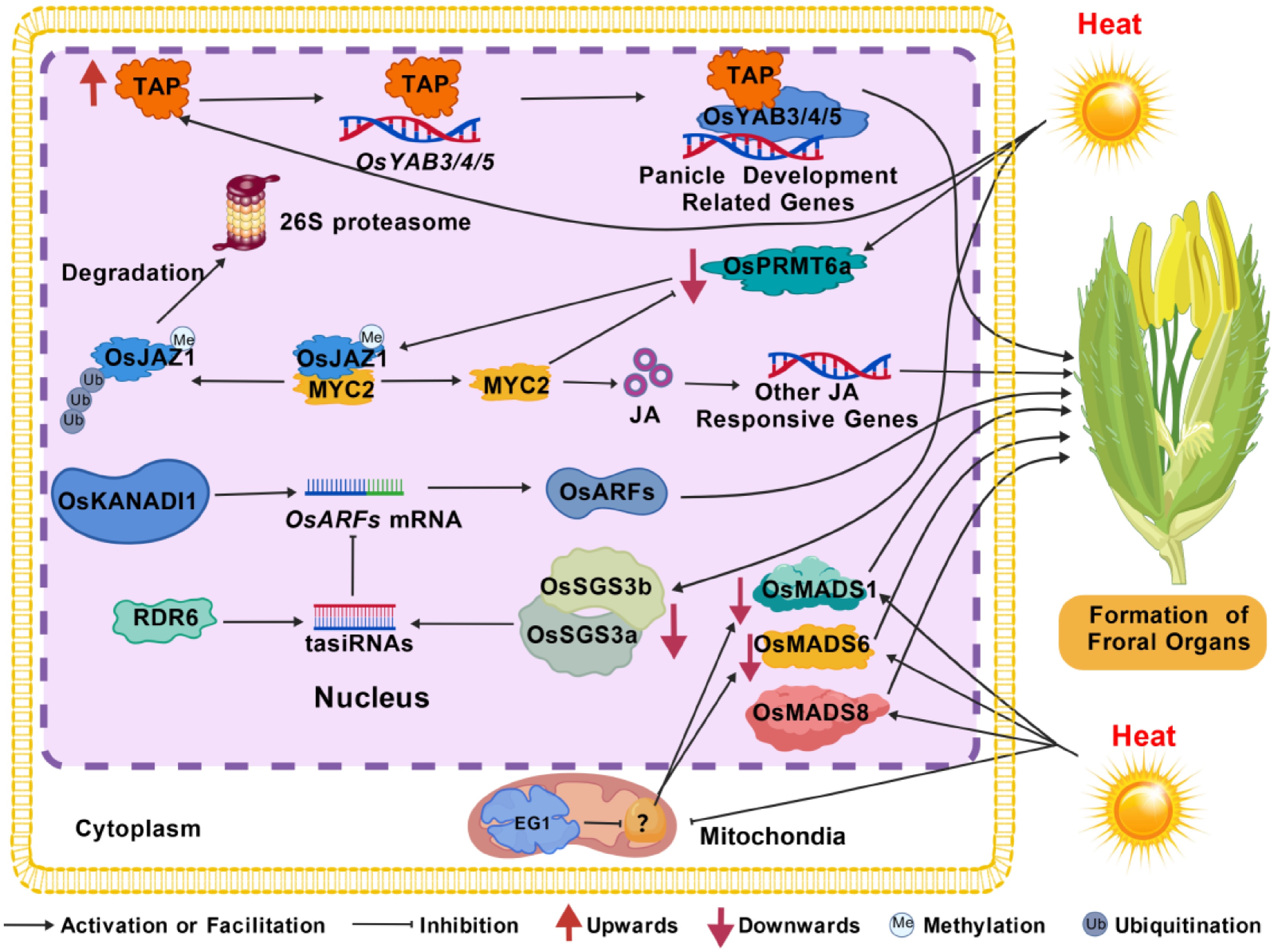
Figure 1.
Molecular regulatory networks governing rice spikelet development under heat stress. Heat stress triggers a cascade of molecular events coordinating spikelet development. Key pathways include: (1) The upregulation of THERMO-SENSITIVE BARREN PANICLE (TAP), which directly regulates and interacts with OsYABBY transcription factors (OsYAB3/4/5) to control inflorescence architecture; (2) Protein arginine methyltransferase OsPRMT6a-mediated methylation promotes the degradation of OsJAZ1, releasing OsMYC2 to activate jasmonic acid (JA)-responsive genes; (3) RNA-dependent RNA polymerase 6 (RDR6) generates trans-acting small interfering RNAs (tasiR-ARFs) that post-transcriptionally regulate AUXIN RESPONSE FACTORS (OsARF2/3), and the transcription of OsARF3A is directly promoted by OsKANADI1; (4) The dsRNA-binding protein OsSGS3a interacts with OsSGS3b to regulate spikelet development via the OsARF pathway; (5) Floral organ identity is maintained by MADS-box transcription factors (OsMADS1/6/8), with the expression of OsMADS1 and OsMADS6 being modulated by the mitochondrial lipase EXTRA GLUME1 (EG1).
Regulation of spikelet development by transcription factors under high temperature
-
Transcription factors, particularly those from the MADS-box and FRS families, play a core role in maintaining spikelet development under high temperature by regulating downstream target genes (Fig. 1). OsMADS8 is critical for maintaining floret meristem determinacy, ensuring normal carpel development, and initiating ovule formation specifically under high temperature[16]. The osmads8 mutants show no obvious abnormalities in floral organs at warm temperature (28 °C), but exhibit abnormal florets at high temperature (34 °C), demonstrating that OsMADS8 regulates floral organ development in a temperature-dependent manner[16]. Two other floral development-related genes, OsMADS1 and OsMADS6, are regulated by EXTRA GLUME1 (EG1), whose transcription, protein stability, and enzymatic activity are all temperature-dependent[17]. Elevated temperatures compromise EG1 function, leading to dysregulated expression of OsMADS1 and OsMADS6, and a consequent disruption of the homeostatic development of floral organs[17]. The FRS transposase family transcription factor THERMO-SENSITIVE BARREN PANICLE (TAP)/Thermo-sensitive Spikelet Defects 1 (TSD1) has been identified as a key thermo-sensitive regulator of spikelet development[18,19]. The expression level and protein abundance of TAP/TSD1 are induced by increasing temperatures and directly regulates the expression of the key panicle and spikelet development related to TOB-like OsYABBY family members (OsYAB3, OsYAB4, and OsYAB5), and forms protein complexes with OsYAB4/5 to coordinately ensure normal inflorescence and spikelet development of rice[18,19].
Regulation of rice spikelet development by plant hormones under high temperature
-
The content and signaling balance of plant hormones, such as JA and auxin, are critical for the adaptive regulation of rice spikelet development under high-temperature stress. Dysfunction of genes related to hormone signaling often leads to spikelet development defects (Fig. 1). Protein Arginine N-Methyltransferase 6a (OsPRMT6a)- responds to environmental stress by mediating arginine methylation of Jasmonate ZIM-Domain 1 (OsJAZ1) to fine-tune JA signaling intensity[20]. RNA-dependent RNA polymerase 6 (RDR6) mediates the production of trans-acting small interfering RNAs targeting AUXIN RESPONSE FACTORs (tasiR-ARFs)[21]. These tasiR-ARFs negatively regulate the expression of OsARF2/3, which are involved in establishing the polarity of rice lemmas (Fig. 1). The osrdr6-1 mutant exhibits normal growth at low temperature but develops abnormal awn-like lemmas and defective spikelets under high temperature. This phenotype arises because elevated temperatures reduce the accumulation of tasiR-ARFs in osrdr6-1, thereby relieving the repression of ARFs and impairing lemma polarity and spikelet development. Furthermore, the MYB transcription factor OsKANADI1 directly binds to the intron of OsARF3A to promote its transcription[22]. Collectively, these findings demonstrate that RDR6 negatively regulates OsARF3A post-transcriptionally via tasiR-ARFs, whereas OsKANADI1 positively regulates OsARF3A expression at the transcriptional level. These two opposing regulatory mechanisms work in concert to maintain OsARF3A transcripts at an appropriate level, thereby ensuring proper lemma development under high temperature stress[21].
In addition, the dsRNA-binding protein SUPPRESSOR OF GENE SILENCING 3a (OsSGS3a) also regulates spikelet development through the OsARF pathway[23]. OsSGS3a interacts with its homolog OsSGS3b, and the OsSGS3a/b complex promotes tasiR-ARF biosynthesis to inhibit the expression of OsARF3s. Simultaneous knockdown of OsSGS3a/b or overexpression of OsARF3s leads to floret defects, sterility, and reduced thermotolerance under high temperature. Conversely, overexpression of OsSGS3a/b or mutation of OsARFs enhances heat resistance. The OsSGS3a/b module itself is thermally responsive: under high temperature, the degradation of OsSGS3a/b protein reduces tasiR-ARF production, leading to the upregulation of OsARF3s and consequently decreased heat tolerance[23].
-
High temperature is one of the major environmental stresses causing the decline in pollen fertility, which impairs key processes such as meiosis of pollen mother cells and anther dehiscence. To counteract these effects, rice pollen employs core protective mechanisms centered on maintaining reactive oxygen species (ROS) homeostasis and regulating heat shock protein activity. Notably. the meiosis process remains highly vulnerable to heat stress even within the protective microenvironment of young anthers[24].
Regulation of pollen fertility under high temperature by ROS
-
During anther development, maintaining a dynamic balance of ROS is crucial for normal reproductive processes (Fig. 2). High temperature disrupts this balance, often leading to reduced floret fertility. Regulating the ROS scavenging system through some molecules such as spermidine, salicylic acid (SA) and abscisic acid (ABA) can effectively alleviate heat-induced damage to pollen fertility. For example, application of exogenous spermidine improves the heat stress resistance by reducing the production rate of superoxide anions and increasing the activity of protective enzymes such as superoxide dismutase (SOD) and catalase (CAT) in rice developing panicles[25,26]. Application of SA during the meiosis stage of rice pollen mother cells enhances pollen viability and seed-setting rate under heat stress by reducing ROS accumulation, and preventing premature degradation of the anther tapetum[27]. Exogenous ABA application prior to heat stress can alleviate heat stress-induced pollen sterility and enhance rice thermotolerance[28]. Overexpression of ABA biosynthesis pathway gene CIS-EPOXYCAROTENOID DIOXYGENASE1 (NCED1) enhances pollen viability and seed-setting rate, improves the activity of antioxidant enzymes such as SOD and peroxidase (POD), and reduces electrolyte leakage and malondialdehyde (MDA) content, thereby improving heat tolerance[29] However, other researchers have shown that higher ABA induced by high temperature stress or exogenous application reduced microspore viability in a dose-dependent manner by triggering ROS generation in the developing anthers[30]. These contradictory findings require further investigation.
