-
Mulberry (Morus spp.), a perennial woody plant belonging to the family Moraceae in the order Rosales[1], is not only the primary food source for silkworms (Bombyx mori) but also possesses significant medicinal value, including blood glucose regulation and antioxidant properties[2,3]. Hybridization serves as a key strategy in plant improvement, facilitating novel gene flow and enhancing desirable traits across diverse plant species[4−6]. In mulberry breeding, hybrids exhibiting potential salt tolerance have been developed[7]. Early hybrid identification is crucial for enhancing breeding efficiency. However, studies on the authentication of Morus hybrids remain limited. Traditional identification methods, such as morphological trait observation[8], are often inadequate because of the pronounced morphological variation observed in mulberry hybrids compared with their parental lines[9]. Therefore, relying solely on morphological characteristics for identifying hybrids is unreliable.
High-throughput molecular DNA markers provide a robust framework for hybrid screening. For instance, 11 single nucleotide polymorphism (SNP) markers have been used to distinguish intraspecific and interspecific hybrids of grain amaranth (Amaranthus spp.)[10]. Similarly, 70 simple sequence repeat (SSR) primers and 8 inter-simple sequence repeat (ISSR) primers have been used to detect genetic diversity and polymorphism in Lilium, facilitating the identification of hybrids and analyses of their genetic relationships[11,12]. In Morus, phylogenetic analysis using internal transcribed spacer and sequence-characterized amplified region markers has enabled the discrimination of putative interspecific hybrids[9,13,14]. However, molecular marker-based hybrid identification faces challenges in polyploid genomes, where the presence of polymorphic loci and varying ploidy levels complicates the correspondence between markers and species[15,16]. This limitation is particularly prominent in autopolyploid species; for instance, SNP markers often fail to differentiate between allelic variants and homologous SNPs arising from genome duplication[17]. The genus Morus exhibits extensive ploidy variation and frequent interspecific hybridization, resulting in a highly complex genetic background[18−22]. Consequently, though molecular DNA markers offer partial insights into hybrid identification, their efficacy is limited in dissecting the precise genomic contributions of the parental lineages.
Cytogenetics, particularly molecular cytogenetic tools such as fluorescence in situ hybridization and genomic in situ hybridization (GISH), is a powerful approach for identifying hybrids and genomic characterization in plants[23]. For instance, hybrid populations in the genus Veronica were successfully identified by comparing the signal numbers of 45S and 5S ribosomal DNA (rDNA) between parents and offspring[24]. Similarly, GISH analysis enabled the accurate identification of intergeneric hybrids between Saccharum officinarum and Narenga porphyrocoma while revealing their chromosomal constitution and n + n inheritance patterns[25]. Recently, the application of GISH has elucidated the genetic relationships among 12 species and three varieties in the genus Morus, leading to the establishment of a cytotaxonomy system comprising five sections and two subsections[20]. Subsequent studies employing comparative GISH (cGISH) with blocking DNA and self-GISH refined this system further, proposing the division of the section Wittiorum into two sections, Wittiorum and Laevigata. Notably, GISH offers several advantages, including reference-genome-free analysis, cost-effectiveness, rapid processing, and intuitive chromosomal visualization. These features make it particularly valuable for studying wild Morus species such as Morus laevigata and Morus wittiorum, which lack reference genomes. Unlike molecular markers (e.g., SSRs or SNPs) that provide fragmented genetic locus information, GISH enables the direct identification of parental chromosome sets and their recombination events in hybrid progenies[26]. Although GISH has limitations in detecting small-fragment genomic structural variations and in discriminating species with extremely close genetic relationships[27], the technique remains critical for analyzing polyploid Morus hybrids, where comprehensive chromosomal insights are essential.
The formation of 2n gametes in plants can occur through pre-meiotic or postmeiotic genome doubling, as well as via meiotic restitution[28−30]. These gametes have been widely documented across diverse plant species, including wild potato (Solanum malmeanum) and lemon (Citrus limon)[31−33]. As a pivotal mechanism for sexual polyploidization, 2n gametes contribute significantly to spontaneous polyploid formation in plants[29,34,35]. For instance, the detection of 2n gametes in cultivated caladium (Caladium × hortulanum) and oil palm (Elaeis guineensis) has provided direct evidence for naturally occurring sexual polyploidization events in these species[36,37]. Furthermore, 2n gametes serve as a crucial tool for artificial polyploid induction, offering significant potential for plant breeding[38]. High-temperature treatment has been shown to induce unreduced 2n pollen, with successful applications reported in Lagerstroemia indica and Camellia oleifera[39,40]. However, no evidence for the occurrence of 2n gametes has been reported in the genus Morus, highlighting a critical gap in polyploidy research and mulberry breeding programs.
The exceptional resistance of M. laevigata to Sclerotinia scleroterum and its elongated fruit phenotype underscore its potential as valuable genetic resources for introgression via hybridization[41]. Moreover, M. wittiorum 'Twcgs' was found to contain significantly higher levels of rutin and nicotiflorin compared with other mulberry cultivars[42]. Previous studies have also shown that hybridization can effectively enhance the content of quercetin 3-(6-malonylglucoside)[43]. On the basis of these findings, six cross combinations were designed as part of breeding programs aimed at improving fruit length, enhancing resistance to S. scleroterum, and increasing flavonoid accumulation. In this study, GISH was used to confirm the hybrid authenticity of seven hybrid progenies derived from these six cross combinations and to characterize the chromosomal composition of two wild mulberry accessions. Comparative analysis against the respective parental lines conclusively verified the hybrid status of all seven progenies. This research provides a reliable method for early identification of mulberry hybrids, which facilitates the effective utilization of mulberry resources and supports advanced genetic breeding programs.
-
Intersectional hybridization experiments were conducted using 10 mulberry accessions (Morus spp.) selected from three taxonomic sections (Alba, Wittiorum, and Laevigata). These accessions were selected because of their superior morphological traits and typical cytotaxonomic characteristics, prioritizing germplasm possessing homologous chromosomes[20]. All accessions, including the wild genotypes and hybrids, were maintained through grafting or seeding in the Mulberry Germplasm Nursery at Southwest University, Chongqing, China. Three representative accessions, namely Morus multicaulis 'Heyebai' (Ma), M. laevigata 'Menghai No. 2' (Ml), and M. wittiorum 'Ailaoshan No. 9' (Mw), were used for genomic DNA extraction. Detailed information regarding the mulberry accessions, including those used for genomic DNA extraction, the parental lines, and the wild specimens, is provided in Supplementary Table S1. Additionally, the seven hybrids from six cross combinations are documented in Supplementary Table S2.
Chromosome preparation
-
Mitotic chromosome preparation followed established protocols[44]. Briefly, fresh leaf and root tip samples were pretreated with 2 mM 8-hydroxyquinoline at room temperature for 3 h, fixed in 3:1 (v/v) ethanol : glacial acetic acid for 4 h, and stored in 70% ethanol at 4 °C. After three washes with distilled water, the tissues were enzymatically digested at 37 °C in a solution containing 2% (w/v) cellulase Onozuka R-10 (YaKult, Japan) and 1% (w/v) pectolyase Y-23 (YaKult, Japan) (pH 5.5). The duration of digestion varied by tissue type: 3 h for leaves and 1 h for root tips. The digested samples were rinsed with 70% ethanol and macerated into a cell suspension. For slide preparation, one drop of the cell suspension was mixed with glacial acetic acid. Chromosome spreads were observed using an Olympus IX73 microscope (Olympus, Japan).
Genomic probe labeling and preparation of blocking DNA
-
Genomic DNA was extracted from young leaves of Ma, Ml, and Mw using the DNAquick Plant System kit (TIANGEN BIOTECH, Beijing, China) according to the manufacturer's instructions. Genomic DNA was then fluorescently labeled by nick translation with either ChromaTide Alexa Fluor 488-5-dUTP (Thermo Fisher Scientific [Invitrogen], Massachusetts, USA) or Texas-Red-5-dCTP (PerkinElmer, Massachusetts, USA)[20]. Blocking DNA (~200-bp fragments) was generated by autoclaving genomic DNA from Ma, Ml, and Mw at 121 °C for 2 min.
