-
The issue of food safety has become a matter of global concern, in which foodborne illnesses represent a significant threat to public health. Thus, the development of highly sensitive detection technologies is crucial to address various biological and chemical contaminants. Taking mycotoxin detection as an example, some researchers have advanced a variety of high-precision methods[1,2]. The breakthroughs in sensitivity and specificity offered by these technological platforms provide direct methodological insights and technical support for the efficient screening of pathogenic bacteria. Although the majority of Escherichia coli (E. coli) strains are non-pathogenic, certain pathogenic E. coli strains have the capacity to induce severe illness. E. coli is classified as a Gram-negative, facultative anaerobic, rod-shaped bacterium. It is widely distributed in the intestines of both humans and animals[3,4]. It is a known pathogen connected to foodborne diseases and serves as a causative agent for various human illnesses, including hemolytic uremic syndrome (HUS), sepsis, meningitis, urinary tract infections (UTIs), wound infections, immune system disorders, and hepatic abscesses[5]. According to the World Health Organization (WHO), approximately 63,000 deaths are caused by pathogenic E. coli annually, and over 5,000,000 disability-adjusted life in five years are recorded[6]. Therefore, timely detection and precise quantification of E. coli are imperative for maintaining food safety. Conventional microbiological detection methodologies, including plate culture techniques, are characterized by their protracted nature and the requirement for substantial labor input. Moreover, these techniques possess a limited capacity to analyze intricate microbial compositions and quantities[7]. In contrast, contemporary techniques such as PCR and immunoassays have achieved substantial progress in terms of detection sensitivity and speed. Nevertheless, their widespread deployment remains impeded by factors such as high costs and operational complexity[8,9]. Recent studies on photoelectrochemical biosensors and nanomaterial-assisted label detection strategies provide valuable technological insights for the rapid detection of pathogens such as E. coli[10,11]. Among these techniques, colorimetric sensors that induce visible color changes in substrates through catalytic reactions have emerged as a promising approach[12,13]. In recent years, the design of novel materials such as nanozymes and the development of biorecognition probes has further enhanced the performance and applicability of such sensors, providing a viable solution for the on-site detection of E. coli[14−16].
Various recognition probes, including antibodies[17], nucleotides[18], and bacteriophages[19], are currently used to detect pathogenic bacteria. However, these probes still generally face issues, including poor stability, complex modification processes, and proneness to inactivation. Aptamers are oligonucleotides (DNA or RNA) that are single-stranded and have high specificity and affinity[20]. They are primarily obtained through a process called Systematic Evolution of Ligands by Exponential Enrichment (SELEX)[21,22]. Aptamers have been demonstrated to form specific three-dimensional (3D) structures. Such structures are complex in nature and are characterized by bulges, loops, hairpins, stems, pseudoknots, triplexes, or quadruplexes. These structures enable their targeting through a variety of mechanisms, including structural compatibility, aromatic ring stacking, hydrogen bonding, electrostatic and van der Waals interactions, or a combination of these effects[23]. Aptamers can be compatible with electrochemical and optical sensing platforms, thus enabling the rapid and portable detection of food hazards[24]. Aptamers thus offer significant advantages over antibodies, including their smaller molecular weight, ease of modification and chemical synthesis, and a lower tendency to elicit immune responses[25].
In the case of food safety detection, aptamers are frequently combined with nano-functional materials, such as gold nanoparticles (Au NPs)[26], quantum dots (QDs)[27], and carbon nanotubes (CNTs)[28], which have been widely used to construct biosensing platforms. However, certain materials continue to encounter difficulties, including restricted catalytic activity, intricate preparation procedures, or poor biocompatibility, which impede their practical utilization. Therefore, the development of novel nanomaterials with high catalytic activity, excellent stability, and ease of functionalization has become a key focus for further improving the performance of aptamer-based sensors. Nanozymes, as a new class of inorganic materials, not only possess the physical and chemical properties of nanomaterials, but also exhibit enzyme-like catalytic functions, making them ideal alternative catalysts in biosensing[29]. The stability, cost-effectiveness, and bio-orthogonal catalytic capabilities of nanozymes have led to their extensive application in the development of various types of biosensors, including colorimetric, electrochemical, and fluorescent sensors[30,31]. In recent years, nanozyme-based colorimetric sensors have gradually emerged as a preferred alternative to traditional chromatographic methods owing to their simplified operation, reduced detection time, cost-effectiveness, and minimal reliance on professional personnel.
Recently, colorimetric sensors that incorporate aptamers and nanozymes have demonstrated significant potential for the detection of E. coli. These methodologies typically capitalize on the high selectivity of aptamers for targeting and the catalytic amplification capability of nanozymes. Consequently, this allows for highly sensitive, quick, and visible detection of the target bacteria. For instance, Li et al.[32] created an aptamer-modified Fe3O4/MWCNTs@Mo-CDs (FMMC) nanozyme, which catalyzes 3,3',5,5'-tetramethylbenzidine (TMB) colorimetry by mimicking the peroxidase activity, enabling rapid and highly sensitive E. coli detection with a limit of detection (LOD) as low as 0.978 CFU·mL−1. Huang et al.[33] advanced a pioneering dual-mode sensing strategy, underpinned by the innovative utilization of Fe3O4 NPs in conjunction with a fluorescein-labelled aptamer (FAM-Apt). The system demonstrates augmented enzyme-like activity at a neutral pH, thereby facilitating the dual detection of E. coli through the mechanisms of fluorescence recovery and colorimetric signal alterations. The detection limit of the system is 6 CFU·mL−1, and 10 CFU·mL−1 in the fluorescence, and colorimetric modes, respectively. Although these achievements demonstrate the promising application prospects of aptamer-nanozyme complexes in E. coli detection, current nanozymes still face challenges, including acidic pH-dependent catalytic activity, complex preparation processes, and limited stability. In contrast, octahedral Ag2O NPs, which have been successfully applied in the detection of small molecules such as pesticide residues[34], heavy metals[35], and pharmaceuticals[36], exhibit excellent oxidase-mimicking activity along with high catalytic efficiency and favorable stability. However, the use of colorimetric sensors that utilize the oxidase-mimicking activity of Ag2O NPs for the purposes of detecting foodborne pathogens such as E. coli remains relatively unexplored.
This study selected the P12-55 aptamer as the specific recognition molecule, employed the oxidase-mimicking activity of octahedral Ag2O NPs as a detection signal, and further leveraged the enhancement effect of aptamers on the oxidase-mimicking activity. This was undertaken to develop a new colorimetric biosensor for the detection of E. coli in food. The constructed biosensor exhibits notable advantages, including strong specificity, high sensitivity, and excellent stability, demonstrating favorable recovery rates in practical samples such as milk and tap water. This colorimetric biosensor provides a convenient and real-time, on-site detection strategy for E. coli in food. Moreover, serving as a modular platform, it can be extended to the detection of additional foodborne pathogens. The objective can be accomplished by means of a straightforward substitution of the aptamer, thus offering a novel approach for the development of straightforward and effective on-site detection techniques for foodborne pathogens.