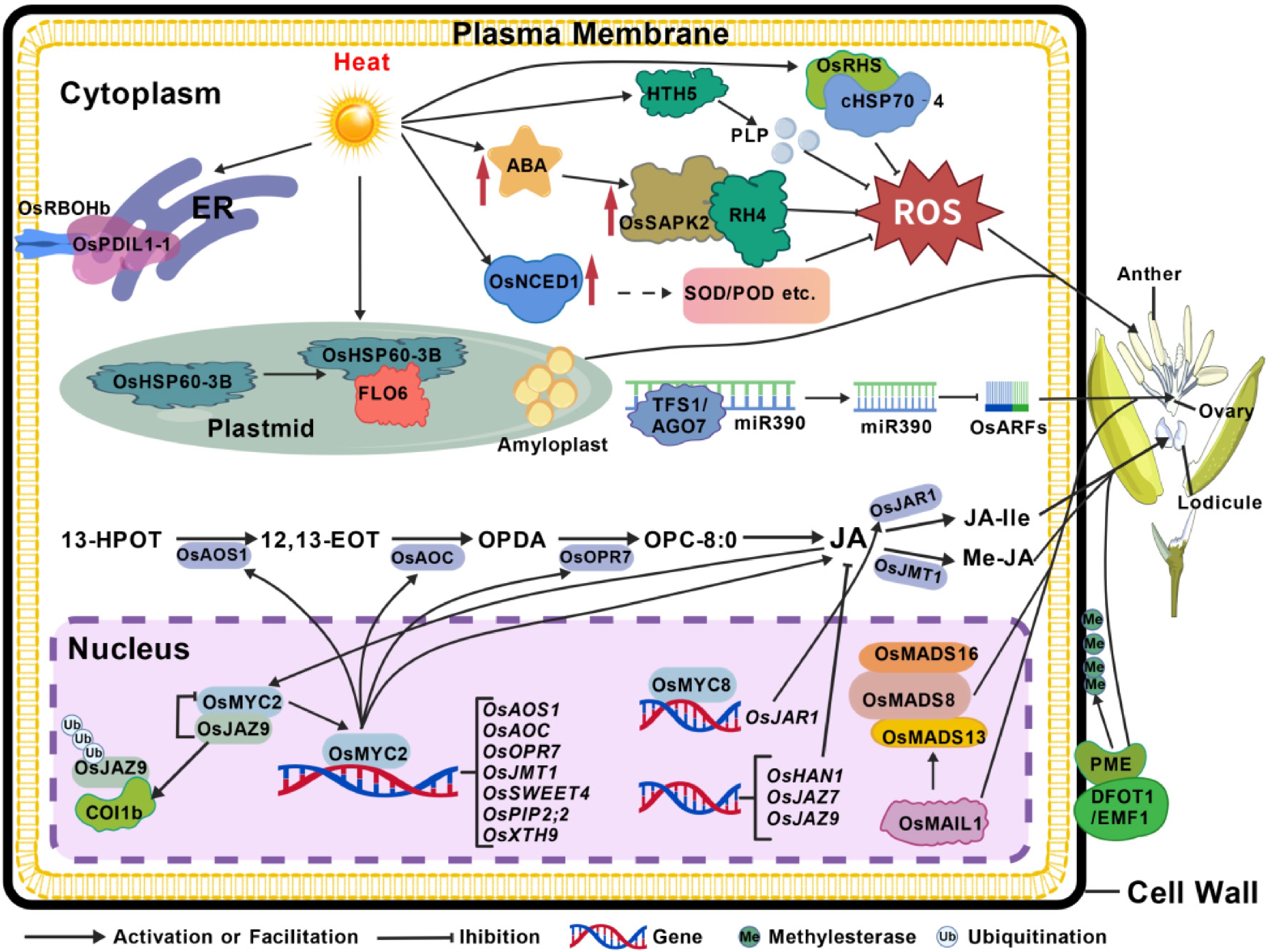
Figure 2.
Molecular networks regulating rice pollen formation, ovary development, and fertilization under heat stress. Heat stress disrupts key reproductive processes, including pollen development, ovary formation, and diurnal flower-opening timing, thereby triggering sophisticated molecular responses to mitigate these damages. The core adaptive mechanisms are outlined below: (1) Pollen development and anther dehiscence: The dynamic balance of reactive oxygen species (ROS) and starch synthesis is crucial for these processes. Multiple mechanisms converge to limit ROS bursts: ABA-activated protein kinase 2 (OsSAPK2) directly interacts with the DEAD-box RNA helicase RH4; CIS-EPOXYCAROTENOID DIOXYGENASE (NCED) expression is upregulated, enhancing the activity of antioxidant enzymes such as SOD and POD; HTH5-encoded pyridoxal 5'-phosphate (PLP) homeostasis protein (PLPHP) increases antioxidant PLP content; REDUCED HEAT STRESS TOLERANCE 1 (OsRHS) inhibits ROS production through interaction with cHSP70-4; and disulfide isomerase-like proteins (PDIL) regulate ROS generation by interacting with RBOHb at the endoplasmic reticulum (ER)-plasma membrane junction. Additionally, rice HEAT SHOCK PROTEIN 60-3B (OsHSP60-3B) interacts with FLOURY ENDOSPERM 6 (FLO6) to ensure proper starch granule formation. (2) Female gametophyte and pistil development: This process involves transcriptional and post-transcriptional regulation of key genes. AGO7 combines with miR390 to form an RNA-induced silencing complex (RISC) that regulates female fertility by inhibiting tasiR-ARFs. MADS8 forms heterodimers with MADS13 and MADS16 to coordinately regulate target genes. Meanwhile, OsMAINTENANCE OF MERISTEM LIKE 1 (OsMAIL1) activates carpel identity genes, including OsMADS13. (3) Diurnal flower-opening timing (DFOT): DFOT1 promotes pectin methylesterase (PME) activity and regulates diurnal flower opening by reducing pectin methylesterification in lodicule cell walls. The JA pathway components OsOPR7, OsMYC2, OsMYC8, OsAOS1, and OsAOC, together with JA inactivation factor OsHAN1 and negative regulators OsJAZ7 and OsJAZ9, jointly fine-tune diurnal flower-opening timing under heat stress.
Some proteins regulating ROS production affect anther fertility. Disulfide isomerase-like protein (PDIL) regulates high-temperature-induced ROS production through interaction with RBOHb at the ER-plasma membrane (ER-PM) junction, thereby affecting rice anther fertility[31]. Furthermore, a quantitative trait locus (QTL) for heading-stage heat tolerance, qHTH5, was cloned from wild rice (Oryza rufipogon), which encodes a mitochondrial-localized pyridoxal 5'-phosphate (PLP) homeostasis protein (PLPHP)[32]. qHTH5 improves the heat tolerance of rice pollen and improves seed-setting by elevating the content of the heat-induced antioxidant PLP and reducing ROS accumulation under high temperature[32].
Regulation of pollen fertility by heat shock proteins (HSPs) under high temperature
-
HSPs function as crucial molecular chaperones in maintaining normal biofunctions of many proteins under high-temperature by stabilizing starch synthesis-related proteins and facilitating ROS scavenging. Among them, OsHSP60-3B represents a key gene essential for maintaining male fertility under high temperature[33]. Knockout of OsHSP60-3B leads to pollen abortion at elevated temperatures, whereas its overexpression enhances pollen thermotolerance. OsHSP60-3B is localized in plastids, and its expression is rapidly upregulated under high temperature. It interacts with the starch synthesis-related protein FLOURY ENDOSPERM 6 (FLO6) and stabilizes it, thereby ensuring proper starch granule formation[33]. Concurrently, OsHSP60-3B mitigates oxidative damage by inhibiting excessive ROS accumulation in anthers[33]. These findings indicates that OsHSP60-3B plays a dual role in regulating pollen starch biosynthesis and ROS homeostasis, highlighting its significant value in safeguarding rice yield stability under high-temperature stress.
In addition, REDUCED HEAT STRESS TOLERANCE 1 (OsRHS1) acts as a negative regulator of rice heat tolerance at the flowering stage[34]. Overexpression of OsRHS1 reduces rice heat tolerance, while its suppression via RNA interference (RNAi) silencing or knockout enhances heat tolerance. OsRHS1 interacts with cHSP70-4, and high temperature strengthens their association. Notably, knockout of cHSP70-4 can rescue the heat-sensitive phenotype cause by OsRHS1 overexpression. In addition, under high-temperature stress, OsRHS1 acts as a transcriptional repressor of multiple stress-responsive genes, including those encoding pectin methylesterase (PMEs), thereby regulating rice heat tolerance at the flowering stage[34].
Mechanism of regulating thermo-sensitive genic male sterility (TGMS)
-
The pollen development of TGMS rice lines is extremely sensitive to temperature variation, resulting in male sterility under high temperature and restored male fertility under low temperature. This characteristic of TGMS lines has been widely used in the two-line hybrid rice breeding system[35,36]. Multiple genes regulating TGMS have been identified, including photoperiod/thermosensitive genic male sterility 12-1 (P/TMS12-1 or PMS3), TMS5, UDP-glucose pyrophosphorylase1 (UGP1), OsMS1wenmin1, TMS10, OsTMS15, OsTMS16, OsTMS18, and OsTMS19[35].
P/TMS12-1 (PMS3) is the first isolated TGMS gene to be applied in two-line hybrid rice. It encodes a long non-coding RNA (lncRNA) termed LDMAR (long-day-specific male-fertility-associated RNA). This transcript is targeted by a 22-nt microRNA miR2118, leading to the production of a series of 21-nt phasiRNAs that regulate P/TGMS in rice[37,38]. TMS5 is the most widely used TGMS gene in two-line hybrid rice breeding in China, accounting for over 95% of TGMS lines[39]. TMS5 encodes a ribonuclease RNase ZS1, which is involved in the repair of 2',3'-cyclic phosphate tRNAs lacking the 3' CCA terminus and containing a 2',3'-cyclic phosphate (cP) group[40,41]. Impaired repair of cP-ΔCCA-tRNAs in tms5 mutant lines under higher temperatures (> 26 °C) leads to a reduction in the abundance of mature tRNAs, causing male sterility, whereas low temperature (23−24 °C) permits sufficient repair and restores fertility of the tms5 mutant lines[41]. OsMS1wenmin1 encodes a PHD-finger transcription factor and regulates TGMS through temperature-dependent transcriptional regulation[42]. TMS10 encodes a leucine-rich repeat receptor-like kinase, and its mutation causes male sterility at high temperature, while functional complementation by its homolog TMS10L restores male fertility at low temperature[43]. Overexpression of UGP1 in rice triggers co-suppression, resulting in a TGMS phenotype[44]. OsTMS15 encodes a leucine-rich repeat receptor-like kinase MSP1 specifically expressed in anthers[45]. Under high-temperature conditions, the tapetum of the ostms15 mutant undergoes progressive vacuolation, failing to support microspore development and leading to sterility. Under low-temperature conditions, the slow development of ostms15 anthers compensates for tapetal development, thereby restoring fertility[45]. OsTMS18 encodes a glucose-methanol-choline (GMC) oxidoreductase, and a Gly-to-Ser point mutation in this gene confers a TGMS phenotype[46]. OsTMS19 encodes a pentatricopeptide repeat protein (PPR) expressed in anther tapetum[47]. A single amino acid substitution in this gene leads to excessive ROS accumulation under high-temperature sterile conditions, resulting in a male sterile phenotype. However, effective ROS scavenging under low-temperature conditions restores fertility[47].