Genomic in situ hybridization
-
GISH analysis was performed according to our previously established protocol[20]. Chromosome slides were crosslinked by ultraviolet light at 1,250 mJ/cm2 for 2 min prior to hybridization. The probe mixture (15 ng/μL of fluorescently labeled DNA in a hybridization buffer: 2 × Saline-Sodium Citrate Buffer (SSC) and 1 × TE; pH 7.0) was denatured with chromosomes at 100 °C for 5 min and then incubated overnight at 42 °C. For GISH with blocking DNA, the hybridization mixture was supplemented with blocking DNA (30× the probe concentration). After being washed in 2× SSC at room temperature for 5 min, the slides were counterstained with 4',6-diamidino-2-phenylindole (DAPI) (1 ng/μL) and sealed with nail polish. For multiple rounds of GISH, the probes were stripped in 50% formamide (containing 2× SSC) at 42 °C for 10 min after removing the coverslips. The slides were dehydrated in an ethanol series (70%, 90%, and 100% for 3 min each at room temperature) before the next hybridization round. Images were captured using an Olympus DP80 CCD camera (Olympus, Japan) under an Olympus IX73 fluorescence microscope and processed using Adobe Photoshop 2021 and Adobe Illustrator 2021 (Adobe, USA).
To ensure accurate and reliable GISH signal quantification, signals were defined as discrete fluorescent foci exceeding an intensity threshold of 50 arbitary units (AU). For each hybrid individual, at least 20 metaphase cells exhibiting a clear chromosome distribution were imaged. Enumeration of the signals within these cells was independently performed by three researchers (Qiming Zhou, Jiacheng Li, and Ziang Li of Nanjing Agricultural University, Nanjing, China), with the results used for subsequent analyses.
-
M. wittiorum and M. laevigata exhibit the longest fruit morphologies in the genus Morus (Supplementary Fig. S1), making them valuable genetic donors for intersectional hybridization programs focused on improving fruit traits. In the present study, 10 mulberry accessions spanning three taxonomic sections (Alba, Wittiorum, and Laevigata) were selected as parental lines for systematic intersectional crosses (Supplementary Table S1), with the crossing scheme detailed in Supplementary Table S2. All designed crosses successfully produced hybrid progeny. Crosses 1−6 yielded 280, 816, 5, 85, 36, and 925 seeds, respectively. For cytological verification of the plants' hybridity, one vigorous individual plant was randomly selected for each cross. To assess their chromosomal composition, GISH was performed using the representative cultivars Ma (Alba), Ml (Laevigata), and Mw (Wittiorum), as either genomic probes or blocking DNA (Supplementary Table S1). This analysis encompassed seven progeny lines derived from six crosses (Supplementary Table S2).
GISH analysis of intersectional hybrids between Morus section Alba and section Wittiorum
-
Two intersectional hybrids were successfully generated between Morus section Alba and section Wittiorum: (1) M. alba 'A-1' × M. wittiorum 'W-1' and (2) M. wittiorum 'W-2' × M. alba 'A-2'. Genomic probes derived from Ma and Mw were used reciprocally as either hybridization probes or blocking DNA to analyze chromosomal compositions. An initial GISH analysis without blocking DNA revealed substantial signal co-localization between the Ma and Mw probes in both the parents and the hybrid progeny, preventing unambiguous chromosomal assignment (Supplementary Fig. S2). To resolve this limitation, cGISH was subsequently performed by using standardized blocking DNA, which enabled precise discrimination of the parental chromosomal contributions in the hybrid offspring.
In the M. alba 'A-1' × M. wittiorum 'W-1' cross, cGISH analysis revealed parental genomic distributions as follows. The Ma probe with Mw blocking DNA identified four hybridization signals in the female parent (Fig. 1a−c). The Mw probe with Ma blocking DNA detected 26 signals in the male parent (Fig. 1j−l). Their hybrid progeny, 'Mp-1', showed differential incorporation with two Ma-derived and 13 Mw-derived signals (Fig. 1d−i). The cross M. wittiorum 'W-2' × M. alba 'A-2' exhibited parallel findings. The Mw probe with Ma blocking revealed 32 signals in the female parent (Fig. 2j−l). The Ma probe with Mw blocking detected four signals in the male parent (Fig. 2a−c). The hybrid 'Mp-2' demonstrated two Ma-specific and 16 Mw-specific signals (Fig. 2d−i).
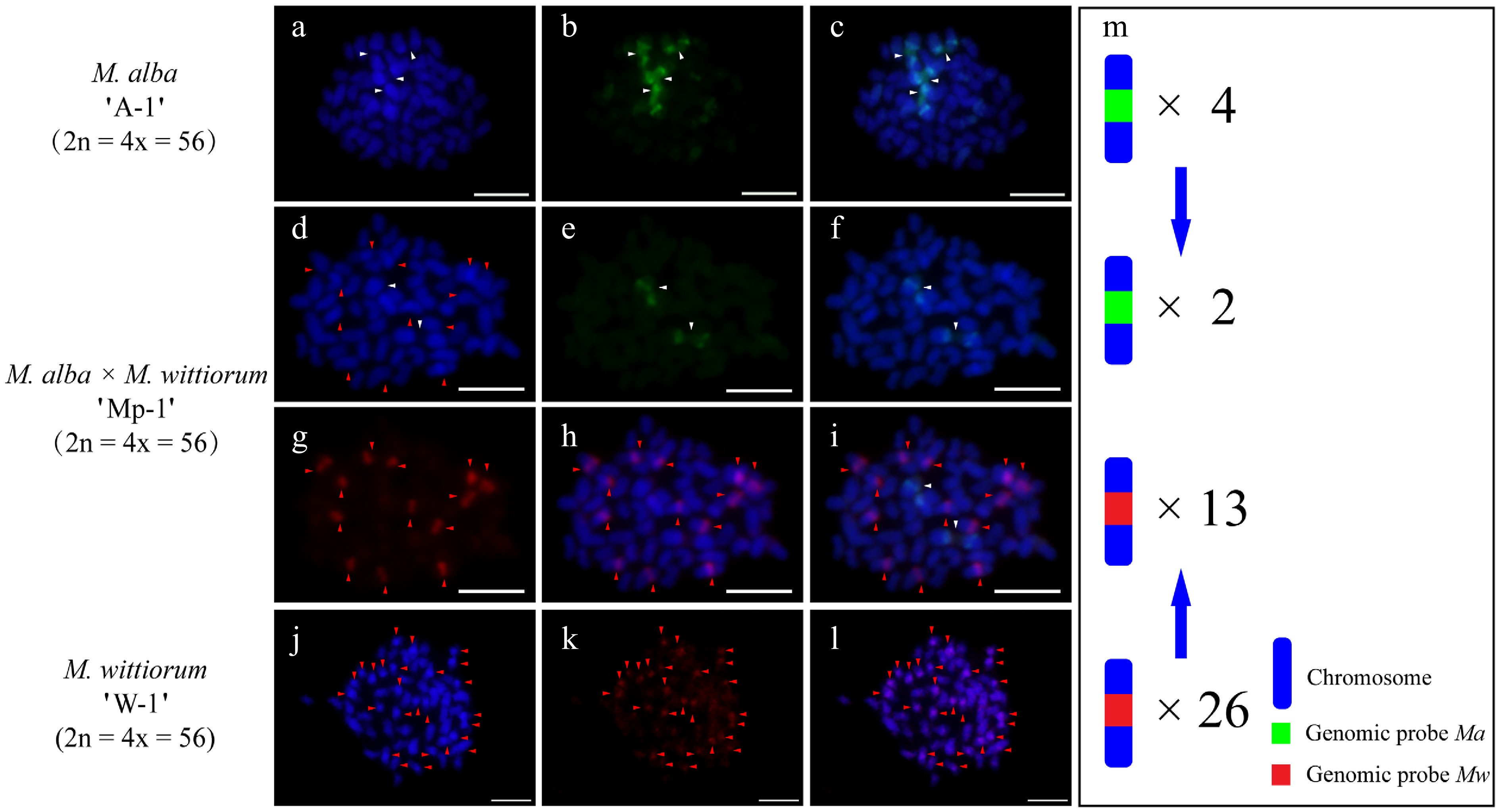
Figure 1.
