-
The genus Diaporthe Nitschke (Sordariomycetes, Diaporthomycetidae, Diaporthales, Diaporthaceae) was described in 1867[1] with the type species D. eres Nitschke. The genus Phomopsis (Sacc.) Bubák was described in 1905[2] with the type species P. lactucae (Sacc.) Bubák. Diaporthe species have a complex pleomorphic life cycle. Previously, Phomopsis species were considered to be the asexual stages of Diaporthe species. According to the International Code of Nomenclature for algae, fungi, and plants and the principle of "one fungus–one name", Diaporthe, being the older generic name, has priority over Phomopsis[3]. Diaporthe encompasses important plant pathogenic, endophytic, and saprobic species, with a diverse host range and a global distribution[4,5].
Species identification in the genus Diaporthe has evolved from host association and morphology to the widespread adoption of DNA sequencing[4−10]. The most taxonomically informative DNA loci more frequently used to reconstruction the molecular phylogeny of Diaporthe species are the internal transcribed spacer (ITS) of the ribosomal DNA and partial calmodulin (cal), histone 3 (his3), translation elongation factor 1α (tef1), and β-tubulin (tub2) genes[4,5,7,9,10]. A polyphasic approach for reliable species recognition should be used that relies on the consolidated species concept (CSC)[11], which involves the incorporation of morphological, biological, and phylogenetic characteristics[9,12,13]. Data on phylogenetic features should be obtained according to the genealogical concordance phylogenetic species recognition (GCPSR)[14] method using a multilocus phylogenetic approach for recognizing fungal species. Certain Diaporthe species are known to produce biologically active compounds with herbicidal activity[15−17]. For example, Diaporthe convolvuli (Ormeno-Nuñez, Reeleder & A.K. Watson) R.R. Gomes, Glienke & Crous, which is host-specific to Convolvulus arvensis L., produces secondary metabolites, namely convolvulanic acid A, convolvulanic acid B, convolvulol, and α-pyrone convolvulopyrone, which possess phytotoxic activity[15]. Fungi with this potential could be explored given their potential application in agriculture as promising candidates for the development of natural herbicides. Field bindweed Convolvulus arvensis is a harmful weed of major agricultural crops that is distributed almost worldwide[18]. In Russia, it is found in the European part, the Caucasus, western and eastern Siberia, and the Far East.
In July 2024, visually healthy leaves of C. arvensis were collected in northwest Russia (Pushkin, Saint Petersburg, Leningrad Region). Two Diaporthe sp. strains were isolated from these leaves. The aim of this study was to identify these strains according to the CSC by their phylogenetic, micromorphological, and cultural features and to assess their pathogenicity.
-
Leaves of C. arvensis were surface-sterilized by consistent rinsing with a 0.1 % AgNO3 and sterile water and incubated on potato sucrose agar (PSA)[19]. The Petri dishes were incubated at 24 °C in the dark and were analyzed after 7–10 days of cultivation. The obtained strains MF-SOR 61.20.1 and MF-SOR 61.20.2 were stored in microtubes on PSA at 4 °C in the mycological collection of pure cultures at the A.A. Jaczewskii Laboratory of Mycology and Phytopathology, All-Russian Institute of Plant Protection (MF, VIZR, Saint Petersburg, Russia).
DNA isolation, polymerase chain reaction, and sequencing
-
Genomic DNA was extracted from mycelia obtained from the cultures incubated on PSA according to the standard cetyltrimethylammonium bromide (CTAB)/chloroform protocol[20].
The ITS region of the rDNA and partial cal, his3, tef1, and tub2 genes were amplified and sequenced for the strains studied. The primers ITS1 and ITS4[21], CAL-228F and CAL-737R[22], CYLH3F[23] and H3-1b[24], EF1-728F and EF1-986R[22], and βtub2Fw and βtub4Rd[25] or T1 and Bt2b[26], were used to amplify the ITS region and the partial cal, his3, tef1, and tub2 genes, respectively. The polymerase chain reaction (PCR) conditions were as follows: 95 °С for 5 min; followed by 35 cycles of 92 °С for 50 s; 55 °C for 40 s, (ITS1/ITS4), 52 °C for 40 s (βtub2Fw/βtub4Rd and T1/Bt2b), 56 °C for 40 s (CAL-228F/CAL-737R), or 58 °C for 40 s (CYLH3F/H31-b and EF1-728F/EF1-986R); 72 °С for 75 s; and a final elongation for 5 min at 72 °С.
Single-strand DNA of amplicons were sequenced by Sanger's method on an ABI Prism 3500 analyzer (Applied Biosystems, Thermo Fisher Scientific, Waltham, MA, USA) with a Bigdye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Thermo Fisher Scientific) according to the manufacturer's instructions. The nucleotide sequences of the ITS, cal, his3, tef1, and tub2 genes were deposited in the GenBank database with the corresponding accession numbers (Table1).
Table 1. GenBank accession numbers of the Diaporthe section Sojae strains included in the study.
