-
Soil nitrogen (N) cycling represents one of the most fundamental biogeochemical processes in terrestrial ecosystems. As a key component of this cycle, nitrification plays a crucial role in regulating N availability, mitigating greenhouse gas (GHG) emissions, and maintaining overall environmental quality. Nitrification is an aerobic, two-step process in which ammonia-oxidizing microorganisms (AOM), including ammonia-oxidizing archaea (AOA) and ammonia-oxidizing bacteria (AOB), oxidize ammonia (NH3) to nitrite (NO2−). This is subsequently followed by the oxidation of NO2− to nitrate (NO3−) carried out by nitrite-oxidizing bacteria (NOB)[1]. Traditionally, these two steps were thought to be performed by distinct functional microbial groups, forming the classical 'two-step nitrification' model. However, the recent discovery of complete ammonia oxidizers (comammox), which are capable of converting NH3 to NO3− within a single organism, has challenged this long-standing paradigm and introduced additional complexity to our understanding of the nitrification process[2,3].
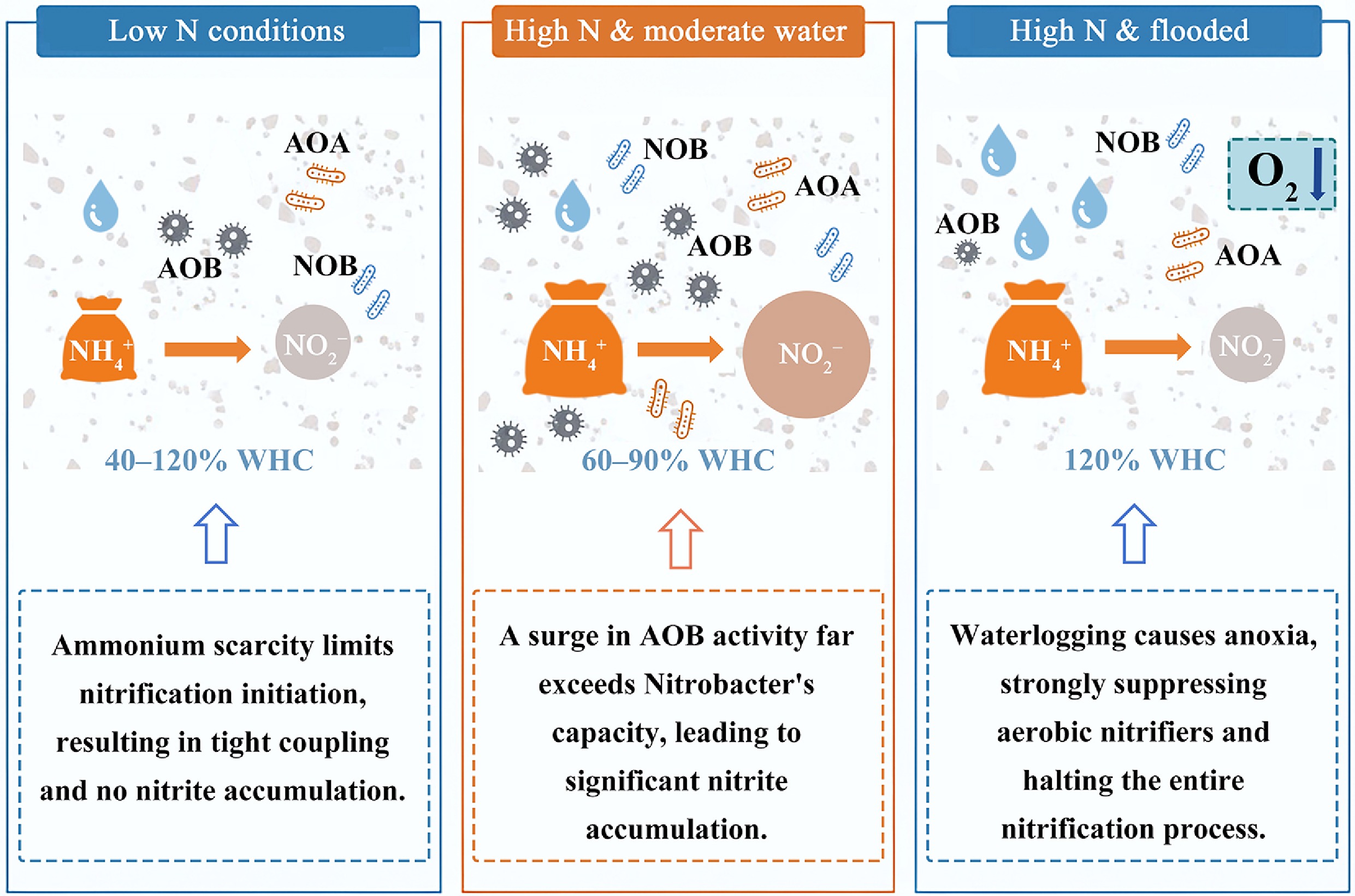
Under normal conditions, the two steps of nitrification are tightly coupled, resulting in minimal accumulation of NO2−. However, environmental disturbances can disrupt this coupling between NH3 and NO2− oxidation, leading to substantial NO2− accumulation[4]. Nitrite acts as a key intermediate and precursor for several harmful nitrogenous compounds, including NO2−, nitric oxide (NO), and nitrous acid[5]. Therefore, elucidating the factors and mechanisms governing the coupling between NH3 and NO2− oxidation is essential for advancing our understanding of N transformation processes and mitigating their environmental consequences.
Soil moisture, as a key environmental factor influencing microbial activity and nutrient diffusion, exerts a profound effect on all stages of the nitrification process. On one hand, soil moisture directly controls oxygen (O2) availability, thereby affecting the growth and activity of aerobic ammonia-oxidizing and nitrite-oxidizing microorganisms. Previous studies have demonstrated that higher water content in inter-aggregate pores can impede O2 diffusion, creating a microenvironment that favors denitrification over nitrification[6,7]. AOB and NOB differ in their affinity for O2; for instance, Nitrobacter winogradskyi (an NOB) is less competitive than Nitrosomonas europaea (an AOB) under low-O2 conditions, placing Nitrobacter winogradskyi at a disadvantage in hypoxic soils[8]. Guisasola et al.[9] further refined the method developed by Wiesmann to simultaneously estimate the O2 affinity constants for AOB and NOB (the O2 affinity constant for AOB (KOA) and O2 affinity constant for NOB (KON), respectively). Their results showed that KOA was 0.74 ± 0.02 mg O2 dm−3, whereas KON was 1.75 ± 0.01 mg O2 dm−3, indicating that NO2− oxidation is more sensitive to fluctuations in dissolved O2 concentrations than NH3 oxidation[9]. Beyond O2 availability, soil moisture also influences nitrification by regulating the diffusion rates of nutrients and substrates. Under either excessively low or high soil moisture conditions, substrate diffusion can be constrained or microbial activity suppressed, thereby reducing nitrification rates[10]. Such constraints may decouple the oxidation of NH3 and NO2−, leading to the accumulation of NO2−.