-
The P12-55 aptamer, which specifically binds to E. coli, was synthesized by Sangon Biotechnology Co., Ltd (Shanghai, China), and subsequent purification was undertaken via high-performance liquid chromatography (HPLC). Its nucleotide sequence is 5'-CATACGATTTAGGTGACACTATAGCCGGAGGGGGGTGAGGTCTGCGGCAGGCTGTGTGGGTGGAATTTCTCCTACTGGGATAGGTGGA-3' (88 nucleotides, nt)[37]. Before use, all aptamers were subjected to rapid denaturation at 95 °C for 5 min, after which they were cooled to room temperature for subsequent experimentation. E. coli (ATCC 25922), Enterobacter sakazakii (E. sakazakii, ATCC 51329), Pseudomonas aeruginosa (P. aeruginosa, ATCC 27853), Bacillus cereus (B. cereus, ATCC 10876), Staphylococcus aureus (S. aureus, ATCC 6538), Salmonella typhimurium (S. typhimurium, ATCC 14028), and Listeria monocytogenes (L. monocytogenes, ATCC BAA-751) were procured from the American Type Culture Collection (ATCC) and stored as magnetic beads at –70 °C. Before utilization, these strains were subjected to activation through cultivation and subsequently stored at −20 °C. Agar powder was procured from Biosharp Biotechnology Co., Ltd (Beijing, China). At the same time, the LB (Luria-Bertani) broth medium was obtained from Shanghai Bio-way Technology Co., Ltd, silver nitrate powder (AgNO3), anhydrous sodium acetate (NaAc), sodium hydroxide (NaOH), glacial acetic acid (HAc), TMB, p-benzoquinone, isopropanol, dextran, starch, sucrose, magnesium sulfate (MgSO4), monopotassium phosphate (KH2PO4), N,N-dimethylformamide (DMF), L-arginine, L-histidine, absolute ethanol, casein, calcium carbonate (CaCO3), and ammonia solution (NH3·H2O) were all procured from Aladdin Co. Ltd (Shanghai, China). The Tetra Pak milk boxes utilized for the actual samples were procured from a proximate supermarket, while water samples were collected from tap water. All glassware utilized in the experiments was meticulously immersed and thoroughly cleaned with a newly composed aqua regia solution (HCl : HNO3 = 3:1). All substances utilized in this study were of analytical grade, and no additional purification steps were deemed necessary. NaAc buffer solutions were adjusted to the desired pH using HAc and NaOH. The preparation of TMB and p-benzoquinone involved their dissolution or dilution in a solution of DMF. At the same time, the other components were dissolved or diluted in water. Double-distilled water (ddH2O) was utilized throughout the experimental process.
Instruments
-
The Ultraviolet-Visible (UV-Vis) spectra and corresponding absorbance values were measured using a Microplate Spectrophotometer (Multiskan Skyhigh, Thermo Scientific, USA). The morphology of E. coli was subjected to observation by means of the BX43 Fluorescence Microscope (Olympus, Japan). Morphological and size characteristics of the octahedral Ag2O NPs were determined by using electron microscopy (SEM) (ZEISS Sigma 300, Germany). In addition, the surface groups and elemental composition of the nanoparticles were examined by means of X-ray Photoelectron Spectroscopy (XPS) (K-Alpha, Thermo Scientific, USA). The crystalline purity of the samples was determined by X-ray diffraction (XRD) (Bruker D8 Venture, Germany).
Preparation of octahedral Ag2O NPs
-
As stated in the preceding report[38], octahedral Ag2O NPs were prepared through the process of hydrothermal reduction. In a 50 mL flask, 25 mL of a solution of NH3·H2O (0.1 M) was slowly added dropwise to 50 mL of AgNO3 (0.1 M), under magnetic stirring at a rate of 600 rpm. Following a period of continuous stirring for a duration of 10 min, a quantity of 2.5 mL of NaOH (0.1 M) was gradually introduced into the mixture, which was stirred continuously, and a substantial brown precipitate was observed to form. The precipitate was then left in the solution for 12 h, after which the complex was subjected to a centrifugation process at 8,000 rpm for a period of 10 min. The collected precipitate was then subjected to a series of washes with ethanol and water until the pH of the final aqueous solution reached a neutral state. Finally, the obtained octahedral Ag2O NPs were dried at 60 °C and stored in a manner that ensured protection from light.
Preparation of the bacterial suspension
-
The bacterial strains employed in the experimental setup were preserved as magnetic beads at −70 °C. E. coli, E. sakazakii, P. aeruginosa, S. aureus, S. Typhimurium, B. cereus, and L. monocytogenes were activated by culturing on LB agar plates. The activated strains were subsequently inoculated into LB liquid culture medium and subjected to shaking conditions at 120 rpm and 37 °C for a period of 18–24 h. The harvesting process was executed during the logarithmic growth phase. For each strain, 1 mL of bacterial suspension was subjected to centrifugation at 4,500 rpm for 5 min at 25 °C. This was followed by several washes with sterile water, and the final resuspension in 1 mL of sterile water. The measurement of the optical intensity at 600 nm (OD600) was made. The experimental procedure was conducted utilizing a microplate reader. Subsequently, a volume of 100 µL from the diluted bacterial suspension was dispensed onto agar plates for the purpose of estimating the bacterial count. This was then expressed in terms of the total number of colony-forming units per milliliter (CFU·mL−1).
Fluorescent imaging of E. coli
-
The binding potential of E. coli and the P12-55 aptamer was observed through the utilization of a BX43 microscope. A FAM-labeled P12-55 aptamer (2 μM) was meticulously amalgamated with a bacterial suspension of E. coli (3 × 108 CFU·mL−1), and then incubated at 37 °C for a duration of 30 min. Consequently, the mixture was applied to a glass slide in order to prepare a smear. In the experiment, the aforementioned smear was observed under a BX43 Olympus fluorescence microscope, with excitation light and natural light being the sources. The microscope was calibrated to a magnification of 1,000×, thereby enabling the observation of individual E. coli.
Determination of reactive oxygen free radicals
-
Using p-benzoquinone and isopropanol as specific radical scavengers, the dynamic trajectory of hydroxyl radicals (·OH) and superoxide anion radicals (·O2−) during the course of the catalytic system was successfully monitored. A volume of 10 μL of Ag2O NPs and 10 μL of P12-55 aptamer solution was introduced into a 1.5 mL centrifuge tube and then thoroughly amalgamated before being subjected to a 37 °C temperature for 15 min. Thereafter, a quantity of 10 μL of TMB, 460 μL of NaAc (140 mM, pH 5.5) buffer, and 10 μL of p-benzoquinone or isopropanol were introduced to the solution, yielding a total of 500 μL. Finally, 200 μL of the resultant mixture was deposited into a microplate well, and the spectroscopic absorption at 652 nm was determined using a microplate reader.
Catalytic kinetics of octahedral Ag2O NPs
-
To establish the catalytic mechanism of octahedral Ag2O NPs towards the substrate TMB, 10 μL of Ag2O NPs, and 480 μL of NaAc (140 mM, pH 5.5) buffer solution was deposited into a centrifuge and mixed until thoroughly amalgamated. Subsequently, 10 μL of TMB substrate at varied concentrations was added and mixed completely, after which absorption of the system at 652 nm was measured at 5 s intervals. To study the effect of the P12-55 aptamer on the catalytic kinetics, 10 μL of Ag2O NPs were initially mixed with 10 μL of varying concentrations of P12-55 aptamer, and incubated at 37 °C for 30 min. Subsequently, 470 μL of NaAc (140 mM, pH 5.5) buffer solution and 10 μL of TMB were deposited, followed by thorough mixing and measurement of the resulting turbidity. The Beer-Lambert Law (A = εcl, where ε is the molar absorption of ox-TMB, 39,000 M−1·cm−1) was utilized to convert the measured absorption to product concentration[39], which was then employed to calculate the rate constant of the reaction. Finally, the catalytic kinetic parameters were evaluated by fitting the Michaelis-Menten equation and the Lineweaver-Burk double-reciprocal graph.
Procedure of E. coli detection
-
Under the optimized reaction conditions, 10 μL of Ag2O NPs dispersion and 10 μL of P12-55 aptamer solution were introduced into a 1.5 mL centrifugation tube. These components were thoroughly mixed, followed by incubation at 37 °C for a duration of 15 min. Next, 10 μL of E. coli suspension at different concentrations was mixed with the above solution and incubated further at 37 °C for 40 min. The control was prepared by the addition of an equivalent volume of sterile water in lieu of the bacterial suspension. Finally, 10 μL of TMB and 460 μL of NaAc (140 mM, pH 5.5) buffer were introduced to attain a total volume of 500 μL. The 200 μL test solution was transferred into a 96-well microplate for the purpose of UV/Vis spectroscopic analysis, which was conducted at 25 °C. The values of both the test sample (A) and the blank sample (A0) at 652 nm were measured. To evaluate the colorimetric detection capability of the method for E. coli, the change in absorption (ΔA = A – A0) was calculated.