The cGISH signal patterns in M. alba 'A-1', M. wittiorum 'W-1', and their hybrid M. alba × M. wittiorum 'Mp-1'. cGISH signals using the genomic probe from M. alba (Ma) (green) with blocking DNA from M. wittiorum (Mw) were detected in (a)−(c) M. alba 'A-1' and (d)−(f) the hybrid 'Mp-1'. cGISH signals using the genomic probe from Mw (red) with blocking DNA from Ma were detected in (g), (h) the hybrid 'Mp-1' and (j)−(l) M. wittiorum 'W-1'. (i) The two-round cGISH showing overlapping signals from Ma (green) and Mw (red) genomic probes with reciprocal blocking DNA. (m) Ideogram summarizing chromosome counts with different genomic probes' signals. White and red arrows point to chromosomes with Ma and Mw signals, respectively. Blue arrows indicate transmission of the parental chromosome to the hybrid. Scale bars represent 5 μm.
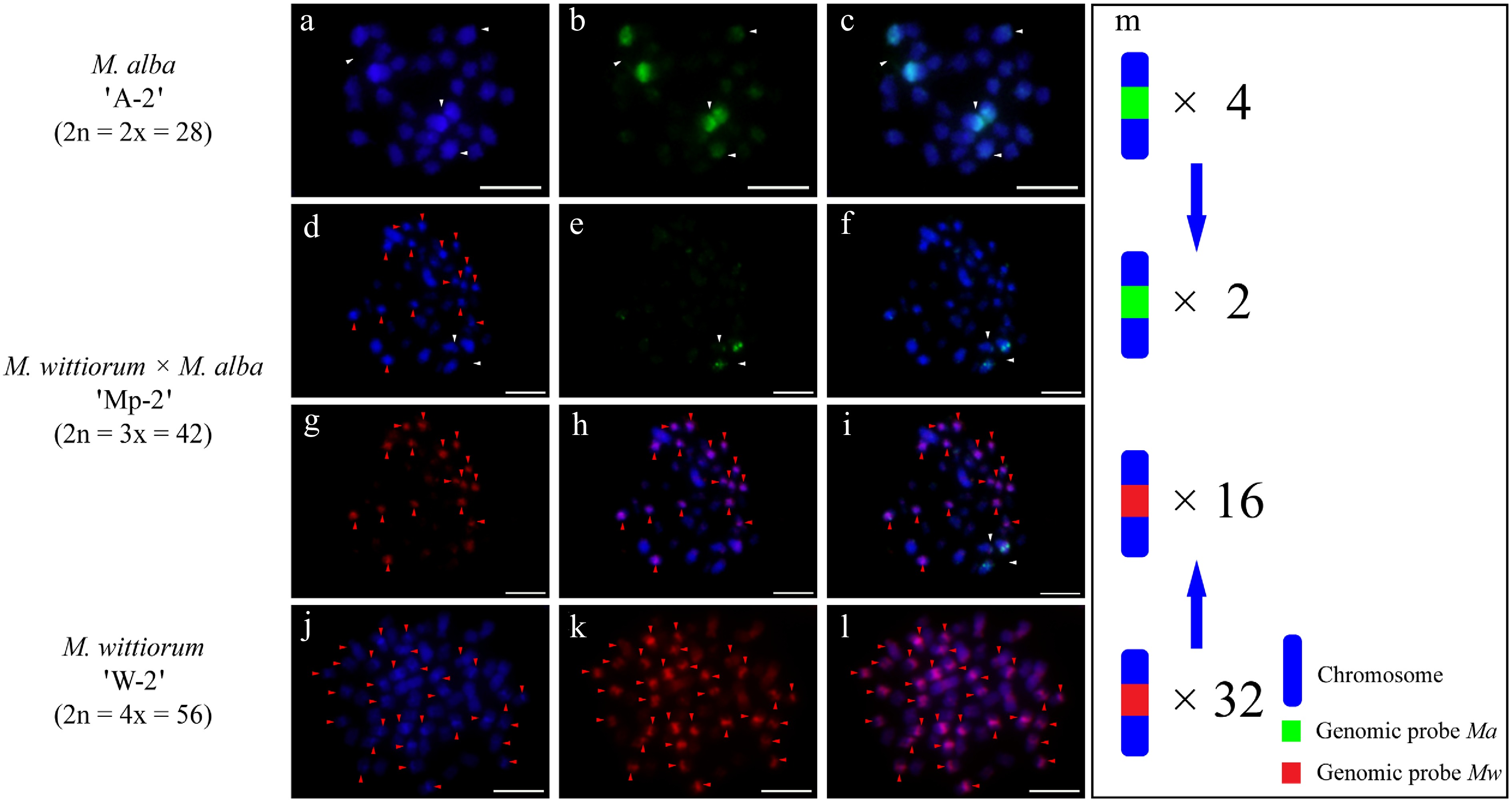
Figure 2.
cGISH signal patterns in M. alba 'A-2', M. wittiorum 'W-2', and their hybrid M. wittiorum × M. alba 'Mp-2'. cGISH signals using the genomic probe from Ma (green) with blocking DNA from Mw were detected in (a)−(c) M. alba 'A-2' and (d)−(f) the hybrid 'Mp-2'. cGISH signals using the genomic probe from Mw (red) with blocking DNA from Ma were detected in (g), (h) the hybrid 'Mp-2' and (j)−(l) M. wittiorum 'W-2'. (i) The two-round cGISH showing overlapping signals from Ma (green) and Mw (red) genomic probes with reciprocal blocking DNA. (m) Ideogram summarizing the chromosome counts with different genomic probes' signals. White and red arrows point to the chromosomes with Ma and Mw signals, respectively. Blue arrows indicate the transmission of parental chromosome to the hybrid. Scale bars represent 5 μm.
GISH analysis of intersectional hybrids between Morus section Alba and section Laevigata
-
Consistent with the cross combinations of section Alba and section Wittiorum, genomic probes targeting Ma and Ml exhibited extensive signal overlap (Supplementary Fig. S3). This ambiguity required the application of cGISH with species-specific blocking DNA to resolve the genomic origins.
cGISH analysis of the M. alba 'A-3' × M. laevigata 'L-1' cross revealed distinct parental genomic distributions. When the Ma probe with Ml blocking DNA was applied, five hybridization signals were detected in the female parent (Fig. 3a−c). The Ml probe with Ma blocking DNA identified eight signals in the male parent (Fig. 3j−l). Their hybrid progeny, 'Mp-3', exhibited differential genomic incorporation, with five signals derived from each parent (Fig. 3d−i). A similar pattern was observed in the M. wittiorum 'L-2' × M. alba 'A-4' cross. The Ml probe with Ma blocking revealed 10 signals in the female parent (Fig. 4j−l), whereas the Ma probe with Ml blocking detected five signals in the male parent (Fig. 4a−c). In the resulting hybrid, 'Mp-4', four Ma-specific and nine Ml-specific signals were observed (Fig. 4d−i).
