-
Haploid refers to an individual whose somatic cells contain only one set of chromosomes. Since the discovery of natural haploids in Datura innoxia by Blakeslee et al. in 1922, researchers have been exploring their potential applications in plant breeding[1]. In 1964, Guha & Maheshwari observed the differentiation of embryoid-like structures within the anther chambers of Datura innoxia through anther culture. In 1966, they confirmed that these embryoids originated from pollen, giving rise to young haploid plants. This discovery established the first experimental system for obtaining haploid plants through in vitro culture, significantly advancing the development of haploid induction technology[2,3].
The core advantage of haploid-related technology lies in its efficiency in achieving genetic homozygosity. Doubled haploid (DH) plants obtained through chromosome doubling possess a completely purified genetic background, resulting in uniform and stable traits in the offspring without segregation. Compared to traditional breeding methods, which require at least six to eight generations of continuous selfing to achieve homozygosity, this technology can shorten the breeding cycle by 50% to 75%[4] (Fig. 1). It serves as a versatile tool, applicable both to specialized tasks like the breeding of self-incompatible plants and to broader breeding strategies, including rapid propagation, mutation, and genetic transformation.
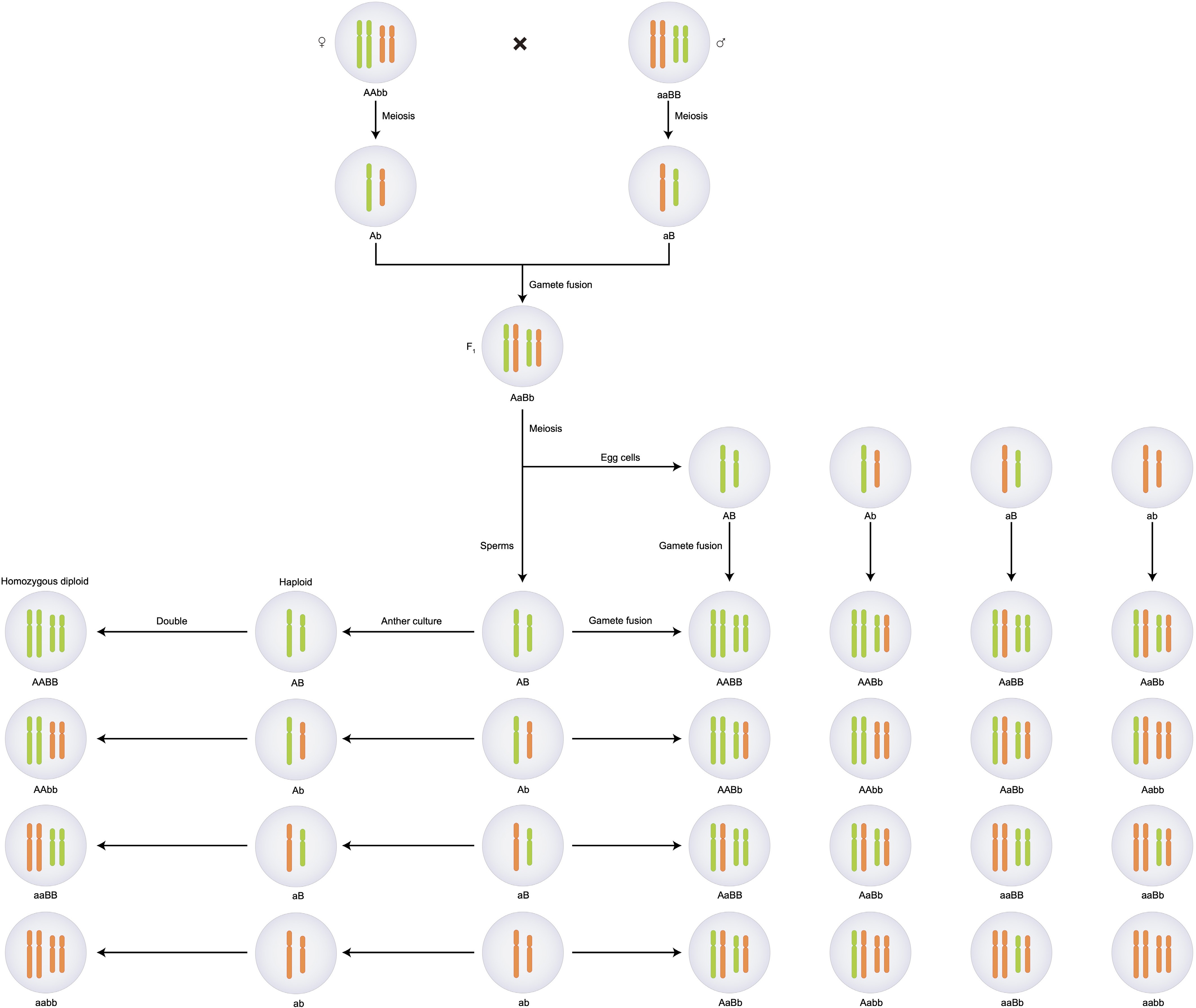
Figure 1.
Genetic schematic diagram of conventional hybrid breeding and haploid breeding methods.
Diploid parents produce gametes through meiosis, and these gametes combine to form the F1 generation. Conventional hybrid breeding: Self-pollination of the F1 generation results in 16 genotype combinations, with a homozygous probability of 1/4; Haploid breeding: Anther culture (in vitro) and chromosome doubling of F1 gametes directly yield homozygous doubled haploid (DH) plants, avoiding the complex screening process of multi-generation self-pollination.Based on their mechanisms, haploid induction technology can be categorized as pollen-related, egg cell-related, or centromere-related methods. The pollen-related induction technology includes three main approaches: (1) In vitro culture of microspores or anthers; (2) Pollen treatment-based embryo rescue; (3) Based on pollen development-associated functional genes. The egg cell-related induction technology also includes three main approaches: (1) In vitro culture of megagametophytes or ovules; (2) Based on egg cell development-associated functional genes; (3) Parthenogenesis. The centromere-related haploid induction technology refers specifically to genetic regulation methods that produce haploids by interfering with the expression of centromere-related functional genes, utilizing mechanisms of abnormal chromosome segregation.
-
Microspores are produced from microsporocytes through meiosis, resulting in a halved chromosome number and carrying the paternal haploid genome. In in vitro culture systems, hormonal regulation can reverse the gametophytic developmental pathway and initiate the sporophytic developmental pathway[5]. Through microspore embryogenesis or organogenesis, these microspores regenerate into haploid plants. Anther culture is currently one of the most widely applied methods for haploid induction. The regenerated population obtained through anther culture may include not only haploid plants derived from pollen but also diploid plants originating from somatic cells such as the anther wall or connective tissue. Therefore, the resulting haploid progeny must be screened and identified to confirm their origin[6].
Pollen treatment-based embryo rescue
-
As the final product of microspore development, pollen can be utilized not only for direct in vitro culture to regenerate haploid plants, but also as a target for physical or chemical treatments—such as γ-irradiation, temperature stress, or EMS mutagenesis—to modulate its viability. Combined with embryo rescue technology, these approaches enable efficient haploid production. This strategy has been successfully applied in multiple crops, such as maize, wheat, and tobacco[7−9]. In 1987, Sauton & Vaulx pioneered the application of this technology to melon, successfully generating haploid plants[10]. Since then, the system has been optimized and systematically extended to other Cucurbitaceae species, including cucumber and zucchini, where it has demonstrated high efficiency in haploid induction[11,12]. At present, this method has become a reliable and stable pathway for obtaining haploids in cucurbit crops, particularly in melon[13].
Based on pollen development-associated functional genes
-
Maize (Zea mays L.), as a typical diploid species (2n = 20), exhibits a natural haploid occurrence rate of only ~0.1%[14]. In 1959, Edward H. Coe discovered that when the maize germplasm Stock 6 was used as a paternal parent in hybridization, the haploid induction rate (HIR) significantly increased to 2%–3%, thus establishing the first maize haploid induction line[14]. Derived lines based on Stock 6, such as RWS, MHI, and PHI, have further improved HIR to 6%–15%[15].
