-
The widespread use of antibiotics in clinical, agricultural, and aquaculture settings has led to the accumulation of antibiotic resistance genes (ARGs) in anthropogenically impacted environments[1−5]. Their cross-species and cross-regional transmission has driven the globalization of antimicrobial resistance (AMR), posing a substantial threat to global public health[6−8]. In 2015, antibiotic-resistant bacteria (ARBs) were responsible for 671,689 infections and over 33,000 annual deaths[9]. In 2021, AMR was associated with 4.7 million deaths, with 1.14 million directly attributable cases[10]. Without coordinated policy intervention, antibiotic consumption is projected to double by 2030, further intensifying the environmental burden of ARGs[11,12].
Anthropogenically impacted freshwater ecosystems are important reservoirs of ARGs in the environment, as they receive continuous inputs from wastewater, agricultural runoff, and aquaculture effluents[13−16]. These systems not only support biodiversity and human life but are also highly susceptible to microbial pollution and ecological disruption[17−19]. Recent studies suggest that cyanobacteria may play a role in the dynamics of ARGs in freshwater ecosystems[20−22]. Under conditions of nutrient enrichment and climate-induced thermal stratification, cyanobacteria have become increasingly dominant in freshwater systems[23,24]. Large-scale cyanobacterial harmful algal blooms (CyanoHABs) have become more frequent and severe, affecting over 20,000 lakes and ~57% of global lake surface area[25,26].
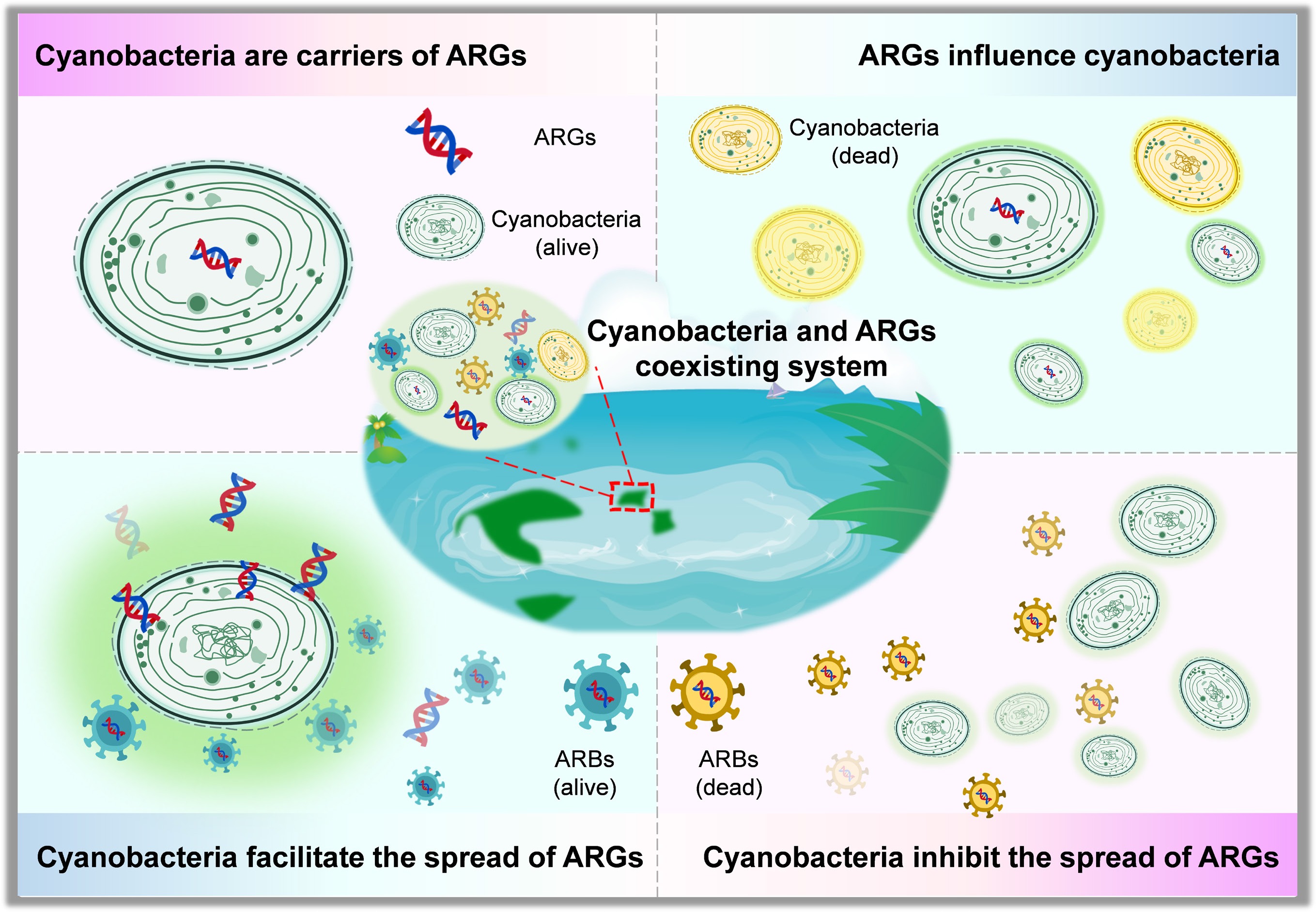
Cyanobacteria share genetic and physiological traits with other bacteria, allowing interactions with ARGs[27,28]. Emerging evidence suggests cyanobacteria play dual roles in ARGs dynamics[29−35]. On the one hand, they can facilitate ARG propagation via mobile genetic elements (MGEs)[36], extracellular polymeric substances (EPS)[37,38], and the formation of microhabitats such as the phycosphere[36,39−43]. Quorum-sensing (QS) systems add another layer of regulation, modulating both ARGs expression and HGT efficiency, thereby potentially accelerating resistance propagation in aquatic environments[44]. On the other hand, they may suppress ARGs' dissemination by altering community structure, inhibiting conjugation, or producing antimicrobial metabolites[45−48]. Additionally, ARGs acquisition may enhance cyanobacterial resistance under antibiotic pressure, further complicating bloom dynamics[49,50].