Determination of E. coli in real samples
-
In light of the prevalence of E. coli in both milk and water, these samples were selected as exemplar matrices for the validation of a biosensor designed for the detection of the aforementioned contaminant in actual samples[40,41]. Pre-treatment of the milk sample was conducted in accordance with methods reported in the extant literature[42]. The milk was initially sterilized, then diluted 20-fold with ddH2O. The sample was then subjected to treatment with a range of concentrations of an E. coli suspension. Each treatment comprised 1 mL of the aforementioned milk sample. For the tap water sample, pretreatment followed established protocols[43]. Tap water underwent a series of filtration and sterilization processes, namely centrifugation and autoclaving, with the objective of eradicating any potential interfering particles or microorganisms. Thereafter, the water was diluted to a ratio of 1:1 with a buffer solution. Finally, 1 mL of the filtered solution was collected, various concentrations of E. coli bacterial suspensions were introduced, then the resulting samples were prepared for detection.
-
SEM observations revealed that the as-prepared Ag2O NPs exhibited a typical octahedral shape (Fig. 1a). A statistical analysis of the distribution of particle sizes was conducted, which indicated that the particle sizes of the Ag2O NPs ranged from 0.7 to 2.3 μm (Fig. 1b). The mean particle size was determined to be 1.37 μm. XRD was used to characterize the crystalline structure of the prepared Ag2O NPs. As demonstrated in Fig. 1c, all diffraction patterns were found to be highly consistent with the Ag2O crystal structure (JCPDS No. 41-1104)[44], and no impurity peaks were detected, confirming the high phase purity and crystallinity of the material. XPS has been utilized to characterize the elemental and chemical composition of surfaces. The full-scan XPS spectrum (Fig. 1d) revealed the existence of two elements, Ag and O. The high-resolution Ag 3d spectrum (Fig. 1e) exhibited two peaks in the binding energy spectrum at 367.7 and 373.7 eV, indicative of Ag 3d5/2 and Ag 3d3/2, respectively. The spin-orbit splitting energy was determined to be 6.0 eV, which was consistent with the characteristic value of Ag+ in Ag2O NPs. The high-resolution O 1s XPS spectrum (Fig. 1f) was then subjected to deconvolution, yielding a characteristic peak for lattice oxygen (Ag-O) at 530.5 eV and a characteristic peak associated with surface-adsorbed hydroxyl groups at 529.2 eV[45].
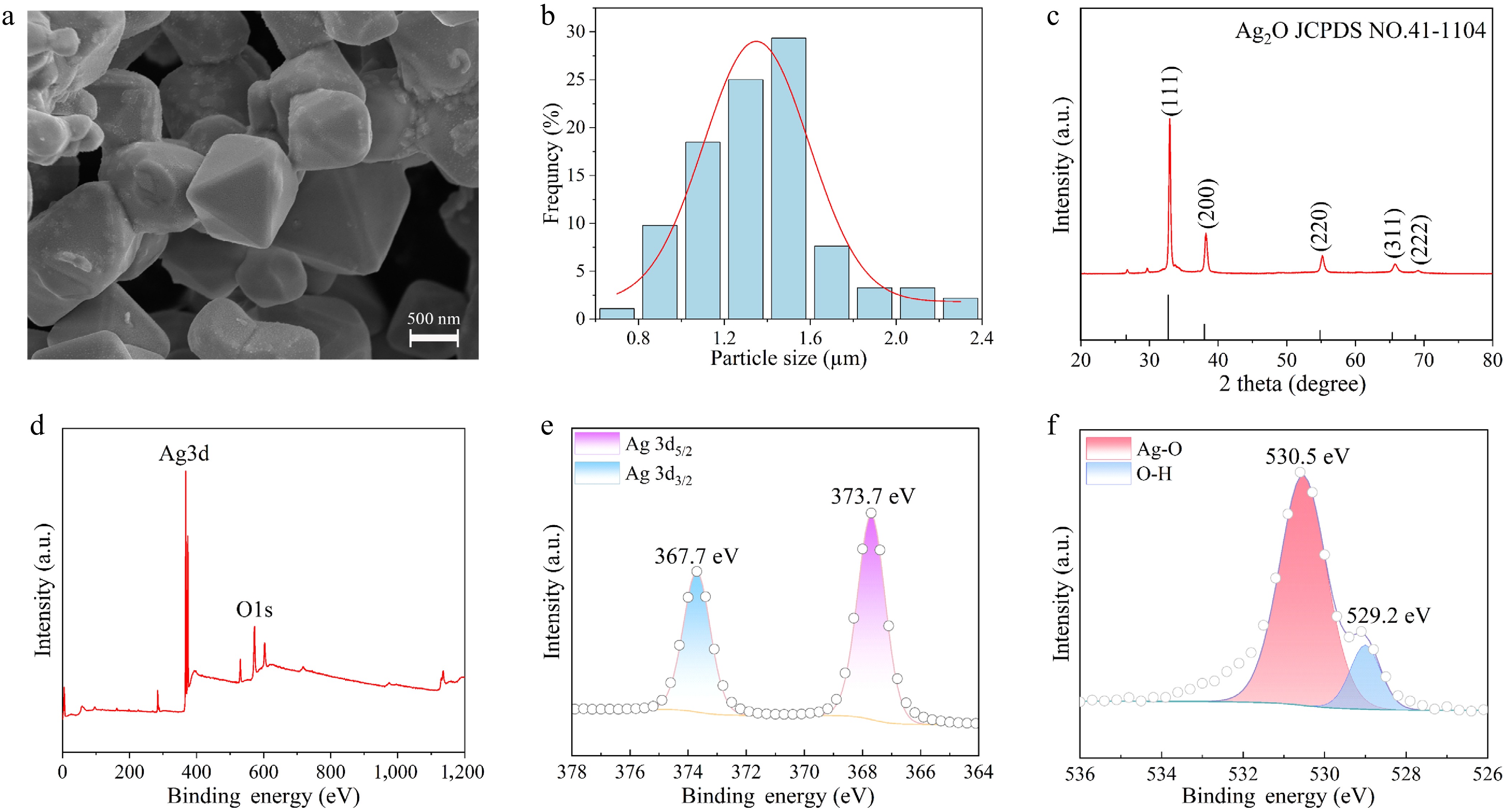
Figure 1.
(a) SEM of octahedral Ag2O NPs. (b) Size distribution characteristics of Ag2O NPs. (c) XRD patterns of Ag2O NPs. (d) XPS survey spectrum of Ag2O NPs. (e) High-resolution Ag3d XPS data of Ag2O NPs. (f) High-resolution O1s XPS data of Ag2O NPs.