The genetic basis of haploid induction in Stock 6 has gradually been elucidated through QTL mapping. The major QTL qhir1, located on chromosome 1 bin 1.04, accounts for ~66% of the phenotypic variation. Its candidate region was narrowed down to 243 kb through fine mapping by Prigge & Dong[16,17]. A minor QTL, qhir8, located on chromosome 9 bin 9.01, explains ~20% of the variation, with a candidate region spanning 789 kb[18]. These two key loci serve as genetic anchors for dissecting the molecular mechanisms of haploid induction.
In early 2017, three independent research teams elucidated the core genetic mechanism underlying maize haploid induction by focusing on the major QTL qhir1. Kelliher et al. first identified that a 4-bp insertion (CGAG) in the maize pollen-specific gene MATRILINEAL (MTL) causes a frameshift mutation, resulting in a truncated protein lacking 29 amino acids that severely disrupts the double fertilization process[19]. Subsequently, Liu et al. confirmed that a homologous 4-bp insertion in the ZmPLA1 gene of inducer lines similarly leads to the deletion of 29 amino acids at the C-terminus of the protein[20]. Shortly thereafter, Gilles et al. identified the NOT LIKE DAD (NLD) gene in the qhir1 region, showing that its loss-of-function mutation induced haploids by affecting pollen development[21]. Despite the different names (MTL/ZmPLA1/NLD), subsequent map-based cloning studies confirmed that all three refer to the same gene. By knocking out orthologs of MTL/ZmPLA1/NLD in monocot crops—such as rice[22], wheat[23,24], foxtail millet[25], barley[26], and sugarcane[27]—researchers have successfully established haploid induction systems in these species.
The phospholipase A protein encoded by MTL/ZmPLA1/NLD is specifically expressed in the vegetative cells of pollen grains, with its subcellular localization at the pollen endomembrane (endo-PM)[28,29]. It plays a role in reproductive development by regulating plant lipid metabolic homeostasis. Jiang et al. integrated multi-omics data and discovered that MTL/ZmPLA1/NLD regulates pollen development by participating in the 'reactive oxygen species (ROS) and stress' pathway. Their study demonstrated that exogenous application of the peroxidase inhibitor methimazole or the ROS inducer phosphatidylcholine could significantly trigger a ROS burst in pollen, leading to DNA fragmentation. Notably, such fragmentation initiates in the pericentromeric regions and finally induces haploid production[30]. This finding aligns closely with Li et al.'s earlier observation of a high frequency of chromosome fragmentation in sperm cells during haploid induction[31], suggesting that ROS-mediated genomic damage might be the core molecular basis of haploid induction. Based on this, Jiang et al. proposed a hypothesis: key genes involved in ROS metabolic regulation may possess haploid induction potential. Through systematic screening, the research team identified that loss-of-function mutants of the peroxidase gene ZmPOD65, which is specifically expressed during trinucleate pollen stages, exhibit significant haploid induction characteristics, with a maximum HIR of 7.7%[30]. This discovery not only validates the critical role of the ROS metabolic pathway in haploid induction but also provides new insights for developing novel haploid induction systems.
In addition, in a systematic screening of the pollen-specific phospholipase A gene family, Li et al. identified ZmPLD3 as the sole phospholipase gene that is specifically expressed in pollen and significantly upregulated in the mtl/zmpla1/nld mutant background[32]. Knockout of ZmPLD3 resulted in a maximum HIR of 0.96%, showing no statistically significant difference compared to the mtl/zmpla1/nld mutant. Intriguingly, analysis of the zmpld3 and mtl/zmpla1/nld interaction uncovered synergy rather than redundancy. The double mutant further elevated the haploid induction rate from 1.2% (observed in the mtl/zmpla1/nld mutant) to approximately 4%, implying that ZmPLD3 functions in a parallel signaling pathway to that of MTL/ZmPLA1/NLD[32]. In addition, the homologous gene of ZmPLD3 in rice, OsPLDα2, also exhibits the same haploid induction ability[33]. Current models propose that PLD knockout disrupts inter-gametophytic communication, culminating in male-specific developmental defects and genome elimination during double fertilization[32] (Fig. 2b).
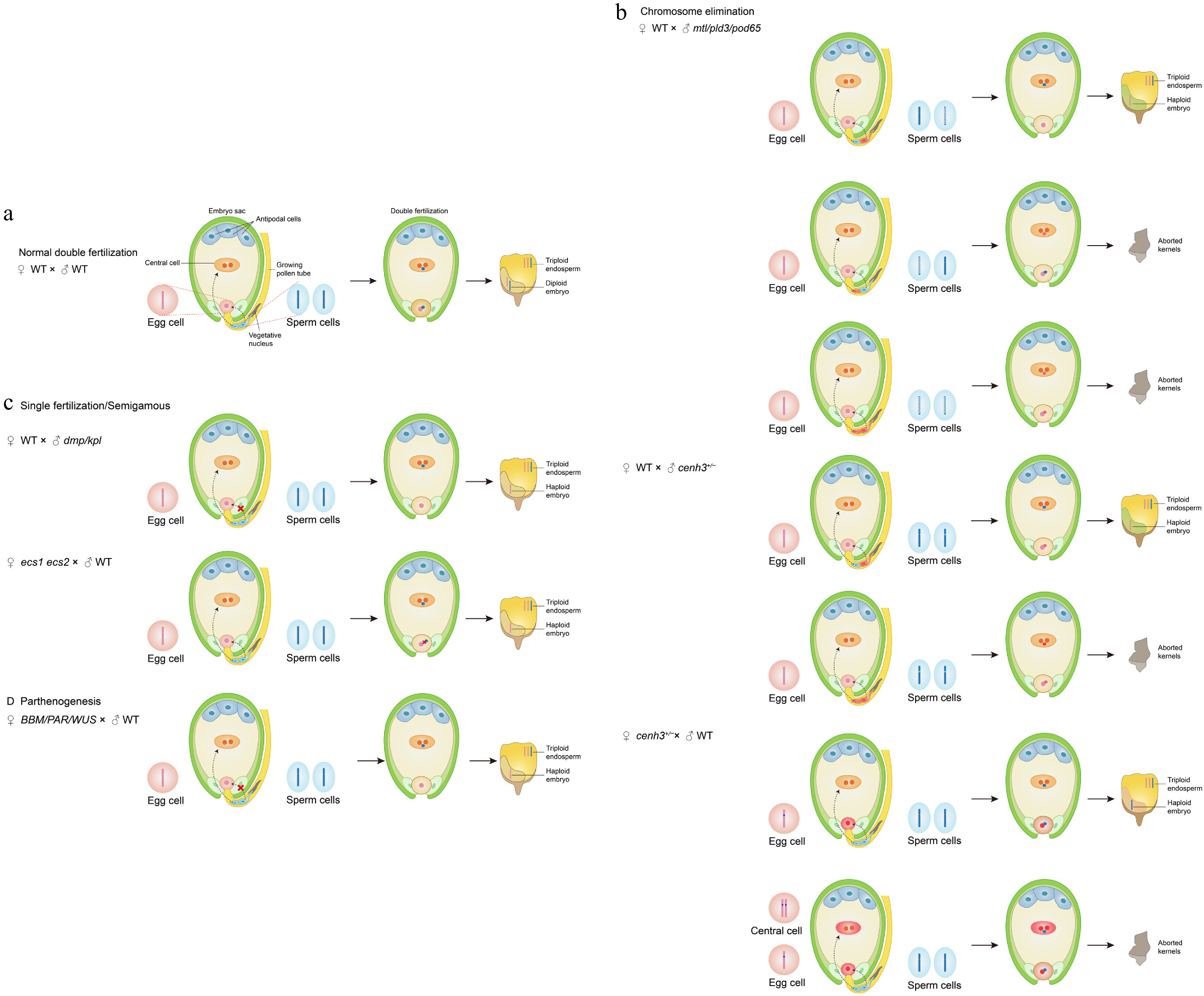
Figure 2.