Existing studies report inconsistent conclusions regarding the relationships between cyanobacteria and ARGs, mainly attributable to differences in spatial scales and environmental contexts. In particular, it remains unclear under which environmental conditions cyanobacteria primarily function as ARG carriers or ecological regulators, or whether they simultaneously perform both roles, and how environmental drivers and microbial interactions jointly shape these patterns. By synthesizing current evidence on environmental drivers and microbial interactions, this review aims to clarify the complex coupling mechanisms between cyanobacteria and ARGs in freshwater ecosystems, identify key sources of uncertainty, and provide an integrated perspective to support future research and management of co-occurring CyanoHABs and ARGs risks.
-
To systematically characterize the current research landscape and emerging trends concerning cyanobacteria-ARGs relationships, a bibliometric analysis was conducted using global publication data retrieved primarily from the ISI Web of Science database, with literature identified through keyword combinations related to cyanobacteria, antibiotic resistance genes, and freshwater environments (detailed literature search strategy and inclusion criteria are provided in Supplementary Texts S1 and S2, with screening criteria summarized in Supplementary Table S1). The analysis emphasized the number of publications, temporal trends, and knowledge gaps within freshwater ecosystems (Fig. 1a−c).
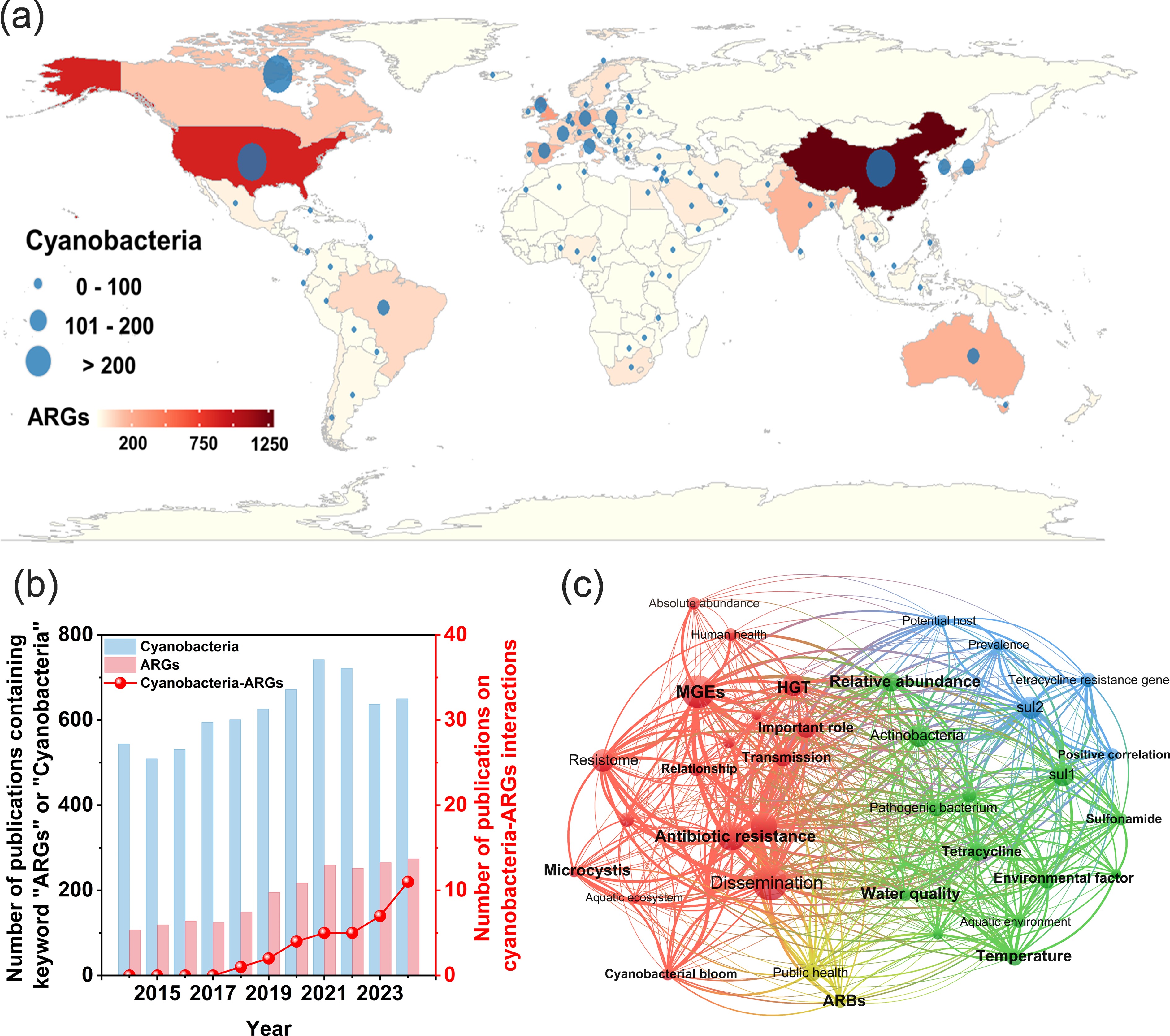
Figure 1.