Design of the aptamer-based biosensor
Principle of E. coli detection
-
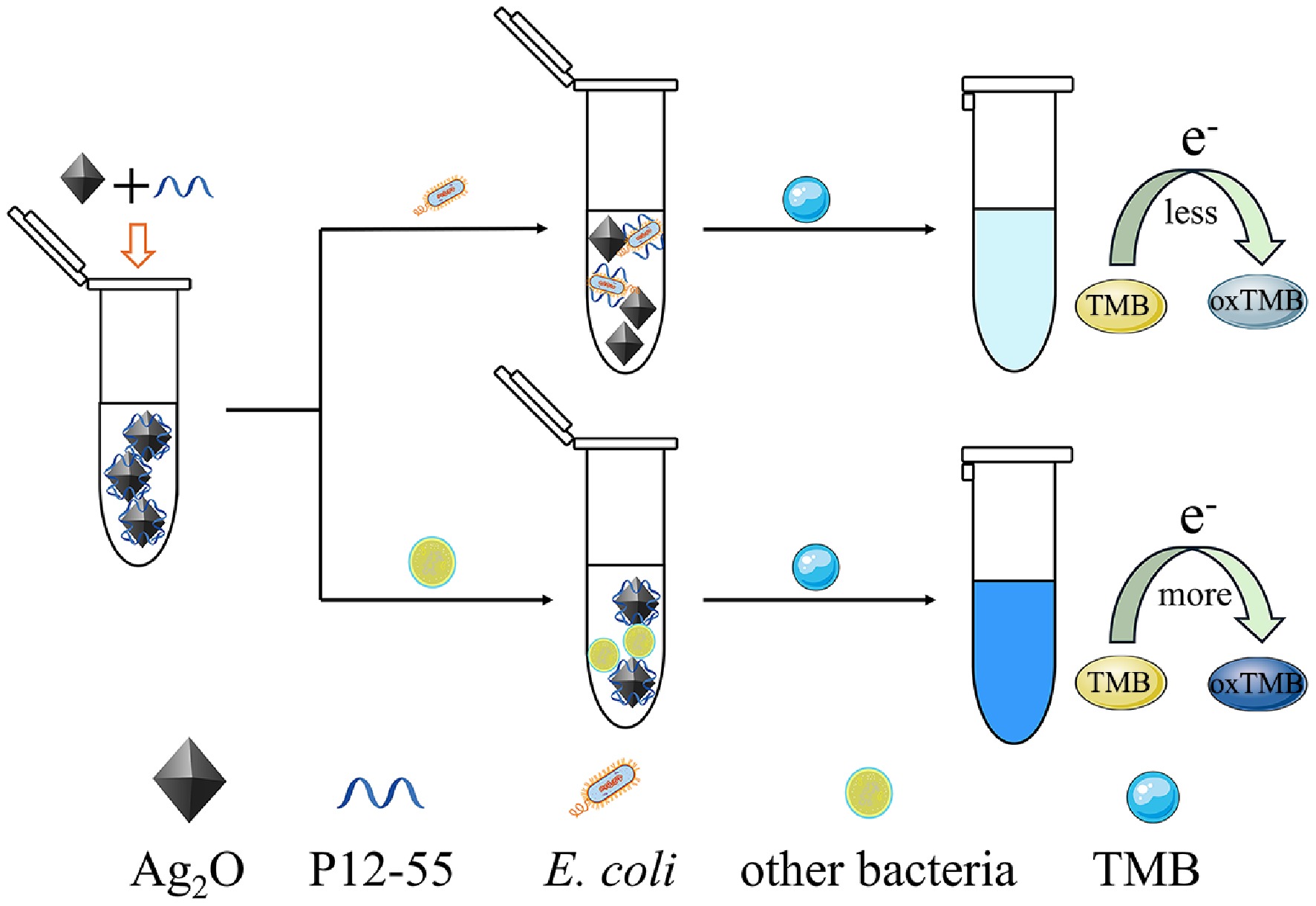
Herein, a newly developed aptamer-based biosensor has been successfully constructed for the highly sensitive detection of E. coli. The biosensor utilizes the improved oxidase-mimicking capability of Ag2O NPs as the sensing signal. Scheme 1 illustrates the detection principle. In instances where E. coli is absent in the biosensor system, the P12-55 aptamer promotes the generation of more ·O2− on the external surface of Ag2O NPs[46], thereby enhancing their oxidase-mimicking activity and leading to the quick oxidation of many TMB molecules into a blue product. When E. coli is present in the system, the P12-55 aptamer preferentially binds to E. coli and is first consumed, thereby preventing the enhancement of the oxidase-mimicking capacity of Ag2O NPs. Consequently, only a small amount of ·O2− is generated on the surface, and the TMB molecules are slowly oxidized to produce a light blue product. By measuring the change in the extinction coefficient of the sensing system at 652 nm, the detection of E. coli can be achieved with great sensitivity and simplicity.
Feasibility of the biosensor
-
The validity and feasibility of the established colorimetric biosensor in detecting E. coli were established through measuring the absorbances at 652 nm, and subsequently observing the color changes of various reaction solutions (Fig. 2). The octahedral Ag2O NPs alone were found to be capable of catalyzing the oxidation of a limited quantity of TMB, producing a light blue product with a negligible peak of absorption at 652 nm (Sample 1). After P12-55 aptamer was adsorbed onto the Ag2O NPs to form a recognition probe, a large amount of TMB was oxidized, yielding a deep blue product, accompanied by a significant peak of absorption at 652 nm (Sample 2). The results confirm that the adsorption of the P12-55 aptamer significantly boosts the catalytic activity of Ag2O NPs, leading to the efficient oxidation of TMB. Following the incorporation of E. coli in the recognition probe, a gradual transition in the color of the sensing mixture from deep blue to light blue was observed (Samples 3–5). Concurrently, the characteristic peak of the absorption spectrum at 652 nm underwent a progressive decline, concomitant with increasing concentrations of E. coli. The potential interactions between octahedral silver oxide, E. coli, P12-55 aptamer, and TMB were further investigated. As demonstrated in Supplementary Fig. S1, there is an absence of mutual influence among these components, thereby confirming that E. coli alone did not affect the oxidase-mimicking activity of octahedral Ag2O NPs. The findings indicate that the aptamer-facilitated catalytic enhancement mechanism can be employed for the highly sensitive-selective detection of E. coli, providing a basis for the construction of stable and reliable colorimetric detection platforms.
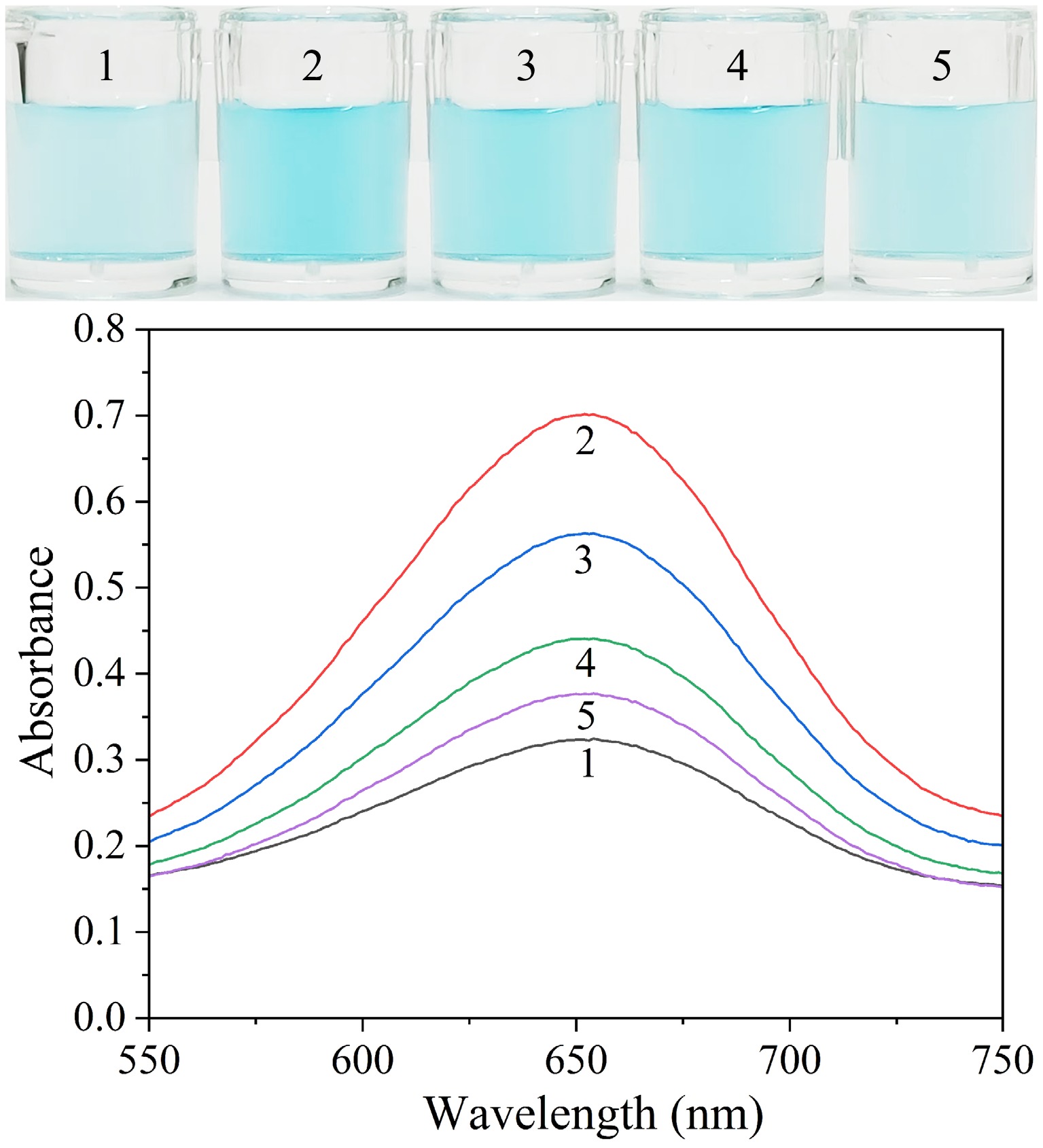
Figure 2.
