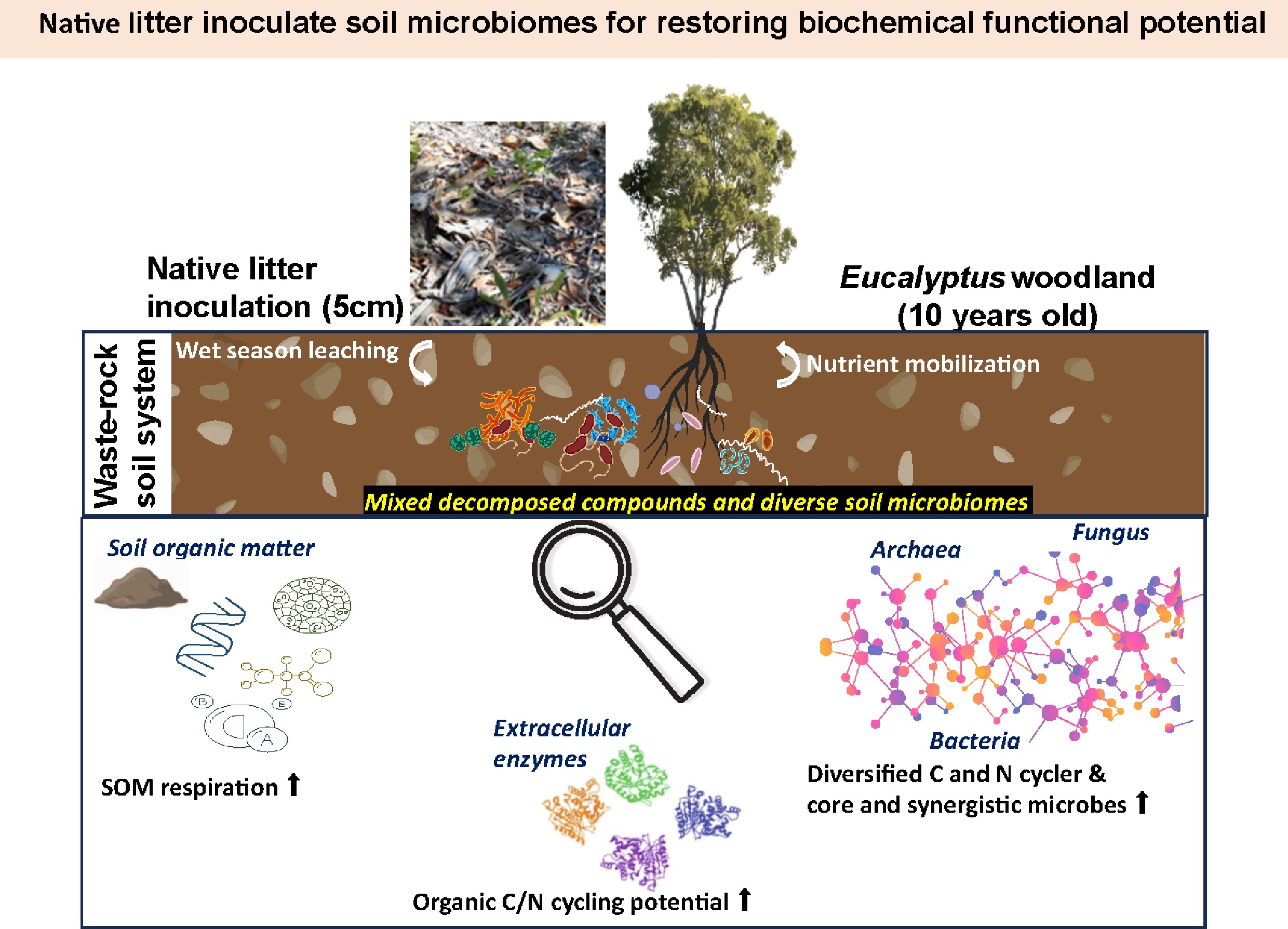
-
Eucalyptus woodland communities are often prioritized for the rehabilitation of remote post-mining landscapes[1,2]. One of the critical aspects for establishing sustainable woodland communities is the development of soil microbiomes capable of decomposing litter with high lignin content and elevated carbon (C) : nitrogen (N) ratios, thereby supporting in situ nutrient cycling and soil organic matter (SOM) formation[3,4]. Soil microbiomes comprise diverse functional groups responsible for litter breakdown and SOM mineralization. In forest soils, organic matter decomposers include bacteria such as Streptomyces, Pseudomonas, Cytophaga, and saprotrophic fungi that are capable of breaking down complex plant polymers[5].
Natural Eucalyptus woodlands generate heterogeneous litter layers of chemically complex organic matter, which require diverse bacterial and fungal communities to collaboratively decompose and mineralize, contributing to SOM formation and nutrient cycling. These microbial communities form intricate ecological networks to process lignin-rich organic matter efficiently[6]. The abundance and interactions of microbiomes in forest soil vary with nutrient availability in soil-litter habitats. For instance, archaea dominate N cycling bioprocesses (i.e., ammonium oxidation, nitrate respiration, and denitrification) in nutrient-poor soils, despite lacking enzymes for cellulose degradation[7]. In contrast, Bacteroidetes thrive in nutrient-rich conditions, whereas oligotrophic taxa like Verrucomicrobia decline with nutrient enrichment[8]. Certain rhizobia (e.g., Mesorhizobium) contribute to litter decomposition and biological N fixation[9]. Among fungi, Basidiomycetes are highly efficient at decomposing woody litter, while arbuscular mycorrhizal fungi (AMF) facilitate plant nutrient uptake through symbiosis but exhibit limited saprotrophic capacity[10].