Mapping the global co-occurrence patterns of eutrophication and ARGs based on cyanobacteria-ARGs-related studies and database integration. Global publication analysis on cyanobacteria and ARGs research: (a) Global distribution of publications related to cyanobacteria and ARGs. (b) Annual publication trends on cyanobacteria, ARGs, and their interactions in freshwater ecosystems. (c) Keyword co-occurrence network highlighting major research themes in the international literature.
As shown in Fig. 1a, among the 164 countries assessed, 117 published 2,089 articles on ARGs, while 118 published 6,829 articles on cyanobacteria. These findings highlight widespread scientific interest in both topics, particularly amid growing concerns about freshwater contamination and public health risks. Figure 1b presents the publication trends from 2010 to 2023. While research on cyanobacteria and ARGs individually continues to rise, studies explicitly addressing their relationship remain limited, with only 35 publications identified. Nonetheless, this number has shown steady growth, suggesting increasing recognition of the topic's significance. Keyword co-occurrence network analysis (Fig. 1c) revealed that current cyanobacteria-ARG research predominantly focuses on tetracycline- and sulfonamide-resistance genes. Environmental variables and anthropogenic activities emerged as key drivers shaping their interaction. However, most existing studies rely on relative abundance data, with limited efforts to quantify both cyanobacteria and ARGs in absolute terms. In addition, the potential influences of climate-related and anthropogenic stressors on their relationship remain insufficiently explored. It should be noted that this bibliometric analysis is inherently influenced by database coverage, keyword selection, and language restrictions, and therefore reflects dominant research trends rather than an exhaustive representation of all relevant studies.
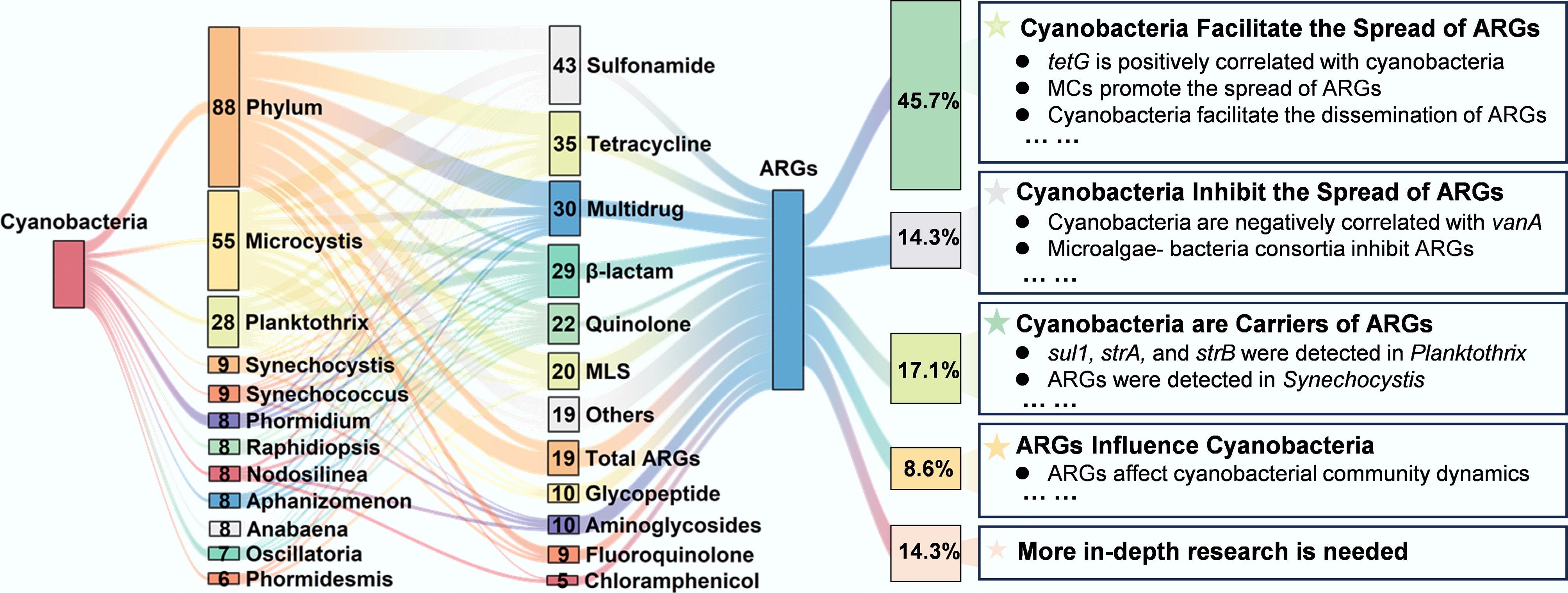
Based on the detailed analysis of 35 existing studies on the relationships between cyanobacteria and ARGs, this study identified and categorized several major interaction patterns (Supplementary Fig. S1). Evidence for cyanobacteria facilitating ARG dissemination was reported in 45.7% of the studies, while 14.3% reported inhibitory effects, and 17.1% classified cyanobacteria as potential ARG hosts. Additionally, 8.6% of studies documented feedback effects of ARGs on cyanobacterial community structure, while 14.3% proposed more complex bidirectional mechanisms that remain to be fully elucidated. In terms of research targets, 53.8% of the studies focused on cyanobacteria at the phylum level. At the genus level, the most frequently studied taxa were toxin-producing cyanobacteria Microcystis (25.6%) and Planktothrix (5.1%) (Supplementary Fig. S2). Figure 2 summarizes co-occurrence patterns between various cyanobacterial genera and ARG types, revealing non-uniform interaction modes that vary depending on cyanobacterial taxonomy and ARGs class (see Supplementary Table S2 for detailed conclusions).