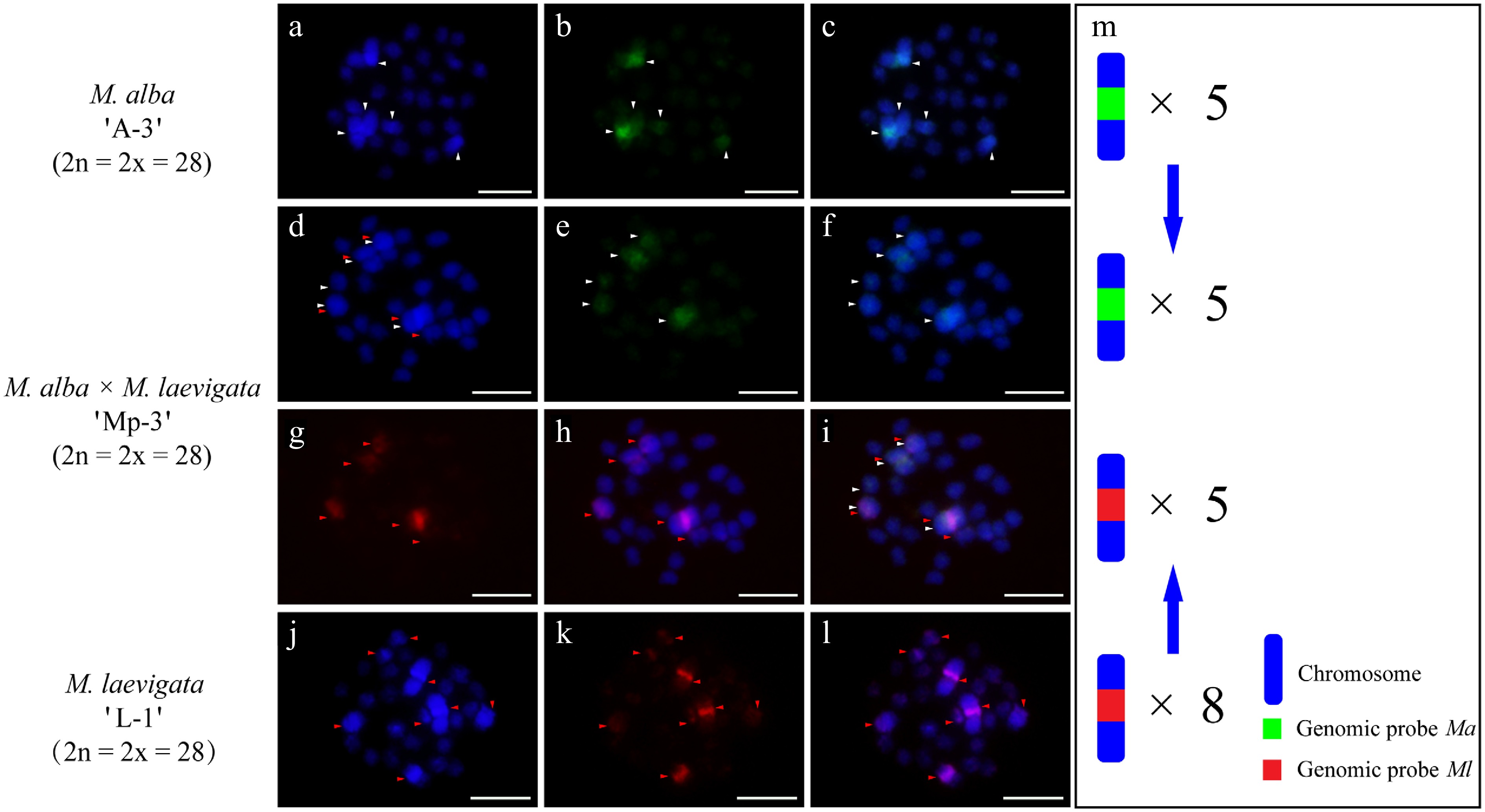
Figure 3.
cGISH signal patterns in M. alba 'A-3', M. laevigata 'L-1', and their hybrid M. alba × M. laevigata 'Mp-3'. cGISH signals using the genomic probe from Ma (green) with blocking DNA from M. laevigata (Ml) were detected in (a)−(c) M. alba 'A-3' and (d)−(f) the hybrid 'Mp-3'. cGISH signals using the genomic probe from Ml (red) with blocking DNA from Ma were detected in (g), (h) the hybrid 'Mp-3' and (j)−(l) M. laevigata 'L-1'. (i) The two-round cGISH showing overlapping signals from the Ma (green) and Ml (red) genomic probes with reciprocal blocking DNA. (m) Ideogram summarizing chromosome counts with different genomic probes' signals. White and red arrows point to the chromosomes with Ma and Ml signals, respectively. Blue arrows indicate the transmission of parental chromosome to the hybrid. Scale bars represent 5 μm.
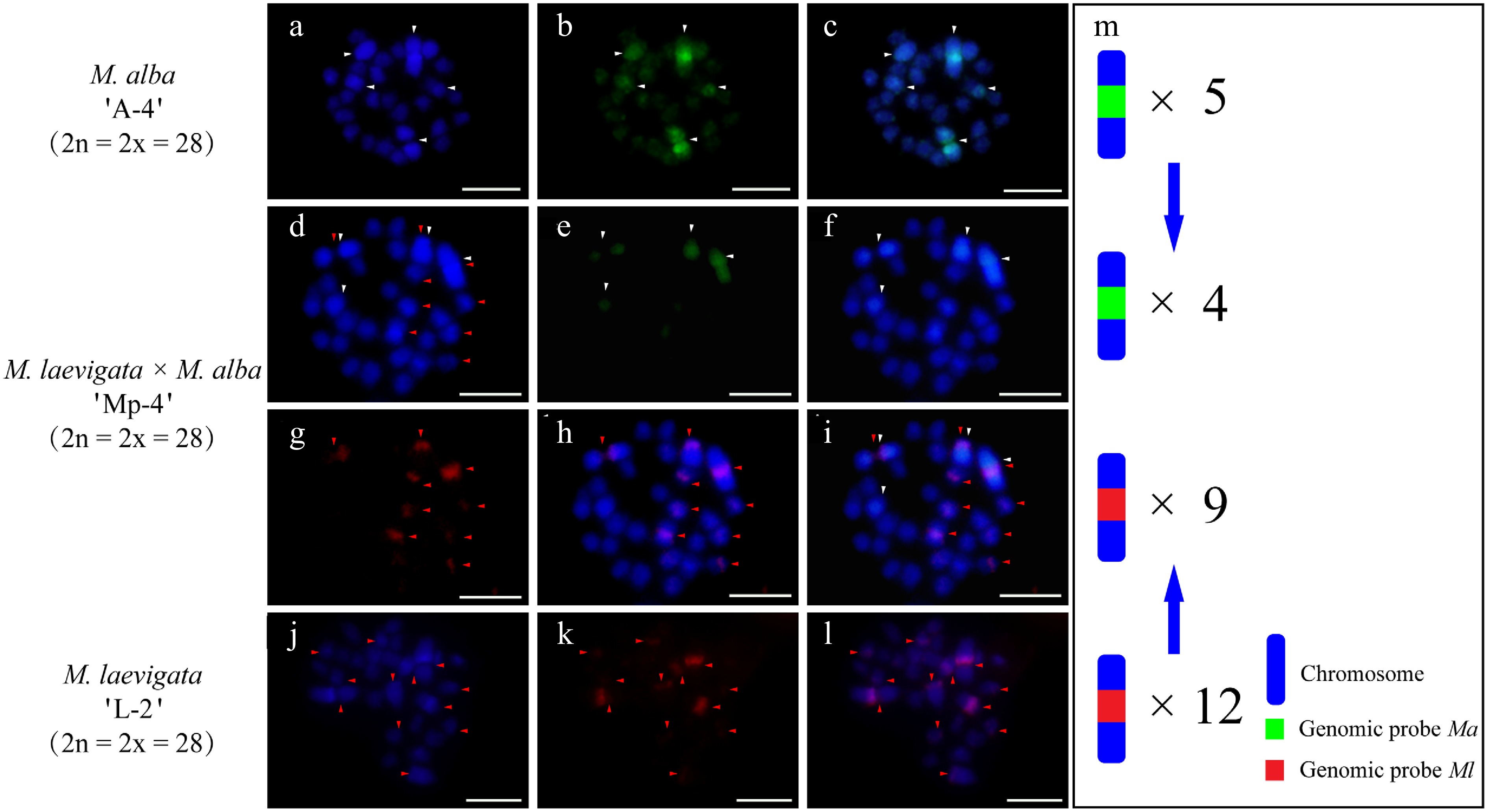
Figure 4.
cGISH signal patterns in M. alba 'A-4', M. laevigata 'L-2', and their hybrid M. laevigata × M. alba 'Mp-4'. cGISH signals using the genomic probe from Ma (green) with blocking DNA from M. laevigata (Ml) were detected in (a)−(c) M. alba 'A-4' and (d)−(f) the hybrid 'Mp-4'. cGISH signals using the genomic probe from Ml (red) with blocking DNA from Ma were detected in (g), (h) the hybrid 'Mp-4' and (j)−(l) M. laevigata 'L-2'. (i) The two-round cGISH showing overlapping signals from the Ma (green) and Ml (red) genomic probes with reciprocal blocking DNA. (m) Ideogram summarizing chromosome counts with different genomic probes' signals. White and red arrows point to chromosomes with the Ma and Ml signals, respectively. Blue arrows indicate the transmission of parental chromosome to the hybrid. Scale bars represent 5 μm.
GISH analysis of intersectional hybrids between Morus section Laevigata and section Wittiorum
-
Intersectional hybridization between Morus section Laevigata and section Wittiorum yielded two hybrid combinations: (1) M. laevigata 'L-2' × M. wittiorum 'W-3' and (2) M. wittiorum 'W-4' × M. laevigata 'L-1'. Genomic probes derived from Ml and Mw were used reciprocally as either hybridization probes or blocking DNA to analyze chromosomal compositions. Consistent with the abovementioned findings, the Ml and Mw genomic probes showed extensive signal overlap (Supplementary Fig. S4), necessitating cGISH with blocking DNA.