The critical sterility-inducing temperature (CSIT) refers to the threshold temperature at which TGMS lines transition from male fertility to complete male sterility, a parameter critical for ensuring the success of two-line hybrid rice production[48]. Elevated or unstable CSITs of TGMS lines are bottlenecks that restrict the breeding of two-line hybrid rice[49]. Two CSIT-related genes, CSIT1/OsLtn1 and CSIT2/Oshel2, were cloned from a TGMS line carrying the tms5 mutation[48−50]. These genes function in ribosome-associated protein quality control (RQC), a pathway activated when protein synthesis or folding is abnormal. Subsequent studies identified additional RQC factors, including Vms1, Rqc1, and Rqc2, that also influence CSIT[41,51,52]. Under RQC activation, CSIT1/OsLtn1 binds to the 80S ribosomes and ubiquitinates aberrant nascent polypeptides for degradation[48]. CSIT2/Oshel2 binds to the large and small ribosomal subunits, ubiquitinating 80S ribosomes to facilitate dissociation and possibly also targeting misfolded proteins for degradation[49]. Impaired CSIT2/Oshel2 function results in partial readthrough of stalled translation sequences, thereby enabling evasion of RQC surveillance[50]. OsRqc1 interacts with the OsVms1 and binds the 60S ribosomal subunit to cleave tRNA-nascent polypeptide conjugates, generating cP-ΔCCA-tRNAs[51]. OsRqc2 recruits tRNAs to the A-site of the 60S ribosome and mediates CATylation of nascent chains for degradation[52]. RNase ZS1 repairs cP-ΔCCA-tRNAs, allowing TRNT1 to add the CCA terminus and regenerate mature tRNAs[41]. Overexpression of specific tRNAs (e.g., tRNA-Ala/Pro/Val) in the tms5 mutant restores male fertility under high temperature[41,50−52]. These findings collectively demonstrate that CSIT of TGMS lines (carrying tms5) is controlled through the RQC pathway. Mutations in CSIT1, CSIT2, Vms1, Rqc1, and Rqc2 reduce the production of cP-ΔCCA-tRNAs at high temperature, thereby increasing the CSIT in these TGMS lines carrying tms5.
-
High temperature causes abnormal pistil development in rice, such as ovule abortion, ovary atrophy, stigma malformation, and failure of pollen tubes to enter the embryo sac. The heat tolerance of pistils is mainly regulated through two major pathways: transcription factors and miRNAs.
Regulation of pistil development by transcription factors under high temperature
-
Transcription factors from the MADS family not only regulate organ development under normal conditions but also play essential roles in pistil development under high-temperature stress (Fig. 2). Loss of MADS8 function leads to abnormal pistil development under high temperature, characterized by increased ovule abortion, ovary atrophy, and significantly reduced female fertility. However, the mads8 mutant shows no obvious defects in female reproductive development at normal temperatures[16]. MADS8 forms heterodimers with proteins such as MADS13 and MADS16 to coordinately enhance cellular protein stability by upregulating the expression of heat shock protein genes and reduces ROS accumulation by activating antioxidant enzyme-encoding genes[16]. Furthermore, OsMAINTENANCE OF MERISTEM LIKE 1 (OsMAIL1), a rice member from the Plant Mobile Domain family, is crucial for female reproductive development under high-temperature stress (Fig. 2). The osmail1 mutant exhibits increased sterility under high temperature due to abnormal pistil development. OsMAIL1 regulates pistil development by activating carpel identity genes such as OsMADS13, thereby maintaining female fertility in rice under high temperature[53].
Regulation of pistil development by miRNAs under high temperature
-
While numerous genes regulating TGMS have been identified, only two thermo-sensitive female sterile (TFS) mutants, tfs1 and tfs2, have been reported so far. Under high temperature, tfs1 exhibits complete female sterility, but it partially regains fertility at low temperature[54] (Fig. 2). The tfs1 phenotype is caused by a point mutation in the ARGONAUTE7 (AGO7) gene. This mutation impairs the ability of TFS1/AGO7 to bind miR390 and form a functional RNA-induced silencing complex (RISC) under high temperature, thereby compromising the recruitment of RNA-dependent RNA polymerase 6 (RDR6). As a result, the production of tasiR-ARFs is reduced, leading to failure of pollen tube entry into the embryo sac and consequent female sterility. At low temperature, the function of mutant TFS1 (mTFS1) is partially restored, improving both the RISC assembly efficiency and RDR6 recruitment. This promotes tasiR-ARF synthesis and partially restores female fertility. The tfs1 mutant has potential application value in hybrid seed production. tfs1 can be used as a male parent line (restorer line) for mixed planting with a TGMS (female parent) line for seed production in high-temperature seasons. By this way the tfs1 line itself does not produce self-pollinated seeds due to female sterility, thus this system can enable fully mechanized hybrid seed production and eliminate the tedious process of manual removal of male parent seeds.
The tfs2 mutant was generated by ethyl methanesulfonate (EMS) mutagenesis of the japonica rice variety Zhonghua 11 (ZH11). It exhibits female sterility under high temperatures above 30 °C, and restored fertility below 27 °C[55]. This phenotype is caused by a point mutation in the HEI10 gene, which functions in meiotic crossover formation. Under high temperature, tfs2 produces abnormal univalents during meiosis, leading to unequal chromosome segregation. In contrast, low temperature restores the dot-like nuclear aggregation of the tfs2 protein, thereby recovering crossover formation and equal chromosome segregation, and partially restoring female gamete fertility[55].
-
The flower-opening stage is considered one of the most heat-sensitive period in the reproductive phase of rice. The 0.5–1 h window of rice flower-opening is critical for pollen release and pollination. High temperature directly inhibits anther dehiscence and pollen adhesion to stigmas, ultimately leading to fertilization failure. The impact of high temperature on fertility during this stage is far greater than that before or after flower-opening, making it the most sensitive period[15]. Some cereal crops, such as sorghum, pearl millet, and wheat, have optimized their diurnal flower-opening times (DFOTs) to cooler early morning or late evening to mitigate heat damage during flower-opening[56]. Analysis of 289 tropical and subtropical crop varieties shows that shifting the peak flowering time to early morning during the dry seasons can reduce the sterility rates by up to 71%[57]. Therefore, developing crops with flower-opening during cooler periods of the day (early morning or late evening) represents a viable strategy to prevent reproductive organs from direct heat exposure, and minimize grain yield losses[58].
Cell wall elasticity regulates rice DFOT
-
Different rice varieties exhibit variations in DFOT, with distinct differences between the indica and japonica subspecies. Generally, indica rice flowers in the morning with cooler temperatures, while japonica rice typically flowers around midday or in the afternoon with higher temperatures. Several genes regulating rice DFOT have been identified (Fig. 2). DFOT1/Early Morning Flowering 1 (EMF1) is the first identified DFOT gene[59]. It is specifically expressed in rice lodicules, with expression gradually increasing as flower-opening approaches, and declining rapidly after flower-opening. Knockout of DFOT1/EMF1 in the japonica variety ZH11 advances its DFOT from 11:00 to 8:30. DFOT1/EMF1 is localized in the cell wall, contains a DUF642 domain, and directly interacts with multiple members of the pectin methylesterase (PME) family. Knockout of PME40 in ZH11 significantly promotes flower-opening time, while overexpression of PME42 delays flower-opening time, even in the dfot1 background. DFOT1 enhances PME activity, thereby reducing the degree of pectin methylesterification in the lodicule cell wall. This process increases the cell wall elasticity modulus and hardens the cell wall, thus restricting water absorption and expansion of lodicule cells. Conversely, loss of function of DFOT1/EMF1 or PME40 leads to decreased PME activity in the lodicule cell wall, and increased pectin methylesterification, which reduces the cell wall elasticity modulus and softens the cell wall. A softened cell wall facilitates water uptake and expansion of the lodicule, resulting in early flower opening. The 5' UTR region of DFOT1/EMF1 shows significant sequence divergence between indica and japonica rice, which may underlie differences in gene expression levels, probably relating to temperatures. This suggests that introducing the indica allele of DFOT1/EMF1 into japonica rice could promote earlier morning flower-opening[59,60].
Regulation of rice DFOT by the JA signaling pathway
-
JA regulates lodicule cell wall remodeling and osmotic pressure, controlling DFOT of rice (Fig. 2). Within a certain range, the number of flower-opening spikelets in rice is positively correlated with methyl jasmonate (MeJA) concentration. Genetic manipulations that enhance JA signaling consistently advance DFOT: overexpressing the JA biosynthesis gene Oxophytodienoate Reductase 7 (OsOPR7), knocking out the JA inactivation gene OsHAN1, or knocking out negative regulators of JA signaling (OsJAZ7 and OsJAZ9), advances the DFOT of the japonica variety ZH11 by 1 h 2 h, 50 min, and 1.5 h, respectively, accompanied by significantly increased JA levels in lodicules[61]. In contrast, knocking out OsAOS1—a key rate-limiting enzyme for JA synthesis (allene oxide synthase)—delays flower-opening after heading[62].
Meanwhile, the JA signaling pathway directly regulates the expression of JA synthesis genes such as Allene Oxide Synthase 1 (OsAOS1), Allene Oxide Cyclase (OsAOC), and OsOPR7 through the core transcription factor OsMYC2 (Fig. 2). Overexpression of OsMYC2 accelerates flower-opening, and its activity is potentially inhibited through interaction with the JA signaling repressor OsJAZ9, forming an OsMYC2-JA feedback loop[63]. Additionally, OsMYC2 directly binds to the promoters of the sugar transporter gene OsSWEET4, the aquaporin gene Plasma membrane Intrinsic Protein 2;2 (OsPIP2;2), and cell wall metabolism-related genes Xyloglucan Endotransglucosylase/Hydrolase 9 (OsXTH9), Glycoside Hydrolase Family 9 Subfamily B Member 13 (OsGH9B13), and Glucan endo-1,3-beta-glucosidase 11 (OsGns11), promoting their expression. This dual action increases osmotic pressure in lodicules and facilitates cell wall relaxation, collectively promoting water absorption and lodicule cell expansion to drive flower opening[61,62].
Another JA-signaling transcription factor, OsMYB8, also contributes to DFOT regulation by directly targeting downstream genes such as the jasmonate synthase gene Jasmonate Resistant 1 (OsJAR1), which catalyzes the production of active jasmonate JA-Ile. OsMYB8 upregulates the expression of osmosis-related genes, promotes water absorption and expansion of lodicules, and thereby regulates rice DFOT. The promoter of OsMYB8 in indica rice contains six single-nucleotide polymorphisms (SNPs) and exhibits higher activity than that in japonica rice, resulting in elevated JA-Ile content and 0.5–1 h earlier flower-opening[64]. Collectively, these studies indicate that the JA pathway fine-tunes rice DFOT by coordinately regulating lodicule cell wall relaxation and osmotic potential.