-
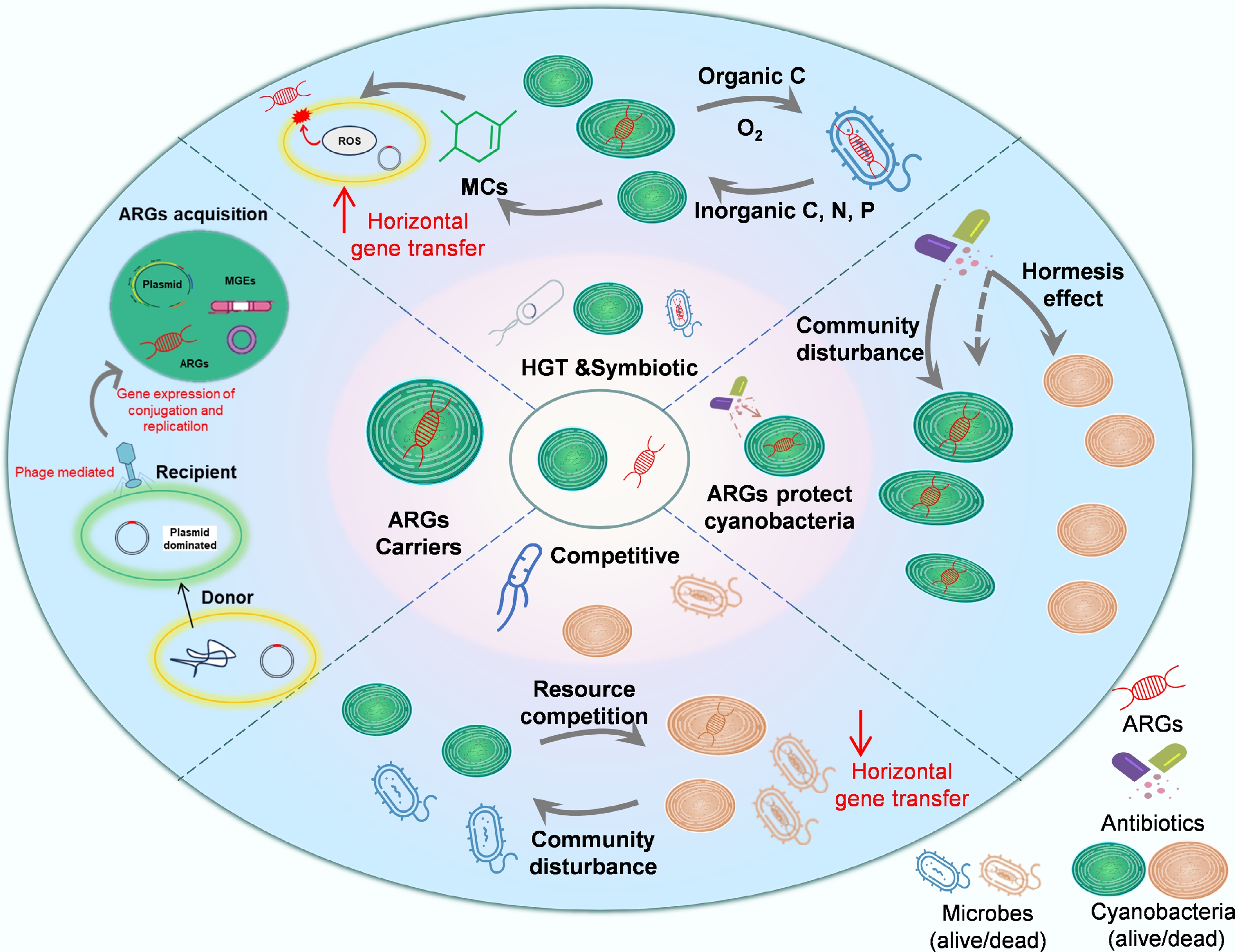
With the intensification of global climate change and anthropogenic disturbances, the interactions between cyanobacteria and ARGs have become increasingly complex. Table 1 summarizes the key physiological and ecological traits of common cyanobacterial genera. These traits not only enhance the environmental adaptability of cyanobacteria in freshwater environments but also play a critical role in the dissemination of ARGs. Figure 3 illustrates the mechanisms of interaction between cyanobacteria and ARGs. For clarity, the mechanisms discussed below are supported by different levels of evidence, ranging from laboratory and mesocosm experiments to field observations and omics-based associations. A summary of the coupling mechanisms and their corresponding levels of evidence is provided in Supplementary Table S3.
Table 1. Common bloom-forming cyanobacterial genera in freshwater ecosystems
Cyanobacterial genus Physiological traits and ecological functions Mechanisms of interaction with ARGs Anabaena Produces microcystins (MCs), anatoxins, or cylindrospermopsin. Capable of nitrogen fixation and can withstand low nitrogen environments[51,52]. 1. MCs can induce oxidative stress in bacteria, leading to the generation of reactive oxygen species (ROS), which subsequently stimulate lysozyme activity and alter bacterial membrane permeability. Increased membrane permeability is considered one of the key factors that enhance the efficiency of ARGs transfer[30,53].
2. Neurotoxins may disrupt bacterial transmembrane potential, thereby increasing membrane permeability and facilitating the transformation of ARGs.
3. Extracellular polysaccharides secreted by toxin-producing strains can serve as carriers for ARGs transfer, while also acting as physical barriers that protect ARBs from antibiotic exposure.
4. During algal blooms, large amounts of dissolved organic matter (DOM) are released, providing carbon sources for heterotrophic bacteria carrying ARGs. Meanwhile, DOM may influence HGT among bacteria through QS mechanisms.