Visual images and absorption spectra of solutions including different compounds. Sample 1: Octahedral Ag2O NPs + TMB; Sample 2: Sample 1 + P12-55 aptamer; Sample 3: Sample 2 + 3 × 102 CFU·mL−1 E. coli; Sample 4: Sample 2 + 3 × 104 CFU·mL−1 E. coli; Sample 5: Sample 2 + 3 × 105 CFU· mL−1 E. coli.
Binding affinity of E. coli to the aptamer
-
The secondary structure of the P12-55 aptamer sequence under its minimum free energy conformation was predicted using RNA secondary structure prediction software (Fig. 3a), yielding a binding free energy value of –9.59 kcal·mol−1. The relatively low binding free energy indicates that this aptamer possesses a stable secondary structure and demonstrates a potentially high affinity for its target, theoretically suggesting that the P12-55 aptamer may exhibit strong binding to E. coli[37]. To establish the binding affinity of the aptamer for E. coli, the 5'-FAM-labeled P12-55 aptamer was incubated with E. coli for 30 min. Under bright-field microscopy, the typical rod-shaped morphology of E. coli was observed (Fig. 3b). Under BW2 excitation light, clear green fluorescent signals were visualized to wrap around the bacterial cells in a ring shape, which perfectly coincided with the bacterial morphology. Notably, the positions of blue fluorescence completely matched those of the bacteria observed under bright-field microscopy (Fig. 3c). The obtained results suggest that the FAM-labeled P12-55 aptamer is capable of binding to the E. coli surface specifically.
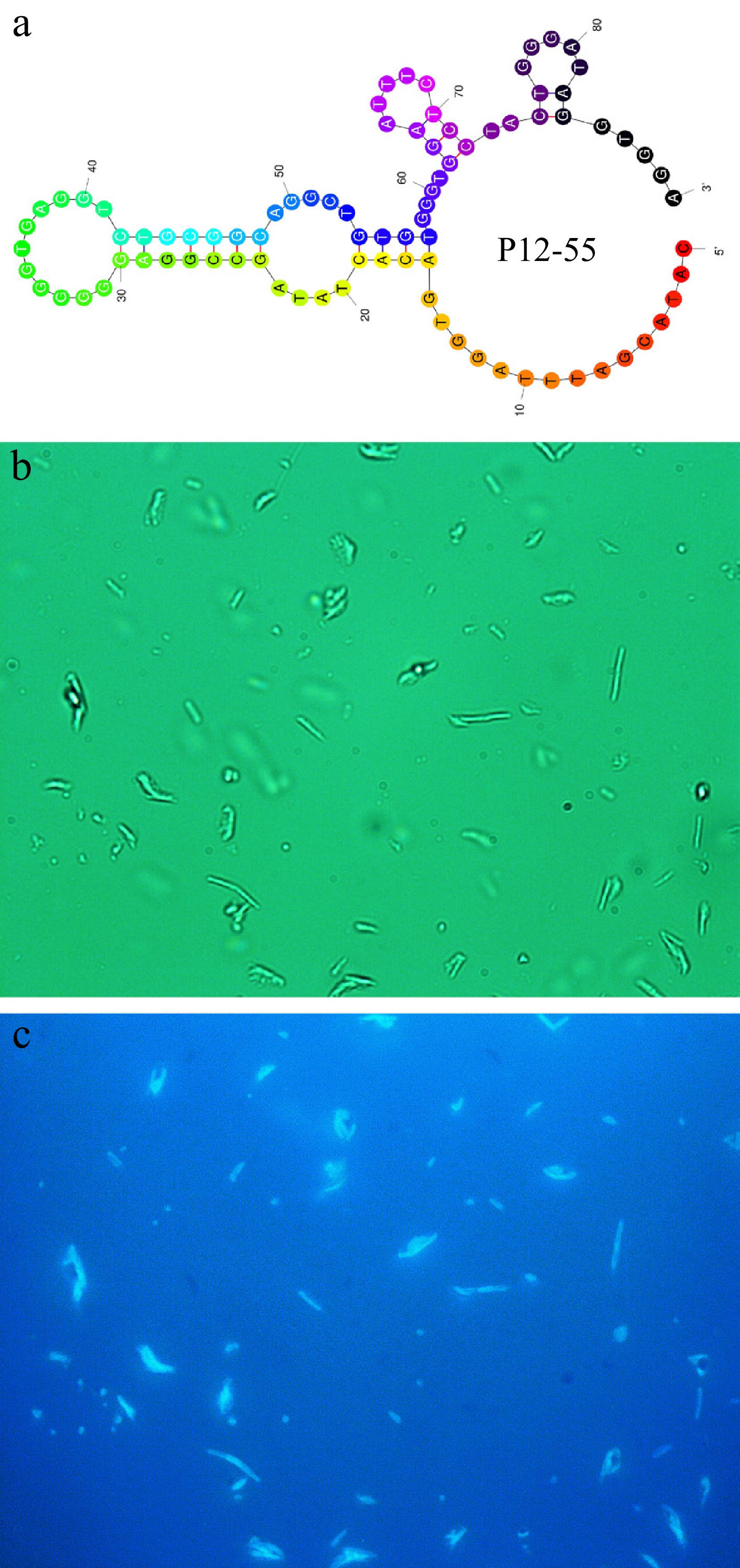
Figure 3.
(a) Secondary structure with the least energy of the P12-55 aptamer. (b) Natural light image of E. coli treated with FAM-labeled P12-55 aptamer. (c) Fluorescent image of E. coli treated with FAM-labeled P12-55 aptamer.
Effect of oxygen on the biosensor
-
As indicated by previous studies, the reaction mechanism underlying the oxidation-mimicking phenomenon exhibited by metal oxide nanomaterials has been shown to be associated with the presence of oxygen free radicals (·O2− and ·OH)[47]. Oxygen free radicals can be generated by dissolved oxygen being catalyzed by nanozymes, and TMB is then oxidized by these radicals to form ox-TMB, a typical blue product. To ascertain the role of oxygen in the Ag2O NPs oxidase-mimicking activity, N2-O2 replacement experiments were conducted. In an O2 saturated buffer, TMB was rapidly oxidized to exhibit a blue color, whereas in an N2 saturated buffer, the color change was significantly decelerated, and the solution eventually only displayed a light blue (Supplementary Fig. S2). The findings indicate that the reaction of TMB oxidation by Ag2O NPs is highly dependent on the presence of dissolved oxygen.
Effect of aptamer on reactive oxygen free radicals
-
Octahedral Ag2O NPs have been demonstrated to possess strong oxidase-mimicking activity[34]. This capacity enables them to catalyze dissolved oxygen, thereby inducing the generation of either ·O2− or ·OH[48]. These oxygen-free radicals have the capacity to oxidize colorless TMB, resulting in the formation of the blue product, ox-TMB. To elucidate the mechanism underlying the enhancement of the oxidase-mimicking catalytic capacity of Ag2O NPs by the P12-55 aptamer, p-benzoquinone, and isopropanol, these substances were introduced into the catalytic system[49,50]. As demonstrated in Supplementary Fig. S3a, no substantial alteration in the values of the measured absorbances was evident following the addition of isopropanol, thereby indicating that the generation of ·OH was not a consequence of the reaction. In contrast, significant decreases in absorbance values were observed in both the aptamer-containing system upon the addition of p-benzoquinone (Supplementary Fig. S3b), confirming the generation of ·O2− in the system. These results suggest that dissolved oxygen, when adsorbed on the surface of Ag2O NPs, underwent an oxidation process to form ·O2−. With increasing concentrations of p-benzoquinone, the catalytic system exhibited a gradual decrease in the absorbance values, thereby demonstrating the effective scavenging of radicals by p-benzoquinone and validating its use as a probe for the tracking of radical dynamics. Importantly, the aptamer-containing system exhibited a smaller decrease in colorimetric signal, further indicating that the binding of P12-55 aptamer to Ag2O NPs catalyzed the generation of more ·O2−, thereby enhancing its oxidase-mimicking activity.