Molecular mechanisms of in vivo haploid induction.
(a) Normal double fertilization. In wild-type plants, two sperm cells fuse with the egg cell and central cell to form a diploid embryo (2n) and a triploid endosperm (3n). (b) Chromosome elimination. Mutations in mtl, pld3, or pod65 induce ROS-mediated chromosomal fragmentation. In cenh3 mutants, impaired centromere function leads to the selective elimination of the paternal genome after fertilization. (c) Single fertilization and semigamy. In dmp or kpl mutants, only the central cell is fertilized, while the egg cell develops autonomously. In ecs1/ecs2 mutants, sperm-egg nuclear fusion fails, triggering haploid embryogenesis. (d) Parthenogenesis induction. Ectopic expression of BBM, PAR, or WUS in egg cells triggers fertilization-independent embryogenesis, producing haploid offspring.Zhong et al. systematically characterized the minor QTL qhir8 and identified the critical role of the DMP gene in haploid induction in non-Stock 6 maize materials for the first time. When this gene is mutated, it induces an HIR of 0.1%–0.3% in maize and 1%–4% in Arabidopsis thaliana. They discovered that ZmDMP and ZmPLA1 act synergistically; the zmdmp mtl/zmpla1/nld double mutant achieved a 5–6-fold increase in HIR[34]. Homologs of ZmDMP have also been used to create haploid induction systems in dicotyledonous species such as Arabidopsis[35], alfalfa[36], tomato[37], tobacco[38], watermelon[39,40], and soybean[41].
DMP encodes a membrane protein containing a DUF679 domain, which is subcellularly localized to the sperm plasma membrane. Double fertilization is a reproductive process unique to angiosperms, during which the egg cell secretes small peptide molecules known as EGG CELL 1(EC1), which are stored in vesicles. When sperm cells approach the egg cell, EC1 is released into the extracellular space via exocytosis. This triggers the redistribution of the fusogenic protein HAPLESS 2(HAP2) or GENERATIVE CELL SPECIFIC(GCS1) from the sperm endomembrane system to the cell surface, thereby activating their fusion capability and promoting sperm-egg binding[42]. In Arabidopsis, loss-of-function mutations in AtDMP8 and AtDMP9, homologs of the maize ZmDMP gene, result in haploid production. Studies have shown that AtDMP8 and AtDMP9 facilitate the translocation of the sperm cell fusogenic proteins HAP2/GCS1 to the plasma membrane in response to EC1 signals secreted by the egg cell during double fertilization, thereby ensuring successful fertilization[43]. Loss-of-function mutations impair the response of HAP2/GCS1 to EC1 signals, significantly affecting sperm-egg fusion, ultimately resulting in fertilization failure, thereby promoting the formation of haploids[44,45]. Based on this mechanistic framework, genes expressed in gametes that affect sperm cell-female gamete fusion have been hypothesized as potential haploid-inducing candidates. Among these, KOKOPELLI (KPL) was identified as a pollen-expressed gene whose mutation causes frequent single fertilization events[46]. Functional validation confirmed that kpl mutants induce haploid formation, achieving a maximum HIR of 0.34%[47] (Fig. 2c).
Androgenesis
-
Androgenesis has long been regarded as a rare biological phenomenon, defined as a mode of asexual reproduction in which a mature male gamete develops directly into an embryo without fertilization. Reports of this phenomenon are limited to a few taxa, including angiosperms, fish, and arthropods[48]. Recently, Shi et al. achieved efficient induction of in vivo androgenesis under laboratory conditions[49]. Their study demonstrated that ectopic expression of the AP2/ERF family transcription factor BABY BOOM (BBM), or its direct downstream effector BBM-activated Androgenesis Regulator 1 (BAR1), in microspores of tobacco and rice was sufficient to reprogram microspore fate and trigger androgenesis within the anther. Notably, this approach enables the high-efficiency production of haploid plants without conventional stress treatment. Further analyses revealed that the conserved BBM-BAR1 regulatory module remodels the chromatin state of the gametophyte, activates cell division and associated DNA demethylation processes, and redirects microspores from the gametophytic to the embryogenic developmental pathway. This work not only deepens our understanding of plant cell totipotency and embryogenesis but also provides a novel technological strategy and theoretical basis for haploid induction, highlighting its significant potential for future crop breeding applications.
Egg cell-related haploid induction technology
In vitro culture of megagametophytes or ovules
-
Research has shown that haploid embryo formation can be induced utilizing megaspores, ovules, egg cells, synergids, or antipodal cells, among others[50]. Since the first report in barley[51], this technique has been successfully extended to other species such as sugar beet[52], onion[53], rice[54], and durum wheat[55]. Although the efficiency of haploid induction through in vitro culture of megagametophytes or ovules is limited by the number of egg cells per ovule, this approach offers distinct advantages in overcoming obstacles such as male sterility, pollen defects, and albino plant regeneration, thus providing an effective alternative strategy for haploid breeding in recalcitrant species[56,57].
Based on egg cell development-associated functional genes
-
The maize indeterminate gametophyte1 (ig1) gene encodes a transcription factor containing a LATERAL ORGAN BOUNDARIES (LOB) domain[58]. In ig1 mutants, the female gametophyte frequently exhibits asynchronous nuclear divisions, aberrant nuclear migration, and defects in microtubule organization during free nuclear division, and the plants also display abnormal leaf morphology[58,59]. Genetic analyses have demonstrated that when ig1/ig1 mutants are used as females, they can induce the production of a certain proportion of paternal haploids[60]. However, because the ig1 gene is pleiotropic, with low haploid induction efficiency and a strong dependence on parental genetic background[61], it has not been adopted for commercial haploid induction systems. Nevertheless, ig1 retains considerable importance in basic research, as it provides critical insights into the relationship between embryo sac development and haploid formation[62]. The mechanism by which ig1 induces haploid formation remains incompletely understood. Current hypotheses suggest that abnormalities in the structure and function of the ig1 mutant embryo sac interfere with karyogamy and early embryonic divisions, which in turn prevent the proper integration of maternal chromosomes into the zygotic nucleus or cause their selective elimination, thereby leading to the formation of paternal haploids[62,63].
Loss of function in egg cell-specific regulatory factors can also trigger haploid induction. The egg cell-specific aspartic endopeptidases ECS1 and ECS2 prevent the entry of supernumerary pollen tubes into the embryo sac, as the ecs1 ecs2 double mutant typically exhibits a polytubey phenotype[64]. Subsequent studies revealed that this double mutant could induce haploid formation either through selfing or by serving as the maternal parent in crosses with other varieties, yielding exclusively maternal-derived progeny with a maximum HIR of 1.1%. Mechanistically, semigamy occurs in the mutant: following sperm-egg cell plasmogamy, the sperm nucleus fails to undergo karyogamy with the egg nucleus, resulting in semigamous zygotes that retain egg cell morphology 24 h post-fertilization. A subset of these zygotes undergoes mitosis and ultimately develops into haploid embryos and plants[65]. The haploid induction capability of the ecs1 ecs2 double mutant has been corroborated by another independent study, which also attributes haploid formation to defects in gamete fusion and the coordination of double fertilization[66].
Phylogenetic analyses show that MTL/ZmPLA1/NLD, which encodes a pollen-specific phospholipase A, belongs to the plant phospholipase A (pPLA) gene family. This family is divided into pPLAI, pPLAII, and pPLAIII. The pPLAII group containing MTL/ZmPLA1/NLD comprises five members (pPLAIIα, -β, -γ, -δ, -ε). Functional characterization showed that T-DNA insertion mutants of pPLAIIα and pPLAIIβ exhibited no detectable haploid induction activity, while pPLAIIδ and pPLAIIε were specifically highly expressed in root tissues with negligible expression in reproductive organs. Crucially, pPLAIIγ, which is specifically expressed in the gynoecium, was identified as a maternal haploid inducer, with RNAi lines and knockout mutants achieving a maximum HIR of 1.07%. Notably, this induction process is strictly dependent on the maternal genotype, and the progeny possess exclusively maternal-derived genomes, exhibiting parthenogenesis-like development[67].