5. Cyanobacteria may suppress the spread of ARGs by competing with ARG-hosting bacteria for resources, thereby reducing the abundance of ARG-hosting bacteria.Microcystis Production of MCs and anatoxins often forms harmful algal blooms[54,55]. Oscillatoria Some species produce MCs or neurotoxins, which are widely distributed[56,57]. Planktothrix Produces MCs, widespread in freshwater lakes[58]. Nodularia Some species produce Nodularin or MCs, and nodularin inhibits protein phosphatase synthesis[59,60]. Aphanizomenon Capable of nitrogen fixation, often forming large-scale algal blooms, can produce MCs[52,61]. Cylindrospermopsis Nitrogen fixation, production of cylindrospermopsin or MCs by some species[62,63]. Gloeotrichia Forms buoyant clusters capable of nitrogen fixation and can produce MCs[64]. Phormidium Some species produce neurotoxins[65]. Raphidiopsis Some species produce toxic substances[66]. Dolichospermum Capable of nitrogen fixation[67]. Lyngbya Some species produce toxic substances[68]. Cyanobacteria are carriers of ARGs
-
In recent years, increasing evidence has shown that cyanobacteria can act as key reservoirs and carriers of ARGs in aquatic environments[29]. For example, in sediment samples collected along the coast of Saudi Arabia, cyanobacteria were identified as a phylum harboring AMR-related genetic sequences, as determined by metagenomic analyses[69]. Typical bloom-forming cyanobacteria, such as Microcystis and Planktothrix, have been found to harbor large numbers of tetracycline- and sulfonamide-resistance genes, which can be disseminated via both extracellular DNA (eDNA) and intracellular DNA (iDNA), as evidenced by metagenomic analyses complemented by quantitative PCR validation[35]. Similarly, multiple ARGs have been detected in cyanobacterial strains isolated from Portuguese reservoirs using culture-dependent molecular screening[70].
The mechanisms by which cyanobacteria act as carriers of ARGs may involve multiple processes. First, under selective pressure from antibiotic contamination, cyanobacteria may activate intrinsic tolerance mechanisms, thereby increasing their ability to acquire or retain ARGs. Second, cyanobacteria have the potential to act as carriers of ARGs, as their genomes harbor MGEs such as transposons and plasmids[22,34]. Certain cyanobacterial species may acquire ARGs from the surrounding environment through HGT[71]; for example, Synechococcus and Phormidium have been shown to acquire ARGs via phage-mediated transfer[45]. These genomic features suggest that cyanobacteria are not only capable of acquiring exogenous ARGs but may also serve as long-term reservoirs.
During dense bloom events such as CyanoHABs, frequent cell-to-cell contact among cyanobacterial populations facilitates HGT processes such as plasmid conjugation, significantly enhancing the intra-population spread of ARGs. This provides further evidence of the host potential of cyanobacteria in the dissemination of ARGs[47]. Additionally, the mechanisms underlying ARGs' carriage in cyanobacteria are influenced by multiple factors, including species-specific traits, physiological states, environmental conditions, and the composition of co-occurring microbial communities[45]. These complex interactions remain poorly understood. Future studies should focus on the species-specific mechanisms by which cyanobacteria acquire and disseminate ARGs in different environments, highlighting their role as carriers of ARGs.
Cyanobacteria facilitate the spread of ARGs
-
During CyanoHABs in certain regions, the abundance and diversity of ARGs have been observed to increase significantly, particularly in eutrophic waters. Bloom-forming cyanobacteria such as Microcystis and Planktothrix play a pivotal role in the dissemination of ARGs, not merely as carriers, but also by facilitating HGT and shaping microbial and environmental conditions conducive to ARG spread.
Cyanobacteria promote the spread of ARGs through multiple mechanisms. Firstly, they can enhance the rate of HGT, thereby facilitating the dissemination of ARGs. For instance, significantly elevated HGT activity has been reported during Planktothrix blooms[50]. MCs produced by Microcystis can disrupt cellular signaling pathways, induce oxidative stress, increase membrane permeability, and cause DNA damage, all of which can promote HGT processes[48]. Metagenomic analyses have revealed that Microcystis-dominated blooms are associated with elevated abundance and diversity of both ARGs and MGEs, suggesting that Microcystis may foster conditions favorable to the dissemination of ARGs in aquatic environments[22].
Secondly, experimentally demonstrated cyanobacteria may indirectly facilitate the spread of ARGs by altering microbial community composition. Cyanobacteria produce a variety of bioactive compounds, including macrolides, alkaloids, indole alkaloids, polyphenols, and terpenes[72], which can exert selective pressure within the microbial ecosystem, enriching ARBs and thereby promoting the accumulation and transmission of ARGs[46]. Certain ARG-carrying taxa, such as Actinobacteriota and Proteobacteria, can also utilize decaying cyanobacterial biomass as a carbon source, leading to increased abundance during the post-CyanoHABs period[73].
Moreover, the metabolic activity and signaling pathways of cyanobacteria may contribute to the dissemination of ARGs. Cyanobacterial metabolites and exudates may form complex chemical signaling networks that mediate interactions between microbes and ARGs, further enhancing ARGs transfer[74]. For example, cyanobacteria may regulate plasmid conjugation and transposon activity through QS. Under environmental stress, acyl-homoserine lactones can activate LuxI/LuxS signaling pathways, increasing the efficiency of ARGs exchange between cyanobacteria and associated microbial communities[44].