Catalytic stability of octahedral Ag2O NPs
-
Under identical concentration and reaction conditions, three independent batches of Ag2O NPs exhibited highly consistent catalytic activity (Supplementary Fig. S4a), demonstrating good reproducibility and batch-to-batch stability of the synthesis protocol. Furthermore, the Ag2O NPs suspension was stored at 25 °C for 30 d, and its catalytic performance was monitored over this period. As shown in Supplementary Fig. S4b, the material retained over 94% of its initial catalytic activity after storage, confirming its excellent long-term stability under storage conditions. These results collectively verify the reliability and stability of the Ag2O NPs for catalytic applications.
Catalytic kinetics of octahedral Ag2O NPs
-
The mechanism underlying the enhancement of oxidase-mimicking capability exhibited by octahedral Ag2O NPs was investigated by determining the catalytic kinetics in the presence and absence of P12-55 aptamer. The adjustment of TMB concentration consequently resulted in the generation of Michaelis-Menten curves (Supplementary Fig. S5a) and Lineweaver-Burk plots (Supplementary Fig. S5b). With the introduction of the P12-55 aptamer, the Km value decreased from 1.920 to 0.843 mM. At the same time, the Vmax increased from 4.33 × 10−8 to 5.77 × 10−8 M·s−1. These results indicate that P12-55 aptamer enhances the catalytic efficiency of Ag2O NPs. The phenomenon under discussion can be explained by the adsorption of P12-55 aptamer onto the surface of Ag2O NPs by means of electrostatic and π-π stacking interactions[51]. This process strengthens the interaction between Ag2O NPs and the TMB substrate, and promotes the generation of more ox-TMB.
Optimization of sensing parameters
Effect of buffer pH
-
The pH of the buffer solution is of critical importance to the performance of the reaction system, as it directly governs the reaction rate, product distribution, and even the reaction pathway by regulating the charge state of reactants, enzyme activity, molecular conformation, and stability. Herein, the pH of the sensing buffer was optimized through the use of 0.4 mM TMB as the catalyst. Within the pH range of 4–5.5, the aptamer biosensor exhibited a gradual increase in the colorimetric signal, reaching a maximal level at pH 5.5. The subsequent increase in pH resulted in a decline in the signal (Supplementary Fig. S6). Therefore, the subsequent experiments were performed in a pH 5.5 buffer solution to achieve optimal colorimetric performance.
Effect of buffer concentration
-
The salt ion concentration affects ionic strength, pH stability, molecular interactions, and overall biosensor performance. Herein, the concentration of NaAc was optimized Supplementary Fig. S7. The colorimetric signal intensity exhibited an increase with rising NaAc concentrations within the range of 100–140 mM, but a decrease beyond this range. This phenomenon may be attributed to the insufficient ionic strength at low concentrations, which reduces reaction efficiency. The excessively high Na+ ions may compete with the metal oxide nanozyme for binding sites, thereby weakening its catalytic capacity of the enzyme.
Effect of aptamer concentration
-
The concentration of the aptamer utilized to modulate the catalytic function of octahedral Ag2O NPs serves as one of the critical parameters for the biosensor. As depicted in Supplementary Fig. S8, the oxidase-mimicking capacity of Ag2O NPs exhibited a gradual increase in response to an increase in the P12-55 concentration. The colorimetric signal attained its zenith at an aptamer concentration of 30 nM. However, further increasing the aptamer concentration instead resulted in a decrease in activity. This phenomenon may be explained by the hypothesis that, at a P12-55 aptamer concentration of 30 nM, the binding between octahedral Ag2O NPs and P12-55 aptamer has approached saturation. Conversely, excessively high aptamer concentrations would coat the surface of the Ag2O NPs, thereby hindering its binding to the substrate and impairing its catalytic activity.
Effect of octahedral Ag2O NPs concentration
-
To investigate the impact of various quantities of octahedral Ag2O NPs on the performance of the biosensor, the quantity of Ag2O NPs was optimized. The colorimetric signal exhibited an initial increase, followed by a subsequent decrease, in response to increased concentrations of Ag2O NPs, reaching a maximum when the Ag2O NPs concentration was 60 μg·mL−1 (Supplementary Fig. S9). This phenomenon may be explained by the fact that the interaction between Ag2O NPs and the aptamer had reached a saturated state at a concentration of 60 μg·mL−1. The subsequent decrease in signal could be attributed to either substrate saturation or nanozyme aggregation, which led to reduced catalytic activity.
Performances of the colorimetric biosensor
Sensitivity
-
To assess the sensitivity of the aptamer-based colorimetric biosensor that has been developed for the detection of E. coli, the response of the biosensor to varying concentrations of E. coli was recorded. As illustrated in Fig. 4, with increasing E. coli concentration, a gradual shift in color from blue to light blue in the catalytic solution was observed, accompanied by a linear upward trend in the biosensor signal intensity. This observation arises because the oxidase-mimicking capability of Ag2O NPs undergoes a restoration following the binding of the P12-55 aptamer to E. coli. The range at which E. coli could be detected in a linear fashion was determined to be 3 × 102 to 3 × 108 CFU·mL−1 (y = 0.128x−0.144, R2 = 0.995). Using the formula 3σ/slope (where σ = 0.00585), the LOD was determined to be 4 CFU·mL−1 (S/N = 3)[52,53]. Notably, when the E. coli count exceeded 3 × 105 CFU·mL−1, a distinct color fading became visually observable. The results obtained demonstrate the sensitivity of the constructed aptamer-based colorimetric biosensor for the detection of E. coli.
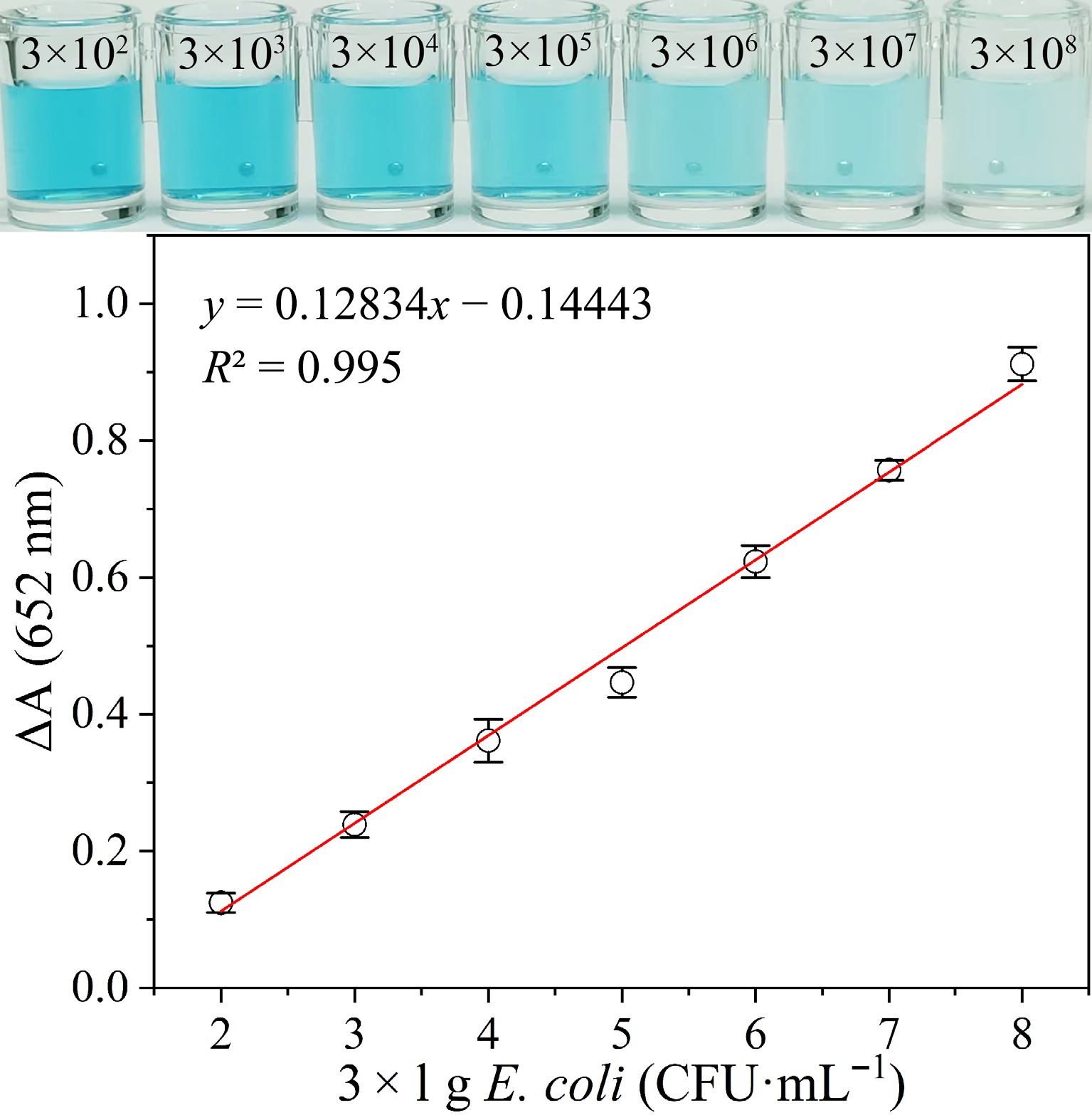
Figure 4.