Recent studies have demonstrated that the heterozygous mutant of the maize gene Gamete Expression protein 1 (ZmGEX1) can function as a maternal parent to induce haploid formation. When crossed with wild-type paternal plants, the zmgex1+/− mutant exhibits an HIR of 1.34%, with progeny retaining the maternal nuclear genome[68]. This phenomenon displays parthenogenesis-like characteristics, wherein the egg cell undergoes autonomous embryogenesis without fusion with the sperm nucleus. Mechanistic analyses suggest that the mutation in ZmGEX1 may interfere with nuclear fusion during fertilization, ultimately leading to haploid induction[68].
Parthenogenesis
-
Parthenogenesis, an asexual reproduction strategy in which a female gamete develops directly into an embryo without fertilization, occurs naturally in certain plants, invertebrates, and vertebrates[69]. In plants, parthenogenesis specifically refers to the differentiation of an egg cell into an embryo-like structure that develops into a haploid embryo[70]. Recent studies have demonstrated that egg cell-specific expression of parthenogenesis-inducing proteins can trigger haploid embryo formation (Fig. 2d). Key proteins identified thus far include the BABY BOOM (BBM), the EAR-motif zinc-finger protein PARTHENOGENESIS (PAR), and WUSCHEL (WUS), a critical regulator of embryonic development (Table 1).
Table 1. List of parthenogenesis genes inducing haploids when specifically expressed.
Gene Source species Recipient species Phenotype PsASGR-BBML Pennisetum squamulatum Pennisetum glaucum 35%–36% parthenogenesis rate1* in T1[71] Oryza sativa 25%–89% parthenogenesis rate2* in T1[72] Zea mays 47%–80% parthenogenesis rate2* in T1[72] Arabidopsis 0[72] Nicotiana tabacum 2.7%–27.3% haploid rate3* in T2[73] BnBBM1 Brassica napus Arabidopsis 0.4% haploid rate3* in T1[74] Brassica napus 1.8 ± 0.6% haploid rate3* in T1[74] Solanum lycopersicon 1.4% haploid rate3* in T1[74] OsBBM1 Oryza sativa Oryza sativa 5%–10% haploid rate3* in T1, 29% haploid rate3* in T2[75] OsBBM4 Oryza sativa Oryza sativa 3.2% haploid rate3* in T1[76] ZmBBM1 Zea mays Zea mays 65% haploid rate3* in T1[77] ZmBBM2 Zea mays Zea mays 0.4%–3.6% haploid rate3* in T1[78] IbBBM Ipomoea batatas Ipomoea batatas 5%–7.1% haploid rate3* in T1[79] ToPAR Taraxacum officinale Lactuca sativa 15% parthenogenesis rate1* in T1[82] Setaria italica 1.4%–10.2% haploid rate3* in T1[83] OsWUS Oryza sativa Oryza sativa 20% parthenogenesis rate1* in T1[84] OsBBM1 Oryza sativa Oryza sativa 15.8% haploid rate4* in T1[49] NtBAR1 Nicotiana tabacum Nicotiana tabacum 3.6% haploid rate5* in tissue culture[49] 1* Parthenogenesis rate = Number of ovaries with parthenogenic embryos/Number of ovaries. 2* Parthenogenesis rate = Number of transgenic lines with haploid embryo production/Number of transgenic lines. 3* Haploid rate = Number of haploids/Total number of seedlings. 4* Haploid rate = Number of androgenesis embryos/Number of mature anthers. 5* Haploid rate = Number of haploids/Number of microspores. The BBM-like gene PsASGR-BBML from Pennisetum squamulatum has exhibited cross-species parthenogenetic induction capabilities. Ectopic expression of PsASGR-BBML in egg cells triggered haploid parthenogenesis in pearl millet[71], rice[72], maize[72], and tobacco[73]. Notably, functional conservation has been observed among its orthologs: BnBBM1 in rapeseed[74], OsBBM1[75] and OsBBM4[76] in rice, as well as ZmBBM1[77] and ZmBBM2[78] in maize, all successfully induced haploid parthenogenesis when expressed ectopically in their respective species. In addition to ectopic expression in egg cells, constitutive activation of the IbBBM gene in Ipomoea batatas using the CRISPR activation (CRISPRa) system, which specifically targets IbBBM, has also been shown to induce the production of haploid progeny[79]. Recent studies have further revealed that the efficiency of BBM-mediated parthenogenesis can be substantially enhanced through co-expression with synergistic factors. For instance, co-expression of OsWOX9A/OsDWT1 with OsBBM1 increased parthenogenetic haploid induction rates to 86%–91%, representing a 4- to 15-fold improvement over OsBBM1 overexpression alone[80]. Similarly, co-expression with cell cycle regulator cyclin delta-2 (CYCD2) boosted induction efficiency by 34%[81].
The PARTHENOGENESIS (ToPAR) gene from Taraxacum officinale, encoding an EAR domain-containing zinc finger protein, has shown conserved functionality across divergent species. Its ectopic expression induced haploid parthenogenesis in Lactuca sativa[82] and Setaria italica[83].
Of particular significance is the discovery that OsWUS from rice, a species not naturally exhibiting parthenogenesis, can induce haploid embryogenesis when specifically expressed in egg cells[84]. This breakthrough challenges the conventional paradigm that parthenogenesis-related genes are exclusively derived from naturally parthenogenetic species, highlighting the untapped potential of crop gene resources for haploid parthenogenesis engineering.
Centromere-related haploid induction technology
-
Since 1970, when Kasha et al. first achieved haploid production by crossing diploid bulbous barley (Hordeum bulbosum) with cultivated barley (Hordeum vulgare), haploid induction via distant hybridization has been progressively developed into an important strategy for generating haploids in wheat and other crops[85]. Subsequent studies revealed that chromosome elimination is closely linked to centromere dysfunction. In the H. bulbosum × H. vulgare hybridization system, the centromeres of the paternal H. bulbosum chromosomes lose functionality due to a significant reduction or complete absence of the Centromeric histone H3 (CENH3) protein. This impairs their proper attachment to the spindle apparatus, ultimately leading to their selective elimination. Consequently, the hybrid progeny retains only the maternal H. vulgare chromosomes, resulting in haploid plants[86].
CENH3 has been identified in recent years as a key gene for the development of haploid inducer lines. CENH3 is uniquely localized to the functional centromeres of eukaryotic cells, where it plays a crucial role in recruiting and stabilizing centromere-associated proteins, ensuring proper chromosome segregation during cell division, and determining the centromere position on chromosomes[87]. CENH3 consists of two domains: an N-terminal tail (which extends from the nucleosome and serves as a target for post-translational modifications) and a C-terminal histone fold domain (HFD)[88].
Ravi & Chan reported in Arabidopsis that mutation of the CENH3 gene leads to the selective elimination of parental chromosomes in the hybrid zygotes, resulting in the production of haploid progeny. When mutant CENH3 plants are crossed with wild-type plants, chromosomes inherited from the mutant parent are selectively eliminated after fertilization due to centromere dysfunction, thereby generating haploids containing only the wild-type genome[89]. Further studies have shown that mutant CENH3 is specifically stripped from chromatin during egg cell maturation, whereas wild-type CENH3 remains stable. After fertilization, in competition for CENH3 loading, the wild-type chromosomes outcompete the mutant chromosomes, leading to preferential CENH3 assembly on chromosomes. Consequently, the mutant chromosomes fail to assemble functional centromeres and are eliminated during subsequent mitotic divisions (Fig. 2b).