In the M. laevigata 'L-2' × M. wittiorum 'W-3' cross, the Ml probe with Mw blocking DNA identified 12 hybridization signals in the female parent (Fig. 5a−c). The Mw probe with Ml blocking DNA detected 29 signals in the male parent (Fig. 5j−l). Their hybrid progeny, 'Mp-5', exhibited 15 Ml-derived and 15 Mw-derived signals (Fig. 5d−i). In the case of M. wittiorum 'W-4' × M. laevigata 'L-1' exhibited parallel findings, as the Mw probe with Ml blocking revealed 28 signals in the female parent (Fig. 6j−l). The Ml probe with Mw blocking detected eight signals in the male parent (Fig. 6a−c). The hybrid 'Mp-6' demonstrated 16 Ml-specific and 14 Mw-specific signals (Fig. 6d−i).
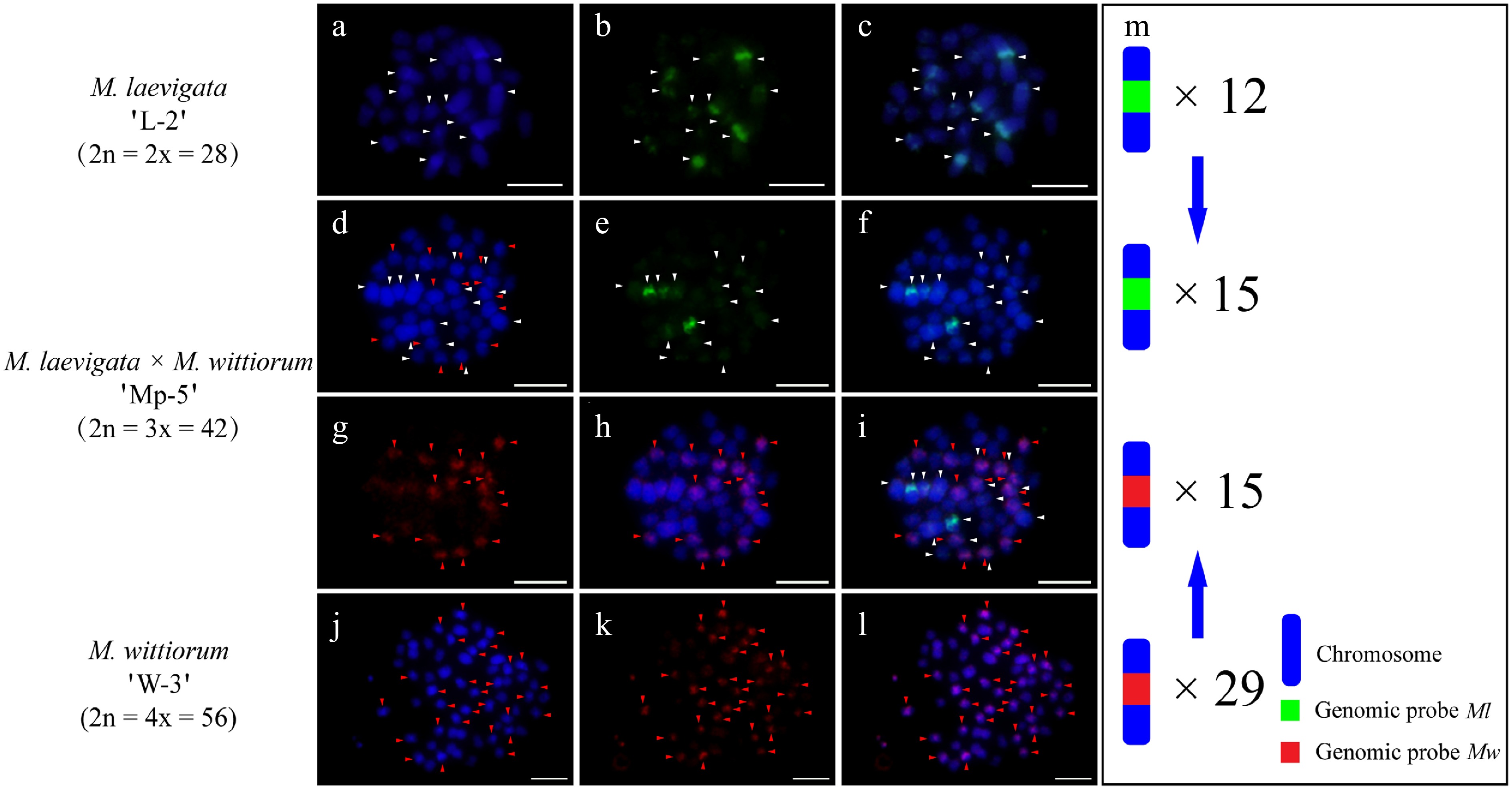
Figure 5.
cGISH signal patterns in M. laevigata 'L-2', M. wittiorum 'W-3', and their hybrid M. laevigata × M. wittiorum 'Mp-5'. cGISH signals using the genomic probe from Ml (green) with blocking DNA from Mw were detected in (a)−(c) M. laevigata 'L-2' and (d)−(f) the hybrid 'Mp-5. cGISH signals using the genomic probe from Mw (red) with blocking DNA from Ml were detected in (g), (h) the hybrid 'Mp-5', and (j)−(l) M. wittiorum 'W-3'. (i) The two-round cGISH showing overlapping signals from the Ml (green) and Mw (red) genomic probes with reciprocal blocking DNA. (m) Ideogram summarizing chromosome counts with different genomic probes' signals. White and red arrows point to chromosomes with the Ml and Mw signals, respectively. Blue arrows indicate the transmission of parental chromosomes to the hybrid. Scale bars represent 5 μm.
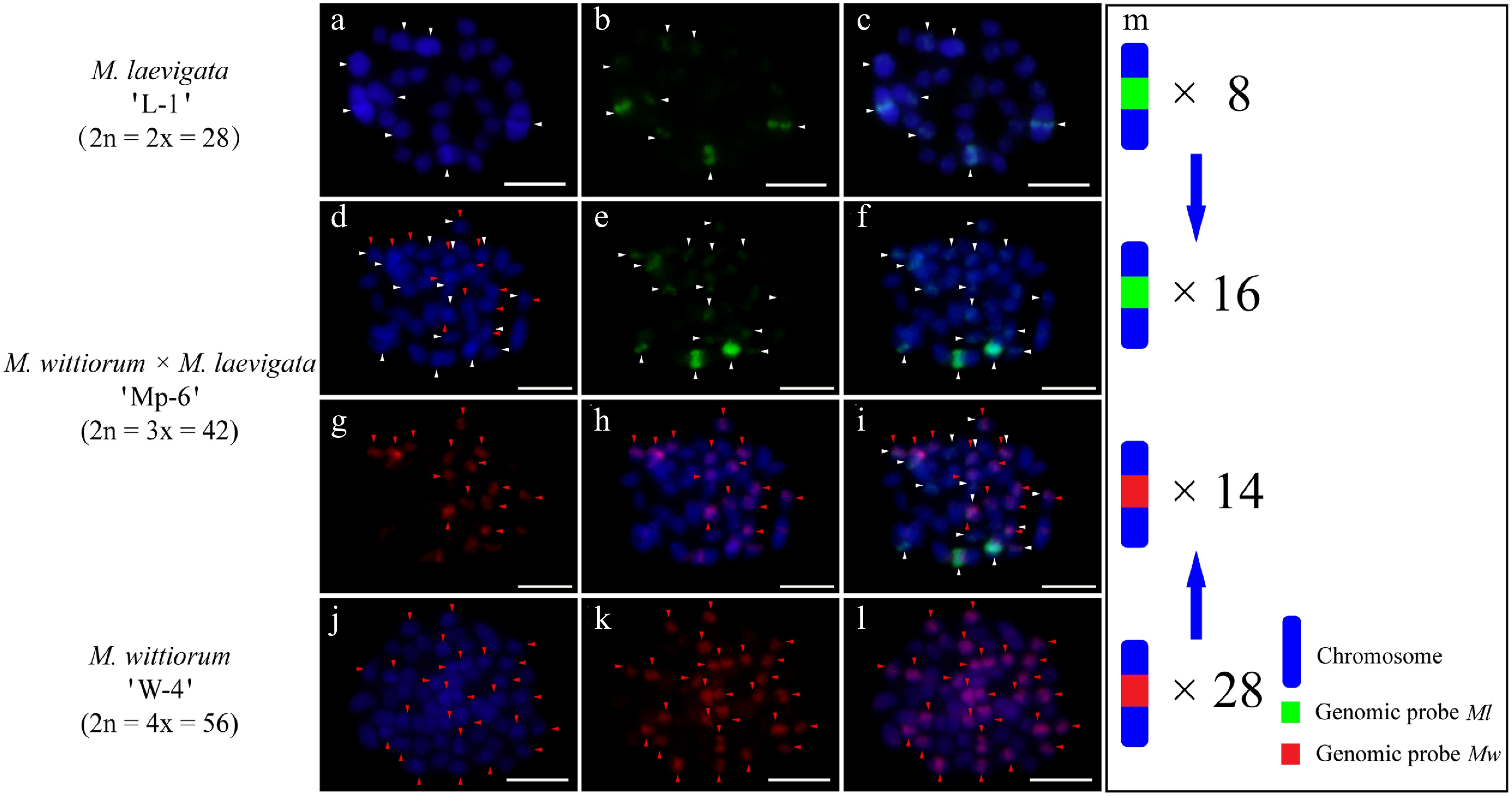
Figure 6.