Diaporthe species Strain number ITS cal his3 tef1 tub2 D. acaciarum CBS 138862, t NR_137939 − KP004504 − KP004509 D. alpiniae CGMCC 3.28221, t PQ321210 PQ336555 PQ336573 PQ336519 PQ336537 D. amaranthophila MAFF: 246900, t LC459575 LC459583 LC459581 LC459577 LC459579 D. ambigua CBS 114015, t KC343010 KC343252 KC343494 KC343736 KC343978 D. angelicae CBS 111592; AR3724, t KC343027 KC343269 KC343511 KC343753 KC343995 D. arctii CBS 139280, t KJ590736 KJ612133 KJ659218 KJ590776 KJ610891 D. arezzoensis MFLU: 19-2880 MT185503 − − MT454019 MT454055 D. atlantica CECT 21217; SP11, t ON159893 ON364019 ON398810 ON398831 ON364040 D. batatas CBS 122.21 KC343040 KC343282 KC343524 KC343766 KC344008 D. beilharziae BRIP 54792 JX862529 − − JX862535 KF170921 D. berteroae BRIP 57900a, t OR019752 − − OR039642 OR039649 D. betae HMPHU 3001; HHL19101101S2D1, t MW882216 MW882219 MW882225 MW882222 MW882228 D. biguttulata CFCC 52584, t MH121519 MH121437 MH121477 MH121561 MH121598 D. brasiliensis CBS 133183, t KC343042 KC343284 KC343526 KC343768 KC344010 D. breyniae CBS 148910, t ON400846 ON409189 ON409187 ON409188 ON409186 D. brideliae CBS 148911, t OR348649 OR468837 OR468807 OR468817 OR468827 D. brumptoniae BRIP 59403a, t OM918702 − − OM960611 OM960629 D. butterlyi BRIP 59194a, t OR019753 − − OR039643 OR039650 D. caatingaensis CBS 141542, t KY085927 − − KY115603 KY115600 D. cameroonensis CBS 148913, t OR348650 OR468836 OR468806 OR468816 OR468826 D. camporesii JZB320143 − − − MN984254 MN561316 D. caryae CFCC 52563, t MH121498 MH121422 MH121458 MH121540 MH121580 D. cerradensis CMRP 4331, t MN173198 MW751655 MW751663 MT311685 MW751671 D. cha SAUCC 1562, t MT199863 MT366912 MT376714 MT376682 MT376652 D. chinensis MFLUCC 19-0101, t NR_172435 MW294199 − MW205017 MW245013 D. chiangraiensis MFLUCC 17-1669, t MF190119 − − MF377598 − D. chimonanthi SCHM 3614, t AY622993 − − − − D. cichorii MFLUCC 17-1023, t KY964220 KY964133 − KY964176 KY964104 D. cinnamomi CFCC 52569, t MH121504 − MH121464 MH121546 MH121586 D. citriasiana ZJUD 30, t JQ954645 KC357491 KJ490515 JQ954663 KC357459 D. convolvuli CBS 124654, t KC343054 KC343296 KC343538 KC343780 KC344022 D. convolvuli FAU649 KJ590721 KJ612130 KJ659210 KJ590765 − D. convolvuli MF-SOR 61.20.1 PX567840 PX585096 PX585101 PX585106 PX585111 D. convolvuli MF-SOR 61.20.2 PX567841 PX585097 PX585102 PX585107 PX585112 D. coracoralinae FCCUFG 38; URM 8912, t PP192078 PP408214 PP421133 PP430449 PP402241 D. cucurbitae DAOM 42078, t KM453210 − KM453212 KM453211 KP118848 D. cuppatae CBS 117499, t KC343057 KC343299 KC343541 KC343783 KC344025 D. cyatheae YMJ 1364, t JX570889 KC465410 − KC465406 KC465403 D. cylindriformispora MFLU 18-2292, t NR_191275 − − OR140382 OR189510 D. desmotis CGMCC 3.28203, t PQ321216 PQ336561 PQ336579 PQ336525 PQ336543 D. destruens ZJUPD02 KP990617 − KP990637 KP990647 KP990627 D. discoidspora CGMCC 3.17255; ZJUD 89, t KJ490622 − KJ490564 KJ490501 KJ490443 D. elaeagni-confertae SAUCC 194.47, t MT822575 MT855656 MT855544 MT855888 MT855772 D. eleutherrhenae 01, t OK017069 − − OK017070 OK017071 D. elizabethblackwelliae BRIP 74769a, t OR290128 − − OR335747 OR352596 D. fici-macrocarpae SAUCC 0412, t PQ321225 PQ336570 PQ336588 PQ336534 PQ336552 D. fici-septicae MFLUCC 20-0178; KUMCC 19-0151, t MW114348 − − MW192211 MW148268 D. fohaiensis SAUCC 194.113, t MT822641 MT855720 MT855608 MT855953 MT855838 D. foliorum CMRP1321, t MT576688 MT584341 MT584338 MT584310 MT584327 D. ganjae CBS 180.91, t KC343112 KC343354 KC343596 KC343838 KC344080 D. goulteri BRIP 55657a, t KJ197290 − − KJ197252 KJ197270 D. griceae BRIP 67014a, t OM918694 − − OM960603 OM960621 D. guangzhouensis CFCC 58151, t PP864725 PP938609 PP938613 PP938601 PP938605 D. gulyae BRIP 54025, t NR111615 − − JN645803 KJ197271 D. gulyae MF-Ha17-043 MK024253 MZ671933 MZ671961 MK039421 MK033489 D. guttulata CGMCC 3.20100, t MT385950 MW022470 MW022491 MT424685 MT424705 D. hazeniae BRIP 75103a, t OR122283 − − OR135527 OR135528 D. helianthi CBS 592.81, t KC343115 KC343356 KC343598 KC343841 KC344083 D. hordei CBS 481.92 KC343120 KC343362 KC343604 KC343846 KC344088 D. iberica CECT 21218; TH77, t ON159902 ON364028 ON398819 ON398841 ON364049 D. infecunda CBS 133812, t KC343126 KC343368 KC343610 KC343852 KC344094 D. infertilis CBS 230.52, t KC343052 KC343294 KC343536 KC343778 KC344020 D. jazanensis PPDU28R, t OQ586408 − − OR137582 OR137586 D. juglandigena GUCC 422.16, t OP581229 − − OP688534 OP688559 D. kyushuensis ch-D-1, t AB302250 − − − − D. leucospermi CBS 111980 JN712460 KY435663 KY435653 KY435632 KY435673 D. longicolla FAU 599, t KJ590728 KJ612124 KJ659188 KJ590767 KJ610883 D. longispora CBS 194.36, t KC343135 KC343377 KC343619 KC343861 KC344103 D. lusitanicae CBS 123212 KC343136 KC343378 KC343620 KC343862 KC344104 D. machili SAUCC 194.111, t MT822639 MT855718 MT855606 MT855951 