Moreover, different ammonia-oxidizing and nitrite-oxidizing microorganisms may exhibit distinct responses to soil moisture, leading to shifts in the dominant microbial communities under varying moisture conditions. These community shifts can, in turn, influence both the rate and pathway of nitrification. For example, some studies have suggested that AOA may have a competitive advantage over AOB under low-O2 conditions, whereas AOB exhibit significantly higher maximum NH3 oxidation rates (Vmax) compared to both AOA and comammox[11]. However, a study by Liu et al.[12] reported that the gene abundances of both AOA and AOB increased with rising soil moisture levels, ranging from 50% to 85% water-filled pore space (WFPS). The effects of soil moisture on AOB abundance remain inconsistent across studies. Hastings et al.[13] observed that increasing soil moisture may enhance AOB abundance by alleviating water stress, whereas Nguyen et al.[14] suggested that excessive soil moisture could restrict O2 diffusion, thereby reducing the abundance of nitrifying bacteria. Additionally, the presence and activity of comammox organisms may also be regulated by soil moisture, potentially altering the dominant nitrification pathways in soils[15,16].
Despite the recognized importance of soil moisture in regulating nitrification, studies explicitly addressing its influence remain limited, and systematic investigations into the coupling between NH3 and NO2− oxidation under varying moisture conditions are still scarce. Most existing research on O2-mediated coupling has been conducted in wastewater treatment systems rather than in natural soils[17]. Although nitrification has been extensively investigated in wastewater treatment systems, the mechanistic understanding derived from aquatic media cannot be directly transferred to soils. Unlike wastewater, soil is a heterogeneous porous matrix in which water content simultaneously affects O2 diffusion, substrate transport, redox gradients, and microbial niche partitioning at the microscale. As a result, the coupling between NH3 oxidation and NO2− oxidation in soils is expected to be more spatially variable and more sensitive to environmental thresholds than in relatively homogeneous aquatic systems. Furthermore, NH4+ availability plays a crucial role in regulating soil nitrification. Higher NH4+ concentrations typically stimulate ammonia oxidation by increasing substrate availability, especially for AOB[18]. Nevertheless, the combined effects of soil moisture and NH4+ availability are still poorly understood.
Therefore, this study aims to quantify the coupling between NH3 and NO2− oxidation under controlled soil moisture and NH4+ conditions. Specifically, we investigate how variations in soil moisture and NH4+ conditions affect the abundance and activity of ammonia-oxidizing and nitrite-oxidizing microorganisms, and how these microbial responses collectively regulate the rate and pathway of soil nitrification. The findings are expected to provide new mechanistic insights into the regulation of N cycling in terrestrial ecosystems.
-
Soil samples were collected from the topsoil layer (0–20 cm) of a peach orchard in Yuncheng City, Shanxi Province, China. The sampling site was an unfertilized agricultural field. To obtain representative samples, a composite sampling strategy was employed by collecting soils from multiple locations within the field. The collected soils were thoroughly mixed in situ, and subsamples were obtained using the quartering method. Samples were transported to the laboratory of Sun Yat-sen University under cold-chain conditions, air-dried at room temperature in the shade, gently ground, and passed through a 2 mm sieve for homogenization. A portion of the processed soil was stored at 4 °C for incubation experiments, while the remaining samples were kept at −20 °C for physicochemical analyses. The basic physicochemical properties of the soil are as follows: total carbon (TC) 10.9 g kg−1, total nitrogen (TN) 1.08 g kg−1, NH4+–N 23.20 mg kg−1, NO2−–N 0.46 mg kg−1, NO3−–N 21.43 mg kg−1, and pH 8.0.
Experimental design
-
Prior to the experiment, soils were adjusted to 30% of water-holding capacity (WHC) with double-distilled water (ddH2O) and pre-incubated at 25 °C in the dark for 7 d to restore microbial activity. Soil moisture was maintained by periodic rewetting during pre-incubation. Afterward, 50 g of fresh soil was placed into 100 mL glass bottles covered with perforated aluminum foil to ensure aerobic conditions, and the incubation temperature was kept at 25 °C for all treatments. Soil moisture was adjusted to four levels of WHC: 40% (W1), 60% (W2), 90% (W3), and 120% (W4). Simultaneously, ammonium chloride (NH4Cl) solutions were added to achieve three NH4+–N levels: 50 mg N kg−1 (A1), 100 mg N kg−1 (A2), and 200 mg N kg−1 (A3). The volume of NH4Cl solution required for each treatment was calculated according to the target N concentration and moisture level, and evenly applied to the soil surface using a micropipette. Each treatment was replicated three times, and soil samples were collected on days 0, 1, 3, 4, 5, 6, 7, and 9 of incubation for subsequent analyses.
Chemical analyses
-
The TC and TN contents of the soils were determined using an elemental analyzer (FlashSmart 11206125, Thermo Scientific, USA). Ammonium and NO3− were simultaneously extracted with 0.01 M CaCl2 at a soil-to-solution ratio of 1:10 (w/v). The soil slurries were shaken horizontally at 200 rpm for 1 h, followed by centrifugation at 690 × g for 15 min, and subsequently filtered through 0.45 μm polypropylene membrane syringe filters (25 mm, VWR Europe). Nitrite was extracted separately using ddH2O at a 1:5 (w/v) ratio, following the same shaking and centrifugation procedure. Nitrite concentrations were determined colorimetrically using the Griess reagent[19]. Nitrate concentrations were quantified indirectly via the vanadium (III) reduction method[20], in which NO3− is reduced to NO2−, allowing the determination of total NO2− + NO3−; the NO3− concentration was then obtained by subtracting the measured NO2− concentration. Ammonium concentrations were determined using the indophenol blue colorimetric method[21]. Soil pH was measured in a 1:2.5 soil-to-water suspension using a pH meter (FE28-Standard, Mettler-Toledo, Switzerland).
DNA extraction and quantitative PCR (qPCR) of functional genes
-
Fresh soil samples (0.5 g) were incubated for 3 d before DNA extraction. DNA was extracted using the DNeasy PowerSoil Kit (Tiangen Biotech, Beijing, China), with a blank control included to monitor potential contamination. DNA quality and concentration were assessed using a NanoDrop spectrophotometer (ND-1000, Thermo Scientific, USA). The abundances of ammonia- and nitrite-oxidizing bacteria were quantified by targeting the amoA and nxrB genes, respectively, while those of denitrifiers were determined using the nirS and nirK genes. Quantitative real-time PCR (qPCR) was performed in triplicate on an ABI Prism 7900 system (Applied Biosystems, USA). Each 20 μL qPCR reaction contained 10 μL of 2× RealStar Green Fast Mixture (GenStar, Cat. No. A301), 0.5 μL each of forward and reverse primers (10 μM), 2 μL of DNA template, and 7 μL of nuclease-free water. The thermal cycling conditions consisted of an initial denaturation at 95 °C for 2 min, followed by 40 cycles of denaturation at 95 °C for 15 s, annealing at 60 °C for 20 s, and extension at 72 °C for 30 s. Standard curves were generated from serial dilutions of plasmid DNA containing known copy numbers of target genes (amoA, nxrB, nirS, and nirK), and gene copy numbers were calculated by comparing the sample Ct values with those of the calibration curves[22]. All primers used in this study are listed in Table 1.