At the Ranger Uranium Mine in Australia's Northern Territory, the rehabilitation landform trial (TLF) was established using a waste rock (WR)-based soil system planted with woodland tree species, and has supported Eucalyptus-dominated vegetation for approximately 10 years. However, WR-based soils are typically low in available nutrients, moisture, and organic matter, resulting in limited microbial abundance and diversity during the early establishment phase[11]. Consequently, additional remediation strategies are required to establish soil microbiomes similar to those in natural woodland soils, to speed up the decomposition of litter returned from colonizing woody plants.
This study investigates a field-based inoculation approach, whereby native litter from natural Eucalyptus woodlands is introduced into WR-based soils to accelerate the establishment of native soil microbiomes. The use of native litter inoculation represents a highly feasible, low-input strategy for industrial-scale mine rehabilitation. In many mining operations, vast quantities of native vegetation and litter are removed during land clearing; repurposing this material as a biological 'inoculant' during the wet season offers a cost-effective alternative to expensive commercial amendments. By utilizing site-specific organic matter, this approach would potentially stimulate native microbial communities and accelerate the establishment of nutrient cycles without the need for high-maintenance soil imports. The impact of litter inoculation on soil microbial communities and functional potential was evaluated during the wet season, a period characterized by peak biological activity[11] and critical for the establishment of framework tree species in this wet–dry tropical system[12].
Specifically, the study addresses three key questions: (1) Can native litter inoculation effectively restore soil microbial communities and their functional potential? (2) Which microbial taxa are most responsive to this intervention? (3) To what extent can this approach drive WR-based rehabilitated soils toward the structural characteristics and functional potential of native woodland soils? These questions were investigated by integrating three complementary microbial indicators, including taxonomic composition to identify the recruitment of key microbial functional groups, co-occurrence microbial network analysis to measure microbial community interactions, and measurable core soil C and N cycling enzymatic activities to provide a direct biochemical link between microbial presence and actual soil biochemical metabolic potentials. Soil samples were collected at the end of the wet season following litter inoculation to characterize shifts in microbial diversity, community composition, network architecture, and core microbial groups in WR-based soils covered with Eucalyptus species. Changes in microbial phenotypes and functional traits were assessed alongside SOM dynamics and C- and N-cycling enzymatic activities and compared with those of non-inoculated WR-based soils and native woodland soils. The findings are expected to inform the integration of litter inoculation strategies into field-scale rehabilitation practices, enhancing the restoration of soil microbial communities and ecosystem functions in post-mining landscapes.
-
The rehabilitation landform trial was established using a non-hazardous and gravel WR–based soil system and has supported Eucalyptus-dominated vegetation for approximately 10 years[11,12]. For comparison, a natural Eucalyptus woodland within the Georgetown Creek Reference Area[13] was selected for soil sampling and for sourcing native litter material used in inoculation. Overall, TLF-1A exhibits a relatively sparse canopy, reduced understory grass cover, and a patchy distribution of Acacia (leguminous) trees. In contrast, the reference woodland is co-dominated by Eucalyptus and Acacia species, displaying a more developed vegetation structure characterized by higher canopy closure, greater understory diversity, and an accumulated litter layer (Fig. 1).
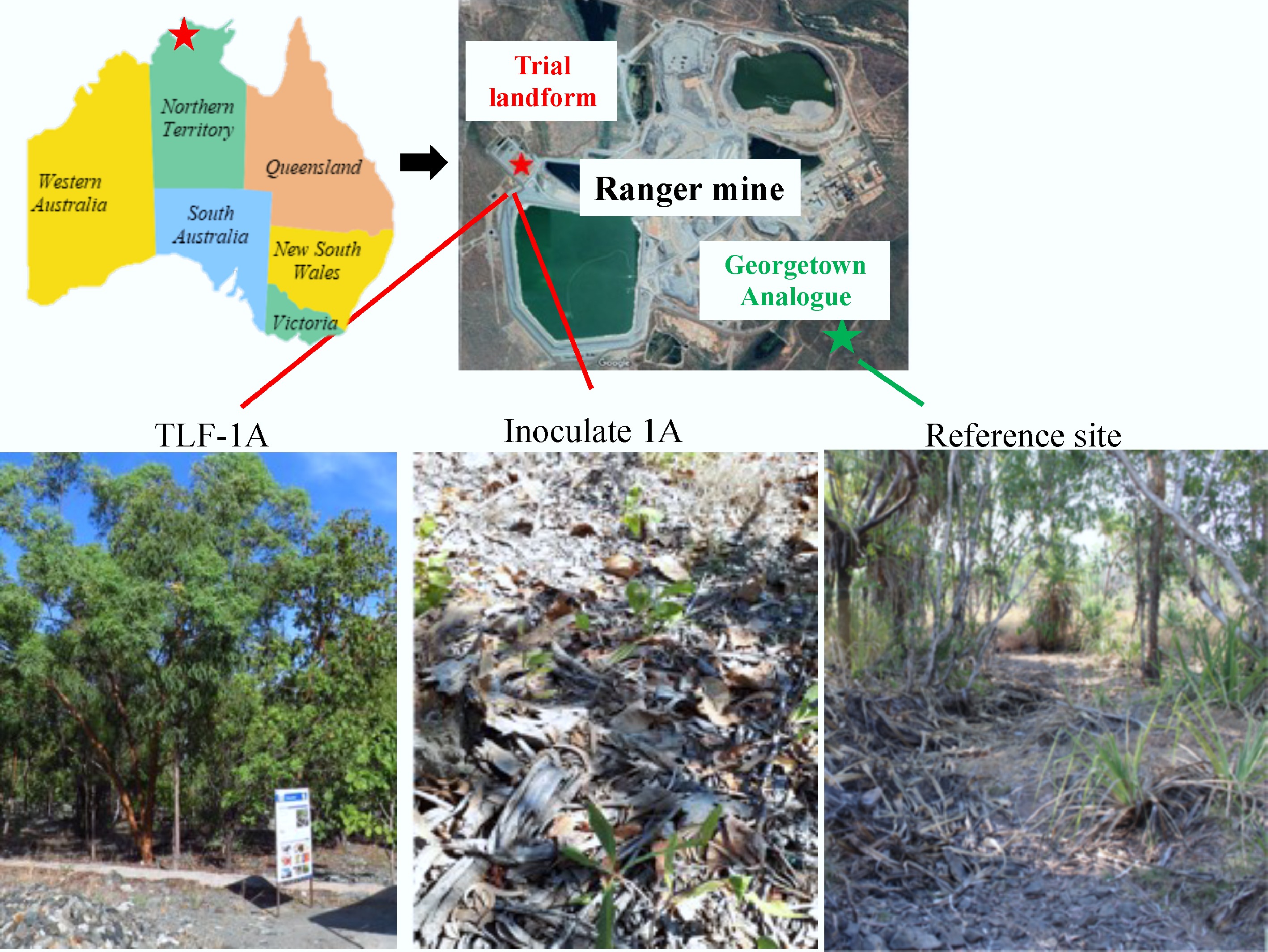
Figure 1.