Regulation of the fertilization process under high temperature
-
The inhibition of pollen germination and pollen tube growth by high temperature is a major cause of fertilization failure. Inositol plays a key role in maintaining rice fertilization under high temperature by regulating Ca2+ signaling and actin filament dynamics (Fig. 2). Pollination is a critical and temperature-sensitive process encompassing pollen adhesion, hydration, germination on stigma, and style-mediated pollen tube growth[65]. However, the molecular mechanisms underlying high-temperature disruption of pollen fertility and fertilization remain poorly understood. Under high-temperature stress, rice myo-inositol-3-phosphate synthase 2 (RINO2) modulates pollen germination and pollen tube growth by regulating Ca2+ signaling and organization of actin filaments in an inositol-dependent manner. Knockout of RINO2 significantly enhances pollen sensitivity to heat stress, as it leads to a sharp decline in inositol and phosphatidylinositol derivative levels (especially PI[4,5]P2), disrupting the normal apical Ca2+ gradient and actin cytoskeleton in pollen tubes, thereby impairing pollen germination and pollen tube growth[66].
-
High temperature during the grain-filling stage leads to the deterioration of rice grain quality, commonly manifested as impaired starch structure, increased chalkiness, and other defects in endosperm development. Multiple physiological and molecular pathways, including starch synthesis regulation, heat shock protein protection, and hormone signal modulation, collectively participate in grain development and quality formation under high-temperature stress (Fig. 3).
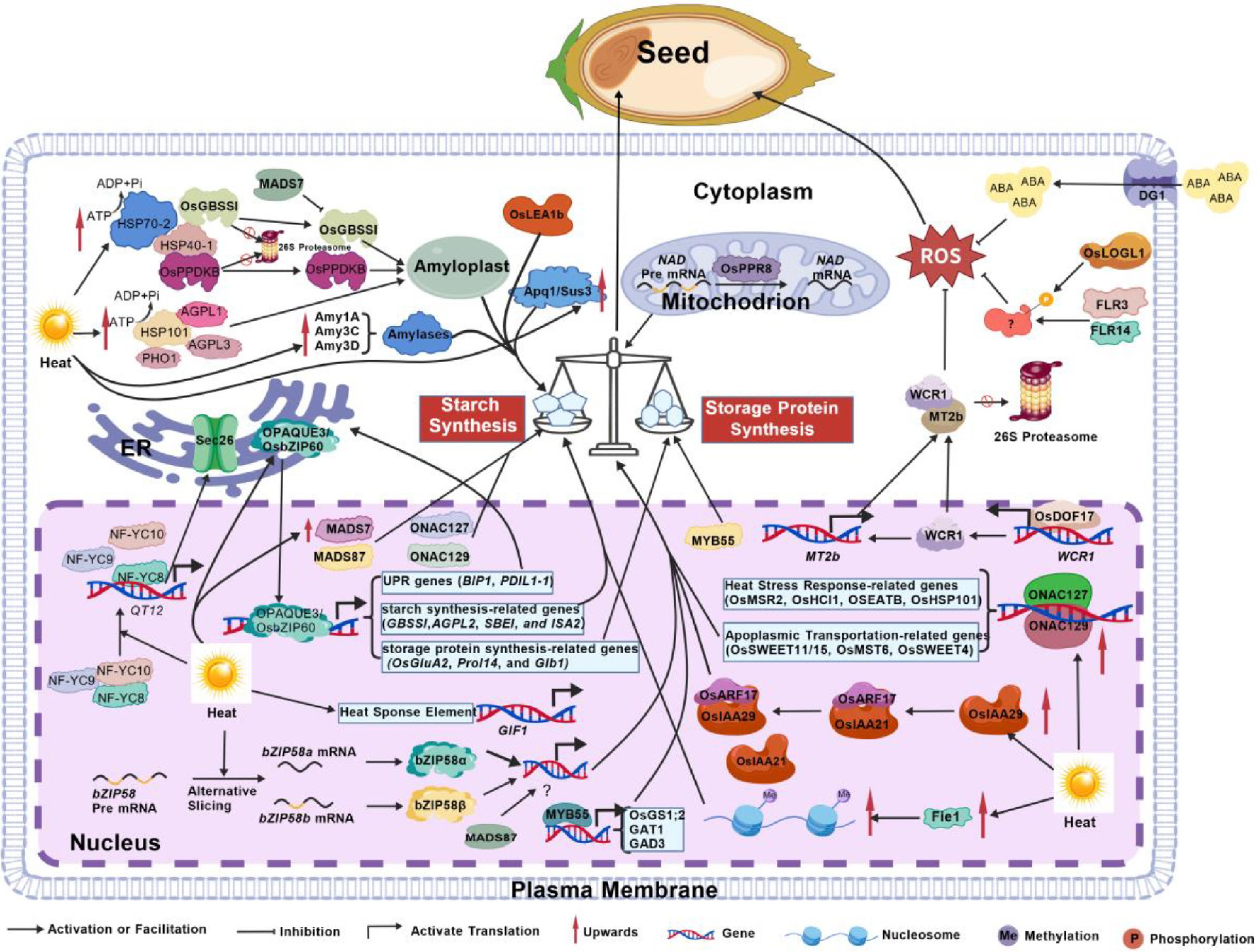
Figure 3.
Molecular mechanisms regulating of rice grain development and quality under heat stress. High temperature affects rice grain development by disrupting the balance between starch and storage protein synthesis and by inducing reactive oxygen species (ROS) accumulation. Key molecular pathways involved in this process are outlined as follows. (1) Starch metabolism and chalkiness formation: α-amylases (including Amy1A, Amy3C, and Amy3D) influence chalkiness formation under thermal stress. Heat shock proteins HSP70-2 and HSP40-1 interact with each other and directly bind key starch synthesis enzymes—GRANULE-BOUND STARCH SYNTHASE (OsGBSSI) and PYRUVATE ORTHOPHOSPHATE DIKINASE (OsPPDKB)—to maintain their stability and activity. HSP101 interacts with starch biosynthesis enzymes such as AGPL1, AGPL3, and PHO1 to regulate starch synthesis and endosperm development. Appearance Quality of Brown Rice 1 (Apq1)/Sucrose Synthase 3 (Sus3) participates in sucrose synthesis, thereby influencing final grain quality. (2) Hormone signaling and redox homeostasis: The transporter DG1 exports abscisic acid (ABA) from vascular tissues into developing caryopses, contributing to stress adaptation. Cytokinin-activating enzyme LONELY GUY-Like 1 (LOGL1), along with FERONIA-LIKE RECEPTORS FLR3 and FLR14, helps maintain redox homeostasis in the endosperm, potentially through phosphorylation of unknown target proteins. (3) Transcriptional and post-transcriptional regulation: OsIAA29 competes with OsIAA21 for binding to OsARF17, thus enhancing the transcriptional activity of OsARF17 and subsequently regulating genes involved in starch and protein synthesis. In addition, transcription factors from multiple families—including MADS-box, MYB, NAC, and bZIP—serve as core regulators of grain quality under high temperature by directly controlling the expression of starch/protein synthesis genes or other heat-responsive genes.
Regulation of grain development and quality by starch and sucrose synthases under high temperature
-
High temperature induces the expression of α-amylases in developing rice endosperm, which contributes to chalkiness formation. The rice genome contains at least eight functional α-amylase genes (Amy1A, Amy1C, Amy2A, Amy3A, Amy3B, Amy3C, Amy3D, Amy3E)[67]. Among them, the promoter activities of Amy1A, Amy3C, and Amy3D are significantly enhanced under high temperature, and their spatial expression patterns align with regions where chalkiness develops. Overexpression of these α-amylase genes leads to a high rate of chalkiness, characterized by loosely packed starch granules with surface pits, reduced starch content, and increased soluble sugar content. These results indicate that the specific upregulation of Amy1A, Amy3C, and Amy3D under high temperature is a major factor driving chalkiness formation. Inhibition of these genes may thus improve rice thermotolerance and reduce the formation of chalky grains[67] (Fig. 3).
In addition, sucrose metabolism also affects grain quality and yield (Fig. 3). Using chromosome segment substitution lines (CSSLs) derived from the indica rice cultivar 'Habataki' in the japonica background of 'Koshihikari', a quantitative trait locus (QTL) termed Appearance Quality of Brown Rice 1 (Apq1) was identified. Apq1 corresponds to the sucrose synthase 3 (Sus3) gene and affects grain quality by participating in sucrose synthesis[68]. The Apq1/Sus3 locus shows functional divergence between these two rice varieties. The Sus3Habataki allele of 'Habataki' is heat-responsive: its expression increases under high temperature (34/26 °C), leading to a significant reduction in grain chalkiness. Introducing the Sus3 allele from 'Habataki' into 'Nipponbare' eliminated chalkiness in all transgenic plants after high-temperature treatment[68]. These results indicate that Apq1/Sus3 represents a natural allelic variant that regulates grain quality under high temperature, highlighting its potential value in breeding.
Cell wall invertases (CWINs) are responsible for decomposing sucrose into glucose and fructose to provide carbon for sink organs, and their expression is inhibited by heat stress (Fig. 3). The fruit-specific invertase (LIN5) in tomato and Grain Incomplete Filling 1 (GIF1) in rice are key genes of the CWIN family, both of which influence yield and carbon allocation. Using an optimized Csy4-PE prime editing system, a 10-bp heat-responsive element (ATTCTAGAAT) was introduced into the promoter regions of tomato LIN5 (452 bp upstream) and rice GIF1 (427 bp upstream)[69]. This modification increased the expression of tomato LIN5 by 2.2-fold and rice GIF1 by 1.96-fold under heat stress. Under heat stress, the yields of edited rice and tomato lines increased by 25% and 33%, respectively, without compromising grain or fruit quality[69]. This demonstrates the potential of using gene editing technology to modify the expression patterns of key metabolic genes for improving heat tolerance while maintaining yield and quality.
Regulation of rice grain filling and quality by heat shock proteins (HSPs) under high temperature
-
Several plastid-localized HSPs have been found to be involved in rice grain filling and quality under high-temperature conditions (Fig. 3). Floury Endosperm 11-2 (FLO11-2) encodes a plastid-localized HSP70-2. A single amino acid change in FLO11-2/HSP70-2 impairs its ATPase activity, compromising its ability to maintain proper protein folding under high temperature. This leads to abnormal starch granule formation and increased chalkiness rate, while the chalkiness rate at low temperature is similar to that of the wild type[70,71]. High temperature upregulates the expression of both HSP70-2 and its co-chaperone, HSP40-1, enhancing their interaction and boosting the ATPase activity of OsHSP70-2 to provide energy for protein stabilization. Additionally, HSP40-1 directly binds to key starch synthesis enzymes Granule-Bound Starch Synthase (OsGBSSI) and Pyruvate Orthophosphate Dikinase (OsPPDKB), thereby inhibiting their heat-induced degradation and maintaining their catalytic activity[71]. Similarly, Floury Endosperm 24 (FLO24) in rice encodes a plastid-localized heat shock protein HSP101. A T296I amino acid substitution in FLO24/HSP101 also significantly reduces its ATP hydrolysis capacity, resulting in white-core endosperm and more severe chalkiness under high temperature[72]. FLO24/HSP101 regulates starch synthesis and endosperm development by interacting with key starch biosynthesis enzymes ADP-glucose Pyrophosphorylase Large Subunit 1 (AGPL1), AGPL3, and Plastidial Starch Phosphorylase 1 (PHO1) to maintain their activity[72−74].