Table 1. Primers used for quantitative real-time PCR (qPCR) analysis
Gene Primer Sequence (5'-3') Amplicon size [bp] Ref. Archaeal amoA Amo19F STAATGGTCTGGCTTAGACG 635 Leininger et al. [45]; Schauss et al.[46] CrenamoA6 GCGGCCATCTGTATGT Bacterial amoA amo41F GGGGTTTCTACTGGTGGT 491 Rotthauwe et al.[47] amoA2R CCCCTCGGGAAGGCTTTTC Nitrobacter-like nxrB nxrB-1F ACGTGGAGACCAAGCCGGG 411 Vanparys et al.[48] nxrB-1R CCGTGCTGTTGAYCTGTTGA Nitrospira-like nxrB nxrB169f TACATGGTGTGGAACA 485 Pester et al.[49] nxrB638r CGGTTCTTGGTCRATCA Comammox amoA ComamoAF AGGNGAYTGGGAYYTTCGG 436 Wang et al.[50] ComamoAR CGGACAWABRTGAABCCCAT nirK copper583F TCATGGTGCTGCGCGKGACGG 326 Yan et al.[51] copper909R GAACTTGCCGGTKGCCCAGAC nirS heme832F TCACACCCCGAGCCGCGCGT 774 Yan et al.[51] heme1606R AGKCGTTGAACTTKCCGGTCGG Soil RNA extraction and reverse transcription
-
Total RNA was extracted from soil samples incubated for 3 d using the RNeasy PowerSoil Total RNA Kit (Qiagen, Hilden, Germany). RNA concentration and purity were immediately assessed using a NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific, USA). The amount of RNA used for reverse transcription was calculated based on its final concentration[23]. Complementary DNA (cDNA) was synthesized with the HiScript III 1st Strand cDNA Synthesis Kit (+gDNA wiper) (Vazyme Biotech, Nanjing, China) according to the manufacturer's instructions. Each 20 μL reaction contained 1 μg of total RNA and 10× random hexamer/oligo(dT) primers. The thermal program consisted of incubation at 25 °C for 10 min, reverse transcription at 45 °C for 1 h, and enzyme inactivation at 94 °C for 5 min, followed by holding at 4 °C. Negative controls lacking either reverse transcriptase or RNA template were included to confirm the absence of DNA or reagent contamination. The synthesized cDNA was stored at −80 °C until further analysis.
Statistical analysis
-
The effects of soil moisture content on the abundances of the AOA-amoA, AOB-amoA, Nitrobacter-nxrB, Nitrospira-nxrB, nirK, and nirS genes, as well as AOR and NOR activities, were assessed using one-way analysis of variance (ANOVA). Prior to ANOVA analysis, the normality and homogeneity of variances were evaluated using Levene's test. Differences among groups were determined by Duncan's post hoc test, with statistical significance set at p < 0.05. Pearson correlation analysis was performed to explore the relationships among net nitrification rate, copy numbers of AOA-amoA, AOB-amoA, Nitrobacter-nxrB, and Nitrospira-nxrB genes, and soil NO2−–N concentrations. All statistical analyses were performed in R programming (version 4.1.3) using the 'stats' and 'ggpubr' packages, and GraphPad Prism (version 10.1.2), and visualizations were created with the assistance of Adobe Illustrator 2018.
-
Ammonia acts as the substrate for NH3 oxidation, producing NO2− as an intermediate. This NO2− then serves as the substrate for the subsequent NO2− oxidation process, which produces NO3−. When the NH4+ concentration was 50 or 100 mg N kg−1, there were no significant differences in the NH4+ decrease rates among the different soil moisture treatments (Fig. 1a, b). However, at a concentration of 200 mg N kg−1, the NH4+ decrease rates in the W2 and W3 treatments were significantly higher than that in W1 (Fig. 1c). In addition, under the same N application level, the NH4+ decrease rate was the slowest in the W4 treatment (Fig. 1a–c). In the W2 and W3 treatments, a pronounced accumulation of NO2− was observed on the third day of incubation (Fig. 1e, f), with the highest NO2− concentration detected in soils receiving 200 mg N kg−1. Significant differences (p < 0.05, n = 3) in NO2− concentration were found among different moisture treatments. At the same N application level, NO3− concentrations did not differ significantly among the soil moisture treatments, except for W4 (Fig. 1g–i).
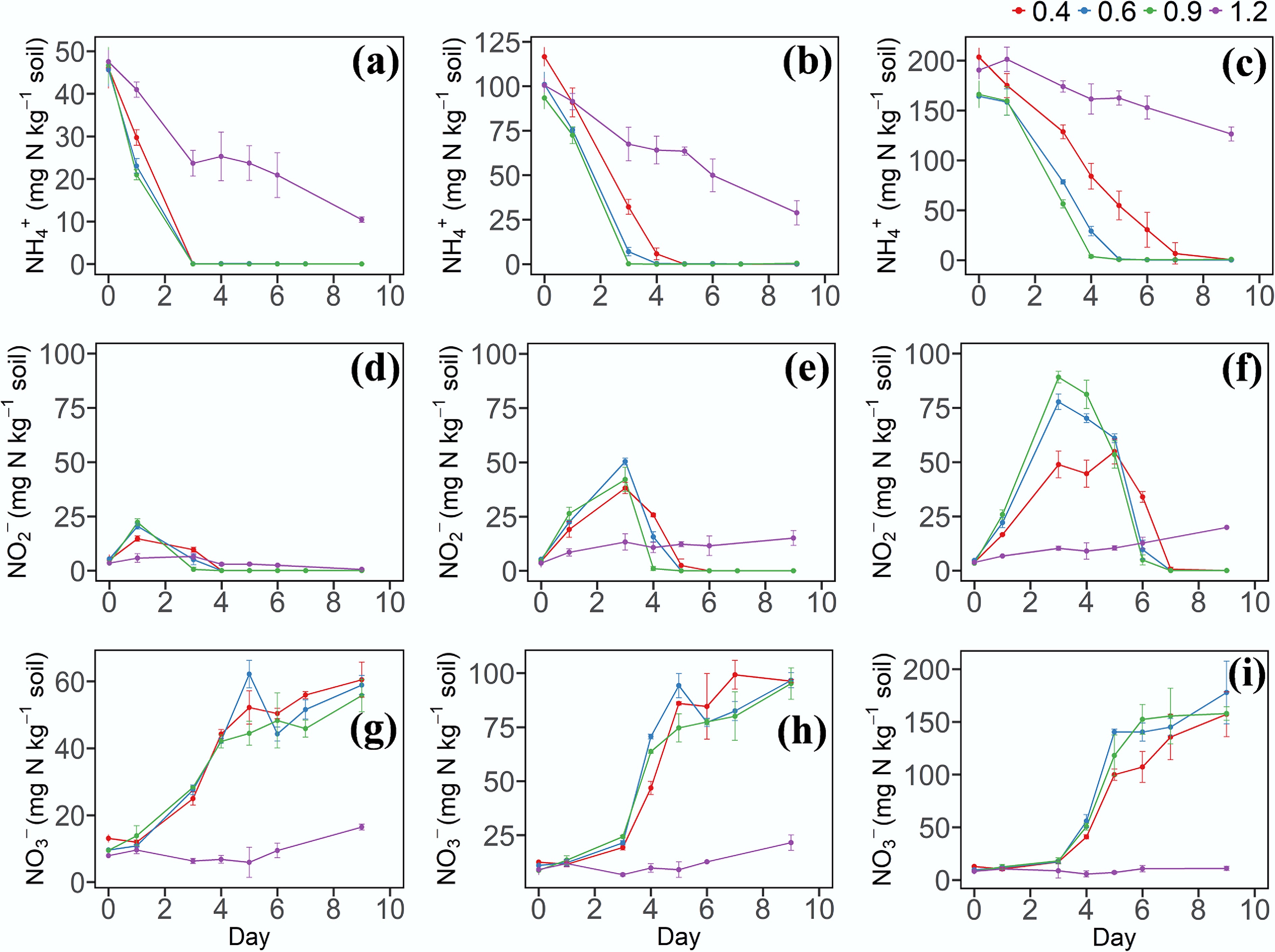
Figure 1.