Map of the Trial Landform (TLF) and Georgetown analogue tropical Eucalyptus open woodland adjacent to the Ranger Mine (Northern Territory, Australia), showing the relative positions of the rehabilitation area and the reference woodland community. Bottom panels show representative field views of sampling sites: TLF-1A, Inoculated-1A, and the reference site.
During the wet season (January), bulk mixed plant litter was collected from the reference woodland dominated by Eucalyptus, with contributions from Acacia species and diverse native grasses. The litter was pre-characterized prior to application and contained 39.5%–45.4% TOC and 0.6%–0.7% N, corresponding to a C:N ratio of 54 to 67. Macronutrient concentrations were 0.03% phosphorus (P), 0.07% potassium (K), 1.4% calcium (Ca), and 0.9% iron (Fe), with moisture content ranging from 7.5%–10.2%. Basal microbial respiration averaged 5.8 mmol CO2/d/kg. Microbial profiling using the amplicon sequencing method revealed that the litter harboured abundant bacterial taxa, dominated by Proteobacteria (41%), Actinobacteria (34.0%), and Acidobacteria (16.5%). The fungal community was dominated by Ascomycota (48.8%) and Basidiomycota (44.2%).
Prior to field application, all litter was thoroughly homogenized to ensure consistency of substrate composition and microbial inoculum. The litter-inoculated treatment was applied to two representative plots in duplicate. This targeted approach was necessary to ensure treatment uniformity, as the high volume of native litter required made it imperative to maintain a consistent application depth and coverage to minimize uncontrolled experimental variance. The homogenized native litter was evenly applied to two 20 m × 20 m plots with two representative plant compositions and coverage located in the central section of TLF-1A. The litter was spread across the soil surface to a thickness of 5 cm, and treated plots were subsequently sampled 15 weeks after application. Over this incubation period, regional climatic conditions included monthly air temperatures ranging from 23.7–37.6 °C and rainfall between 161 and 398 mm (Bureau of Meteorology, Station 014090, Rangers NT).
At the end of the wet season incubation, surface soils (0–5 cm) were collected after carefully removing the litter layer from three sites: (1) untreated TLF-1A plots; (2) litter-inoculated plots within TLF-1A (hereafter 'Inoculated 1A'); and (3) the reference natural woodland (Reference Site). To account for the high natural heterogeneity of established systems, five plots (with three quadrats each, n = 15) were sampled for both the reference woodland and the untreated WR-based soils. This provided a comprehensive landscape-scale baseline. Each sampling plot containing three 20 m × 20 m quadrats, was established in both the untreated TLF-1A and the reference woodland, positioned near trees of similar age with relatively uniform litter cover. For the inoculated treatment, two sampling plots with duplicate quadrats were sampled (n = 4). These surface soils comprised weathered rock fines mixed with organic particulates and small fragments of decomposing litter. Samples were sealed in heavy-duty polyethylene bags and stored at 4 °C for overnight transport to the laboratory. Subsamples designated for molecular analyses were immediately frozen at –80 °C for subsequent microbial DNA extraction.
Soil biochemical analysis and microbial activities
-
Air-dried soils were sieved (< 2 mm) before analysis. Soil pH and electrical conductivity (EC) were determined in a 1:5 (w/v) soil-to-water suspension using a pH electrode (TPS 900-P) and an EC meter (TPS 2100), respectively. Total organic carbon (TOC) and total nitrogen (TN) were measured by dry combustion using a LECO 928 Series Macro Determinator (LECO Corporation, USA) after samples were finely ground. Water-soluble organic carbon (WSOC) was quantified from a 1:10 (w/v) soil-to-water extract using a Shimadzu TOC analyzer. For mineral N, soils were rehydrated to 60% field capacity and pre-incubated in the dark at 25 °C for 7 d. Extracts (1:2 soil-to-solution ratio) were analyzed for NH4+, NO2−, and NO3− using a continuous-flow autoanalyzer (Skalar Analytical, Breda, The Netherlands). Available P was extracted with 0.5 M NaHCO3 (pH 7.0; 1:30 soil-to-solution ratio) and determined colorimetrically using the molybdate blue method. Total P and sulfur (S) in the samples were determined following digestion with the aqua regia method, and both the digest and water-soluble S were subsequently analyzed using inductively coupled plasma optical emission spectroscopy (ICP-OES) (iCAP 7000 Series, Thermo Scientific, Waltham, MA, USA).