Visual color and absorption variation (ΔA) of the catalytic solution added with varying concentrations of E. coli.
The efficacy of the established colorimetric biosensor was then juxtaposed with that of previously documented E. coli detection methodologies, the findings of which are delineated in Table 1. Although noteworthy sensitivity is exhibited by electrochemical methods in the detection of E. coli, their sensing signals are prone to interference from a variety of factors, including sample temperature, pH, and ionic strength. This may impose limitations on the applicability of these methods in complex food samples. Despite fluorescence methods having high accuracy, they typically require complex material modification, high costs, and tedious operational procedures. In contrast, colorimetric technology represents the simplest strategy, enabling visual distinction of E. coli without the need for complex instruments. The developed biosensor exhibits high sensitivity, low interference, and excellent stability, thus obviating the necessity for sophisticated instrumentation or complex modifications. This study reports, for the first time, a novel colorimetric biosensor utilizing oxidase-mimicking activity for E. coli detection, offering a novel approach for the convenient detection of E. coli in food samples.
Table 1. Comparison of the established colorimetric biosensor in detecting E. coli with other methods.
Method Materials Linear range (CFU·mL−1) LOD (CFU·mL−1) Actual sample Ref. Electrochemistry Ag NPs 3.4 × 101−3.4 × 106 34 Tap water, artificial urine, milk [54] MXene-GO 3.0–3.0 × 106 3.0 Serum [55] Fe3O4@COF-Au NPs 1.0 × 102−1.0 × 109 10 Orange juice, milk, serum [56] Fluorescence UCNPs-WS2 8.5 × 101−8.5 × 108 17 Tap water, green tea powder [57] TPE-(COOH)4Na 6.5–6.5 × 107 2.8 Milk [58] Fe3O4 + FAM-Apt 1.0 × 101−1.0 × 108 6.0 Milk [33] Colorimetry K, Na-PHI 1.0 × 102−1.0 × 104 85 Egg white, tofu, soya milk [59] Au NRs 1.0 × 101−1.0 × 106 7.0 Water [43] Apt-Au@Fe3O4 NPs 1.0 × 101−1.0 × 108 3.0 Tap water, milk, orange juice [60] Ag2O NPs 3.0 × 102−3.0 × 108 4.0 Tap water, milk This work MXene-GO: MXene and graphene oxide composite; COF: covalent organic framework; UCNPs: upconversion nanoparticles; TPE-(COOH)4Na: sodium tetrakis (phenylethynyl) benzenesulfonate; K, Na-PHI: potassium/sodium poly (heptazine imide); NRs: nanorods. Selectivity
-
The evaluation of specificity of the colorimetric biosensor for a number of competing bacteria, including P. aeruginosa, E. sakazakii, L. monocytogenes, S. aureus, S. typhimurium, and B. cereus were separately introduced into the sensing system. Additionally, considering potential interference from complex components in food samples (e.g., casein, carbohydrates, amino acids, and metal ions), some competitive targets such as dextran, starch, sucrose, Ca (II), Mg (II), K (I), L-arginine and L-histidine were chosen to confirm the selectivity of the colorimetric biosensor. By observing the color change of the sensing mixture (Fig. 5a), only the solution containing E. coli exhibited a light blue color, whereas solutions containing other competitive targets remained dark blue. The recording of the variations in absorbance at 652 nm (Fig. 5b) showed that only the sample containing E. coli demonstrated a substantial decline in absorbance, while the solutions added with other competitive targets displayed no significant change in absorption. These results demonstrate that the P12-55 aptamer is capable of binding specifically to E. coli, even in the presence of competing bacteria or complex interfering components, confirming that the biosensor exhibits excellent selectivity and anti-interference capability.
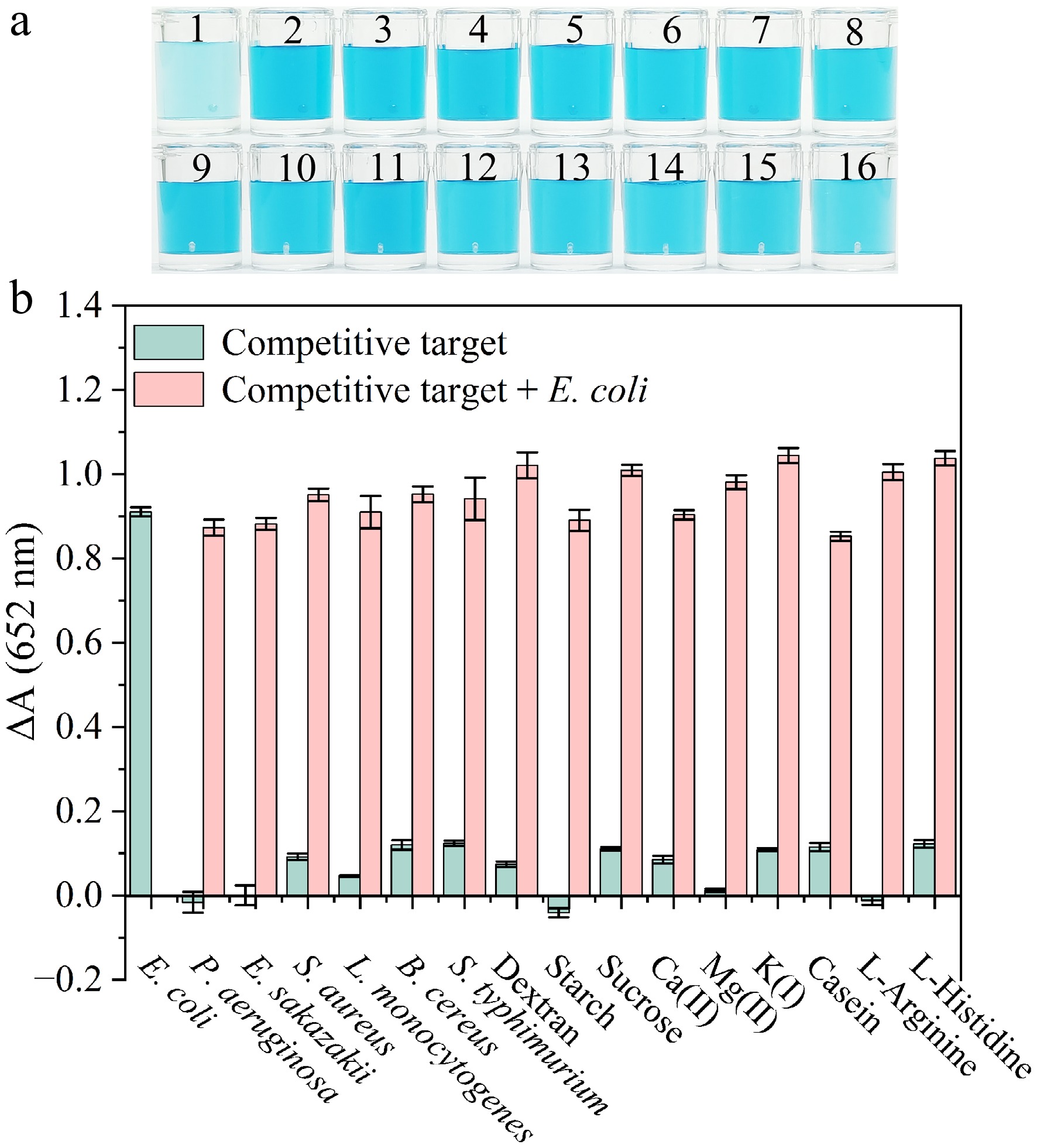
Figure 5.