cGISH signal patterns in M. laevigata 'L-1', M. wittiorum 'W-4', and their hybrid M. wittiorum × M. laevigata 'Mp-6'. cGISH signals using the genomic probe from Ml (green) with blocking DNA from Mw were detected in (a)−(c) M. laevigata 'L-1' and (d)−(f) the hybrid 'Mp-6'. cGISH signals using the genomic probe from Mw (red) with blocking DNA from Ml were detected in (g), (h) the hybrid 'Mp-6', and (j)−(l) M. wittiorum 'W-4'. (i) The two-round cGISH showing overlapping signals from the Ml (green) and Mw (red) genomic probes with reciprocal blocking DNA. (m) Ideogram summarizing chromosome counts with different genomic probes' signals. White and red arrows point to the chromosomes with Ml and Mw signals, respectively. Blue arrows indicate the transmission of parental chromosomes to the hybrid. Scale bars represent 5 μm.
A pentaploid hybrid, designated 'Mp-7', was identified in the cross of M. wittiorum 'W-4' × M. laevigata 'L-1'. The genomic probes of Ml and Mw (no blocking DNA) hybridized to nearly all chromosomes (Fig. 7a−d). The Ml probe with Mw blocking DNA detected 9 clear signals, whereas the Mw probe with Ml blocker identified 30 signals in 'Mp-7' (Fig. 7e−h).
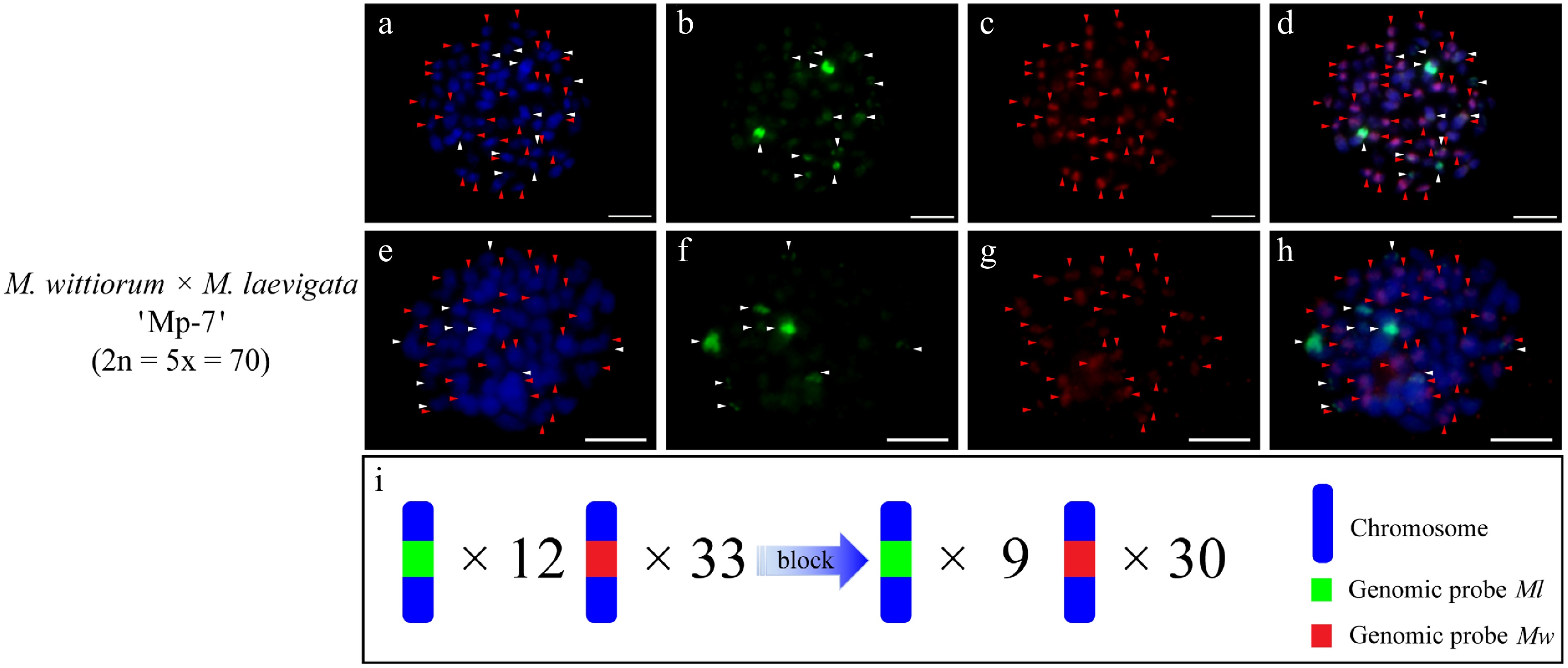
Figure 7.
cGISH signal patterns in the hybrid 'Mp-7' from the cross M. wittiorum 'W-4' × M. laevigata 'L-1'. (a)−(d) Dual-color GISH signals using the genomic probes from Ml (green) and Mw (red) were detected in the hybrid 'Mp-7'. (e)−(h) The two-round cGISH signals using the genomic probes from Ml (green) and Mw (red) with reciprocal blocking DNA were detected in the hybrid 'Mp-7'. (i) Ideogram summarizing chromosome counts with different genomic probes' signals. White and red arrows point to chromosomes with the Ml and Mw signals, respectively. The blue gradient arrow indicates the changes in signal number after the application of blocking DNA. Scale bars represent 5 μm.
GISH analysis of putative wild hybrids
-
On the basis of prior work demonstrating the utility of the Ma and Mw genomic probes for rapid mulberry genotyping[20], we used these two probes to analyze the wild accessions M. laevigata 'Wg-1' and M. australis 'Wg-2' (Supplementary Fig. S5). The Ma probe hybridized to five chromosomal loci, whereas the Mw probe detected four clear signals in M. laevigata 'Wg-1' (Supplementary Fig. S5a1−a4). In M. australis 'Wg-2', the Ma probe marked chromosome 1 and chromosome 2, whereas Mw detected 31 signals (Supplementary Fig. S5b1−b4). According to the established cytotaxonomic criteria for mulberry[20], this pattern suggested that 'Wg-L1' is a hybrid between sections Laevigata and Alba, and 'Wg-2' is a hybrid between sections Alba and Wittiorum. cGISH validation with blocking DNA was performed. The Ma probe with Ml blocking DNA produced 8 signals in M. laevigata 'Wg-1', whereas the Ml probe with Ma blocking DNA yielded 10 signals (Fig. 8a−d). In M. australis 'Wg-2', the Ma probe with Mw blocking detected 3 signals, whereas the Mw probe with Ma blocker revealed 29 signals (Fig. 8e, f).