(a)–(c) Dynamic changes of NH4+–N, (d)–(f) NO2−–N, and (g)–(i) NO3−–N in soil under different soil water contents and N levels. 0.4, 0.6, 0.9, and 1.2 represent W1, W2, W3, and W4 WHC, respectively. (a), (d), and (g) represent soil with a N content of 50 mg N kg−1. (b), (e), and (h) represent soil with a N content of 100 mg N kg−1. (c), (f), and (i) represent soil with a N content of 200 mg N kg−1. Error bars indicate standard deviations (n = 3). Abbreviations: NH4+: ammonium, NO2−: nitrite, NO3−: nitrate, WHC: water holding capacity.
Variation of AOM and NOB abundance affected by N dose and WHC
-
The gene abundances of AOA-amoA, AOB-amoA, Nitrobacter-nxrB, Nitrospira-nxrB, nirK, and nirS were quantified using quantitative PCR (qPCR) on the third day of incubation (Fig. 2). Although AOA-amoA abundance increased at W1 and W4 WHC, its response to N addition lacked a consistent trend across treatments (Fig. 2a). The gene abundance of AOB-amoA was regulated by both N dose and soil moisture, showing a positive synergistic effect. Specifically, higher soil moisture (W3/W4) promoted gene abundance to a greater extent under high N addition levels than under low levels, highlighting their significant interaction (Fig. 2b). Moreover, the abundance of comammox-amoA shared a similar trend with that of AOA-amoA, and it was low across all soil samples (Fig. 2c).
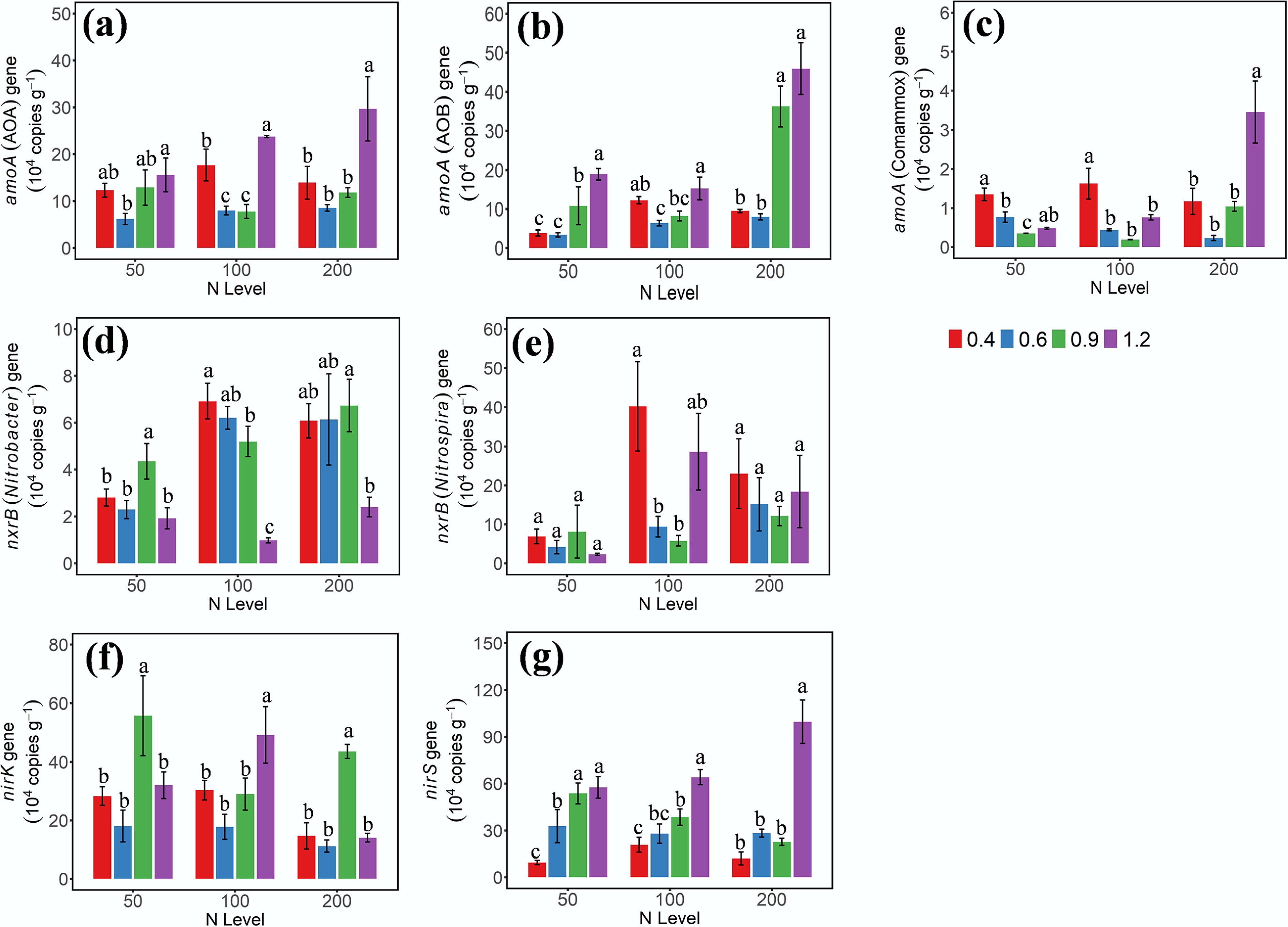
Figure 2.
Gene abundances of (a) AOA-amoA, (b) AOB-amoA, (c) amoA-comammox, (d) nxrB-Nitrobacter, (e) nxrB-Nitrospira, (f) nirK, and (g) nirS on day 3 under different soil moisture and N treatments. 0.4, 0.6, 0.9, and 1.2 represent W1, W2, W3, and W4 WHC, respectively. Error bars indicate standard deviations (n = 3). Different lowercase letters indicate significant differences (p < 0.05) between moisture content treatments within the same N application level.