Sucrose-induced respiration (SIR) was measured as an indicator of potential microbial activity. Fresh soil (50 g) was adjusted to 60% field capacity with 100 mM sucrose solution, and CO2 evolution was immediately quantified using the Qubit Q-box SR1LP Soil Respiration Package (Qubit Systems Inc., Canada) under a closed-flow configuration. The N mineralization rate (NMR) was calculated from the difference in mineral N between two extractions (1:2 ratio) performed after 7 and 31 d of incubation (25 °C, dark, 60% field capacity).
Activities of β-1,4-glucosidase (βG), cellobiohydrolase (CB), and L-leucine aminopeptidase (LAP)—key enzymes in C and N cycling—were assayed using fluorogenic substrates labelled with 4-methylumbelliferone (MUB) or 7-amino-4-methylcoumarin (MUC)[14]. Specifically, 4-MUB-β-D-glucopyranoside, 4-MUB-β-D-cellobioside, and L-leucine-7-AMC were used for βG, CB, and LAP, respectively. For each assay, 5 g of pre-incubated fresh soil was suspended in 45 mL of 50 mM sodium acetate buffer (pH 6.5). Aliquots (800 µL) of soil suspension were dispensed into 1.2 mL deep-well 96-well plates with 200 µL of substrate solution (200 µM) or standard (0–100 µM). Plates were incubated at 25 °C for 3 h, centrifuged, and 200 µL of supernatant was transferred to black 96-well microplates. Fluorescence was measured at 365 nm excitation and 445 nm emission using a CLARIOstar Plus plate reader (BMG Labtech, Germany).
Soil microbiomes phylogenetic profiling
-
Soil DNA (stored at −80 °C) was extracted using the DNeasy PowerSoil Kit (Qiagen, Germany) following the manufacturer's protocol. Extracted DNA was submitted to the Australian Centre for Ecogenomics (University of Queensland) for paired-end Illumina MiSeq sequencing. As previously described[11], the V6–V8 region of the 16S rRNA gene was amplified with primers 926F and 1392R, while fungal ITS regions were amplified with primers ITS3 and ITS4. All raw reads are available in the NCBI SRA under BioProject PRJNA675659. Sequence quality was assessed using FastQC, and paired-end reads were processed in QIIME2 (v2022.4) as previously described, with adapters and primers, and low-quality sequences trimmed and removed. Amplicon sequence variants (ASVs) were inferred using Deblur, which also removed chimeric and artefactual reads. ASVs representing < 0.1% relative abundance were excluded to minimize potential cross-sample contamination. Taxonomic assignment was performed using UCLUST against the Greengenes2 (2022.10) database for 16S rRNA, and the UNITE v8 database (February 2019) for ITS sequences. Quality-filtered ASV tables were rarefied to 15,142 (16S) and 5,943 (ITS) reads per sample and imported into R for diversity analysis. α-diversity (ASV richness) was calculated using the vegan package. β-diversity was assessed using Bray–Curtis dissimilarities on square root–transformed relative abundances and visualized via principal coordinate analysis (PCoA). Abundant microbial families were visualized with the ComplexHeatmap package.
Soil core microbiomes and co-occurrence network analyses
-
Core archaeal, bacterial, and fungal taxa were defined as ASVs present in > 80% of all samples. Core ASV numbers were counted, and their cumulative relative abundances were calculated per sample. Taxonomic identities of core members were refined using BLASTN searches against reference databases. Shared core ASVs among sites were visualized using the VennDiagram package[15].
To infer potential microbial co-occurrence patterns, site-specific networks were constructed using the top 250 most abundant prokaryotic and fungal ASVs using pairwise Spearman's rank correlations to generate correlation and significance matrices. Only robust (Spearman's correlation coefficient |ρ| > 0.6) and statistically significant correlations after 'FDR' for false discovery rate correction (p < 0.05) were retained. Network visualization and topological analyses were performed with igraph[16]. Randomized networks were also generated (using a fixed seed of 123) for comparison. Randomized networks (seed = 123) were generated for structural comparison. Network modules (≥ 5 nodes) were identified using the fast-greedy modularity optimization algorithm, and module-wise relative abundances were calculated. Key topological indices—average degree, clustering coefficient, average path length, and graph density—were derived in igraph to characterize network structure and node-level connectivity.
Statistical analysis
-
All statistical analyses were performed in R. Both univariate and multivariate approaches were used to assess site effects. For univariate comparisons, homogeneity of variances among treatments was assessed for each response variable using Levene's test. When variances were heterogeneous, treatment effects were evaluated using Welch's ANOVA, followed by Games–Howell post-hoc tests. When the assumption of homogeneity was met, a standard one-way ANOVA with Tukey's HSD was applied. For taxa where Levene's test could not be computed due to zero variance or low prevalence, Welch's ANOVA was used by default. Variance and significance of site effects on microbial community heterogeneity were evaluated using PERMANOVA (adonis function, vegan package) and PERMDISP. P-values were adjusted for multiple comparisons using the Benjamini–Hochberg false discovery rate (FDR) procedure. Relationships among microbial community structure, geochemical parameters, and soil functional responses to litter inoculation were examined using distance-based redundancy analysis (db-RDA). Statistical significance was determined at p < 0.05.