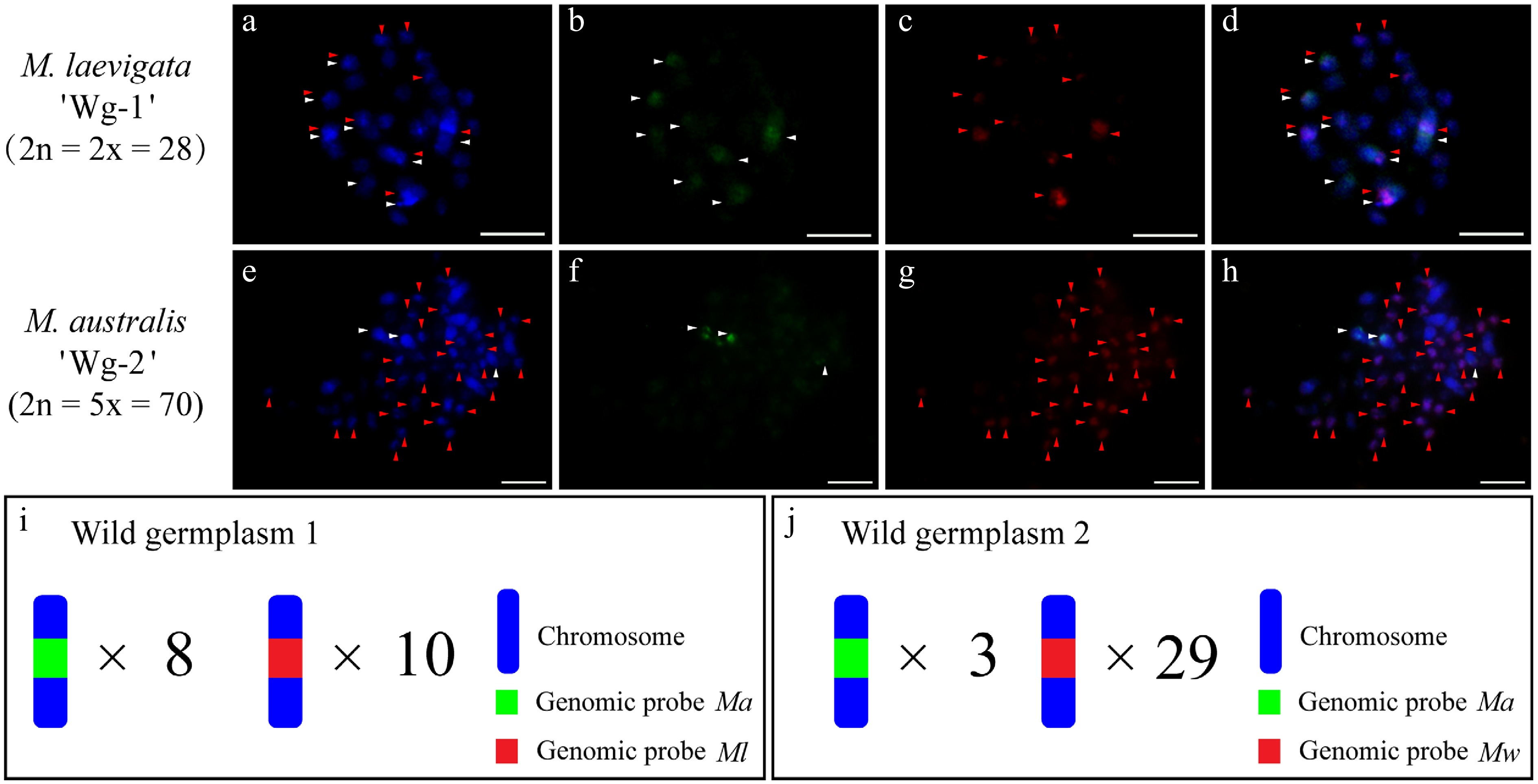
Figure 8.
cGISH signal patterns in the wild accessions M. laevigata 'Wg-1' and M. australis 'Wg-2'. (a)−(d) The two-round cGISH signals using genomic probes from Ma (green) and Ml (red) with reciprocal blocking DNA were detected in M. laevigata 'Wg-1'. (e)−(h) Corresponding two-round cGISH signals using genomic probes from Ma (green) and Mw (red) with reciprocal blocking DNA were detected in M. australis 'Wg-2'. (i), (j) Ideograms summarizing chromosome counts with different genomic probes' signals. White arrows point to chromosomes with Ma signals, whereas red arrows point to chromosomes with (a)−(d) Ml signals or (e)−(h) Mw signals. Scale bars represent 5 μm.
-
Cytology-based early hybrid authentication has been widely adopted as an efficient and reliable method for identifying hybrids in various plant species, including Lilium and Capsicum[45−47]. However, GISH faces challenges in Morus species because of the highly similar chromosome size and the cross-hybridization of genomic probes, which generate nonspecific signals across different mulberry sections. This limitation has been successfully overcome in other crops, such as rice (Oryza sativa) and kiwifruit (Actinidia chinensis), through the incorporation of blocking DNA, which suppresses repetitive DNA interference[48,49]. In this study, cGISH with blocking DNA enabled precise chromosomal delineation in hybrids derived from six cross combinations. Hybrid status was confirmed through a comparative analysis of quantitative signals, where the progeny exhibited approximately half the number of cGISH signals relative to their parents. For instance, when probed with Ma (blocked with Mw DNA), the hybrid 'Mp-1' displayed two distinct signals, contrasting with the four signals observed in its female parent M. alba 'A-1'.
Early hybrid authentication at the seedling stage is crucial for minimizing resource expenditure on false hybrids[50]. Given the prolonged juvenile phase in perennial species like Morus, rapid detection significantly accelerates breeding programs[51,52]. Our results demonstrate that GISH is a robust tool for identifying intersectional mulberry hybrids, facilitating efficient mulberry breeding programs. Moreover, the successful development and precise identification of intersectional hybrids provide a clear direction for advancing distant hybridization strategies in mulberry.
Cytogenetic characterization of wild mulberry accessions using GISH with blocking DNA
-
As a wind-pollinated species with weak interspecific reproductive barriers, mulberry has generated numerous wild interspecific hybrids[13,14,19,21,22]. However, the parental origins and ploidy variability of these wild accessions remain poorly resolved[20], significantly impeding their utilization in breeding. Although cGISH previously predicted 15 accessions as hybrids, their precise chromosomal composition was unresolved[20]. In this study, cGISH with blocking DNA outperformed dual-color GISH in clarifying the genomic constitution of wild mulberries, yielding unambiguous chromosomal compositions. Specifically, it was confirmed that M. laevigata 'Wg-1' (derived from sections Laevigata and Alba) and M. australis 'Wg-2' (from sections Alba and Wittiorum) are intersectional hybrids. M. laevigata 'Wg-1' exhibited a balanced Laevigata–Alba inheritance, whereas M. australis 'Wg-2' showed asymmetry, with Wittiorum being dominant.
Although dual-color GISH enables rapid hybrid screening, cGISH with blocking DNA facilitates precise chromosomal identification. Their complementary use provides a robust framework for dissecting the genetic complexity of wild mulberries. Furthermore, these validated wild hybrids represent valuable germplasm for trait introgression (e.g., via backcrossing), offering a faster alternative to conventional breeding by bypassing lengthy generation cycles.
The genetic relationship of three mulberry sections revealed by GISH
-
Previous studies using cGISH with blocking DNA have identified distinct chromosomal signal patterns among Morus sections Alba, Wittiorum, and Laevigata[20]. In this study, we applied this approach to 10 mulberry accessions and seven intersectional hybrids derived from these sections, providing multidimensional insights into their genetic relationships and taxonomic boundaries. Variations in the probes' specificity and signal patterns were detected. For instance, the Ml probe produced differences in both signal numbers and intensities in M. laevigata 'L-2' under different blocking DNA conditions. Despite the presence of blocking DNA, the Ml probe consistently generated signals on chromosomes corresponding to its parental origin. This effect was further demonstrated in the hybrids 'Mp-3' and 'Mp-4', where overlapping two-round GISH signals were detected. Consequently, progeny derived from laevigata crosses exhibited a pronounced abundance of GISH signals, suggesting that laevigata genomes harbor diverse repeat sequences and share closer genetic affinity with section Alba than previously recognized. These results align with earlier observations in hybridization studies of Erianthus rockii × N. porphyrocoma[53].
Furthermore, clear intersectional genetic divergence was observed. In the hybrids 'Mp-1' to 'Mp-6', differential signal intensities generated by different genomic probes confirmed substantial genomic differentiation among the three sections. Previously sequenced genomic data indicated a significant difference in the divergence time between M. wittiorum and M. alba[54]. Phylogenetic analyses based on plastomes and single-copy nuclear genes also demonstrated that the two species occupy distinct phylogenetic positions[19]. These results provide improved phylogenetic resolution for the three mulberry sections, refining their taxonomic boundaries and offering a framework for future studies on the dynamics of intersectional hybridization.