MT855836 D. mayteni CBS 133185, t KC343139 KC343381 KC343623 KC343865 KC344107 D. megalospora CBS 143.27 KC343140 KC343382 KC343624 KC343866 KC344108 D. melongenicola CGMCC 3.27978, t PQ321221 PQ336566 PQ336584 PQ336530 PQ336548 D. melonis CBS 507.78, t KC343142 KC343384 KC343626 KC343868 KC344110 D. middletonii BRIP 54884e, t KJ197286 − − KJ197248 KJ197266 D. minusculata CGMCC 3.20098, t MT385957 MW022475 MW022499 MT424692 MT424712 D. monetii MF-Ha18-049, t MW008494 MZ671939 MZ671965 MW008516 MW008505 D. moorei BRIP 61500b, t OR019755 − − OR039645 OR039652 D. morindendophytica ZHKUCC 22-0069, t ON322897 − ON315027 ON315054 ON315087 D. moriniae BRIP 60190a, t OM918698 − − OM960607 OM960625 D. myracrodruonis URM 7972, t NR_163320 MK205290 − MK213408 MK205291 D. neoarctii CBS 109490, t KC343145 KC343387 KC343629 KC343871 KC344113 D. novem CBS 127270, t KC343156 KC343398 KC343640 KC343882 KC344124 D. nepheliicola MFLU 23-0067, t PP915805 PP911610 − PP911611 PP911608 D. orixiae KUNCC 21–10714; HKAS 121465, t OK283041 OK484485 OK484486 OK432279 OK432278 D. ovalispora CGMCC 3.17256; ZJUD 93 KJ490628 − KJ490570 KJ490507 KJ490449 D. oxe CBS 133186, t KC343164 KC343406 KC343648 KC343890 KC344132 D. pachirae CDA 728, t MG559537 MG559535 − MG559539 MG559541 D. paranensis CBS 133184, t KC343171 KC343413 KC343655 KC343897 KC344139 D. passiflorae GZCC 22-0014 OP056703 − OP150773 OP150542 OP150620 D. passiflorae CBS 132527, t JX069860 − KY435654 − − D. pedratalhadensis FCCUFG 48; URM 8304, t PP192073 − PP421129 PP430438 PP402232 D. phaseolorum AR4203; CBS 139281, t KJ590738 KJ612135 KJ659220 KJ590739 KJ610893 D. pseudobiguttulata CGMCC 3.17248; ZJUD 47, t KJ490582 − KJ490524 KJ490461 KJ490403 D. pseudofoliicola HNCM045, t OR647680 − OR671932 OR671940 OR671948 D. pygmaeae CDP 1370, t PP577992 PP579348 − PP579317 PP579332 D. quercicola CSUFTCC104, t ON076567 ON081670 ON081667 ON081659 − D. racemosae CPC 26646 MG600223 MG600219 MG600221 MG600225 MG600227 D. raonikayaporum CBS 133182, t KC343188 KC343430 KC343672 KC343914 KC344156 D. rizhaoensis CFCC 57562, t OP955930 OP959782 OP959785 OP959767 OP959773 D. rosae MFLUCC 17-2658, t NR_172401 MG829273 − − MG843878 D. rosiphthora COAD 2914 MT311197 MT313691 − MT313693 − D. sackstonii BRIP 54669b, t KJ197287 − − KJ197249 KJ197267 D. schini CBS 133181, t KC343191 KC343433 KC343675 KC343917 KC344159 D. schoeni MFLU 15-1279, t KY964226 KY964139 − KY964182 KY964109 D. sclerotioides CBS 296.67, t KC343193 KC343435 KC343677 KC343919 KC344161 D. serafiniae BRIP 54136, t KJ197273 − − KJ197235 KJ197253 D. shawiae BRIP 64534a, t OM918701 − − OM960610 OM960628 D. siamensis MFLUCC 10_0573a JQ619879 − − JX275393 JX275429 D. sojae FAU635; CBS 139282, t KJ590719 KJ612116 KJ659208 KJ590762 KJ610875 D. stewartii CBS 193.36 FJ889448 JX197415 − GQ250324 JX275421 D. submersa CGMCC 3.24297; GZCC 19-0129, t OP056717 OP150710 OP150786 OP150556 OP150633 D. subordinaria CBS 464.90, t KC343214 KC343456 KC343698 KC343940 KC344182 D. talong MCC-MNH 2727, t OQ123545 − − OR943637 OR943674 D. tarchonanthi CBS 146073 MT223794 − MT223759 − MT223733 D. tecomae CBS 100547 KC343215 KC343457 KC343699 KC343941 KC344183 D. tectonendophytica MFLUCC 13-0471 KU712439 KU749354 − KU749367 KU743986 D. terebinthifolii CBS 133180, t KC343216 KC343458 KC343700 KC343942 KC344184 D. thunbergiicola MFLUCC 12–0033, t KP715097 − − KP715098 − D. trevorrowii BRIP 70737a, t OM918703 − − OM960612 OM960630 D. tulliensis BRIP 62248a, t KR936130 − − KR936133 KR936132 D. ueckerae FAU 656, t KJ590726 KJ612122 KJ659215 KJ590747 KJ610881 D. unshiuensis ZJUD50, t KJ490585 − KJ490527 KJ490464 KJ490406 D. vangoghii MF-Ha18-046, t MW008492 MZ671937 MZ671963 MW008514 MW008503 D. vargemgrandensis FCCUFG 58; URM 8784, t PP192069 PP421068 PP421135 PP430456 PP421092 D. vexans CBS 127.14, t KC343229 KC343471 KC343713 KC343955 KC344197 D. vochysiae LGMF1583, t MG976391 MK007528 MK033323 MK007526 MK007527 D. yunnanensis CGMCC 3.18289; LC6168, t KX986796 KX999290 KX999267 KX999188 KX999228 Diaporthella corylina CBS 121124, t KC343004 KC343246 KC343488 KC343730 KC343972 The studied Diaporthe strains are indicated in bold. t, the ex-type strain. Acronyms of culture collection: AR: isolates in culture collection of Systematic Mycology and Microbiology Laboratory; BRIP: Plant Pathology Herbarium, Department of Employment, Economic, Development and Innovation, Queensland, Australia; CBS: Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands; CDP: culture collection of D.S. Pereira, housed at the Lab Bugworkers | M&B-BioISI | Tec Labs – Innovation Centre, Faculty of Sciences, University of Lisbon, Lisbon, Portugal; CECT: Spanish Type Culture Collection at University of Valencia, Valencia, Spain; CFCC: China Forestry Culture Collection Center, Beijing, China; CMRP: Microbiological