-
Across all sites, surface soils were non-saline and nutrient-poor, reflecting the inherently oligotrophic characteristics of the WR-based soil system (Supplementary Table S1). Soils at the TLF site exhibited several biochemical properties comparable to those of reference woodland, but with distinct deficiencies in key functional attributes. For instance, TOC was substantially lower in the TLF soils (1.9% ± 1.1%) than in the reference site (4.5% ± 2.1%), indicative of limited SOM in the WR-based soils. TOC was closely associated with WSOC, mineral N, and microbial activity metrics, including SIR, NMR, and C- and N-degrading enzyme activities (Fig. 2). Collectively, these patterns highlight the constrained microbial functionality in the surface layers of both TLF-1A and Inoculated 1A soils.
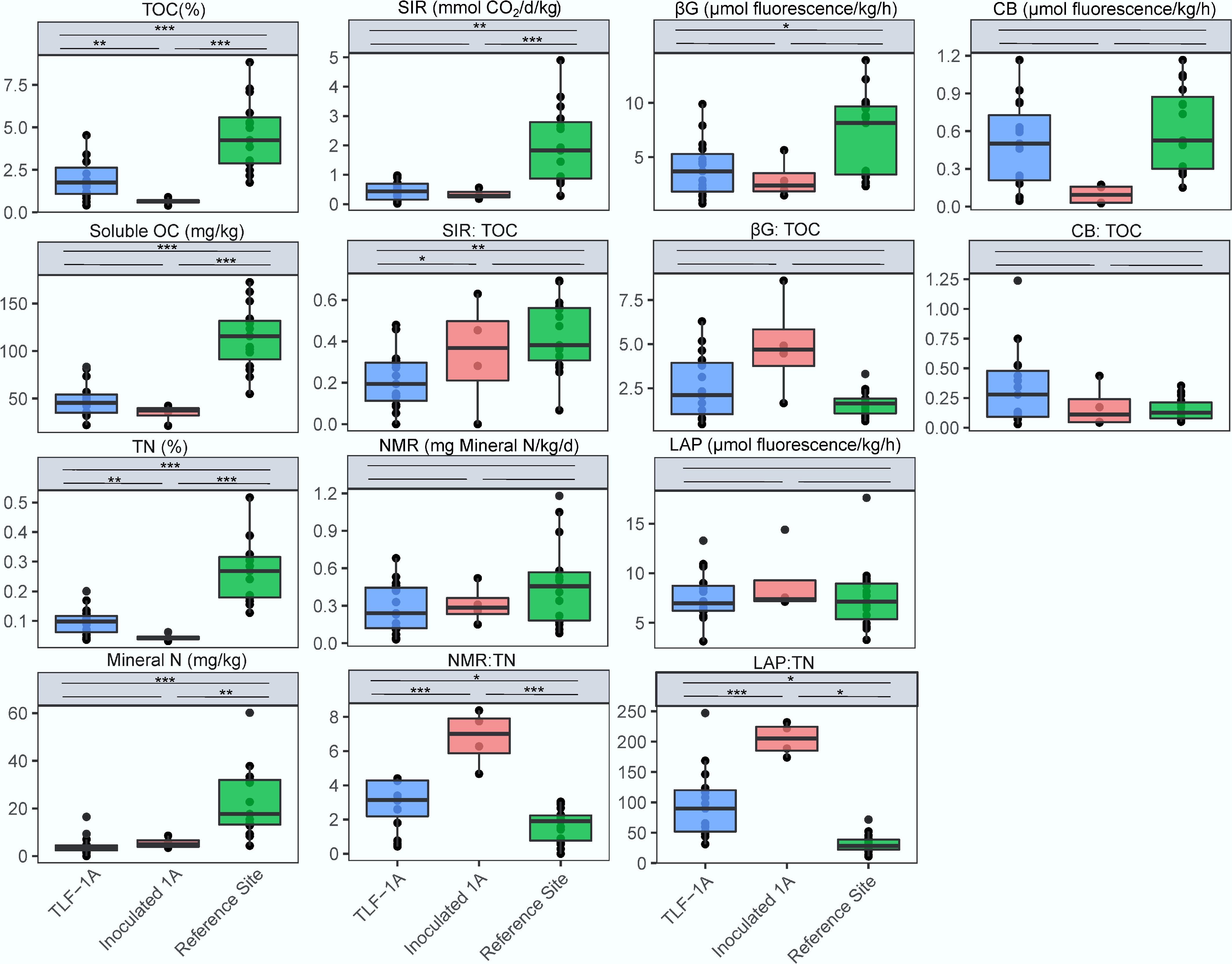
Figure 2.
Total organic carbon (TOC), sucrose-induced respiration rate (SIR), and enzymatic activities of soils for β-1,4-glucosidase (βG), cellobiohydrolase (CB) and the ratio of SIR:TOC, βG: TOC and CB: TOC. Total nitrogen (TN), mineral N, net N mineralization rate (NMR), L-leucine aminopeptidases activities (LAP) and the ratio of NMR:TN and LAP: TN. Enzymatic activities were measured following the MUB/MUC-substrate fluorometric procedure and reported in mmol fluorescence produced per hour per kg of soil. Significant differences between treatment means were determined using post-hoc comparisons with ***, **, * above the line indicating p < 0.001, p < 0.01, and p < 0.05, respectively. For data meeting the assumption of homogeneity of variance, Tukey's Honestly Significant Difference (HSD) test was employed. For heteroscedastic data where Welch's ANOVA was used, the Games-Howell post-hoc test was applied to account for unequal variances and sample sizes.
Litter inoculation significantly and rapidly modified soil C dynamics, reflecting a pulse of microbial metabolic responsiveness at the end of the wet season. Following litter inoculation, the WR-based soils exhibited a notable decline in TOC (Fig. 2). When normalizing to this total soil C pool, the SIR-to-TOC ratio increased markedly, indicating an elevated respiratory demand per unit of soil OC. Enzymatic profiles corroborated these patterns: the absolute βG and CB activities remained low, and the βG-to-TOC ratio increased in WR-based soils after litter inoculation, while the CB-to-TOC ratio remained relatively stable across sites. These findings suggest a clear divergence in the response of key microbial groups and metabolic functions to short-term litter inputs.