Transmission patterns of parental chromosomes in mulberry hybrids
-
Although mulberry has traditionally been characterized by n + n inheritance, reports of abnormal meiotic behavior[55,56] suggested potential deviations from this pattern. However, prior to this study, 2n gametes had never been documented in Morus. The standard n + n chromosomal inheritance pattern was consistently observed in the identified hybrids 'Mp-1' to 'Mp-6'. Here, we provide the first evidence of teh formation of 2n gametes in the female parent M. wittiorum 'W-4' (2n = 4x = 56), which generated the pentaploid hybrid 'Mp-7' (2n = 5x = 70). As is well known, there are three primary mechanisms underlie the formation of 2n gametes: the omission of one meiotic division, alterations in spindle morphology during meiotic division (e.g., parallel spindles), or defects in the cytokinesis of meiosis[57−59]. According to the genetic composition, 2n gametes are classified into two types: first division restitution (FDR) and second division restitution (SDR). FDR 2n gametes contain complete homologous chromosome pairs, comprising unreduced sets derived from both parents[60]. Consequently, SDR 2n gametes consist of sister chromatid pairs derived from a single parent, resulting in significantly lower heterozygosity compared with FDR gametes[61]. In this study, direct cytological evidence of abnormal female meiotic behavior in the maternal parent M. wittiorum 'W-4' was unobtainable because of the production of a 2n egg. Therefore, the specific mechanism responsible for the formation of 2n gametes in 'W-4' could not be determined. Further investigations into this mechanism will be conducted using cytogenetic analysis with other male mulberry accessions.
Moreover, a chromosome constitution similar to that of the pentaploid hybrid 'Mp-7' was detected in M. australis 'Wg-2', suggesting it may also be a pentaploid accession derived from the formation of 2n gametes. The findings indicate that ploidy variation in the genus Morus may be driven by the occurrence of 2n gametes, a phenomenon that is more prevalent in wild accessions than in cultivated accessions, which typically exhibit lower ploidy levels. Beyond their role in natural polyploidization, 2n gametes could have significant implications for breeding. They may stabilize the chromosomal composition in progeny, mitigate dosage effects, and overcome the reproductive barriers associated with odd-ploidy levels, thereby preserving breeding potential. This parallels their well-documented roles in polyploid breeding systems in species such as Lilium, Eucommia, and Populus[62−66]. Further research should focus on protocols for inducing 2n gametes and elucidating their underlying mechanisms in Morus. Such studies would advance our understanding of natural sexual polyploidization and support the development of targeted polyploid breeding strategies.
-
In this study, cGISH with blocking DNA proved to be highly effective for early hybrid identification and precise analysis of the chromosomal composition across wild mulberry accessions. We present the first documented evidence of 2n gametes in Morus, showing that a maternal 2n egg from M. wittiorum 'W-4' (2n = 4x = 56) produced the pentaploid hybrid 'Mp-7' (2n = 5x = 70). These results establish GISH as a robust cytogenetic tool for authentication of species and hybrids in mulberry, while highlighting the critical role of 2n gametes in polyploidy-based breeding strategies. This work not only advances our understanding of natural polyploidization mechanisms in mulberry but also introduces innovative approaches for accelerating the genetic improvement of this agronomically significant species. Future studies should optimize GISH-based screening to improve breeding efficiency and explore the molecular pathways underlying the formation of 2n gametes.
-
All experimental methods, including the collection of plant materials, complied with the relevant institutional, national, and international guidelines and legislation. The voucher specimens of all plants are deposited at the Mulberry Germplasm Nursery at Southwest University, Chongqing, China.
-
The authors confirm their contributions to the paper as follows: conceptualization: He N, data curation, methodology: Zhou Q, Li J; visualization: Zhou Q, Qiu J; resources: Yuan J, Li Z, Chen S; funding acquisition: Xuan Y, He N; writing – original draft: Zhou Q; writing – review & editing: Zhou Q, Li J, Xuan Y, He N. All authors reviewed the results and approved the final version of the manuscript.
-
The plant materials reported in the manuscript are included in Supplementary Tables S1 and S2 and are freely available to all the readers upon reasonable request.
-
This work was supported by the National Key Research and Development Program of China (Grant No. 2024YFD2201302), the Science and Technology Pilot Project—Key Research and Development Project (Grant No. SWU-XDZD22008), the Fundamental Research Funds for the Central Universities (SWU-KQ24048), Chongqing Mulberry Germplasm Repository at Beibei (Grant No. ZWZZ20240002), and Zhonghuai Xiang Academician Workstation Project, Yunnan Province, China (Grant No. 202305AF150166). We extend our gratitude to Ziang Li from Nanjing Agricultural University for his assistance in counting the fluorescent signals on the chromosomes and to Jianglian Yuan for her assistance in operating the fluorescence microscope (Olympus IX73). The authors acknowledge the support from the Naban River Watershed National Nature Reserve Administration, Zhenyuan Management and Protection Bureau of Ailao Mountain National Nature Reserve Administration in Yunnan Province, and Baoshan Agricultural Science Research Institute (Baoshan Agricultural Technology Extension Center) for their assistance with and permission for sample collection.
-
The authors declare that they have no conflict of interest.
-
# Authors contributed equally: Qiming Zhou, Jiacheng Li, Yahui Xuan
- Supplementary Table S1 List of the mulberry accessions used in this study.
- Supplementary Table S2 List of seven mulberry accessions from six hybrid combinations.
- Supplementary Fig. S1 Fruit morphological characteristics of M. laevigata and M. wittiorum.
- Supplementary Fig. S2 cGISH signal patterns in M. alba 'A-1', M. wittiorum 'W-1', their hybrid M. alba × M. wittiorum 'Mp-1', M. wittiorum 'W-2', M. alba 'A-2', and their hybrid M. wittiorum × M. alba 'Mp-2'.
- Supplementary Fig. S3 cGISH signal patterns in M. alba 'A-3', M. laevigata 'L-1', their hybrid M. alba × M. laevigata 'Mp-3', M. laevigata 'L-2', M. alba 'A-4', and their hybrid M. laevigata × M. alba 'Mp-4'.
- Supplementary Fig. S4 cGISH signal patterns in M. laevigata 'L-2', M. wittiorum 'W-3', their hybrid M. laevigata × M. wittiorum 'Mp-5', M. wittiorum 'W-4', M. laevigata 'L-1', and their hybrid M. wittiorum × M. laevigata 'Mp-6'.
- Supplementary Fig. S5 cGISH signal patterns in M. laevigata 'Wg-1' and M. australis 'Wg-2'.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhou Q, Li J, Xuan Y, Qiu J, Yuan J, et al. 2026. Precise identification of intersectional hybrids in Morus using genomic in situ hybridization (GISH). Forestry Research 6: e010 doi: 10.48130/forres-0026-0009
Precise identification of intersectional hybrids in Morus using genomic in situ hybridization (GISH)
- Received: 23 October 2025
- Revised: 28 January 2026
- Accepted: 11 February 2026
- Published online: 03 April 2026
Abstract: Mulberry (Morus spp.) includes ecologically important tree species that are highly valued for their exceptional economic and medicinal properties. Among its diverse species, Morus wittiorum and Morus laevigata are particularly valuable genetic resources because of their resistance to Sclerotinia, their relatively high content of specific flavonoids, and elongated fruit morphology. In this study, hybridization experiments were conducted using 10 mulberry accessions spanning three taxonomic sections (Alba, Wittiorum, and Laevigata). All six attempted intersectional crosses successfully yielded hybrid progeny. Using genomic in situ hybridization with blocking DNA, we detected distinct chromosomal signal patterns among the three sections, enabling precise identification of hybrid and wild-type chromosomal constitutions. Notably, this study provides the first documented evidence of 2n gamete formation in the genus Morus, where 2n eggs from M. wittiorum 'W-4' produced a pentaploid hybrid, 'Mp-7'. This discovery not only rewrites the chromosomal inheritance patterns of Morus but also unveils untapped polyploid breeding potential. These findings provide an efficient approach for identifying hybrids and offer novel polyploid breeding strategies, thereby promising to reshape global mulberry breeding and creating new opportunities for the genetic improvement of this agronomically important species.
-
Key words:
- Mulberry /
- Intersectional hybrids /
- Hybrid identification /
- GISH /
- 2n gametes