Collections of Paraná Network, Paraná, Brazil; COAD: Coleção Octávio Almeida Drummond, Universidade Ferderal de Viçosa, Viçosa, Brazil; CPC : Culture collection of Pedro Crous in Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands; CSUFTCC: Central South University of Forestry and Technology Culture Collection, Hunan, China; DAOM: Canadian Collection of Fungal Cultures, Ottawa, Canada; FAU: Florida Atlantic University, Biological Sciences Department, Davie, Florida, USA; FCCUFG: Coleção de Culturas de Fungos da Universidade Federal de Goiás, Laboratório de Micologia, Instituto de Patologia Tropical e Saúde Pública (IPTSP), Universidade Federal de Goiás, Goiânia, Brazil; GUCC: Culture Collection at the Department of Plant Pathology, Agriculture College, Guizhou University, China; GZCC: Guizhou culture collection, Guiyang, China; CGMCC: China General Microbiological Culture Collection Center, Beijing, China; HMPHU: Herbarium of Microbiology and Phytopathology, Heilongjiang University, Harbin, China; HUMCC: Heilongjiang University Microbiological Culture Collection Centre, Harbin, China; JZB: Beijing Academy of Agriculture and Forestry Sciences Culture Collection, China; KUNCC: Kunming Institute of Botany Culture Collection, Kunming, China; LC: Personal Culture Collection Lei Cai, State Key Laboratory of Mycology, Institute of Microbiology, Chinese Academy of Sciences, China; LGMF: Culture collection of Laboratory of Genetics of Microorganisms, Federal University of Parana, Curitiba, Brazil; MAFF: MAFF Genebank Project, Ministry of Agriculture, Forestry and Fisheries, Tsukuba, Japan; MF: collection of pure cultures of A. A. Jaczewskii Laboratory of mycology and phytopathology of All-Russian Institute of Plant Protection, Saint Petersburg, Russia; MFLUCC: Mae Fah Luang University Culture Collection, Chiang Rai, Thailand; MFLU: Herbarium of Mae Fah Luang University, Chiang Rai, Thailand; SAUCC: Shandong Agricultural University Culture Collection, Shandong, China; SCHM: Mycological Herbarium of South China Agricultural University, Guangzhou, China; ZHKUCC: culture collection of Zhongkai University of Agriculture and Engineering, Guangzhou, China; ZJUD, ZJUPD: Diaporthe strains deposited in Zhejiang University, Zhejiang, China. Phylogenetic analysis
-
Sequences were assembled using Vector NTI advance v. 11.0 (Invitrogen, Thermo Fisher Scientific) and aligned with ClustalX 1.8[27]. The alignments were concatenated using Sequence Matrix[28]. To implement GCPSR and estimate the possibility of combining the set of five loci, single locus trees were generated and compared to detect conflict (data not shown). A multilocus phylogenetic analysis was based on the combined ITS, cal, his3, tef1 and tub2 sequences for all strains studied and all species currently accepted in the Diaporthe section Sojae sensu Dissanayake et al.[4] and Zhang et al.[5], 89 species in total. Additionally, sequences for 34 species published in 2023–2025 that were omitted by these authors were included. The tree was rooted using the ex-type Diaporthella corylina Lar.N. Vassiljeva (CBS 121124). Sequences of representative Diaporthe strains and type species were obtained from GenBank (Table 1).
Phylogenetic analysis of the combined aligned data consisted of maximum likelihood (ML) and Bayesian inference. ML analysis was implemented in the IQ-TREE[29]. Modeltest in IQ-TREE determined the most suitable nucleotide substitution model for the nucleotide dataset according to the Bayesian information criterion (BIC) model TIM2e+F+R5. Bootstrap values with 100,000 replicates were calculated for tree branches. Bayesian inference was performed by MrBayes v. 3.2.1. in ARMADILLO v. 1.1[30] using a Markov chain Monte Carlo (MCMC) sampling method. The general time-reversible model of evolution, including estimation of invariable sites and assuming a gamma distribution with six rate categories was used for Bayesian inference analyses. Four MCMC chains were run simultaneously, starting from random trees for 107 generations and sampled every 1,000 generations. The first 25% of the generations were discarded as burn-in, and Bayesian posterior probabilities (BPP) were calculated from the remaining trees.
Morphology
-
Pure cultures of the studied strains were incubated on PSA, oatmeal agar (OA)[31], and Czapek agar (CZ)[19]. Petri dishes (90 mm in diameter) were placed for 10 days under 12 h near-ultraviolet light/12 h dark to stimulate sporulation. Colony morphology and diameter were examined after 10 days. Observations and measurements of 100 conidia for each strain were conducted with an Olympus SZX16 stereomicroscope (Olympus, Tokyo, Japan) and an Olympus BX53 microscope. Images were captured with a PROKYON camera (Jenoptik, Jena, Germany) with Nomarski differential interference contrast.