The function of the CENH3 gene is conserved across species. CENH3 is indispensable for plant growth and development due to its essential role in both mitosis and meiosis; consequently, complete disruption of this gene leads to homozygous lethality[89,90]. Leveraging its conserved function, partial modification of CENH3 has enabled haploid induction, a strategy that has led to key breakthroughs in several crops such as maize, wheat, cucumber, and melon[91−93]. In maize, Kelliher et al. developed a transgenic complementation line, ZmCENH3-tailswap, by replacing the N-terminal tail domain of the CENH3 protein. This line was capable of inducing maternal haploids, with an HIR of up to 3.6%[92]. Wang et al. generated heterozygous Zmcenh3 frameshift mutants (+/−) using CRISPR/Cas9 and found that when used as the paternal, the HIR reached 5%, significantly higher than the 0.5% observed when used as the maternal, while the mutant plants exhibited normal growth[94]. To overcome hybridization barriers in allohexaploid wheat, Lv et al. developed a paternal haploid inducer line by editing the TaCENH3α-A gene to create a frameshift repair heterozygote (+/r) combined with complete knockouts of the B and D genome copies. This approach achieved a stable HIR of 7%–8%. The findings suggested that partial CENH3 function must be retained in the A genome (+/r heterozygosity), while complete loss of function in the B and D genomes is necessary for highly efficient selective chromosome elimination[93]. In cucurbit crops, Van Dun et al. employed EMS mutagenesis to generate CENH3 mutants, achieving HIR of approximately 1% in cucumber and 1.5% in melon, thereby providing preliminary evidence for haploid induction in this crop group[91]. In addition to genetic modifications, environmental conditions also affect the efficiency of CENH3-mediated haploid induction. Wang et al. reported that in Arabidopsis thaliana, the CENH3-GFP-tailswap inducer is highly sensitive to ambient temperature: lower temperatures restore pollen viability but reduce the HIR, whereas higher temperatures markedly increase HIR but are accompanied by seed abortion. By temporally manipulating the growth temperature before and after pollination, the maternal HIR could be increased to approximately 24.8%[95]. Balancing HIR and normal inducer plant growth is crucial. Yang et al. achieved this by utilizing a miRNA-induced gene silencing (MIGS) approach. They targeted CENH3 using a promoter from the RPS5a (ribosomal protein S5a) gene, which is specifically expressed during gametophyte- and embryo-development. This strategy generated inducer lines exhibiting both high HIR and normal growth characteristics[96].
The centromere licensing factor Kinetochore Null2 (KNL2) co-localizes with CENH3 throughout the cell cycle, except during metaphase to mid-anaphase[97]. Its C-terminal domain contains a conserved CENPC-k motif, which mediates the interaction between KNL2 and CENH3. Functional analyses demonstrate that either deletion of the CENPC-k motif or mutation of a single conserved residue completely abolishes the centromeric localization of KNL2[98]. Critically, point mutations in the CENPC-k motif alone are sufficient to establish haploid inducer lines. Furthermore, transient heat stress applied to knl2 mutants increases the HIR 10-fold, likely due to heat-induced aggravation of meiotic and mitotic defects[99].
The loss-of-function mutation of the methylcytosine-binding protein VARIANT IN METHYLATION 1 (VIM1) induces hypomethylation of centromeric repeats, centromeric decondensation, and reduced CENH3 density at the centromeres. These changes significantly enhance the HIR of CENH3-based inducer lines[100].
DNA Ligase 4 (LIG4) repairs DNA double-strand breaks through the non-homologous end joining (NHEJ) pathway. When the lig4 loss-of-function mutant is used as the paternal parent, its inability to efficiently repair chromosomes may lead to the elimination of its own genome, thereby significantly enhancing the CENH3-mediated paternal haploid induction rate (HIR)[101].
Haploid induction can be achieved through various approaches, including in vitro culture of male or female gametophytes, development of haploid inducer lines via targeted mutations in genes associated with pollen development, egg cell development, or centromere (Table 2), and ectopic expression of parthenogenesis-related genes to artificially trigger haploid embryogenesis (Table 1).
Table 2. List of genes inducing haploids when knocked out.
Gene Source species Function Haploid inducer parent Haploid induction rate (HIR) (%) ig1 Zea mays LOB domain transcription factor, expressed in embryo sac Paternal up to 3.3[60] MTL/ZmPLA1/NLD Zea mays Phospholipase A, expressed in endo-PM of the vegetative cell Paternal 2–6.7[19−21] POD65 Zea mays Peroxidase, expressed in pollen Paternal 1–7.7[30] PLD3 Zea mays Phospholipase D3, expressed in pollen Paternal 0.85–0.96[32] PLDα2 Oryza sativa Phospholipase D, expressed in pollen Paternal 0.3–0.6[33] DMP Zea mays DUF679 domain membrane protein, expressed in pollen Paternal 0.1–0.3[34] KPL Arabidopsis Natural cis-antisense siRNA-encoding gene, expressed in pollen Paternal 0.07–0.34[47] ECS1/2 Arabidopsis Aspartic endopeptidases, expressed in egg cell Maternal ~1[65,66] pPLAIIγ Arabidopsis Phospholipase A, expressed in gynoecium Maternal 0.93–1.12[67] ZmGEX1 Zea mays Versatile membrane protein, expressed in gamete Maternal 0.49–1.94[68] CENH3 Arabidopsis Centromeric histone 3, expressed in centromere Maternal ~30[89] Paternal ~4[89] KNL2 Arabidopsis Centromere licensing factor Maternal 1–10[99] Haploid screening
-
With the progressive elucidation of key haploid induction genes such as MTL/ZmPLA1/NLD, DMP, and CENH3, the efficient and accurate screening of haploids has become a central challenge in engineered breeding systems. Traditional identification methods primarily rely on plant phenotypic traits, cytogenetic analysis, and genetic marker technologies[24,102]. Among these, morphological identification is limited by long growth periods of plants, strong environmental sensitivity, and the need for extensive land use, while cytological methods based on chromosome counting are labor-intensive and cumbersome. Consequently, the development of novel molecular markers and high-throughput screening technologies has become a major research focus in haploid screening.
In genetic marker systems, the anthocyanin biosynthesis gene R1-nj has been widely used due to its embryo- and endosperm-specific coloration. When maize haploid inducer lines homozygous for the R1-nj marker are used to pollinate Zhengdan 958, the haploid embryos of the progeny exhibit colorless characteristics[103]. However, the coloration stability of this marker is often affected by genetic background and environmental factors, resulting in an identification accuracy of only 60%–90%[104]. To overcome these limitations, Chen et al. developed the Maize Anthocyanin Gene Inducer (MAGIC) system by co-expressing transcription factors ZmC1 and ZmR2 to create an embryo-specific purple coloration in the MAGIC1 system, enabling haploid identification with 99.1% accuracy just 12 d after pollination. However, the germination rate of seeds under this system was only 34.8%. To address this issue, the MAGIC2 system replaced the promoter with the seed-specific bidirectional promoter 2R5SGPA, which not only maintained the high identification accuracy of MAGIC1 but also successfully increased the seed germination rate to 85.3%[104].
In recent years, the introduction of the betacyanin biosynthesis pathway has provided a new paradigm for haploid identification. Wang et al. integrated the RUBY system, containing the Cytochrome P450 enzyme (CYP76AD1), the DOPA dioxygenase (BvDODA1), and the 5-O-glucosyltransferase (cDOPA5GT), into maize inducer lines, allowing for a gradual visible coloration in hybrid embryos as early as 10 d after pollination. After 24 h of in vitro culture, betacyanin accumulation was sufficient to achieve 100% accuracy in haploid identification[105].
Differences in oil content between hybrid diploid and haploid seeds produced by crossing high-oil inducers and regular maize lines can be exploited using nuclear magnetic resonance (NMR) or near-infrared (NIR) spectroscopy, significantly reducing detection time compared to manual identification[103]. A high-oil maize haploid inducer line, CHOI4, was developed by crossing the haploid inducer CAU2 with the high-oil population BHO. CHOI4 exhibited an average kernel oil content of 11% and a mean HIR of 15.78%[106]. The application of instrument-based detection allowed for rapid and accurate discrimination between haploid and diploid kernels, thereby minimizing subjective errors associated with manual assessment through the quantification of physiological and biochemical traits.