Soil N cycling processes also responded strongly to litter inoculation. Across all sites, TN and mineral N pools (NH4+ and NO3−) remained significantly lower in TLF and inoculated soils than in the reference woodland (Fig. 2). NMR and LAP activities—key indicators of organic N mineralization—were comparable between TLF soils and the reference site. When normalized to TN, both the NMR-to-TN and LAP-to-TN ratios were substantially higher in the inoculated soils, indicating greater microbial potential in organic N mineralization and nutrient mobilization.
Taken together, these findings demonstrate that litter inoculation enhanced microbial functional potential by triggering biological SOM decomposition, increasing C turnover, and improving N mineralization relative to available nutrient pools, despite limited accumulation of bulk organic matter in the WR-based soils.
Pattern of soil microbial diversity and co-occurrence network
-
Sequencing depth was sufficient to capture the diversity of soil microbiomes across all sites (Supplementary Fig. S1). In total, 308,079 high-quality 16S rRNA and 2,498,992 ITS rRNA sequences were obtained, clustering into 232 archaeal, 8,907 bacterial, and 2,195 fungal ASVs. Bacterial richness was highest, followed by fungi and archaea (Fig. 3a).
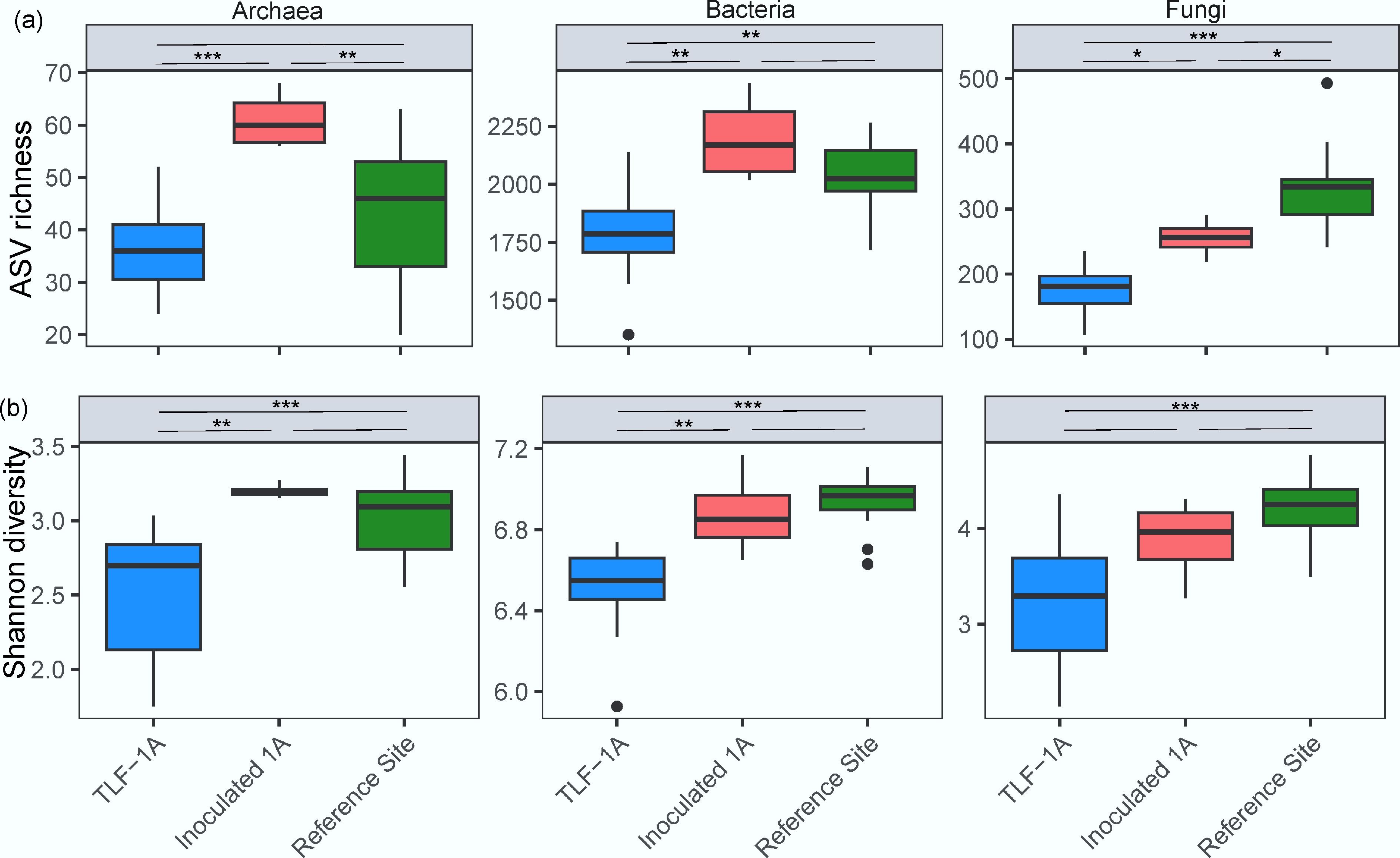
Figure 3.
Box plots of (a) ASV richness, and (b) Shannon index of archaeal, bacterial, and fungal communities in surface soil from TLF-1A, Inoculated 1A and the reference site. Box plots show quartile values. Significant differences between treatment means were determined using post-hoc comparisons with ***, **, and * above the line indicating p < 0.001, p < 0.01, and p < 0.05, respectively. For data meeting the assumption of homogeneity of variance, Tukey's Honestly Significant Difference (HSD) test was employed. For heteroscedastic data where Welch's ANOVA was used, the Games-Howell post-hoc test was applied to account for unequal variances and sample sizes.