Regulation of grain filling and quality by ROS and hormones under high temperature
-
The fine regulation of ROS homeostasis and hormone signaling, including abscisic acid (ABA), cytokinin, and auxin, constitutes an important physiological basis for maintaining grain filling efficiency and quality under high temperature (Fig. 3). Exogenous ABA pretreatment activates the ABA signaling pathway, increases the activity of antioxidant enzymes such as superoxide dismutase (SOD), peroxidase (POD), catalase (CAT), and ascorbate peroxidase (APX), upregulates the expression of ROS-scavenging genes, and reduces the accumulation of ROS content (O2- and H2O2) in rice grains under high temperature[75]. These changes improve the activity of starch synthesis-related enzymes, reduce the rate and degree of chalkiness, and improves the cooking quality of rice grains by increasing starch content and gel consistency, while decreasing amylose percentage under high temperature[75]. In addition, the Grain-Filling 1 (DG1) gene encodes a MATE transporter that facilitates ABA export from vascular tissues to caryopses through nodes and spikelet axes[76]. DG1-mediated long-distance ABA transport and seed development are thermosensitive: under high temperature, DG1 transports more ABA to developing seeds, activating the expression of grain filling-related genes. In contrast, the dg1 mutant shows temperature-sensitive grain filling defects, with reduced filling efficiency under high temperature[76].
The F-box protein White-Core Rate 1 (WCR1), a negative regulator of grain chalkiness, reduces ROS production and delays endosperm programmed cell death (PCD) by upregulating the metallothionein gene MT2b and interacting with MT2b to inhibit 26S proteasome-mediated degradation, ultimately reducing chalkiness degree[77]. The upstream transcription factor OsDOF17 binds to an AAAAA cis-element in the WCR1 promoter to activate its expression. An A/G polymorphism in the WCR1 promoter affects OsDOF17 binding affinity: the WCR1A allele, prevalent in tropical japonica rice, shows stronger binding, higher WCR1 expression, and lower chalkiness degree, whereas the WCR1G allele, common in temperate japonica rice, exhibits weaker binding, lower WCR1 expression, and higher chalkiness degree. This natural variation in the OsDOF17-WCR1 module provides a potential genetic resource for improving grain quality[77].
Rice FERONIA-LIKE RECEPTOR 3 (FLR3) and FERONIA-LIKE RECEPTOR 14 (FLR14) regulate grain quality by maintaining redox homeostasis in the endosperm[78]. Knockout of FLR3 or FLR14 significantly increases the proportion of white-core grains, causes abnormal starch and protein accumulation, increases ROS content, and accelerates endosperm PCD. In contrast, overexpression of FLR3 or FLR14 reduces the chalkiness rate and improves grain quality. Additionally, mutation of FLR3 or FLR14 also exacerbates the chalkiness rate under high temperature, indicating that both receptors function as positive regulators of thermotolerance by mitigating heat-induced oxidative stress[78].
Cytokinin and auxin pathways also regulate rice grain development under high-temperature stress. The cytokinin-activating enzyme LONELY GUY-Like 1 (LOGL1) affects the response to high night temperatures by regulating circadian rhythm-related genes and the thiamine biosynthesis pathway[79]. The logl1 mutant shows about 9% higher grain weight than the wild type under high night temperatures, while LOGL1-overexpressing lines are more sensitive to high night temperatures, with a more significant decrease in grain weight[79]. Indole-3-Acetic Acid 29 (OsIAA29) is an atypical Aux/IAA protein preferentially expressed in seeds, and its expression is induced by high-temperature stress. The loss-of-function mutant osiaa29 exhibits increased grain shrinkage, higher chalkiness rates, reduced grain weight, and significantly lower starch and glutelin content under high temperature, while showing wild-type grain phenotype at normal temperatures. OsIAA29 competes with OsIAA21 for binding to OsARF17, thereby enhancing OsARF7-mediated transcriptional activation of genes involved in starch and protein synthesis, ultimately supporting rice grain filling under high temperature[80].
Regulation of grain quality by transcription factors under high temperature
-
Transcription factors from multiple families, including MADS-box, MYB, NAC (NAM, ATAF1/2, CUC2), bZIP (basic Leucine Zipper), function as core regulators of grain quality under high temperature by directly regulating starch/protein synthesis genes or heat-responsive genes. Their alternative splicing or natural genetic variations represent important molecular bases for thermotolerance divergence between indica and japonica subspecies (Fig. 3).
OsMADS7 and OsMADS87 act as negative regulators of grain quality. OsMADS87, a type I MADS-box gene specifically expressed during the syncytial stage of seed development, regulates seed size and heat sensitivity by inhibiting the initiation of endosperm cellularization[81]. Reduced OsMADS87 expression accelerates cellularization, resulting in smaller seeds with reduced sensitivity to moderate heat stress, whereas its overexpression leads to larger seeds and increased heat sensitivity. Another MADS-family member, OsMADS7, is significantly induced under high temperature and may reduce amylose content by negatively regulating the transcription and splicing efficiency of the granule-bound starch synthase gene GBSSI[81].
In contrast, OsMYB55, ONAC127, and ONAC129 positively regulate grain filling and quality (Fig. 3). OsMYB55 expression is induced by high temperature, promoting rice vegetative growth and mitigating yield losses under heat stress[82]. OsMYB55 binds to promoter regions of target genes and directly activates the expression of genes such as glutamine synthetase (OsGS1;2), glutamine amidotransferase (GAT1), and glutamate decarboxylase 3 (GAD3)[82]. The activation of these genes increases the total amino acid content, as well as the contents of individual amino acids such as L-glutamate, GABA, and arginine, thereby improving rice stress tolerance[82].
Two seed-specific NAC transcription factors, ONAC127 and ONAC129, are preferentially expressed during grain filling and further induced by heat stress (Fig. 3). Knockout or overexpression of either ONAC127 or ONAC129 leads to incomplete grain filling and grain shrinkage, with more severe defects under heat stress. ONAC127 and ONAC129 can form heterodimers and directly target genes, including the monosaccharide transporter gene Monosaccharide Transporter 6 (OsMST6) and sugar transporter genes OsSWEET4 and OsSWEET11/15, thereby regulating apoplastic sugar transport and starch translocation from the pericarp to the endosperm. Additionally, ONAC127 and ONAC129 also directly bind to heat stress-responsive genes such as the calmodulin-like protein gene Multiple Stress Responses 2 (OsMSR2), APETALA2/Ethylene-Responsive Factor (AP2/ERF) transcription factor genes ERF protein Associated with Tillering and Panicle Branching (OsEATB), OsHSP101, and Heat and Cold Inducible 1 (OsHCI1), coordinating maintaining material transport and thermal response during grain filling[83].
The bZIP transcription factor bZIP58 serves as a key integrator of starch and storage protein accumulation under high temperature (Fig. 3). It regulates the expression of storage protein genes Glutelin type-A (GluA) and 14-kDa Prolamin (Prol14) and starch synthesis genes through alternative splicing under high temperature, while inhibiting starch-hydrolyzing α-amylase genes to maintain the accumulation of storage substances under high temperature[84] (Fig. 3). High-temperature stress induces alternative splicing of OsbZIP58, producing a truncated isoform, OsbZIP58β, which retains the 5th intron. Compared with the full-length OsbZIP58α, OsbZIP58β exhibits lower transcriptional activation ability for downstream target genes, leading to insufficient expression of storage substance synthesis genes and overexpression of hydrolysis genes. Ultimately, this imbalance results in floury and shrunken grains with reduced quality. Notably, OsbZIP58 shows subspecies-associated splicing divergence: heat-tolerant indica varieties (e.g., 9,311) exhibit lower alternative splicing ratios under high temperature (the ratio of OsbZIP58β/OsbZIP58α increases above 2-fold, whereas heat-sensitive japonica varieties (e.g., Dongjin) increase the ratio 3.5-fold. This suggests that modulating OsbZIP58 alternative splicing could offer a breeding strategy to enhance rice thermotolerance during grain filling[84].
Regulation of grain quality by endoplasmic reticulum (ER) homeostasis under high temperature
-
ER homeostasis is crucial for balancing starch and storage protein synthesis in rice grains. The normal function of ER stress sensors and their transcriptional regulation under high temperature are essential for maintaining grain quality (Fig. 3). Rice OPAQUE3/OsbZIP60 is an ER-localized transmembrane basic leucine zipper (bZIP) transcription factor that functions at the intersection of starch/protein synthesis and ER homeostasis[85]. On one hand, OPAQUE3/OsbZIP60 directly binds to and activates starch synthesis-related genes (GBSSI, AGPL2, SBEI, and ISA2) and storage protein synthesis-related genes (OsGluA2, Prol14, and Glb1), thereby promoting starch and protein accumulation. On the other hand, it targets and activates ER chaperone genes (e.g., OsBIP1 and PDIL1-1) to facilitate proper protein folding and maintain ER homeostasis. Under high-temperature stress, OPAQUE3/OsbZIP60 translocates to the nucleus and enhances the regulation of its target genes, ensuring a balance between storage synthesis and ER stress adaptation. opaque3/osbzip60 mutants show opaque endosperm, over-accumulation of 57 kDa proglutelin, and significant reductions in starch and protein contents. These defects are further exacerbated under high temperature (35 °C/28 °C) as the mutants fail to effectively cope with proteotoxic stress, leading to aggravated protein misfolding and transport disorders, increased proglutelin retention, and further downregulation of starch synthesis genes, ultimately resulting in more severe grain shrinkage and chalkiness[85].
In addition, QUALITY-THERMOTOLERANT GENE 12 (QT12), a negative regulator of grain quality and yield under heat stress, was identified through analysis of 533 global rice core germplasm accessions[74]. QT12 encodes the β subunit of the ER-localized Sec61 translocon. Under normal temperatures, the transcription factors NF-YB9, NF-YC10, and NF-YA8 form a complex, so as inhibiting NF-YA8's binding to the CCAAT-box in the QT12 promoter, resulting in low expression of QT12; this maintains the ER homeostasis and balances starch/protein synthesis, collectively conferring thermal insensitivity. Under high temperature, the interactions among NF-YB9, NF-YC10, and NF-YA8 are weakened, thus releasing NF-YA8 to bind the QT12 promoter and activate its expression. This leads to QT12 protein over accumulation, which inhibits the ER stress sensor IRE1 and triggers sustained ER stress and overactivation of the unfolded protein response (UPR). Consequently, the balance between starch and storage protein synthesis is disrupted in the endosperm, resulting in increased grain chalkiness and reduced grain quality[74].
Regulation of grain quality by mitochondria under high temperature
-
As the central organelle for energy metabolism, mitochondria play a direct role in supplying energy and supporting grain quality formation under high temperature. Mutations in genes regulating mitochondrial function can exacerbate heat-induced damage to rice quality (Fig. 3). Rice PENTATRICOPEPTIDE REPEAT PROTEIN 8 (OsPPR8), which localizes to both mitochondria and the nucleus, encodes a typical P-type PPR protein containing eight PPR motifs[86]. OsPPR8 regulates the cis-splicing of mitochondrial NAD2 intron 3, thereby affecting mitochondrial complex I activity and ATP production. This pathway supports starch synthesis in developing grains and helps maintain energy supply under high-temperature stress. Mutation of OsPPR8 leads to decreased mitochondrial complex I activity and ATP content under heat stress, resulting in reduced plant thermotolerance and poor grain quality[86].