Based on DNA abundance, Nitrospira was the predominant NOB across all soil samples (Fig. 2e). At the 50 mg NH4+–N addition level, gene abundance of Nitrospira-nxrB showed no significant differences among soils with varying moisture contents (p < 0.05). At the 100 mg NH4+–N level, the gene abundance of Nitrospira-nxrB was higher in soils at W1 and W4 WHC. The gene abundance of Nitrobacter remained consistently low across all treatments, with the lowest values observed under saturated conditions (W4) at the same N level (Fig. 2d). The gene abundance of the nirS increased with increasing soil moisture across all N treatments, whereas the gene abundance of nirK did not show a clear trend, indicating a greater sensitivity of nirS-containing denitrifier populations to soil moisture variation (Fig. 2f, g).
Active nitrifying microorganism abundance affected by N dose and WHC
-
The expression levels of functional genes involved in NH3 and NO2− oxidation were assessed using cDNA-based quantification (Fig. 3). The cDNA abundance patterns of AOB-amoA and AOA-amoA differed from their respective DNA-level profiles. Specifically, at the 50 mg NH4+–N addition level, AOB-amoA cDNA abundance was significantly higher under W4 WHC compared to the other moisture treatments (p < 0.05), while no significant differences were observed under the other two N levels. For AOA-amoA, higher cDNA abundance was observed only under W2 and W4 at the high N addition level. Notably, the AOB-amoA cDNA abundance was markedly higher than that of AOA-amoA under W4 conditions. Compared to the DNA-based results, the cDNA/DNA ratio of Nitrobacter-nxrB was higher than that of Nitrospira-nxrB, indicating that although Nitrospira exhibited higher relative abundance at the DNA level, only a small fraction of the population was transcriptionally active.
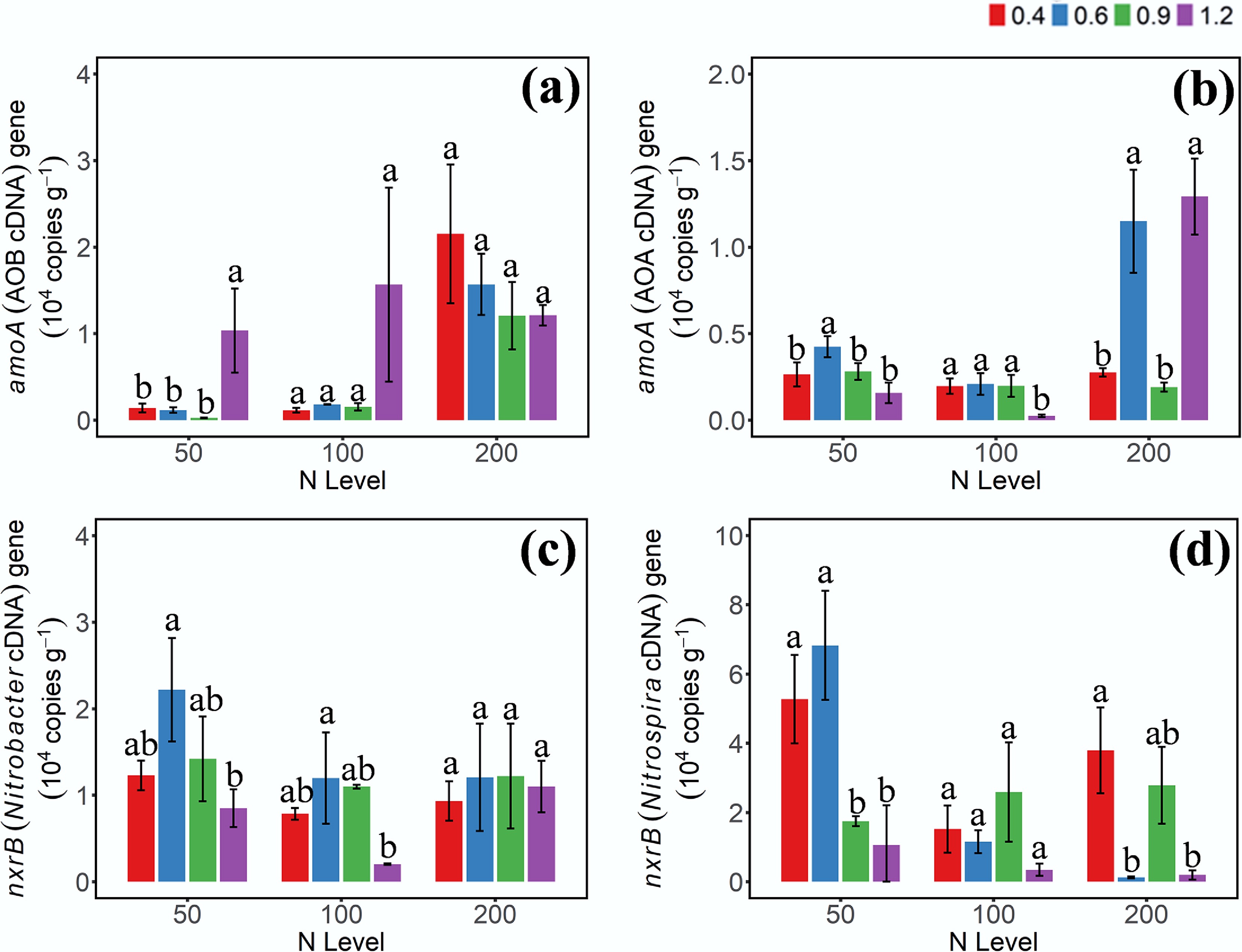
Figure 3.
Gene abundances of (a) AOB-amoA, (b) AOA-amoA, (c) Nitrobacter-nxrB, and (d) Nitrospira-nxrB cDNA on day 3 under different moisture and N addition treatments. 0.4, 0.6, 0.9, 1.2 represent W1, W2, W3, and W4 WHC, respectively. Error bars indicate standard deviations (n = 3). Different lowercase letters indicate significant differences (p < 0.05) between moisture content treatments within the same N application level.
Correlations among nitrite concentration and nitrification processes
-
Soil moisture significantly affected the coupling between NH3 and NO2− oxidation (Fig. 4). NAOR (net NH3 oxidation rate from day 0 to 3) was positively correlated with the AOB : AOA gene abundance ratio (R2 = 0.54, p < 0.05), reflecting differences in the contributions of bacterial and archaeal NH3 oxidizers. NNOR1 (net NO2− oxidation rate from day 0 to 3) also showed a positive correlation with the Nitrobacter : Nitrospira ratio (R2 = 0.24, p < 0.05), suggesting shifts in nitrite-oxidizing community structure. Moreover, NO2− concentration was positively correlated with both the NAOR : NNOR1 ratio (R2 = 0.40, p < 0.05) and the AOB : Nitrobacter gene abundance ratio (R2 = 0.56, p < 0.05), indicating that imbalances between functional groups may contribute to NO2− accumulation.