Microbial diversity in TLF soils remained lower than in the reference woodland, particularly for bacteria and archaea. Fungal richness also declined in the WR-based soils, though to a lesser extent. Litter inoculation increased microbial richness and evenness across domains. In Inoculated 1A soils, archaeal and bacterial richness rose significantly to levels comparable to the reference site, while fungal richness also increased modestly. Shannon indices followed similar trends, suggesting that litter addition enhanced both richness and evenness (Fig. 3b). Overall, these results indicate that litter inoculation partially restored microbial diversity, narrowing the difference between WR-based and native reference soils.
Litter inoculation also influenced microbial co-occurrence patterns. Network analysis revealed distinct topological shifts among sites and microbial domains (Fig. 4). In the reference soils, the archaeal–bacterial network exhibited high connectivity (average degree = 18.3), a short path length (2.1), and moderate clustering (0.59), consistent with a well-integrated community. The TLF-1A network showed similar modularity (4) but higher clustering (0.68) and density (0.11), with a slightly longer path length (2.4) associated with tighter local associations and lower overall integration. Following litter inoculation, the prokaryotic network became more modular (23) but less connected (average degree = 6.7; density = 0.04), indicating increased compartmentalization and reduced global linkage (Supplementary Table S2). Fungal networks exhibited broadly similar patterns. The reference network was moderately connected (average degree = 11.1; clustering = 0.55), while the TLF-1A network was less connected (average degree = 9.7) and more modular (9). After litter inoculation, modularity (20), connectivity (12.6), and clustering (1.00) all increased, reflecting denser local interactions among fungal ASVs. These structural shifts likely reflect niche differentiation and localized microbial interactions under nutrient-limited WR-based soils associated with litter inoculation.
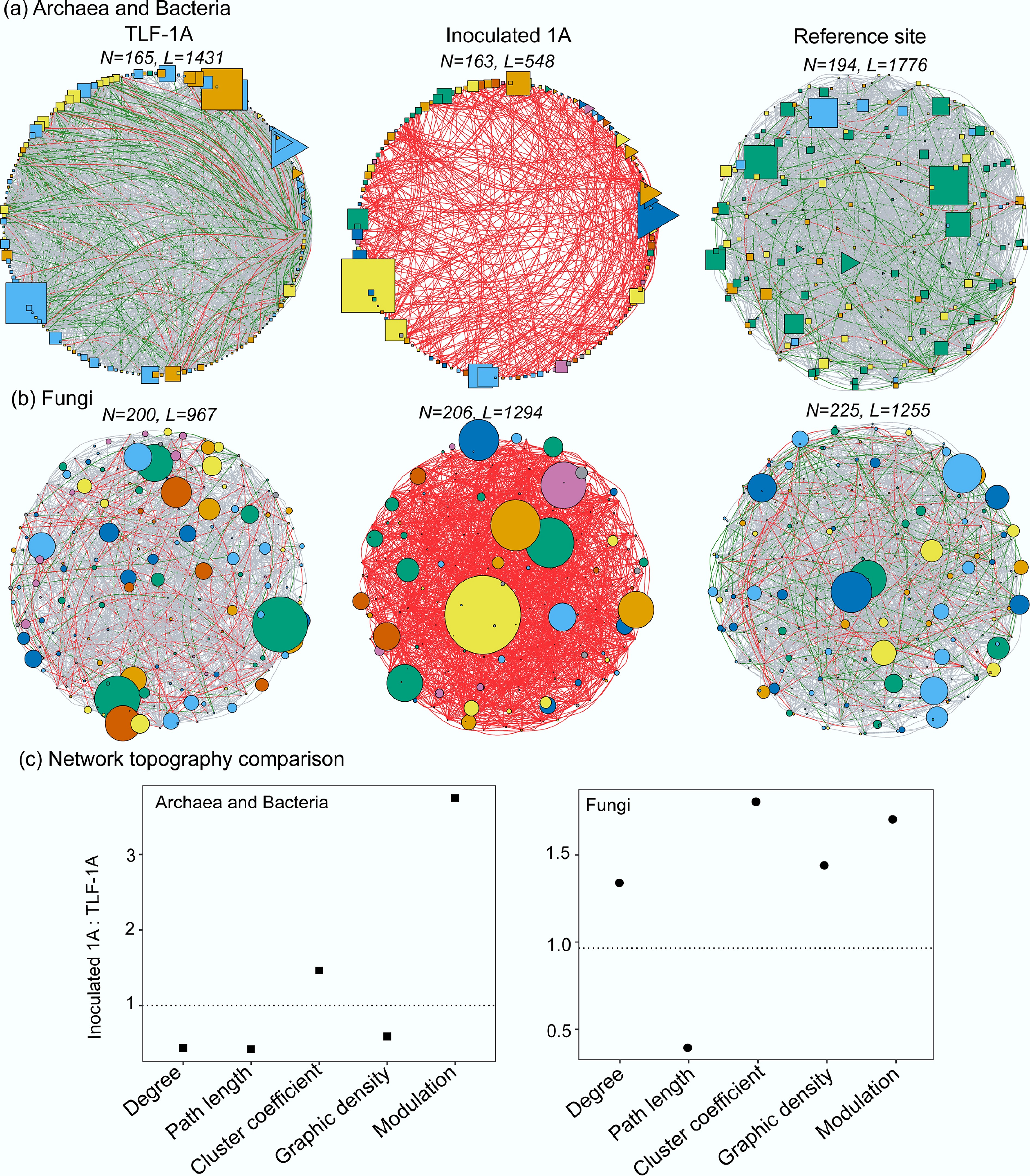
Figure 4.