Pathogenicity test
-
For the pathogenicity tests, leaf segments of C. arvensis were inoculated with all the studied strains. The mycelial suspension was used as the inoculum. For inoculation, strains were grown in a liquid soybean (Glycine max) nutrient medium with the following composition: KH2PO4 (0.2%), (NH4)2SO4 (0.1%), MgSO4 (0.1%), glucose (2%), and soy flour (1%). In total, 50 mL of the medium in 250-mL flasks was inoculated with three 5-mm mycelial discs cut from 2-week-old colonies grown on PSA and kept on an orbital shaker (200 rpm) for four days at 24 °C. The mycelium was separated from the liquid culture by filtering and then dried with filter paper and ground in sterile water to prepare a mycelial suspension of 100 mg/mL. Visually healthy C. arvensis leaves were collected from their natural habitat and cut into segments (1.5 cm × 1.5 cm) then placed in Petri dishes. Six inoculated leaf segments (replicates) were used per strain. To create a moisture chamber effect, filter paper premoistened with sterile water was added to each Petri dish. A drop of the mycelial suspension (10 µL) was applied to the center of leaves which had previously been wounded with a needle (diameter of damage: 0.5 mm) or intact leaves on the abaxial surface of the leaf. In the control, the wounded and nonwounded leaf segments were treated with 10 µL of sterile distilled water. After inoculation, closed Petri dishes with the leaf segments were kept at room temperature and evaluated for the presence of symptoms at 3, 5, and 7 d post treatment (dpt). Subsequent reisolation of the fungus from the leaf segments and identification were carried out to fulfill the Koch postulates.
-
A multilocus phylogenetic analysis based on the ITS, cal, his3, tef1, and tub2 sequences inferred intraspecific relationships within all species (123) currently accepted in the Diaporthe section Sojae for which nucleotide sequences are available. The total analysis included 129 strains, 2 of which were our strains and 127 were references. The total combined matrix consisted of 2,900 characters with gaps (520 bp, ITS region; 560 bp, cal; 498 bp, his3; 452 bp, tef1; and 870 bp, tub2), among which 616 (21%) characters were conserved, 1,480 (51%) were informative, and 804 (28%) were uninformative.
The analysis of the single-gene phylogenies revealed no conflicts. The individual gene trees, as well as the combined tree (Fig. 1), showed that the studied strains MF-SOR 61.20.1 and MF-SOR 61.20.2 with the ex-type CBS 124654 and representative Florida Atlantic University (FAU) 649 strains clustered in distinct monophyletic clade corresponding to D. convolvuli species.
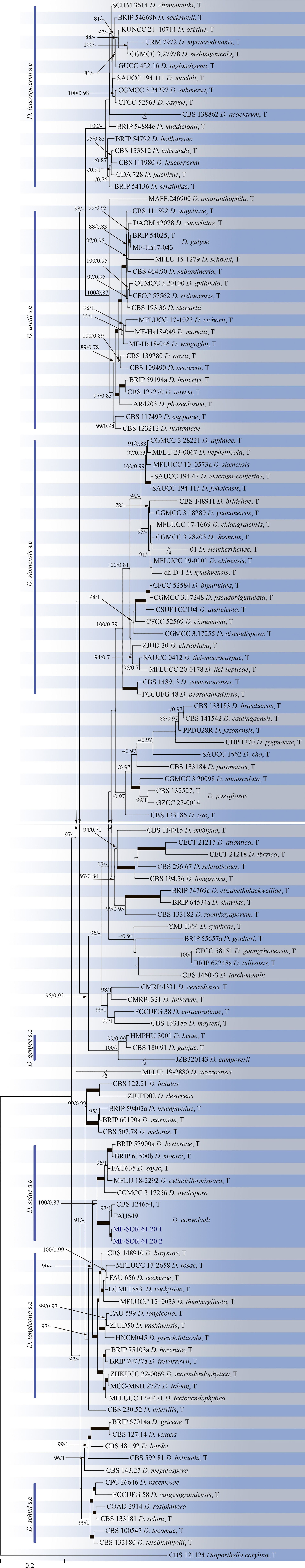
Figure 1.
Phylogenetic tree of Diaporthe section Sojae inferred from a ML analysis based on a concatenated alignment of the ITS region, and partial cal, his3, tef1, and tub2 genes. The ML bootstrap support values (MLBS ≥ 70%) and Bayesian posterior probabilities (BPP ≥ 0.70) are given at the nodes (MLBS/BPP). The studied strains are given in blue. The ex-type strains are marked with T. Purple lines indicate the boundaries of species complexes.
Morphology
-
On all tested nutrient media, the colonies of the strains reached the edge of the Petri dish within 10 days. Colonies on PSA (Fig. 2a) had sparse, silvery-white aerial mycelia generally at the colony periphery. On OA (Fig. 2b), aerial mycelia were absent. On CZ (Fig. 2c), aerial mycelia that were abundant, silvery-white, and floccose covered the entire colony, forming dense cushions. Conidiomata were pycnidial, abundant on all media, and the colonies on PSA and OA appeared mottled because of the presence of numerous pycnidia. Pycnidia were distributed evenly across the colony on all media and formed concentric rings at the periphery on PSA, and were superficial and immersed, solitary or within large conglomerates (Fig. 2d, e), often arranged in long (5–10 pycnidia on PSA and OA; Fig. 2f) or short (on CZ 3–5 pycnidia) chains. Pycnidia (Fig. 2g, h) were subglobose or globose, oval or pyriform, glabrous, ostiolate, and pappilate or with a short neck. On PSA, they were 210–386 (292 ± 14) × 140–359 (249 ± 14) μm; on OA, they were 88–364 (273 ± 17) × 95–277 (200 ± 12) μm; on CZ, they were 230–454 (343 ± 13) × 218–499 (312 ± 17) μm. The pycnidial wall was pseudoparenchymatous, composed of isodiametric cells, with the outer layers pigmented. Conidiophores reduced to phialidic conidiogenous cells (Fig. 2i, j) formed from the inner cells of the pycnidial wall, and were hyaline, smooth, and bottle-shaped, measuring 5.64–15.89 (9.69 ± 0.87) × 3.69–8.97 (5.65 ± 0.47) μm. Only alpha conidia were observed. Alpha conidia (Fig. 2k) were smooth, hyaline, guttulate, and ellipsoidal. On PSA, they were 8.1–16.2 (12.3 ± 0.1) × 3.2–4.7 (3.8) μm; on OA, they were 8.8–14 (11.5 ± 0.1) × 3.2–4.5 (3.8) μm; on CZ, they were 7.5–16.2 (11.8 ± 0.2) × 3.3–5 (4.1) μm. Ascomata were absent.