Epigenetic regulation of grain size and quality
-
Epigenetic modification contributes to grain development and quality under high temperature by controlling the expression of key genes. Allelic variations in related genes are closely associated with cultivar differences in thermotolerance (Fig. 3). Fertilization Independent Endosperm 1 (Fie1) is a key regulator of grain width and quality under high night temperatures[87]. Fie1-knockout lines produce larger grains under normal conditions, but under high night temperatures, rice grains become significantly smaller with reduced quality, indicating that Fie1 deficiency enhances sensitivity to high night temperatures. Fie1 encodes a component of the Polycomb Repressive Complex 2 (PRC2), and its expression is regulated by temperature-sensitive epigenetic modifications, including reduced DNA methylation and decreased H3K9me2 repressive modification; severe heat stress inhibits its expression. Among different rice varieties, those carrying the major Fie1 allele (CC) exhibit wider grains and higher Fie1 transcription levels under high night temperatures, while varieties with the minor Fie1 allele (GG) show narrower grains and lower transcription levels[87].
In addition, LATE EMBRYOGENESIS ABUNDANT (OsLEA1b), a protein of previously unknown function localizing to the nucleus and cytoplasm, has been identified as influencing stress tolerance and grain quality[88]. OsLEA1b-knockout mutants show abnormal starch granule structure in endosperm cells, high chalkiness degree, significantly reduced grain weight and grain number per panicle, and significantly lower starch, protein, and lipid contents compared to the wild type; these effects are more pronounced under high temperature. OsLEA1b is also induced by drought stress, and its mutants show significantly reduced survival rates and increased ROS levels under drought stress. These results indicate that OsLEA1b affects rice grain quality under high temperature by regulating starch biosynthesis[88].
-
Currently, research has expanded beyond the cloning of genes that specifically regulate spikelet, pollen, pistil, flowering, fertilization, grain development, or quality formation under high-temperature stress. A growing number of genes that simultaneously influence both vegetative and reproductive development under thermal stress have been identified. These regulatory processes mainly encompass phytohormones and signal transduction, chloroplast development and photosynthesis, RNA processing, protein synthesis and degradation, ROS regulation, etc. (Fig. 4).
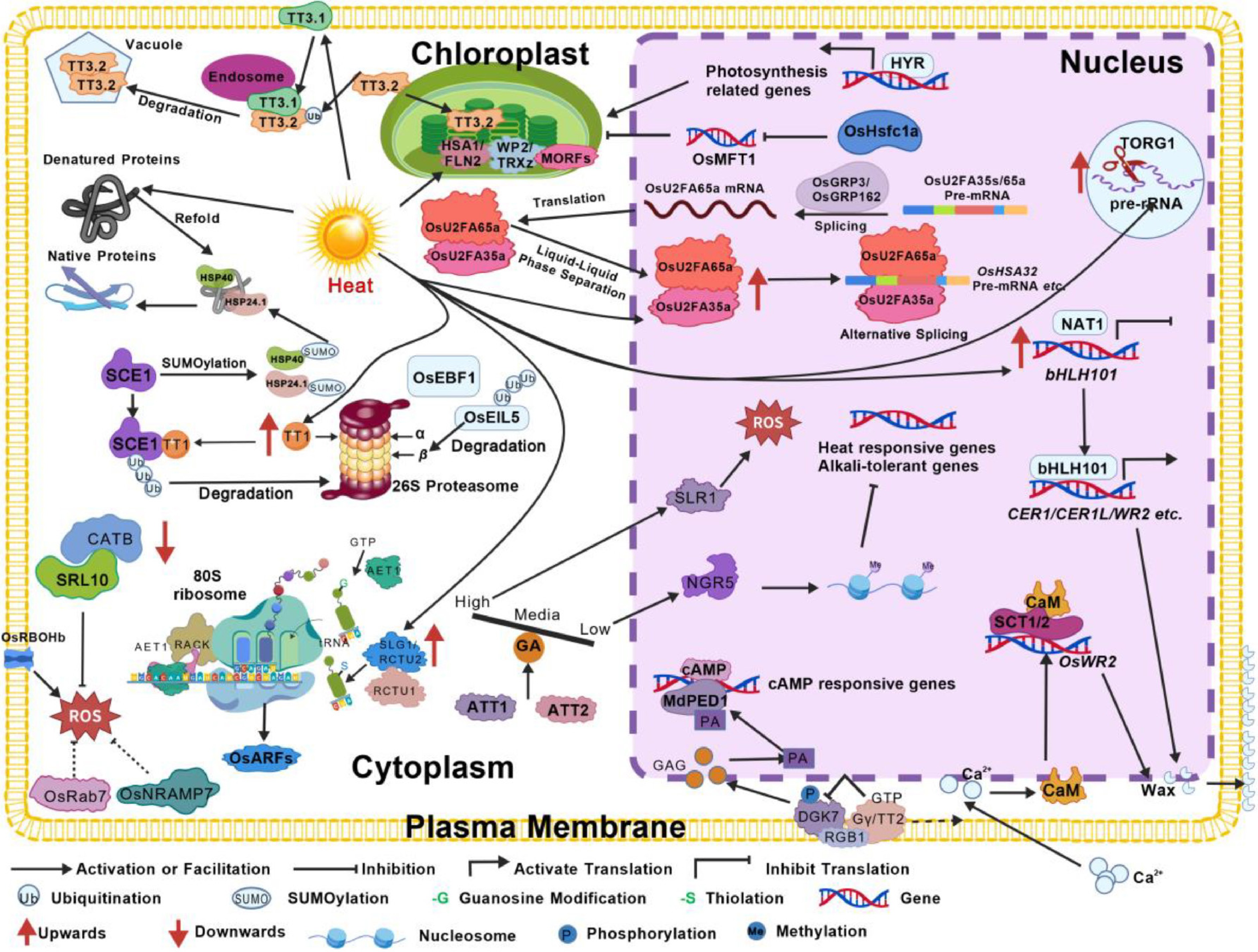
Figure 4.
Molecular networks of entire growth period of rice under high temperature. High-temperature stress triggers multi-level molecular responses that coordinately regulate rice development during both vegetative and reproductive phases. The key mechanisms can be summarized as follows: (1) Phytohormones and signal transduction: ATT1/2 encode GA20-oxidases that fine-tune gibberellin levels, balancing ROS scavenging and epigenetic regulation via the SLR1-NGR5-LC2 complex. TT2 promotes CaM-SCT1 interaction, inhibiting wax synthesis gene OsWR2 expression. TT2 inhibits diacylglycerol kinase 7 (DGK7) activity by dephosphorylation, locking metal-dependent phosphodiesterase (MdPDE1) activity and nuclear translocation, regulating the expression of the second messenger cyclic adenosine monophosphate (cAMP)-responsive genes and enhancing the heat tolerance of rice. NEGATIVE REGULATOR OF THERMOTOLERANCE 1 (NAT1) inhibits bHLH110 expression that promotes the expression of genes such as ECERIFERUM1 (CER1) and CER1L, regulating rice wax synthesis and positively enhancing rice heat resistance; (2) Chloroplast development and photosynthesis: TT3.1 ubiquitinates the chloroplast precursor protein TT3.2 for degradation, and YIELD RICE (HYR) is a key regulator of rice photosynthesis. Heat shock transcription factor OsHsfc1a enhances thermotolerance by regulating MOTHER OF FT AND TFL1 (OsMFT1) and preserving chloroplast structure under heat stress; (3) RNA processing: OsU2AF35a interacts with OsU2AF65a and undergoes liquid-liquid phase separation to promote correct pre-mRNA splicing. OsGRP3 and OsGRP162 interact with spliceosome components to regulate alternative splicing, while TOGR1 maintains rRNA homeostasis. HEAT-STRESS SENSITIVE ALBINO 1 (HSA1)/fructokinase-like protein 2 (FLN2) interacts with thioredoxin z (TRXz) regulates plant chloroplast RNA editing by controlling the redox states of MORFs, thereby contributing to the accurate translation of chloroplast-encoded proteins; (4) Protein synthesis, modification and degradation: AET1 and SLG1/RCTU2 coordinate tRNA modification and translation efficiency. TT1 encodes the α2 subunit of the 26S proteasome, clearing cytotoxic denatured proteins under heat stress and promoting degradation of SCE1 to regulate HSP accumulation by mediating their SUMOylation; (5) ROS and antioxidant regulation: OsRbohB positively regulates ROS generation, while OsRab7, OsNRAMP7, and the SRL10-CATB complex enhance antioxidant capacity and inhibit ROS production.
Phytohormones and signal transduction
-
Phytohormones and signal transduction control rice heat tolerance by regulating downstream heat response genes, ROS, and wax synthesis, etc. (Fig. 4). Exogenous application of salicylic acid (SA) and fine-tuning gibberellins (GAs) levels can enhance stress tolerance at the seedling and reproductive stages[27,89]. Recent studies show that fine-tuning gibberellin levels can increase rice stress resistance[89]. Through chromosome segment substitution lines (CSSLs), ALKALI-THERMAL TOLERANCE1/2 (ATT1/2), which encodes GA20-oxidases, are shown to regulate combined alkali-thermal tolerance by adjusting gibberellin concentrations[89]. High GA levels promote degradation of the DELLA protein SLENDER RICE1 (SLR1), activate NADPH oxidase, leading to excessive ROS accumulation and reducing seedling survival rate under alkali-thermal stress. Lower GA levels stabilize SLR1, which forms a complex with NITROGEN-MEDIATED TILLER GROWTH RESPONSE5 (NGR5), and a component of the Polycomb Repressive Complex 2 (PRC2) to mediate H3K27me3 methylation, inhibiting the expression of heat-responsive and alkali-tolerant genes and increasing stress sensitivity. Moderate GA levels can balance ROS scavenging capacity and H3K27me3 modification levels while activating the expression of stress-tolerance genes, enabling coordinated improvement of stress resistance and yield[89].
Calcium and lipid signaling molecules play an important role in heat stress response in rice. THERMOTOLERANCE 2 (TT2), which encodes a γ subunit of GTP-binding proteins, negatively regulates rice heat tolerance[90]. TT2 facilitates calcium influx to cytoplasm, which promotes the interaction between calmodulin (CaM) and sensing Ca2+ Transcription factor 1 (SCT1), and inhibits the transcription of Wax Synthesis Regulatory 2 (OsWR2), and reduced cuticular wax deposition, resulting in a heat-sensitive phenotype[90]. In addition, TT2 also regulates heat stress by lipid signaling molecules. The plasma membrane-located protein diacylglycerol kinase 7 (DGK7) catalyzes the conversion of diacylglycerol (DAG) to phosphatidic acid (PA), thereby converting physical heat signals into lipid signals[91]. And then PA binds to metal-dependent phosphodiesterase (MdPDE1), translocates to the nucleus, and degrades the second messenger cyclic adenosine monophosphate (cAMP), activating the expression of cAMP-responsive genes (such as antioxidant and heat shock-related genes) and enhancing the heat tolerance of rice[91]. TT2 inhibits DGK7 activity by Ser477 dephosphorylation, blocking MdPDE1 activity and nuclear translocation[91]. These findings reveal a new signal transduction pathway in which physical heat signals are transduced to physiological lipid signals and nuclear cAMP signaling to regulate the heat tolerance of rice. Moreover, overexpression of DGK7 and MdPDE1 results in a maximum yield increase of 108.67% in field trials, providing a potential application for the development of heat-tolerant rice[91]. In addition, NEGATIVE REGULATOR OF THERMOTOLERANCE 1 (NAT1) is a CCCH-type zinc finger protein transcriptional repressor that directly inhibits the expression of the transcription factor bHLH110[71]. In turn, bHLH110 directly promotes the expression of genes such as ECERIFERUM1 (CER1) and CER1L, which regulate rice wax synthesis and positively enhance rice heat resistance at both the seedling and reproductive stage[71].