Cyanobacteria inhibit the spread of ARGs
-
Although many studies have demonstrated the role of cyanobacteria in promoting the dissemination of ARGs, opposing evidence suggests that cyanobacteria may also suppress the spread of ARGs through various pathways. On one hand, the rapid proliferation of cyanobacteria in aquatic systems can significantly alter physicochemical parameters and microbial community structures, creating environmental stress that negatively affects ARBs. For example, during CyanoHAB events, the abundance of ARGs in both free-living and particle-attached bacteria was significantly reduced, with ARG-carrying functional groups in particle-attached bacteria being more strongly affected[75]. This indicates that bacterial lifestyle (free-living vs particle-attached) may mediate differential responses to cyanobacterial blooms.
On the other hand, in contrast to Microcystis, some cyanobacterial genera exhibit a negative correlation with MGEs, suggesting that these taxa may suppress HGT and reduce ARGs abundance in aquatic environments[22]. Additionally, it has been reported that algal–bacterial consortia can secrete large quantities of maleimides, which inhibit ARGs dissemination by interfering with the replication of exogenous plasmids. However, the environmental thresholds that control this process remain unclear[76]. A microbial interaction modeling approach proposed by Bao et al. provides a quantitative tool for assessing such processes[77]. By quantifying the degree of cyanobacterial interference with microbial communities, this approach may support more predictive assessments of ecological risks associated with ARGs.
These findings suggest that different cyanobacterial taxa, under varying environmental conditions, may influence ARGs dynamics through mechanisms such as community disruption, resource competition, and metabolic inhibition. Future studies should examine diverse nutrient regimes and environmental stressors across different water types and cyanobacterial community structures, while differentiating bacterial lifestyles. Such efforts may help uncover key regulatory pathways and critical thresholds for cyanobacteria-mediated ARGs dissemination.
ARGs influence cyanobacteria
-
Certain antibiotics have been reported to exert algicidal effects on cyanobacteria, acting as inhibitors under specific conditions[78,79]. However, cyanobacteria can display remarkable environmental adaptability in response to such stressors[80]. Studies have shown that cyanobacteria may acquire ARGs from surrounding ARBs, enhancing their survival under antibiotic pressure[50]. Antibiotic-resistant cyanobacterial populations may expand through several adaptive strategies: (1) leveraging resistance advantages to inhibit or outcompete susceptible algal species[81−83]; (2) absorbing intracellular organic matter released from dead competitors as nutrient sources; and (3) selectively enriching symbiotic bacteria capable of vitamin biosynthesis and pollutant degradation, thereby forming more stable microbial consortia that further facilitate interspecies ARGs transfer[81]. This resistance-driven ecological advantage not only increases the frequency of CyanoHAB outbreaks but also promotes the evolution of resistance traits through sustained gene accumulation and positive selection.
The effects of antibiotics on cyanobacteria vary substantially with concentration. Owing to the hormesis effect, low concentrations of antibiotics may stimulate cyanobacterial growth and toxin production while simultaneously enhancing ARGs expression and transfer, thereby accelerating the development of AMR in cyanobacterial populations[81,84−88]. In contrast, high antibiotic concentrations can cause photosystem-specific damage in Microcystis, inhibiting cell growth and protein synthesis, potentially suppressing bloom formation[75,89,90]. This nonlinear dose-response relationship suggests the existence of ecological thresholds in cyanobacteria-ARGs interactions, beyond which the direction of ecological feedback may shift. However, systematic studies are lacking to unravel the coupled and cascading effects among antibiotic types and concentrations, cyanobacterial community structures, and ARGs dynamics.
The ecological advantage conferred by acquired ARGs not only enhances the competitive and dispersal capacities of cyanobacteria but also contributes to more frequent and persistent harmful algal blooms. The coexistence of ARGs and cyanobacteria may pose emerging threats to downstream ecosystems and human populations in urbanized regions[51,91]. Although the potential role of ARGs in enhancing cyanobacterial environmental resistance has been recognized, their underlying molecular mechanisms and regulatory networks remain to be fully elucidated.
-
The interaction between cyanobacteria and ARGs is a complex environmental issue. Investigating the roles of environmental factors, anthropogenic activities, and climate change in shaping this interaction is essential to elucidate the underlying mechanisms across different ecological levels. Such insights can inform the development of targeted prevention and control strategies, reduce the occurrence of CyanoHABs and ARGs pollution, and ultimately contribute to protecting aquatic ecosystem health and public safety.
Environmental factors
-
Natural factors such as nutrient concentrations, temperature, pH, light intensity, and hydrodynamic conditions form the fundamental basis for regulating both cyanobacterial proliferation and the dissemination of ARGs. These parameters not only independently affect cyanobacteria and ARGs, but also shape the ecological context in which their interactions occur.
Nitrogen and phosphorus are key determinants of eutrophication levels and cyanobacterial biomass in freshwater ecosystems[26]. When TN:TP < 25, cyanobacteria tend to dominate, whereas higher ratios favor the growth of green algae and diatoms[23]. Such shifts in community composition can indirectly regulate ARGs dynamics by altering microbial assemblages, interspecies interactions, and the prevalence of cyanobacterial genera with differing ARGs association potentials. Nutrient levels can also influence the composition of cyanobacterial genera; in eutrophic waters, Microcystis, Planktothrix, Raphidiopsis, and Dolichospermum often dominate, potentially increasing their association with ARGs[92]. Eutrophication has been shown to significantly enhance the dissemination potential of ARGs within microbial communities[50].
Temperature is another major driver of both cyanobacterial growth and ARGs behavior. Beyond promoting bloom formation, elevated temperatures increase microbial metabolic activity and the frequency of gene exchange events, thereby enhancing ARG dissemination in aquatic environments[26,52]. Moreover, rising temperatures can accelerate the evolution of AMR in cyanobacteria; for example, when the temperature increases from 16 to 36 °C, the level of genotypic resistance may increase by up to 295-fold[88].