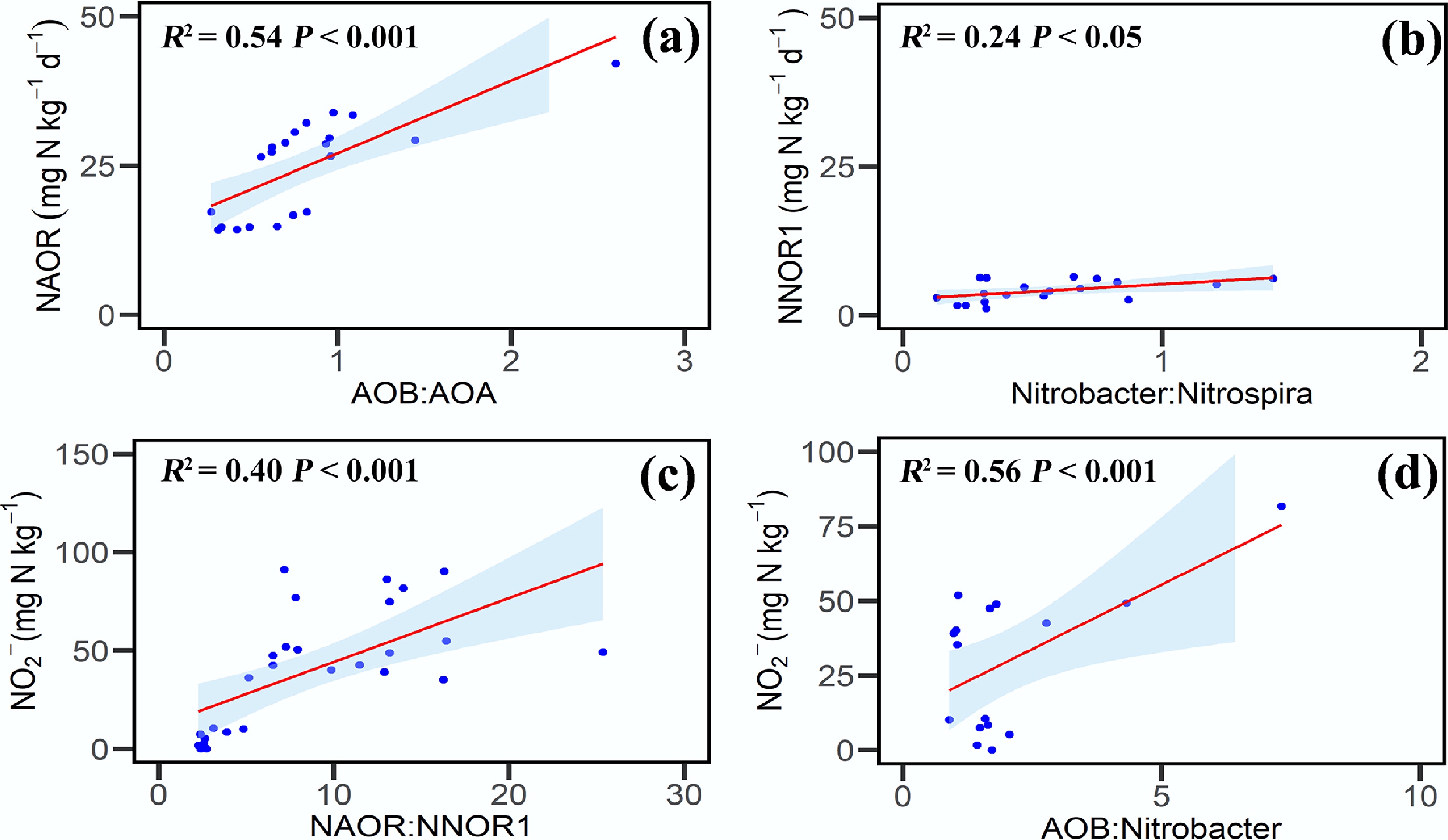
Figure 4.
Relationships between net nitrification rate, nitrite concentration, and gene abundance. (a) NAOR and the AOB : AOA gene abundance ratio. (b) NNOR1 and the Nitrobacter : Nitrospira ratio. (c) NO2− concentration and the NAOR:NNOR1 ratio. (d) NO2− concentration and the AOB : Nitrobacter ratio. Symbols represent individual soil samples; lines denote significant correlations; and asterisks indicate statistically significant correlations (p < 0.05). Abbreviations: NAOR is the net ammonia oxidation rate from day 0 to 3; NNOR1 is the NO2− oxidation rate from day 0 to 3.
-
Net NH3 oxidation rates did not differ significantly among the W1, W2, and W3 WHC treatments in soils amended with 50 mg of N, indicating that under relatively low N input, soil moisture had little effect on NH3 oxidation. However, when the N input was increased to 100 mg NH4+–N, net NH3 oxidation rates under W2 and W3 WHC were significantly higher than those under W1 WHC. This shift suggests that under high NH4+ concentrations, NH4+ is no longer the limiting factor for NH3 oxidation, and soil moisture becomes the dominant factor regulating the oxidation rate. This finding is consistent with previous studies; for instance, Cheng et al.[24] observed that the total nitrification rate was significantly higher at 90% WHC compared to 70% WHC in forest soils. In fact, numerous studies have reported that mineralization and nitrification rates tend to increase with rising soil moisture across arable land, grasslands, and forest soils[25,26]. This enhancement may be attributed to two complementary mechanisms. First, increasing soil moisture can alleviate water limitation and improve the mobility of dissolved substrates such as NH4+, thereby increasing substrate availability for ammonia oxidizers. Second, moderate soil moisture may still maintain sufficient O2 supply for nitrification. However, this effect should be interpreted cautiously, because further increases in soil moisture generally reduce soil aeration and restrict O2 diffusion, which can ultimately suppress nitrification[27]. In contrast to this trend, net NH3 oxidation rates were consistently lowest under W4 WHC across all N addition treatments. The decline in nitrification under saturated conditions was most likely associated with O2 limitation caused by reduced soil aeration. These observations are consistent with the findings of Wang et al.[27], who reported that soil NO3−–N concentrations accumulated more rapidly when WFPS was ≤ 80% than when WFPS ≥ 90%.
The effect of soil moisture on net NH3 oxidation varied with the level of NH4+ addition. At low NH4+ concentrations, changes in moisture had little impact on net NH3 oxidation. However, as NH4+ availability increased, the effect of moisture became more pronounced, showing a significant positive correlation at higher NH4+ levels. Notably, excessive moisture (120% WHC) consistently suppressed net NH3 oxidation across all NH4+ treatments.
Consistent with the net NH3 oxidation rates, no significant differences were observed in soil NO2− concentrations among the 40%, 60%, and 90% WHC treatments under low NH4+ input (50 and 100 mg N kg−1). However, when NH4+ was elevated to 200 mg N kg−1, soils at 60% and 90% WHC exhibited significantly higher NO2− accumulation compared to the 40% WHC treatment, whereas at 120% WHC, NO2− levels remained comparatively low for all NH4+ levels. Further, soil NO3− concentrations remained relatively low under the highest soil moisture treatment (W4) for all NH4+ levels.
This interpretation is consistent with previous studies. For example, a modeling and incubation study demonstrated that as soil saturation increases beyond an optimal threshold, nitrification rates drop sharply while denitrification becomes the dominant pathway, with the greatest N loss observed under near-saturated conditions[28]. Similarly, a global synthesis found that denitrification rates are strongly positively correlated with increases in soil water-filled pore space and soil NO3− content, and negatively correlated with soil O2 availability—supporting the view that in high-moisture soils, O2 limitation shifts the balance from nitrification toward denitrification (and gaseous N losses)[29]. This pattern suggests that under saturated or near-saturated moisture conditions, nitrification is inhibited—likely due to limited O2 diffusion—and concurrently, enhanced denitrification or other anaerobic N-transformation processes may consume NO2−/NO3−, preventing their accumulation.