Breakthroughs in fluorescence marker technology have significantly advanced the precision of haploid identification. A dual-fluorescence marker system, constructed using embryo- and endosperm-specific promoters, enabled differential coloration between the embryo and endosperm. In this system, both diploid and haploid kernels exhibit red fluorescence in the endosperm, whereas green fluorescence in the embryo is observed exclusively in diploid kernels[107]. The FAST-Red fluorescence method, employing a two-step strategy of seed immersion and root tip detection, enables non-destructive identification of tomato haploids[37]. Further innovation involves the naked-eye-visible marker MoDsRed, which, when combined with a handheld light source, allows 100% accurate sorting of maize kernels[108]. These technological innovations offer broadly applicable solutions for haploid breeding across multiple crop species.
Chromosome doubling
-
Chromosome doubling is a crucial step in all haploid utilization technologies. Over the past decades, various chromosome doubling methods have been developed. Currently, the most commonly used approaches involve treatment with colchicine or herbicides such as amiprophos-methyl (APM), trifluralin, pronamide, and oryzalin. These chemical agents induce chromosome doubling via microtubule depolymerization, which disrupts spindle assembly during mitosis[56]. Kato & Geiger pioneered the nitrous oxide (N2O) treatment for maize chromosome doubling[109]. Subsequent studies in Lilium species demonstrated that the mechanism of this method also relies on microtubule depolymerization[110]. However, chemical treatments may cause cellular damage. Spontaneous genome doubling offers a chemical-free alternative and has been reported in crops such as maize, barley, wheat, and onion during microspore or anther culture. Furthermore, its doubling efficiency can reach up to 70%[111−113].
However, spontaneous chromosome doubling cannot meet the demands of large-scale production, and colchicine-based methods remain the predominant approach for haploid doubling in current agricultural practices[114]. Common techniques for colchicine application include seed soaking, bud immersion, root immersion, apical meristem droplet treatment, stem injection, and tissue culture. Recently, Lv et al. optimized a chromosome doubling technique by delivering colchicine to the shoot apical meristem using a semi-solid fluorinated oil carrier, resulting in a doubling efficiency from 31% to 55%[70].
The application of haploid technology in combination with genome editing techniques
HI-edit/IMGE technology: overcoming the bottleneck of genetic transformation
-
Although the CRISPR/Cas system has revolutionized plant genome editing, its application in commercial crops remains significantly constrained by the species- and genotype-dependence of conventional genetic transformation. To overcome this limitation, haploid inducer-mediated genome editing (HI-Edit/IMGE) has emerged as a promising solution[115,116]. By integrating haploid induction with genome editing, this strategy enables broad applicability across diverse genetic backgrounds. Specifically, CRISPR/Cas components are introduced into haploid inducer lines, which are then crossed with the recipient material. Haploids carrying the desired mutations can be obtained in the progeny by taking advantage of genome elimination from the paternal or maternal side during haploid induction. Subsequent chromosome doubling can rapidly produce homozygous doubled haploid lines harboring the desired edits within two generations, significantly shortening breeding cycles and enhancing the efficiency of genome editing applications (Fig. 3a). To date, HI-Edit has been successfully implemented in maize[115−117], Arabidopsis[115], and wheat[115]. Its core advantages lie in bypassing the need for genotype-dependent transformation systems and enabling the generation of non-transgenic edited materials through transgenerational segregation of editing components. However, several challenges remain for its practical deployment in crop breeding programs. These include limitations in haploid induction rates and editing elements transmission efficiency. Strategies such as increasing the editing frequency at target sites, employing alternative promoter elements, and improving chromatin accessibility at target sites have been proposed to optimize overall system performance[117].
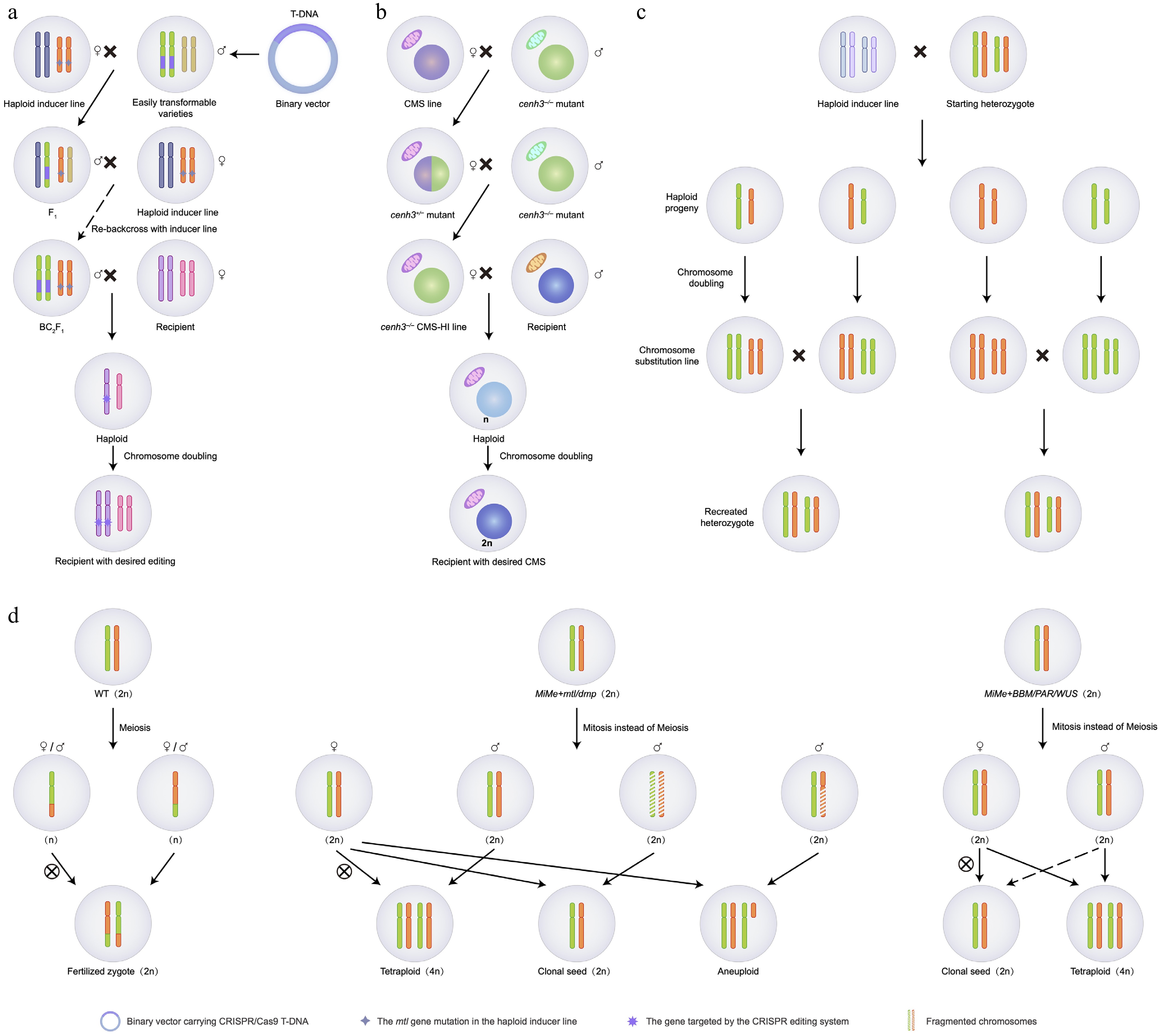
Figure 3.
Applications of haploid induction in crop breeding.