Physicochemical parameters such as pH, conductivity, and water transparency not only influence cyanobacterial growth, but also modulate the microenvironmental conditions that shape cyanobacteria–ARGs interactions. For example, weakly alkaline conditions (pH 8.0–8.5) favor cyanobacterial proliferation and may enhance extracellular polymer production, creating niches conducive to ARGs retention and HGT[93]. Elevated conductivity, reflecting higher ionic strength, can alter microbial community structure and increase cell membrane permeability, potentially facilitating the exchange of ARGs. Reduced water transparency, by enhancing light-use efficiency, may promote dense bloom formation and microbial aggregation, thereby stabilizing ARGs reservoirs[94,95]. Additionally, low wind speed, hypoxia, and stratified water columns limit environmental disturbance and oxygenation, creating stable conditions that may further promote ARGs persistence and interaction with CyanoHABs[96].
Anthropogenic activities
-
Anthropogenic activities, particularly agriculture, industrialization, and urbanization, have emerged as key drivers of intensified interactions between cyanobacteria and ARGs in freshwater ecosystems[97]. Agricultural non-point source pollution and municipal wastewater discharge have markedly increased nitrogen and phosphorus concentrations in surface waters, providing a nutrient-rich foundation for rapid cyanobacterial expansion[98−100]. The proliferation of cyanobacteria creates highly active ecological niches that facilitate close interactions with microbial communities, enhancing the HGT of ARGs among microorganisms[101].
The direct release of antibiotics and ARGs is another major pathway through which human activities influence cyanobacteria-ARGs interactions[102−106]. Antibiotics used in livestock farming, medical treatment, and household applications often enter aquatic systems through wastewater discharge. These inputs not only exert selective pressure on cyanobacteria but also lead to the rapid accumulation of ARGs in the environment[36]. Uncontrolled sources such as septic tank leakage, damaged sewage pipelines, and leachate from landfills further contribute to a complex network of ARGs inputs into aquatic environments[106,107]. In response to antibiotic and chemical stressors, bacteria have evolved diverse resistance mechanisms and can transfer ARGs via HGT to other microorganisms, including cyanobacteria[29].
In summary, anthropogenic activities accelerate the environmental accumulation of ARGs not only through direct pollutant inputs but also by altering environmental conditions that favor cyanobacterial growth and ARG dissemination. This dual impact poses a significant risk to the stability of aquatic ecosystems and public health, warranting greater scientific and regulatory attention.
Climate change
-
Global climate change, particularly rising temperatures and increasingly frequent extreme weather events, has emerged as an important driver influencing the interaction between cyanobacteria and ARGs[88,108]. Temperature increases can reshape aquatic microbial community composition, often leading to shifts in the relative abundance of cyanobacteria, heterotrophic bacteria, and antibiotic-resistant bacteria, thereby altering ARGs dynamics at the community level. Metagenomic analyses from temperature-controlled experiments have shown that warming can significantly reduce ARGs diversity while increasing their overall abundance, suggesting heightened resistance risks under global warming scenarios[50,109].
Under high greenhouse gas emission scenarios, the mean annual surface temperature of lakes is projected to rise by approximately 2.2 °C by the end of the century[110]. This warming trend is expected to exacerbate eutrophication, increase algal biomass, and degrade water quality, thereby enhancing both cyanobacterial proliferation and ARGs dissemination[111]. In many freshwater systems, warming favors cyanobacteria over other phytoplankton and microbial groups, increasing their relative dominance and the frequency of bloom events, which in turn strengthens their role as potential ARG carriers and interaction hubs. Even moderate warming (2–5 °C) has been found to promote the spread of toxic cyanobacteria and antimicrobial resistance[112]. Furthermore, global warming indirectly intensifies oxidative stress among aquatic microorganisms, facilitating cyanobacterial growth and increasing ARGs transfer, thus compounding ecological risks to freshwater systems[52,112]. Climate change may also accelerate the HGT of ARGs within ARB populations[113,114]. These processes suggest that increases in cyanobacterial abundance represent a key pathway linking warming to ARGs dynamics, alongside temperature-driven changes in resistant bacterial populations and microbial interaction networks.
High-temperature conditions also favor the production of cyanotoxins, the proliferation of ARGs and aquatic pathogens, and longer-lasting, more intense CyanoHABs[115−117]. Importantly, climate change may indirectly influence ARGs transmission pathways by altering agricultural practices, including increased pesticide and antibiotic use[118]. In naturally multidrug-exposed environments, elevated temperatures may further accelerate the evolution of antimicrobial resistance in cyanobacterial populations[88].
In the face of the intertwined 'Climate-CyanoHABs-ARGs' challenge, quantitative assessments from local to global scales are urgently needed. Priority should be given to integrated management strategies that yield both ecological and societal co-benefits, such as enhanced nutrient control, ecological restoration, and stricter regulation of pesticide and antibiotic usage.
Microbial interactions and ecological relationships
-
The complex interactions among microorganisms, particularly involving symbiosis, competition, and metabolic cooperation, play a critical role in shaping microecological environments and influencing the dynamics between cyanobacteria and ARGs[50]. Studies have shown that HGT of ARGs between free-living microalgae and bacteria occurs at low frequencies. In contrast, mutualistic interactions between microalgae (e.g., green algae) and their associated bacteria can stimulate the production of antimicrobial compounds and create chemically selective environments that favor the enrichment of ARGs. Similar mechanisms are hypothesized to exist in cyanobacterial symbiotic networks, potentially altering the accumulation and dissemination of ARGs[119].