Chloroplast development and photosynthesis
-
Chloroplast development and photosynthesis are easily affected by heat stress, contributing to a shortened life cycle and reduced yields[1]. A QTL TT3 was identified from heat-tolerant African cultivated rice variety CG14[92]. TT3 constitutes a genetic module comprising two antagonistic genes—the positive regulator TT3.1 and its substrate TT3.2. TT3.1 is a plasma membrane-localized E3 ubiquitin ligase that translocates from the cell surface to endosomes under heat stress, where it ubiquitinates the chloroplast precursor protein TT3.2. This modification targets TT3.2 for degradation through the multivesicular body-vacuole pathway, reducing the amount of mature TT3.2 protein entering chloroplasts, and thereby alleviating heat-induced chloroplast damage caused by TT3.2 accumulation. The TT3 locus from African cultivated rice (CG14) confers stronger high-temperature resistance than its Asian cultivated rice (WYJ) counterpart because the African rice TT3.1 allele encodes a protein with stronger E3 ubiquitin ligase activity, enabling more efficient degradation of TT3.2. TT3.1 is considered the first identified potential high-temperature sensor in crops, capable of transducing temperature signals into biological responses[92]. Similarly, heat shock transcription factor OsHsfc1a enhances thermotolerance, preserving chloroplast structure under heat stress by regulating MOTHER OF FT AND TFL1 (OsMFT1)[93]. The expression of OsHsfc1a is induced by heat stress, and overexpressing OsHsfc1a exhibits enhanced heat tolerance, with the yield per plant increased by 18%–21%[93]. OsHsfc1a directly binds to OsMFT1 and represses its transcription, which maintains chloroplast integrity, reduces reactive oxygen species (ROS) accumulation, enhances the heat tolerance of rice seedlings, and increases grain yield[93]. In addition, the AP2/ERF transcription factor HIGHER YIELD RICE (HYR) is a key regulator of rice photosynthesis and photosynthetic carbon metabolism[94]. It enhances photosynthetic performance under normal, drought, and high-temperature conditions, promotes chlorophyll synthesis, increases chloroplast number and starch accumulation, optimizes root architecture, improves water use efficiency and the soluble sugar content such as sucrose, and thereby improves both heat tolerance and grain quality[94].
RNA processing
-
RNA processing, including mRNA splicing, organellar RNA editing, and rRNA/tRNA processing and modification, plays an essential role in maintaining rice heat tolerance during the reproductive stage (Fig. 4). Recent studies highlight the critical importance of accurate mRNA splicing under high temperature. The rice splicing cofactor U2 snRNP auxiliary factor 35a (OsU2AF35a) interacts with OsU2AF65a to splice pre-mRNAs such as OsHSA32 by liquid-liquid phase separation (LLPS), thereby enhancing rice heat tolerance[95]. GLYCINE-RICH RNA-BINDING PROTEIN 3 and 162 (OsGRP3 and OsGRP162) interact with spliceosome components OsU1-70K and OsU2AF35a/b, and are involved in regulating alternative splicing of mRNA and regulating rice heat tolerance at both seedling and reproductive stages[96]. HEAT-STRESS SENSITIVE ALBINO 1 (HSA1), encoding a plastid-localized fructokinase-like protein 2 (FLN2), interacts with thioredoxin z (TRXz) in rice chloroplasts and regulates plant chloroplast RNA editing by controlling the redox states of MORFs, thereby contributing to the accurate translation of chloroplast-encoded proteins[97]. The nucleolus-localized DEAD-Box RNA Helicase Thermotolerant Growth Required1 (TOGR1) binds to the small subunit (SSU) precursor rRNA processing complex and helps maintain rRNA homeostasis under high temperature to ensure cell division and growth[98]. In addition, microRNA biosynthesis also affects plant thermo-tolerance. The RNA-binding protein SEMI-ROLLED LEAF 10 (SRL10) influences rice leaf morphology by regulating microRNA biosynthesis. It also interacts with catalase isozyme B (CATB) to enhance the stability of both proteins, thereby improving hydrogen peroxide scavenging capacity and rice heat tolerance[99]. The srl10 mutant shows partially-rolled leaves and heat sensitivity, with decreased seedling survival and seed-setting rates under high-temperature stress, while overexpression of SRL10 increases both seedling survival and seed-setting rates[99].
Protein synthesis, modification, and degradation
-
High temperatures easily affect protein synthesis, modification, and degradation. Multiple key regulatory genes and pathways involved in this process have been identified so far (Fig. 4). ADAPTATION TO ENVIRONMENTAL TEMPERATURE 1 (AET1), a tRNAHis guanylyltransferase, and tRNA 2-thiolation protein 2 (RCTU2)/Slender Guy 1 (SLG1) participates in the rice high-temperature response through coordinated tRNA modification and translational regulation[100]. AET1 maintains tRNA homeostasis by catalyzing the addition of a guanosine modification at the 5' end of pre-tRNAHis. AET1 also interacts with RACK1A and eIF3h and directly binds mRNAs containing upstream open reading frames (uORFs) (such as OsARF19 and OsARF23) to regulate their translation efficiency[100]. A major heat-tolerance QTL, THERMOTOLERANCE1 (TT1), was identified using chromosome segment substitution lines (CSSLs) derived from heat-tolerant African rice (Oryza glaberrima), and heat-sensitive Asian rice WYJ[101]. TT1 encodes the α2 subunit of the 26S proteasome, and the TT1 allele from African rice confers stronger heat tolerance due to an amino acid substitution (Arg99His), which enhances the proteasome activity. This enables more efficient clearance of cytotoxic denatured proteins under heat stress, thereby maintaining cellular homeostasis[101]. TT1 also interacts with the negative thermotolerance regulator SUMO-conjugating Enzyme 1 (SCE1), and promotes its degradation via the ubiquitination pathway. SCE1 affects the accumulation of HSPs (e.g., Hsp24.1, Hsp40) by mediating their SUMOylation, and loss of SCE1 function enhances HSP accumulation, and improves heat tolerance[102]. These findings indicate that TT1 and SCE1 regulate rice heat tolerance through ubiquitination and SUMOylation pathways.
ROS homeostasis
-
High temperatures easily induce excessive accumulation of ROS. Regulating ROS production and scavenging through related genes to maintain ROS homeostasis represents an important physiological mechanism for rice to resist high-temperature stress across growth stages (Fig. 4). OsRBOHb is a key gene regulating ROS production. Knocking out OsRBOHb reduces ROS accumulation in seeds, seedlings, anthers, and grains, significantly improving rice tolerance to high temperature, along with increased growth, grain yield, and quality. In contrast, OsRBOHb-overexpression lines show the opposite phenotypes. These results indicate that inhibiting OsRBOHb provides an effective strategy to alleviate rice heat damage and improve yield and quality[103]. Overexpression of the rice small GTP-binding protein gene OsRab7 (OsRab7) significantly upregulates the expression of antioxidant enzyme genes (e.g., SOD, APX, POD), ROS-scavenging related genes (e.g., OsCATA, OsCATB), and stress-responsive genes (e.g., Dehydration-Responsive Element-Binding Protein 2A, DREB2A), Late Embryogenesis Abundant (OsLEA3)), thereby enhancing rice tolerance to both drought and high-temperature stresses and increasing grain yield[104]. In addition, Natural resistance-associated macrophage protein 7 (OsNRAMP7), which localizes to the ER and Golgi apparatus, shows significantly increased expression under under heat stress[105]. It mediates rice heat tolerance by regulating the antioxidant system, water balance, and ribosome-photosynthesis coordination network, functioning positively at both seedling and reproductive stages[105]. These findings collectively demonstrate that eliminating abnormal ROS accumulation is a key mechanism for enhancing plant heat tolerance.
-
In recent years, groundbreaking progress has been made in research on rice heat tolerance with the rapid advancement of molecular biology, genome, various gene cloning methods, and gene editing, crop cultivation science, and so on. A series of heat tolerance-associated genes have been cloned and functionally characterized, and various efficient strategies for heat tolerance improvement have been gradually applied in practice. We propose several key strategies to enhance rice heat tolerance, so as to provide a reference for subsequent relevant research and application.
Gene cloning for heat tolerance
-
Molecular breeding has become a core approach for rice heat tolerance improvement, with the advantages of high precision and efficiency. Mapping and cloning heat tolerance-related QTLs and functional genes can rapidly breed new heat-tolerant rice germplasms and varieties with superior agronomic traits via gene editing or molecular marker-assisted selection enables the accumulation of multiple elite heat-tolerant alleles. For example, several important QTLs, including TT1, TT2, TT3, and qHTH5, have been cloned from wild rice (O. rufipogon) or heat-tolerant cultivars such as African rice (O. glaberrima) using approaches including genome-wide association study (GWAS), map-based cloning, chromosome segment substitution lines (CSSLs), and near-isogenic lines (NILs), etc[32,90−92,101]. These naturally occurring QTL variants can improve rice heat tolerance without compromising other agronomic traits, and can be used for molecular marker-assisted screening of heat-tolerant haplotypes to enhance yield stability under high-temperature conditions. The introduction of heat-tolerant TT2, TT3, or QT12 into the elite rice cultivar increased the grain yield per plant by 54.7%, 20%, and 77.9%, respectively, compared with the control under high-temperature stress[74,90,92]. Therefore, cloning heat-tolerance genes with no obvious adverse effects from different heat-tolerant rice cultivars or wild rice resources, and then introgressing these genes into elite commercial cultivars with superior agronomic traits, enables the targeted improvement of rice heat tolerance. The improved cultivars exhibit strong adaptability and are thus amenable to large-scale promotion and application in agricultural production.
Regulation of the core pathways of heat tolerance
-
High-temperature stress impairs various reproductive and developmental processes of rice. To counteract high-temperature damage, rice activates its endogenous heat signal transduction pathways and initiates the expression of a series of heat tolerance-associated genes. The key heat signal transduction pathways in rice provide important molecular targets for improving rice heat tolerance.
First, the heat tolerance of rice at the reproductive stage can be enhanced by upregulating the expression of positive regulators of heat tolerance and downregulating or knocking out negative regulators. For example, TT1 acts as a positive regulator of heat tolerance, and its overexpression can increase the seed setting rate of rice by approximately 30% under high-temperature stress[101]. TT2 negatively regulates rice heat tolerance through the DGK7–PA–MdPDE1–cAMP signaling pathway[91]. Single-gene modification of DGK7 or MdPDE1 achieved a 50%–60% increase in grain yield compared with the control, while double-gene modified lines of TT2 and DGK7 nearly doubled the grain yield with significantly improved rice quality[91].