(a) HI-Edit/IMGE technology. A CRISPR/Cas9-expressing HI line is crossed with a target recipient. The Cas9 cassette edits the recipient genome before the HI genome is eliminated, yielding gene-edited haploids that maintain the recipient’s genetic background after chromosome doubling. (b) Cytoplasmic replacement. A cenh3-mediated CMS-HI line is used as the maternal parent to cross with a recipient. This generates paternal haploids with the donor CMS cytoplasm and recipient nuclear genome, accelerating CMS line development. (c) Reverse breeding. Suppression of meiotic recombination allows the generation of gametes with intact parental chromosomes. Subsequent DH production and crossing of complementary lines enable the reconstruction of the original F1 hybrid genotype. (d) Synthetic apomixis. Combining the MiMe genotype (mitosis instead of meiosis) with chromosome elimination (mtl, dmp) or parthenogenesis (BBM, PAR, WUS) enables clonal reproduction of maternal genomes through seeds.Rapid development of cytoplasmic male sterile (CMS) lines
-
Cytoplasmic male sterile (CMS) lines play a critical role in hybrid seed production by inducing pollen abortion to prevent self-pollination and maintain hybrid purity. However, conventional CMS line development requires six to eight generations of backcrossing to introgress sterile cytoplasm into elite parental lines; it is a time-consuming and labor-intensive process. Haploid inducer-mediated cytoplasmic replacement has emerged as a revolutionary strategy. By eliminating the maternal nuclear genome selectively while retaining its sterile cytoplasm, this approach enables the direct transfer of the CMS trait from the donor to the target line[118].
The core procedure involves using a haploid inducer line that carries CMS cytoplasm as the maternal parent and the target recipient line as the paternal parent. The resulting haploid progeny inherit the nuclear genome from the paternal parent and the cytoplasm from the maternal parent. Following chromosome doubling, new CMS germplasm can be rapidly obtained, featuring a homozygous nuclear background and the desired sterile cytoplasm. This strategy shortens the CMS conversion process to just two to three generations, significantly reducing the breeding cycle and improving production efficiency (Fig. 3b).
In maize, Bortiri et al. developed a CENH3-based paternal haploid inducer line carrying a dominant color marker gene, and successfully applied this line to mediate cytoplasmic substitution, enabling the rapid conversion of any maize inbred line into the corresponding CMS line[119]. A similar strategy was applied in broccoli, where Han et al. established a CENH3-based paternal haploid inducer line named HI-CMS, which was likewise used to mediate cytoplasmic substitution, allowing for the efficient conversion of any broccoli inbred line into the corresponding CMS line[120].
Although haploid induction technology has greatly improved the efficiency of CMS line development, its practical application is still constrained by factors such as HIR and the success rate of chromosome doubling. With continued refinement of this technology, it is anticipated that this technology will be applied to more species, thereby accelerating the efficiency and modernization of hybrid seed production.
Reverse breeding
-
In conventional plant breeding, the inherent randomness of genetic recombination during meiosis results in unpredictable combinations of alleles in each offspring, requiring breeders to invest significant time and effort in identifying individuals with desirable traits. In contrast, reverse breeding offers a fundamentally different approach. Starting from a phenotypically superior F1 hybrid, this strategy utilizes molecular and cellular engineering techniques to reconstruct its parental lines, enabling the controlled re-creation and sustained use of elite hybrid genotypes.
The reverse breeding process generally involves the following steps: (1) Selection of a F1 hybrid with superior agronomic traits as the starting material; (2) Suppression of meiotic recombination to preserve parental haplotypes; (3) Obtain haploids: ensure that each gamete carries a non-recombined haploid genome; (4) Conversion of these haploid plants into homozygous DH lines; (5) Screening of these homozygous lines to identify two complementary genotypes that recapitulate the original hybrid genotype upon crossing (Fig. 3c).
Among these steps, suppression of recombination and haploid induction are considered the two most critical bottlenecks. To suppress recombination, several strategies have been explored, including RNA interference (RNAi) and virus-induced gene silencing (VIGS), which target genes critical for homologous chromosome pairing, synapsis, or crossover formation, such as ASY1, PAIR2, DMC1, PTD, SPO11, and SDS[121]. Alternatively, exogenous application of recombination-suppressing chemical agents, such as Mirin, has also been shown to be effective[122]. For haploid induction, established methods include anther or ovule culture, as well as the use of haploid inducer lines (e.g., cenh3 mutants) to generate haploid plants.
This concept represents a disruptive innovation in modern breeding, providing a precise and efficient means to stabilize and propagate superior hybrid genotypes. Proof-of-concept studies have demonstrated its feasibility in the model plant Arabidopsis thaliana[123−125]. However, its application in major crop species has not yet been reported. This is mainly due to several technical challenges: the suppression of recombination is significantly difficult in complex crop genomes; haploid induction efficiency remains low in many crop species; and chromosome doubling, a prerequisite for doubled haploid production, is often inefficient or genotype-dependent, further limiting the widespread application of the technique in crop breeding.
Apomixis
-
Apomixis, an asexual reproductive strategy enabling clonal seed formation, naturally occurs in over 400 angiosperm species[126], including sorghum, dandelion, citrus, apple, beet, and so on. However, functional apomictic systems have not yet been identified in major cereal crops. Extensive research has focused on identifying key genes, molecular mechanisms, and embryogenesis processes involved in apomixis. Early studies aimed to introduce apomixis-related genes into target species, but stable expression and functional realization in major crops have not yet been achieved (Table 3).
Table 3. List of methods inducing apomixis.
Species Method Setting percentage (%) Clone seed ratio (%) Arabidopsis Crossing MiMe plants with GEM line ND 33[130] Oryza sativa Simultaneous editing MiMe genes in BBM1 transgene plants (MiMe + OsBBM1-ee) ND 11–29[75] Oryza sativa Simultaneous editing MiMe genes and MTL in hybrid rice (Fix1) 3.7–5.2 4.7–9.5[134] Oryza sativa Simultaneous editing MiMe genes and expressing egg-cell specific BBM4 (Fix2) 80.9–86.1 1.3–2.4[76] Oryza sativa Simultaneous editing MiMe genes and expressing egg-cell specific PAR 74.4–82.8 42.9–67.7[132] Arabidopsis Simultaneous editing MiMe genes, AtDMP8 and AtDMP9 (AtFix1) 10–24 3.8[131] Oryza sativa Editing MiMe genes, and expressing egg-cell specific enhanced BBM1 (MiMe-ECA1-AZP2) Up to 89.7 95–100[133] Oryza sativa Simultaneous editing MiMe genes and expressing egg-cell specific OsWUS (Fix3) 72.0–85.2 0.5–21.7[84] Oryza sativa Simultaneous editing MiMe genes and OsPLDα2 in hybrid rice (Fix4) 82.1–85.0 1.2[33] 'ND' indicates not shown in the literature. For hybrid crops, apomixis offers a transformative potential: fixing heterosis by producing clonal seeds that retain hybrid vigor across generations, thereby eliminating the need for repeated hybridization. This concept was first proposed as early as the 1930s and has long been regarded as a holy grail in agriculture. However, due to technological limitations, progress in this field remained slow until recent years, when significant breakthroughs began to emerge.
The current strategy for engineering apomixis aims to convert sexual reproduction into a gametophytic apomictic process. This involves three key steps: (1) suppressing meiosis in megaspore mother cells and inducing mitosis instead; (2) replacing sexual reproduction with parthenogenesis, allowing an embryo genetically identical to the maternal parent to develop without fertilization; and (3) enabling autonomous endosperm development (Fig. 3d).
For the first step, the conversion of meiosis into mitosis can be achieved by disrupting homologous chromosome pairing and recombination, promoting the premature separation of sister chromatids, and skipping the second meiotic division. This can be accomplished by knocking out SPO11-1, REC8, and OSD1 in Arabidopsis[127], or their homologs in rice and tomato[128,129], thereby generating 'Mitosis instead of Meiosis' (MiMe) genotypes.
For the second step, there are two main strategies to induce parthenogenesis and produce clonal seeds.