(a) Color change of the colorimetric sensor in the presence of competitive targets. Samples 1 to 16 are individually E. coli, P. aeruginosa, E. sakazakii, S. aureus, L. monocytogenes, B. cereus, S. typhimurium, dextran, starch, sucrose, Ca (II), Mg(II), K(I), casein, L-Arginine and L-Histidine. (b) Variation in absorbance in response to E. coli and other targets. The concentration of each bacterial strain was 3 × 108 CFU·mL−1. Concentrations of carbohydrates, metal ions, and amino acids of 100 μg·mL−1.
Analysis of E. coli in real samples
-
To evaluate the application performance of the aforementioned biosensor in relation to the detection of E. coli in actual samples, a series of spiking recovery trials were carried out using milk and tap water as matrices. As presented in Table 2, the recuperation rates of E. coli exhibited a range from 99.1% to 104%, with relative standard deviation (RSD) of 4.57% to 6.11%, all of which were below 10%. These results demonstrate that the established colorimetric biosensor demonstrates adequate accuracy, and is appropriate for the effective detection of E. coli in actual samples.
Table 2. Determination of E. coli in the spiked real food samples.
Sample Addition
(CFU·mL−1)Detection
(CFU·mL−1)Recovery
rate (%)RSD (%) Milk 0 0 None None 3.00 × 108 3.04 × 108 101 5.10 3.00 × 107 3.06 × 107 102 4.57 3.00 × 106 2.97 × 106 99.1 4.88 Water 0 0 None None 3.00 × 108 3.13 × 108 104 5.91 3.00 × 107 3.03 × 107 101 6.11 3.00 × 106 3.08 × 106 103 5.19 -
This study demonstrates that the oxidase-mimicking catalytic capacity of octahedral Ag2O NPs is significantly increased under the influence of P12-55 aptamer, thus providing a feasible approach for the efficient detection of E. coli in milk and tap water. The constructed aptamer-based colorimetric biosensor not only enables specific recognition of E. coli, but also exhibits favorable recovery rates in real samples, highlighting its promising practical application potential. Additionally, the biosensor demonstrates a high level of sensitivity, with an LOD of 4 CFU·mL−1. Furthermore, the favorable recovery rates were obtained in the detection of dairy products and tap water samples, further confirming the reliable nature of the method. The colorimetric biosensor represents a simple, reliable, and low-cost method for E. coli detection.
This work was financially supported by the Scientific Research Projects of General Administration of Customs (2023HK129), the National Natural Science Foundation of China (32360621, 32160603), and the Guizhou Provincial Key Technology R&D Program (QKHZC[2026]318).
-
The authors confirm their contributions to the paper as follows: methodology: Huang X, Zhang L, Zhou C, Wu Y; investigation: Huang X, Zhang L, Lu T, Li J, Wang M, Dai W, Zhou C, Xu J; data curation: Huang X, Zhang L, Lu T, Wang M, Xu J; formal analysis: Zhang L, Lu T, Li J, Wu Y; validation: Huang X, Lu T, Li J, Dai W, Zhou C, Xu J; Supervision: Zhang L, Wu Y; writing-original draft: Huang X; funding acquisition, project administration, writing-review & editing: Wu Y. All authors reviewed the results and approved the final version of the manuscript.
-
The data supporting the findings of this study are available upon reasonable request from the corresponding author. All relevant data generated or analyzed during this study are included in this published article and its supplementary information files.
-
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
-
accompanies this paper online at: https://doi.org/10.48130/fia-0026-0014.
-
Authors contributed equally: Xiaohuan Huang, Ling Zhang
- Supplementary Fig. S1 Visual images and absorption spectra of solutions including different compounds.
- Supplementary Fig. S2 Comparison of TMB colorimetric reactions catalyzed by Ag2O NPs in air or N₂-saturated buffer solutions.
- Supplementary Fig. S3 (a) Impact of ·OH inhibitors on the comparative activities of Ag2O NPs in disparate systems. (b) Impact of ·O2− inhibitors on the relative activities of Ag2O NPs in different systems.
- Supplementary Fig. S4 (a) Differences in the catalytic activity of Ag2O NPs obtained from various batches. (b) Effect of long-term storage on catalytic activity of Ag2O NPs.
- Supplementary Fig. S5 Steady-state kinetic analysis and the enzymatic mechanism of octahedral Ag2O NPs catalyzed by P12-55 aptamer. (a) Michaelis-Menten plot. (b)Lineweaver-Burk plot.
- Supplementary Fig. S6 Effect of buffer pH on the colorimetric biosensor for E. coli detection.
- Supplementary Fig. S7 Effect of salt ion concentration on the colorimetric biosensor for E. coli detection.
- Supplementary Fig. S8 Effect of P12-55 aptamer concentration on the colorimetric biosensor for E. coli detection.
- Supplementary Fig. S9 Effect of Ag2O NPs concentration on the colorimetric biosensor for E. coli detection.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of China Agricultural University, Zhejiang University and Shenyang Agricultural University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Huang X, Zhang L, Lu T, Li J, Wang M, et al. 2026. A colorimetric biosensor for Escherichia coli detection in food based on aptamer-enhanced oxidase-mimicking activity of octahedral Ag2O nanoparticles. Food Innovation and Advances 5(2): 185−194 doi: 10.48130/fia-0026-0014
A colorimetric biosensor for Escherichia coli detection in food based on aptamer-enhanced oxidase-mimicking activity of octahedral Ag2O nanoparticles
- Received: 27 November 2025
- Revised: 20 January 2026
- Accepted: 25 January 2026
- Published online: 30 April 2026
Abstract: Escherichia coli (E. coli) poses a grave threat to food safety, underscoring the need for expeditious and precise detection methodologies. Conventional colorimetric approaches are simple, but they frequently lack accuracy and stability. This study constructed an aptamer-based colorimetric biosensor, leveraging the enzyme-mimicking activity of octahedral Ag2O nanoparticles (NPs), mediated by oligonucleotides, aiming for highly sensitive foodborne E. coli detection. The P12-55 aptamer can be adsorbed onto the octahedral Ag2O NPs surface, thereby significantly enhancing their oxidase-mimicking activity. The aptamer enhances activity by promoting ·O2− generation, and accelerating electron transfer to the 3,3',5,5'-tetramethylbenzidine (TMB) substrate. In the presence of E. coli in the sensing system, the aptamer exhibits a preferential binding affinity for the bacteria, thereby restoring the oxidase-mimicking activity and enabling the transduction of the detection signal. The biosensor exhibited a range of linear detection of 3 × 102–3 × 108 CFU·mL−1, and the limit of detection (LOD) was as low as 4 CFU·mL−1. Recovery rates of 99.1% to 104% were achieved in milk and tap water samples, demonstrating outstanding practicality and reliability. This study proposes a novel and efficient method for the rapid detection of harmful bacteria in food, which is of significant importance for food safety assurance.
-
Key words:
- Pathogenic bacteria /
- Aptamer /
- Nanozyme /
- Colorimetric detection /
- Sensor