Second, fine-tuning the expression of heat tolerance-associated genes is an effective way to improve rice heat tolerance. For example, both high and low concentrations of GA are unfavorable for rice heat tolerance, whereas a moderate GA concentration achieves a synergistic improvement in stress resistance and grain yield[89]. With the development of gene editing technology, editing the promoter or 5' untranslated region (5' UTR) of GA synthesis-related genes to screen lines with an optimal GA level has great potential for practical application in rice production.
Finally, heat-inducible elements can be inserted into the promoters of key yield-associated genes by gene editing technology to acquire heat-tolerant traits. For example, a 10-bp heat-responsive element was introduced into the promoter regions of tomato LIN5 and rice GIF1 by using an optimized Csy4-PE prime editing system, resulting in the yields of the edited rice and tomato lines being increased by 25% and 33% under heat stress, respectively, without any compromise to grain or fruit quality[69]. In conclusion, dissecting the regulatory network of rice heat tolerance and integrating it with modern molecular breeding technologies can greatly improve the heat tolerance of rice.
Alleviating high-temperature damage
-
Alleviating high-temperature damage is an agronomic measure to counteract high-temperature stress in current rice production. Adjustment of sowing date, which changes the planting time of rice to make its key growth stages (heading and flowering stage, grain-filling stage) avoid the high-temperature stress period, fundamentally reduces high-temperature damage. Rice varieties with earlier DFOT can avoid damage from high midday temperatures by editing DFOT-related genes such as DFOT1/EMF1 and PMEs[59]. In addition, application of exogenous growth regulators is one of the effective measures to alleviate high-temperature stress and improve spikelet fertility. For example, exogenous application of plant growth hormones such as salicylic acid can enhance the stress tolerance of rice at the seedling and reproductive stages[27]. Exogenous application of some chemical molecules, such as spermidine, can improve heat stress resistance[26]. In summary, these hormones or chemical molecules can be formulated into agents for single or mixed spraying, which can effectively reduce yield loss under high-temperature stress.
AI-based molecular breeding
-
With the rapid development of artificial intelligence (AI), AI-based molecular breeding may be a promising trend in crop improvement. First, AI can efficiently analyze massive multi-omics data, including genomics, transcriptomics, and proteomics, to obtain candidate genes significantly associated with heat tolerance[106,107]. Second, AI models (e.g., AlphaFold) can accurately predict the three-dimensional (3D) structure of proteins, RNA, and DNA[108]. Meanwhile, AI can predict the functions and interaction relationships of genes and proteins through sequence features, which greatly reduces the trial-and-error cost of subsequent wet experiments and accelerates the dissection of molecular mechanisms underlying heat tolerance[109]. For example, a SNP mutation in the 26S proteasome subunit encoded by TT1 enhances proteasome activity under heat stress. Novel 26S proteasomes with higher heat tolerance can be designed based on generative AI models, and then introduced into rice for functional validation. In this way, heat tolerance genes are not only screened from natural variations but also created through AI-based molecular design breeding, which enriches the heat-tolerant germplasm resource bank and improves breeding efficiency.
Intelligent environmental regulation strategies
-
Against the backdrop of intensifying global climate change and frequent extreme weather events, traditional agricultural production is severely constrained by environmental stresses and can hardly ensure sustained and stable crop yields. Therefore, developing intelligent strategies for high and stable yield based on digital technologies has become a core direction of modern agricultural transformation. Centered on intelligent environmental perception, precise regulation, and smart decision-making, this strategy establishes a full-coverage environmental monitoring system via the Internet of Things, to collect real-time multi-dimensional data on soil, meteorology and air. It intelligently regulates key environmental factors, including water, fertilizer, temperature, humidity, and light, according to the requirements of crops at different growth stages, and realizes accurate early warning and control of pests and diseases with artificial intelligence. Meanwhile, it selects high-quality varieties adapted to regional environments, optimizes cultivation patterns, and establishes an intelligent early warning and emergency stress-resistance mechanism for extreme weather, achieving visualized and intelligent management of the whole production process through a smart decision-making platform. Supported by an all-round guarantee system of technology, policy, talent, and standards, this strategy effectively breaks the limitation of traditional agriculture 'relying on weather conditions'. It not only improves the stability of crop yields and safeguards food security, but also promotes the green and low-carbon development of agriculture, driving the transformation of modern agriculture toward intelligence, high efficiency, and sustainability.
-
The research on heat tolerance in rice began with early agronomic observations of heat damage, gradually progressing to in-depth analyses of physiological and biochemical mechanisms. With advances in genetics and molecular biology, the focus shifted to screening heat-tolerant germplasm, mapping QTLs, and cloning and functionally validating key genes. Recently, genomics, multi-omics integration, and gene editing technologies have driven the systematic analysis of heat tolerance regulatory networks, enabling a leap from mechanistic exploration to molecular design breeding. However, substantial research gaps and challenges remain to be addressed. In the future, we propose that future research on rice thermotolerance may focus on cloning important QTLs, enclosing their complex multi-level regulatory networks, applying in real-world field, and AI-based breeding.
Analysis of network regulation and field application
-
Early research on rice thermotolerance primarily focused on identifying and validating the functions of individual key genes through laboratory experiments. However, this single-gene approach struggled to explain the complexity of quantitative traits and often showed diminished effects in field conditions. With the rapid advancement of omics technologies—including genomics, transcriptomics, proteomics, and metabolomics—the research focus has shifted from searching for single key genes to deciphering complex multi-level regulatory networks. These networks encompass signal transduction, transcriptional regulation, protein synthesis and modification, and epigenetic mechanisms, emphasizing their cross-talk with developmental stages and responses to other stresses. In particular, cloning QTL from wild rice or thermotolerant varieties and elucidating their regulatory networks holds significant applied value. Concurrently, the validation framework for rice thermotolerance is transitioning from controlled laboratory environments to real-world field scenarios, where thermotolerance is evaluated for its practical agricultural value under dynamic and compounded stress conditions. These two major shifts—from single gene to network and from laboratory to field—are jointly advancing the design of climate-resilient rice varieties with enhanced thermotolerance.
Limited exploration and utilization of germplasm resources
-
Most successfully cloned genes so far are associated with heat-sensitive phenotypes or basal heat tolerance—where loss of function leads to sensitivity under heat stress. In contrast, few genes conferring active heat resistance have been identified, resulting in an incomplete understanding of the genetic basis of rice heat tolerance and a very limited pool of superior alleles available for breeding. Excellent heat-tolerance gene alleles present in wild rice and diverse local rice germplasm resources remain underexplored and underutilized. Additionally, current genetic improvement for heat tolerance still mostly relies on traditional breeding and limited QTLs, with the scale and efficiency of applying new technologies such as gene editing and synthetic biology needing to be improved.
A potential solution is to combine population genetics with functional genomics to 'mine for gold' in germplasm banks with rich diversity. Whole-genome resequencing has been performed on diverse rice populations, including local Asian cultivars, African rice cultivars, and wild rice relatives. Combined with multi-year, multi-location heat-tolerance phenotype data, genome-wide association studies (GWAS) may be conducted to identify new heat-tolerance QTLs/alleles. Furthermore, establishing a heat-tolerance evaluation system for core germplasm will enable large-scale, phenotypically precise identification and classification of germplasm resources.
Challenges in precise phenotyping
-
Rice heat tolerance is a complex quantitative trait influenced by multiple genes and environmental factors. Conventional phenotyping methods, often conducted in greenhouses and artificial climate chambers, or using treatments at the seedling stages, fail to simulate complex field conditions involving diurnal temperature fluctuations, combined stresses (e.g., high light, drought), and other environmental variables. This can lead to discrepancies between controlled environment screenings and actual field performance. A gene conferring heat tolerance under constant temperature may be completely ineffective under realistic stress combination in the field. Although some teams have attempted to develop high-throughput field identification methods, standardizing and accurately quantifying heat-tolerance phenotypes (e.g., seed-setting rate, chalkiness degree) at different growth stages (especially heat-sensitive flowering and grain-filling stages) remains a major challenge. Most growth models underestimate the impact of short-term extreme heat on yield, partly due to inaccurate parameterization of sensitive stages and temperature thresholds. To address these issues, integrating remote sensing technology, hyperspectral imaging, infrared thermal imaging, light detection and ranging (LiDAR), and artificial intelligence (AI) can enable non-destructive, automated, and precise c field phenotyping. Developing image-based algorithms to automatically quantify traits such as seed-setting rate, pollen viability, and chalkiness degree will improve efficiency and accuracy. Furthermore, establishing a multi-ecological-site, multi-year field experiment network will enable large-scale identification of heat tolerance in germplasm resources across different rice-growing regions. Combining meteorological data to evaluate material performance during naturally occurring heat waves ensures that gene-mediated heat tolerance is expressed in real agricultural environments, thereby screening gene resources with practical application potential.
In summary, the core shifts in future rice heat tolerance research will be: (1) From identifying 'laboratory heat-tolerance genes' to discovering 'field heat-tolerance genes'; (2) From studying responses to 'single high-temperature stress' to analyzing mechanisms of 'combined stresses'; (3) From focusing on 'single-trait improvement' to pursuing 'synergistic enhancement of grain yield, quality, and heat tolerance'. The ultimate goal is to breed novel rice varieties capable of withstanding complex climate challenges, and ensuring global food security.
This work was supported by grants from the Guangdong Basic and Applied Research (Grant No. 2022B1515120036), National Natural Science Foundation of China (Grant No. 32472215), and Guangdong Provincial Special Innovation Projects for Regular Institutions of Higher Education (Grant No. 2024KTSCX023).
-
The authors confirm their contributions to the paper as follows: prepared the draft manuscript: Peng G, Li K, Li L, Li S; modified the draft manuscript: Liu Z; conceived and modified the draft manuscript: Liu Y, Zhou H. All authors reviewed and approved the final version of the manuscript.
-
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of Hainan Yazhou Bay Seed Laboratory. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Peng G, Li K, Li L, Li S, Liu Z, et al. 2026. Molecular basis of heat tolerance in rice. Seed Biology 5: e016 doi: 10.48130/seedbio-0026-0013
Molecular basis of heat tolerance in rice
- Received: 09 December 2025
- Revised: 13 February 2026
- Accepted: 24 February 2026
- Published online: 06 May 2026
Abstract: Rice is one of the most important food crops and global warming, and extreme high temperatures severely threaten rice production. Rice is highly sensitive to heat stress during the reproductive stage, resulting in pollen sterility, a reduced seed-setting rate, and poor grain quality. Therefore, identifying key genes and elucidating the molecular mechanisms responding to heat stress can be leveraged to develop climate-resilient rice varieties, ultimately enhancing adaptation to the impact of future climate change on global rice production. Considerable progress has been made in elucidating the mechanisms of heat stress in rice. This review summarizes recent advances in our understanding of the molecular mechanisms underlying heat tolerance in rice, with a focus on the reproductive stage, including spikelet development, pollen fertility, pistil development, diurnal flower-opening time, grain filling, and grain quality. Additionally, we propose integrated strategies for mitigating high-temperature damage, addressing persistent challenges, and outlining future research directions for breeding heat-tolerant rice varieties.
-
Key words:
- High temperature /
- Heat tolerance /
- Reproductive stage /
- Rice