During bloom events, cyanobacteria and their associated microbes often form complex interspecies aggregates, such as algal flocs and biofilms, which provide a stable microhabitat for ARBs[36]. These microenvironments promote ARBs survival and increase the likelihood of ARGs exchange via HGT by enhancing physical contact among microbial cells. Microbial aggregation and metabolic cooperation within these micro-niches contribute to ARGs persistence and facilitate their transfer[43].
However, microbial ecological relationships are not limited to cooperation; competitive interactions also occur. Such contradictory outcomes may arise from interspecific differences among cyanobacteria (e.g., Microcystis, Planktothrix, and Anabaena exhibit distinct environmental responses) as well as from variations in microbial community composition, nutrient levels, and the intensity of antibiotic pollution.
Overall, microbial ecological interactions are likely to be dynamic, context-dependent, and shaped by multiple biotic and abiotic factors. Future research should focus on elucidating the mechanisms underlying cyanobacteria-ARG host interactions, identifying key environmental drivers that modulate symbiosis-competition transitions, and uncovering the true influence of these complex microbial networks on the potential for ARG dissemination.
-
Experimental studies and field observations indicate that cyanobacteria and antibiotic resistance genes (ARGs) interact in complex ways within freshwater ecosystems. The nature of these interactions differs among cyanobacterial taxa and ARG types, and is further influenced by climate change, environmental factors, and microbial community succession. A deeper mechanistic understanding of cyanobacteria–ARG dynamics is essential for public health protection, effective water resource management, and mitigation of antimicrobial resistance. Despite recent advances, significant knowledge gaps remain, necessitating targeted research in the following areas:
(1) Current studies predominantly rely on relative ARGs abundance analyses, with limited attention to precise, taxon-specific correlations between cyanobacterial genera and absolute ARGs abundances. As a result, most existing evidence reflects spatiotemporal co-variation rather than direct causal relationships. Quantitative, spatiotemporally matched investigations across local to global scales are needed to disentangle the combined effects of climate change, human activities, and microbial community restructuring on cyanobacteria–ARGs interactions.
(2) The effects of ARGs on cyanobacterial population dynamics, community structure, and ecological functions remain poorly understood. Future work should integrate long-term monitoring with multifactorial experiments to clarify these bidirectional interactions.
(3) The lack of comprehensive databases encompassing diverse geographic regions, climatic conditions, and waterbody types hinders mechanistic insights into their coupling effects and potential cascading impacts. Constructing multidimensional knowledge frameworks linking environmental parameters (e.g., water quality, anthropogenic pressures), cyanobacterial traits, and ARGs profiles could reveal their coevolutionary patterns and inform management strategies.
(4) Given the stark differences in cyanobacterial community composition, environmental adaptability, and microbial interactions across habitats (e.g., marine systems, wetlands), systematic comparisons of cyanobacteria-ARGs dynamics in disparate ecosystems are essential to generalize their environmental behaviors.
-
It accompanies this paper at: https://doi.org/10.48130/biocontam-0026-0001.
-
The authors confirm their contributions to this review as follows: Yang Yu: data collection and cleaning, generated all figures and tables and wrote the first draft of the manuscript; Zhiguang Niu: developed the initial concept and provided research funds; Yifan Zhao: contributed to the revision and proofreading of this paper; Shuaiyi Li: contributed to the revision and proofreading of this paper; Ying Zhang: contributed to the revision and proofreading of this paper; Chenchen Wang: developed the initial concept. All authors reviewed the results and approved the final version of the manuscript.
-
Raw data used in this study have been made publicly available via Environmental Data Initiative (https://doi.org/10.6073/pasta/f777b79289993b4676e458913cf78690).
-
This study was supported by the National Natural Science Foundation of China (Grant Nos 42477409 and 42277374).
-
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
-
Full list of author information is available at the end of the article.
- The supplementary files can be downloaded from here.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Yu Y, Niu Z, Zhao Y, Li S, Zhang Y, et al. 2026. An integrated conceptual framework of coupling mechanisms between cyanobacteria and antibiotic resistance genes in freshwater ecosystems. Biocontaminant 2: e004 doi: 10.48130/biocontam-0026-0001
An integrated conceptual framework of coupling mechanisms between cyanobacteria and antibiotic resistance genes in freshwater ecosystems
- Received: 10 December 2025
- Revised: 13 January 2026
- Accepted: 23 January 2026
- Published online: 11 March 2026
Abstract: Cyanobacteria are integral components of freshwater ecosystems. As prokaryotes, they can harbor antibiotic resistance genes (ARGs); however, the co-occurrence patterns between specific ARGs and cyanobacteria observed in metagenomic data remain challenging to interpret. This review synthesizes current evidence by integrating qualitative assessment, bibliometric trend analysis, and a conceptual framework to examine cyanobacteria–ARG interactions and their modulating drivers, such as climatic variation, anthropogenic pressures, environmental factors, and microbial interdependencies. Two principal findings are identified: cyanobacteria function both as carriers of ARGs and as ecological regulators, and many documented cyanobacteria–shared environmental drivers influence ARG relationships. The review highlights the need for taxon-resolved quantitative datasets, mechanistic studies of ARG-mediated cyanobacterial resilience, and cross-system comparisons utilizing multi-omics and predictive modeling. Advancing these research directions will improve understanding of cyanobacteria–ARG linkages and enhance risk assessment in the context of global warming and increasing anthropogenic impact.
-
Key words:
- Cyanobacteria /
- ARGs /
- Relationships /
- Driving factors /
- Freshwater ecosystems