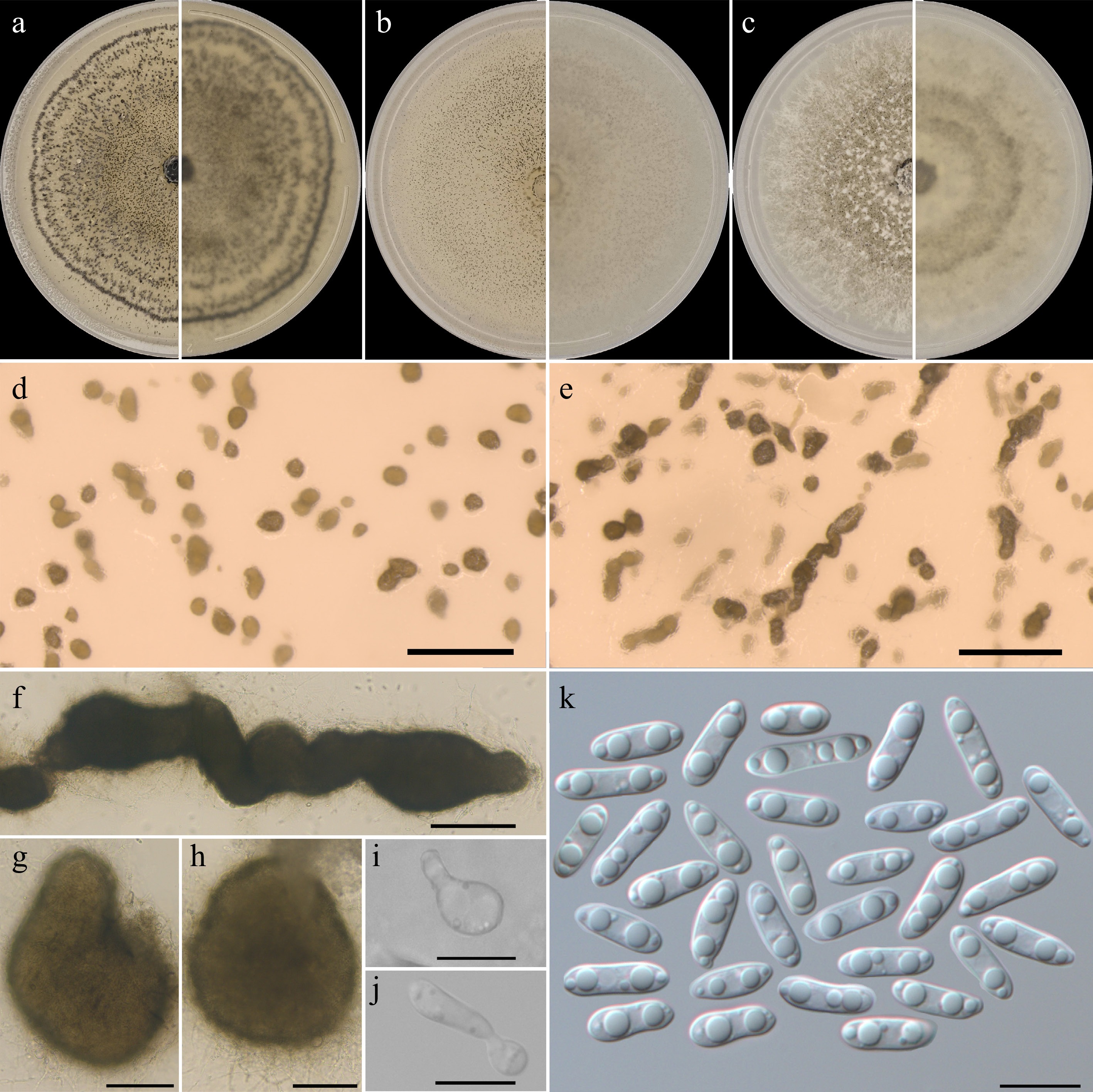
Figure 2.
Diaporthe convolvuli strain MF-SOR 61.20.1 at 10 days of growth. (a)–(c) Pure cultures: left half, front; right, reverse. (a) PSA. (b) ОА. (c) CZ. (d), (e), (g), (h) Pycnidia on ОА. (f) Pycnidia on OA aggregated in chains. (i) Conidiogenous cell. (j) Conidia on a conidiogenous cell. (k) Conidia. Scale bars: (d), (e) = 1 mm; (f) = 200 μm; (g)–(h) = 100 μm; and (i)–(k) = 10 μm.