Responses of AOB and AOA to soil moisture content
-
AOB and AOA play critical roles in soil nitrification processes; however, their responses to soil moisture content exhibit distinct patterns. In this study, under the 50 mg NH4+–N treatment, AOB-amoA cDNA abundance was significantly higher at the W4 WHC than at other moisture conditions (p < 0.05), whereas no significant differences were observed under the other two NH4+ levels. This response may be closely related to the residual NH4+ concentrations in the soil. In contrast, AOA-amoA cDNA abundance declined under higher moisture levels, except in soils treated with 200 mg NH4+–N. Interestingly, although AOA-amoA DNA abundance was relatively high at both W1 and W4 WHC, the corresponding AOA-amoA cDNA abundance remained low, suggesting that transcriptional activity did not parallel population size. This observation does not necessarily contradict previous studies reporting a competitive advantage of AOA under high-moisture or low-O2 conditions, but instead suggests that the response of AOA activity to moisture may be nonlinear and strongly dependent on NH4+ availability and soil-specific environmental conditions[30,31].
Ample research suggests that AOB have a competitive advantage under high NH4+ conditions due to their efficient ammonia-oxidizing capacity. This finding aligns with our observations, as the elevated NH4+ concentration at W4 WHC likely created favorable conditions for AOB growth and activity[32]. In our experimental system, substrate availability, alongside O2 availability, played a key role in driving the nitrification process. This was evidenced by significant differences in NAOR and NO2− accumulation in soils amended with 200 mg NH4+–N (p < 0.05), whereas such differences were not observed under the 50 and 100 mg NH4+–N treatments. These findings suggest that sufficient NH4+ supply is an important condition for enhanced AOB-driven nitrification, particularly under high moisture levels.
Differential responses of Nitrobacter and Nitrospira to soil moisture conditions
-
Nitrospira and Nitrobacter represent the two major lineages of NOB, yet their ecological niches and physiological strategies differ markedly, resulting in contrasting responses to N input and soil moisture conditions. In our study, Nitrospira-nxrB gene abundance was substantially higher than Nitrobacter-nxrB across all treatments, and increased consistently with rising NH4+ levels. Moreover, Nitrospira abundance peaked under both low (40% WHC) and excessively high moisture (120% WHC), suggesting that this lineage possesses a broader tolerance to substrate and O2 fluctuations. These findings align with previous reports describing Nitrospira as an oligotrophic, K-selected NOB with high affinity for NO2− and superior ability to persist under nutrient limitation or microaerobic conditions[33,34].
In contrast, Nitrobacter-nxrB abundance increased from 50 to 100–200 mg N kg−1 NH4+ addition but did not rise further at higher N levels. Moreover, unlike Nitrospira, Nitrobacter abundance decreased sharply under 120% WHC, especially at high NH4+ inputs. This pattern is consistent with its classification as a fast-growing, r-strategist NOB that prefers well-aerated, nitrite-rich environments[35]. The suppression of Nitrobacter under saturated soil likely reflects O2 limitation, because this lineage relies strictly on O2 as the terminal electron acceptor and exhibits poor tolerance to anoxia or reduced redox potential[36]. In addition, micro-scale acidification under high water saturation, which is known to inhibit Nitrobacter more strongly than Nitrospira, may further constrain its growth and activity[37].
The transcriptional responses revealed even more pronounced lineage-specific differences. Nitrospira-nxrB cDNA abundance was highest under low N addition and under 40%–60% WHC, indicating that although Nitrospira is numerically dominant at the DNA level, its active fraction declines when substrate concentrations become excessive. This supports the view that many Nitrospira cells in soils may remain in a dormant or low-activity state, activating only under conditions of moderate NO2− supply that match their oligotrophic lifestyle. In contrast, Nitrobacter-nxrB cDNA showed limited sensitivity to N or moisture treatments, except for a notable decline under 100 mg N kg−1 at 120% WHC, reinforcing its vulnerability to O2 constraints and redox instability. The generally higher cDNA/DNA ratios observed in Nitrobacter than in Nitrospira suggest that, although less abundant, Nitrobacter maintains a larger proportion of metabolically active cells, consistent with its copiotroph strategy and rapid-response physiology[38].
These findings highlight that soil moisture and N availability regulate NO2− oxidation not only through their effects on O2 diffusion and substrate supply, but also by shifting the competitive balance between K-selected Nitrospira and r-selected Nitrobacter. Moderate soil moisture promotes the activity of Nitrobacter, whereas Nitrospira gains a competitive advantage under both drier and more saturated conditions, reflecting its greater ecological plasticity and ability to tolerate fluctuating microaerobic environments. These lineage-specific responses ultimately shape NO2− oxidation dynamics and may influence nitrification bottlenecks under variable hydrological regimes.
Mechanistic drivers of soil nitrite accumulation
-
The strong positive correlations between peak NO2− concentrations and both the net NH3 oxidation rate (NAOR) and the net NO2− oxidation rate (NNOR) highlight the central role of process imbalances in regulating NO2− accumulation. Such imbalances arise when the capacity of NO2− oxidizers cannot match the production of NO2− from NH3 oxidation, a phenomenon widely reported as a key driver of NO2− accumulation in soils[39]. Consistent with this mechanism, NO2− accumulation in our soils was positively associated with the relative abundance ratio of AOB to Nitrobacter, suggesting that a dominance of fast-growing AOB over Nitrobacter can lead to transient NO2− buildup under conditions where NO2− oxidation becomes rate-limiting. Previous work has also shown that NO2− oxidation is particularly sensitive to environmental stressors such as high NH4+ levels and reduced O2 availability, both of which can disproportionately suppress Nitrobacter activity and thereby promote NO2− accumulation[40,41].
The observation that NAOR was strongly correlated with the AOB : AOA amoA ratio further confirms that AOB were the principal drivers of NH3 oxidation in this system, aligning with the broader consensus that AOB generally dominate under elevated NH4+ concentrations and in moderately moist soils[31]. Likewise, NNOR was positively correlated with the Nitrobacter : Nitrospira ratio, indicating that Nitrobacter played a leading role in NO2− oxidation despite the well-known oligotrophic lifestyle and high substrate affinity of Nitrospira[42]. This pattern may reflect the higher growth rate and substrate tolerance of Nitrobacter, enabling it to respond more rapidly to increasing NO2− levels under moderate moisture conditions.