The first strategy involves haploid induction-mediated chromosome elimination. For example, using MiMe lines as the maternal parent and CENH3 mutant lines (GEM lines) as the paternal parent, paternal chromosomes are eliminated, resulting in seeds that retain only the maternal nuclear genome and develop into diploid clonal seeds[130]. Alternatively, the simultaneous knockout of the three MiMe genes along with the haploid induction gene MTL/ZmPLA1/NLD or DMP can also lead to apomictic reproduction[75,131].
The second strategy involves synergistic activation of parthenogenesis. Clonal seeds can be generated by knocking out the three MiMe genes in lines overexpressing the BBM gene[75]. Similarly, ectopic expression of BBM, PAR, or WUS, combined with the knockout of the three MiMe genes, has also been shown to induce clonal seed formation[76,84,132,133].
These findings have not only established a solid theoretical foundation for achieving efficient apomixis in crops but also provided innovative technological avenues for fixing hybrid vigor and enabling rapid clonal propagation. Although engineered apomictic systems have been demonstrated as proof of concept in the model plant Arabidopsis thaliana and in rice, their commercial deployment still faces major challenges, such as seed abortion caused by defective autonomous endosperm development; elevated genetic instability risks from aneuploidy induced by the MiMe system, and technical limitations in multiplex gene editing efficiency.
-
In recent years, several key genes associated with haploid induction have been identified. However, research in this field remains at an early stage overall. The regulatory mechanisms of many genes involved are still not fully elucidated. Current understanding of haploid induction is largely centered on genes related to pollen, egg cells, and centromeres.
Among these, the study of pollen-related haploid induction genes has shown the most significant progress. Notably, the phospholipase A gene MTL/ZmPLA1/NLD in maize has been most extensively investigated. Its mutation confers high-frequency maternal haploid induction in Stock 6 and its derivatives[19−21]. Subsequent research has shown that mutations in MTL/ZmPLA1/NLD trigger a ROS burst in pollen, resulting in fragmentation and elimination of the paternal genome post-fertilization, thereby producing maternal haploids[30]. Additionally, mutations in the pollen-specific genes such as ZmPOD65, ZmPLD3, and OsPLDα2 also confer haploid induction capability[30,32,33], suggesting crucial roles for phospholipid metabolism and oxidative stress pathways in haploid formation. However, the absence of known orthologs of MTL/ZmPLA1/NLD in dicots limits its broader applicability. In contrast, the DMP gene is widely conserved in dicots, and its mutation effectively induces maternal haploids across multiple crops[37,38,40,41,43]. Studies indicate that DMP mutation impairs pollen function, preventing sperm-egg cell fusion and triggering autonomous development of the egg cell into an embryo, ultimately generating haploid plants with only the maternal genome.
Egg cell-related haploid induction genes have also gained increasing attention. For example, double mutants of ECS1 and ECS2 fail to complete karyogamy after fertilization, resulting in maternal haploid formation[65,66]. Furthermore, mutations in the gynoecium-specific phospholipase gene pPLAIIγ and the gamete-expressed protein gene ZmGEX1 likely prevent nuclear fusion during fertilization, ultimately inducing haploid formation. The mutants of both genes act as maternal haploid inducers, producing progeny with genomes exclusively derived from the maternal parent and exhibiting parthenogenesis-like characteristics[67,68]. Another group of genes is directly associated with parthenogenesis mechanisms. Normally, the BBM gene is primarily expressed in sperm cells and activates zygote division post-fertilization. However, its specific expression in the egg cell enables autonomous initiation of division without fertilization, resulting in haploid embryo formation[75]. Similarly, the PAR gene, which is predominantly expressed in egg cells of the apomictic plant dandelion, can induce embryo development without fertilization when ectopically expressed in egg cells of heterologous species[82,83]. Unlike BBM and PAR, WUS does not originate from a natural parthenogenesis system but also demonstrates haploid induction potential, suggesting that non-natural parthenogenesis systems may harbor valuable gene resources for haploid induction[84]. These discoveries offer new strategies for haploid induction via the parthenogenesis pathways, although the functional effects and regulatory networks of these genes across species remain unclear, requiring further validation and engineering optimization.
Research on centromere-associated haploid induction primarily focuses on the variant histone H3, known as CENH3. CENH3 replaces canonical histone H3 in centromeric nucleosomes, defining centromere position and recruiting kinetochore proteins[135]. During hybridization, chromosomes with mutant CENH3 fail to effectively recruit kinetochore proteins, leading to the formation of weak centromeres. These chromosomes are subsequently eliminated during embryogenesis, leaving only chromosomes from the normal parent, thus producing haploid progeny[136]. Interestingly, the HIR is higher when the cenh3 mutant serves as the maternal parent rather than the paternal. This is likely because egg cells undergo three rounds of meiotic division, whereas sperm undergo two, resulting in lower CENH3 levels in egg cells and increased likelihood of maternal chromosome elimination, thereby forming paternal haploids[94]. Additionally, mutations in KNL2, a loading factor for CENH3, can also induce haploids[98], indicating that not only CENH3 itself but also its assembly and regulatory factors influence haploid formation. However, current CENH3-based haploid induction strategies still rely heavily on genetic engineering—such as fusion protein creation or targeted mutagenesis—and induction efficiencies remain suboptimal. Therefore, identifying more effective regulatory factors and optimizing induction conditions are urgently needed to enhance practical utility.
In summary, haploid induction is a complex process driven by multiple genetic mechanisms, including pollen development, fertilization failure of egg cells, activation of parthenogenesis, and chromosomal stability. Continued discovery of novel haploid induction genes will not only contribute to the development of more efficient induction systems but also deepen our understanding of the underlying biological processes.
-
The authors confirm their contributions to the paper as follows: study conception and design, literature search, data extraction, draft manuscript preparation: Zhao J, Li Y; literature collection, figure preparation: Zhao Q, Zhang K; data analysis: Xu Z, Xu L; manuscript revision: Xu Z, Xu L, Li Y; reference management, proofreading: Xiang S, Xia X, Tong Y; supervision, strategic guidance: Li Y. All authors reviewed the results and approved the final version of the manuscript.
-
We thank Dr. Yi Zheng, Dr. Jianhang Guo, Dr. Shu Yuan, and Dr. Mengmeng Fu, and other colleagues for their insightful discussions and suggestions that greatly improved this review.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of Yunnan Agricultural University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhao J, Zhao Q, Zhang K, Xu Z, Xu L, et al. 2026. Overview of haploid technologies and opportunities in plant breeding. Agrobiodiversity 3(2): 41−54 doi: 10.48130/abd-0026-0004
Overview of haploid technologies and opportunities in plant breeding
- Received: 04 February 2026
- Revised: 20 March 2026
- Accepted: 23 March 2026
- Published online: 22 April 2026
Abstract: Haploid breeding technology serves as a pivotal approach to accelerate plant breeding by enabling the generation of homozygous doubled haploid (DH) lines within a single generation, thereby significantly shortening breeding cycles. We systematically categorize current haploid induction strategies into pollen-mediated, egg cell-mediated, and centromere-mediated approaches, based on their underlying mechanisms. Pollen-related approaches include in vitro microspore/anther culture, pollen treatment combined with embryo rescue, and mutations in genes regulating pollen development. Egg cell-related strategies encompass in vitro female gametophyte culture, mutations in genes involved in egg cell development, and the induction of parthenogenesis. Centromere-related techniques primarily achieve selective chromosome elimination through the manipulation of CENH3 and its associated factors, leading to haploid formation. We also highlight recent advances in haploid screening and chromosome doubling methods, along with the integration of gene editing and haploid induction technologies. These combined methods have facilitated the generation of edited lines in any genetic background (HI-Edit/IMGE), the development of cytoplasmic male sterile lines in diverse backgrounds, the sustainable reproduction and utilization of elite hybrid genotypes (reverse breeding), and the fixation of heterosis through clonal reproduction (synthetic apomixis). Haploid induction technology is becoming a cornerstone in the development of precise genetic improvement systems that are deeply integrated with modern breeding, making it possible to provide robust support for crop molecular design breeding.