Pathogenicity
-
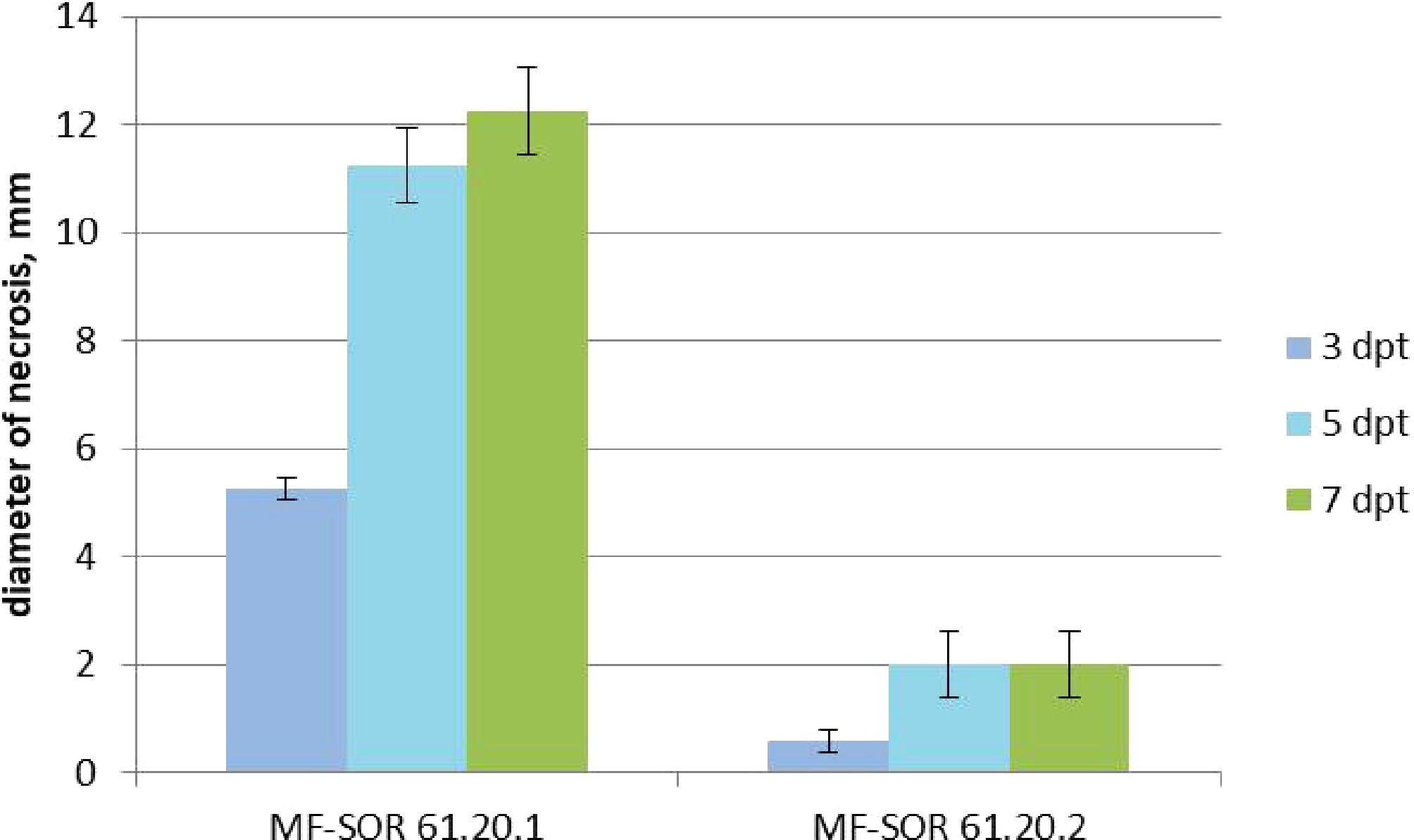
The pathogenicity of D. convolvuli was tested using C. arvensis leaf segments (Fig. 3). No distinct necrosis was observed on the intact sides of the segments. The first significant symptoms, namely necrosis, on the wounded abaxial surfaces of the leaf segments were observed at 3 dpt. At 3 dpt, inoculation with the strain MF-SOR 61.20.2 led to the development of tan to brown necrotic lesions measuring 0.6 ± 0.3 mm in diameter on leaf segments. At 5 dpt, the lesions measured 2.0 ± 0.5 mm. By 7 dpt, lesion size did not increase. In contrast, the strain MF-SOR 61.20.1 induced brown to black necrotic lesions. At 3 dpt, the lesions measured 5.2 ± 0.4 mm in diameter. A marked increase was observed by 5 dpt, with the lesions reaching 11.9 ± 1.3 mm. Subsequently, lesion expansion slowed, and at 7 dpt, the lesions measured 12.25 ± 0.81 mm (Fig. 4). To confirm Koch's postulates, fungal cultures were reisolated in all cases from the necrosis formed on the leaf segments. These isolates had morphological characteristics identical to those of all the strains tested.