However, under high soil moisture (120% WHC), the regulatory framework governing NO2− accumulation shifted markedly. In these nearly saturated soils, NO2− and NO3− concentrations remained low despite active NH3 oxidation, indicating that denitrification masked nitrification signals by consuming NO2− and NO3− as electron acceptors. Hence, the pathway responsible for NO2− dynamics appeared to vary with soil moisture. At 40%–90% WHC, NO2− accumulation was more consistent with nitrification-derived buildup caused by incomplete coupling between NH3 oxidation and NO2− oxidation. At 120% WHC, however, enhanced denitrification likely altered or consumed NO2− intermediates, thereby suppressing their net accumulation. The sharp increases in nirS gene abundance with rising moisture content, together with nitrogen-dependent responses of nirK communities, support the activation of oxygen-sensitive denitrifiers under high moisture. This observation is consistent with the established ecological differentiation of these functional groups: nirS-type denitrifiers typically dominate under low-oxygen or intermittently anoxic conditions, whereas nirK-bearing microorganisms include taxa associated with both denitrification and nitrification pathways[43]. The enhanced nirS abundance under saturated conditions, therefore, aligns with classical models of moisture-driven denitrification[44], explaining the observed suppression of inorganic N pools. The pathway responsible for NO2− dynamics appeared to vary with soil moisture. At 40%–90% WHC, NO2− accumulation was more consistent with nitrification-derived buildup caused by incomplete coupling between NH3 oxidation and NO2− oxidation. At 120% WHC, however, enhanced denitrification likely altered or consumed NO2− intermediates, thereby suppressing their net accumulation.
Environmental implications
-
This study provides new mechanistic insight into how soil moisture and NH4+ availability jointly regulate the coupling between NH3 oxidation and NO2− oxidation, thereby shaping the fate of reactive N in terrestrial ecosystems. By integrating process-based measurements with functional gene and transcript analyses, our results demonstrate that the balance between AOB- and AOA-driven NH3 oxidation and the differential contributions of Nitrobacter and Nitrospira to NO2− oxidation are key determinants of whether NO2− accumulates or is efficiently oxidized to NO3−. The finding that NO2− accumulation is strongly associated with microbial community composition—particularly high AOB : Nitrobacter ratios—highlights the importance of microbial ecological interactions in governing transient N intermediates that influence nitrification efficiency and downstream N losses.
Limitations
-
Despite providing important insights into the moisture-N regulation of nitrification pathways, this study has several limitations that should be acknowledged. First, the study did not employ selective inhibitors such as acetylene, 1-octyne, or chlorate to differentiate the contributions of AOA, AOB, and specific NOB groups. As a result, causal links between gene abundance/expression and process rates remain inferential rather than experimentally verified. In addition, the incubation experiment was conducted under controlled laboratory conditions, which inevitably simplifies the physical and biological complexity of natural soils. Factors such as heterogeneous pore structure, fluctuating redox conditions, plant-microbe interactions, and microscale O2 dynamics were not fully represented, potentially limiting the extrapolation of our results to field environments. Moreover, denitrification was inferred from nirS/nirK abundances and N-balance patterns; direct measurements of denitrification rates, N2O/N2 production, or NO3− reduction were not conducted. Future studies combining selective inhibition, isotope tracing, and direct gaseous N flux measurements will be necessary to quantitatively separate the contributions of nitrification and denitrification pathways under varying moisture and N regimes.
-
This study demonstrates that soil moisture and N availability jointly regulate the coupling efficiency between NH3 oxidation and NO2− oxidation by reshaping nitrifier community structure, functional gene expression, and microbial activity. Increasing N inputs and moderate moisture levels (60%–90% WHC) enhanced AOB-dominated NH3 oxidation, often exceeding the nitrite-oxidizing capacity of Nitrobacter, thereby promoting NO2− accumulation. The strong positive correlations between NO2− accumulation and both the NAOR : NNOR ratio and the AOB : Nitrobacter abundance ratio further confirm that imbalances between NH3 oxidation and NO2− oxidation are a primary driver of NO2− buildup. Under waterlogged conditions (120% WHC), the apparent accumulation of NO2− and NO3− was substantially reduced despite active NH3 oxidation. This pattern reflects the increased importance of denitrification under oxygen-limiting, high-moisture conditions, as indicated by the sharp rise in nirS abundance. These findings highlight that at high soil moisture, NO2− consumption through denitrification masks the accumulation that would otherwise result from decoupled NH3 and NO2− oxidation, demonstrating a shift from a nitrification-dominated system toward a more tightly integrated nitrification-denitrification regime. However, this study did not employ selective nitrification inhibitors (e.g., acetylene, 1-octyne, chlorate), limiting our ability to definitively attribute measured NH3 and NO2− oxidation rates to specific functional groups. Further work should incorporate inhibition assays and isotope tracing approaches to directly quantify the contributions and interactions of AOA, AOB, NOB, and denitrifiers under dynamic moisture-nitrogen regimes.
-
The authors confirm their contributions to the paper as follows: All authors contributed to the study conception and design. Zhijie Li: writing – original draft. Jiani Ma: writing – original draft. Miao Chen: writing – original draft. Weiqi Kuang: conceptualization. Gaochao Cai: material preparation, data collection, and formal analysis. Yubin Wang: material preparation, data collection, and formal analysis. Yunyun Cao: material preparation, data collection, and formal analysis. Shurong Liu: funding acquisition and writing – review and editing. All authors reviewed the results and approved the final version of the manuscript.
-
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
-
This research was supported by the Provincial Natural Science Fund of Guangdong (Grant No. 2022A1515010786) and the National Natural Science Foundation of China (Grant No. 42407418).
-
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
-
# Authors contributed equally: Zhijie Li, Jiani Ma
Full list of author information is available at the end of the article. - Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Li Z, Ma J, Chen M, Kuang W, Cai G, et al. 2026. Soil moisture-driven changes in the balance of ammonia and nitrite oxidation. Environmental and Biogeochemical Processes 2: e010 doi: 10.48130/ebp-0026-0005
Soil moisture-driven changes in the balance of ammonia and nitrite oxidation
- Received: 16 October 2025
- Revised: 31 March 2026
- Accepted: 16 April 2026
- Published online: 29 April 2026
Abstract: Soil moisture profoundly regulates nitrification by influencing oxygen (O2) diffusion, substrate mobility, and microbial activity. However, the mechanisms underlying moisture-driven shifts in the coupling between ammonia (NH3) and nitrite (NO2−) oxidation remain poorly understood. In this study, a laboratory incubation experiment was conducted using agricultural soils under four water-holding capacities (WHC: 40%, 60%, 90%, and 120%) and three ammonium (NH4+) levels (50, 100, and 200 mg NH4+–N kg−1). Under low nitrogen (N) input, soil moisture exerted minimal influence on NH3 oxidation. At higher NH4+ concentrations, however, oxidation increased substantially at moderate moisture (60%–90% WHC) and dropped under waterlogged conditions (120% WHC) due to O2 limitation. Quantitative PCR and cDNA analyses indicated clear divergences in the responses of ammonia-oxidizing archaea (AOA), ammonia-oxidizing bacteria (AOB), and nitrite-oxidizing bacteria (NOB) to soil moisture. AOB activity was highest under conditions of high N availability and moderate soil moisture. In contrast, Nitrobacter activity declined sharply when soils became saturated, whereas Nitrospira showed a much broader tolerance to moisture extremes. Correlation analyses revealed that NO2− accumulation was driven by an imbalance between NH3 and NO2− oxidation: a situation particularly pronounced under high N and high moisture conditions, where rapid NH3 oxidation by AOB outpaced the oxidation capacity of Nitrobacter. Together, the results highlight that soil moisture and N availability co-determine the coupling of NH3 and NO2− oxidation by driving microbial community changes and functional gene responses, yielding new insights into moisture-driven regulation of soil N cycling.