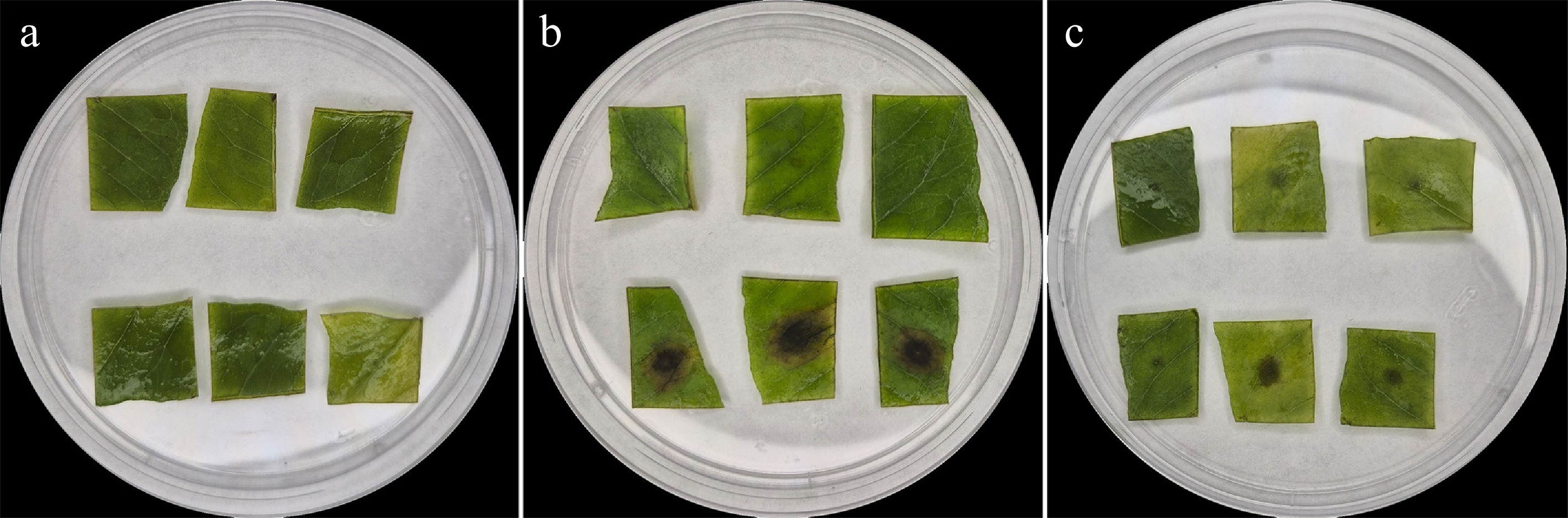

Figure 3.
Pathogenicity of Diaporthe convolvuli on leaf segments of Convolvulus arvensis at 5 dpt. The first line in each set (Petri dish) contains the inoculated intact abaxial part of the leaf segments; the second line is the abaxial part wounded with a needle. Inoculation variants: (a) Inoculation with sterile water; (b) mycelial suspension of MF-SOR 61.20.1; (c) mycelial suspension of MF-SOR 61.20.2.
-
Two studied strains isolated from C. arvensis, MF-SOR 61.20.1 and MF-SOR 61.20.2, were definitively identified as D. convolvuli. Phylogenetically, these strains were closely related to the ex-type (CBS 124654) and representative (FAU 649) D. convolvuli strains, and their morphological features were consistent with that of D. convolvuli[32]. Diaporthe convolvuli was originally described as Phomopsis convolvuli Ormeno-Nunez, Reeleder & A.K. Watson in 1988[32] as the causal agent of leaf spot and anthracnose in field bindweed in Montreal (Canada). To our knowledge, this is the first finding of this fungus in Russia (Saint Petersburg, Pushkin) and the third in the world. The first report from Canada[32] was of a strain that was later patented for the biological control of C. arvensis[33]. The second was from Turkey[34]. Furthermore, this study provides the first description, refinement, and comprehensive illustration of the fungus's cultural and micromorphological characteristics since 1988.
Both strains exhibited pathogenicity to field bindweed leaves under laboratory conditions. This observation suggests that they might be worth investigating further as potential candidates for biological weed control. The fact that these strains were isolated from visually healthy leaves but demonstrated pathogenicity under laboratory conditions indicates a latent infection in the original plant material.
This work was financially supported by the Russian Science Foundation, grant number 25-74-00050.
-
The authors confirm their contributions to the paper as follows: study conception and design, analysis and interpretation of results: Gomzhina MM, Gasich EL; data collection, manuscript review and editing: Gasich EL; draft manuscript preparation: Gomzhina MM. Both authors reviewed the results and approved the final version of the manuscript.
-
The data generated and analyzed during this study are available in this article. DNA sequence data are available in the GenBank database, and the accession numbers are provided in Table 1. The Diaporthe convolvuli MF-SOR 61.20.1 and MF-SOR 61.20.2 strains have been deposited in the the mycological collection of pure cultures at the A.A. Jaczewskii Laboratory of Mycology and Phytopathology, All-Russian Institute of Plant Protection (MF, VIZR, Saint Petersburg, Russia).
-
The authors declare that they have no conflict of interest.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Gomzhina MM, Gasich EL. 2026. First report of Diaporthe convolvuli on field bindweed (Convolvulus arvensis) in Russia. Studies in Fungi 11: e013 doi: 10.48130/sif-0026-0016
First report of Diaporthe convolvuli on field bindweed (Convolvulus arvensis) in Russia
- Received: 24 November 2025
- Revised: 07 April 2026
- Accepted: 14 April 2026
- Published online: 11 May 2026
Abstract: Convolvulus arvensis (field bindweed) is a widely distributed and harmful weed in Russia. The fungal strains MF-SOR 61.20.1 and MF-SOR 61.20.2 were isolated from the visually healthy leaves of C. arvensis collected in northwest Russia (Saint Petersburg, Pushkin) and initially identified by morphological features as Diaporthe sp. The aim of this study was to identify these strains using molecular genetic, micromorphological, and cultural characteristics, and to assess their pathogenicity. Multilocus phylogenetic analysis, based on sequences of the internal transcribed spacer (ITS) of rDNA, as well as partial calmodulin (cal), histone 3 (his3), translation elongation factor 1α (tef1), and β-tubulin (tub2), allowed definitive identification of these strains as Diaporthe convolvuli. Pathogenicity assays demonstrated that strains were pathogenic to C. arvensis, causing necrosis on leaf segments under laboratory conditions. The pathogenicity was confirmed by reisolation of D. convolvuli from infected leaves. To our knowledge, this is the first finding of this fungus in Russia (Saint Petersburg, Pushkin). This study provides the first description, refinement, and comprehensive illustration of the fungus's cultural and micromorphological characteristics since 1988.
-
Key words:
- Diaporthe /
- Phomopsis /
- Field bindweed /
- Identification /
- Molecular phylogeny /
- Pathogenicity