Patterns of co-occurrence networks for (a) archaeal and bacterial ASVs, and (b) fungal ASVs from TLF-1A, Inoculated 1A and the reference site. The N and L above the networks represent the nodes and edge numbers. The shapes of the network present archaeal (triangle), bacterial (circle) and fungal (square) ASVs. The colors of nodes indicate different network modules with at least five nodes detected using the greedy modularity optimization method. The size of a node is proportional to the relative abundance. The edge connection stands for a strong (Spearman's ρ > 0.6) and significant (p < 0.05) correlation. The color of the edges represent the correlationship coefficient (r > 0.9, red; r: 0.8–0.9, green; and r: 0.6–0.8, grey, respectively). (c) Network topological properties of soil microbiomes change in TLF after litter inoculation.
Soil core microorganisms and functional groups
-
Across all sites, the soil microbiome encompassed three archaeal, 38 bacterial, and ten fungal phyla, collectively representing ~40% of currently recognized microbial phyla[17]. A subset of ASVs formed the core microbiome (> 80% occurrence), representing ecologically and functionally significant taxa. As summarized in Table 1, core microbial ASVs varied among sites and domains, with the lowest richness and relative abundance in rehabilitated TLF-1A soils. Litter inoculation consistently expanded the core microbial pool, particularly for archaea and bacteria. Specifically, in TLF-1A soils, archaeal core ASVs comprised 8.3% of the total community, similar to the reference site; following inoculation, the core archaeal pool increased to 27 ASVs (44.3%). For bacteria, TLF-1A soils shared 101 core ASVs (5.6% of richness), which increased more than six-fold to 677 ASVs (30.8%) after litter inoculation, higher than the reference site (469 ASVs, 23.2%). Fungal core ASVs also rose from 16 (9.2%) in TLF-1A soils to 62 (24.2%) post-inoculation.
Table 1. Summary of the number and percent of core microbial ASVs richness within soil archaeal, bacterial, and fungal communities across the rehabilitated TLF-1A, Inoculated 1A, and reference site soils
Site Archaea Bacteria Fungi No. Richness No. Richness No. Richness TLF-1A 3 8.3% 101 5.6% 16 9.2% Inoculated 1A 27 44.3% 677 30.8% 62 24.2% Reference site 6 14.0% 469 23.2% 30 9.1% Venn diagram analysis identified 33 ASVs shared across all sites, including two archaeal, 25 bacterial, and six fungal ASVs (Supplementary Fig. S2). These taxa accounted for a low overall relative abundance (average 0.46%, range 0.01%–8.5%), yet their functional roles suggest importance for soil biochemical processes. As summarized in Table 2, the two core archaeal ASVs comprised ammonia-oxidizing archaea (AOA, Candidatus Nitrososphaera) and methanogenic archaea, reflecting complementary functions in C and N cycling. Specifically, Candidatus Nitrososphaera is known to mediate ammonia oxidation, a key step in N cycling in acidic to neutral, N-limited soils. The methanogenic archaea, while they participate in the terminal stages of SOM degradation, rely on syntrophic interactions with hydrolytic, fermentative, and acetogenic bacteria[18]. The list of the core bacteria demonstrated their capacity to drive SOM turnover and mineral transformation. More than half of core bacterial ASVs were affiliated with Actinobacteria and Proteobacteria. Several taxa were adapted to extreme or variable soil conditions, including thermophilic Chloroflexi (tolerant to thermal stress)[19], highlighting persistence under fluctuating environmental stress. Other core bacteria, Geodermatophilus[20], Acidimicrobidae[21], and Sulfuriferula plumbiphila[22], are known to be actively involved in chemolithoautotrophic S oxidation. The core fungal ASVs were predominantly Ascomycota, with species such as Aspergillus saccharolyticus, Aspergillus tubingensis, and Curvularia guangxiensis showing high relative abundances and likely contributing to SOM degradation and P solubilization[23].
Table 2. A compiled list of core microbial ASVs, their relative abundance and putative functions in the surface soils of TLF-1A, Inoculated 1A and the reference site
Domain/phyla Closest blast
speciesTLF-1A
(%)Inoculated
1A (%)Reference
site (%)Putative functions Ref. Archaea (total) 7.7 8.9 10.4 Crenarchaeota Candidatus nitrososphaera 3.3 6.1 1.9 Ammonium oxidizer [24] Euryarchaeota Methanogenic archaeon 4.4 2.8 8.5 Methane producer [18] Bacteria (total) 2.63 3.07 4.62 Acidobacteria Arenimicrobium luteum 0.2 0.4 0.7 Obligate aerobic soil organic matter decomposer [25] Acidobacteria bacterium 0.04 0.06 0.09 Soil organic matter decomposer [25] Acidobacteria bacterium 0.11 0.07 0.29 Soil organic matter decomposer [25] Actinobacteria Acidimicrobidae bacterium 0.05 0.03 0.03 Iron reducer [26] Rhabdothermincola sediminis 0.22 0.33 0.07 Thermophilic, aerobic organic matter decomposer [27] Aciditerrimonas sp. 0.04 0.01 0.02 Iron-reducing thermoacidophile [21] Geodermatophilus sp. 0.42 0.27 0.36 Gamma-ray resistant and aerobic rock degrader [20] Geodermatophilus sp. 0.28 0.18 0.87 Stress-tolerant rock degrader [20] Actinomycetospora sp. 0.02 0.07 0.06 Aerobic organic matter decomposer [28] Solirubrobacter sp. 0.18 0.13 0.17 Aerobic soil organic matter decomposer [29] Bacteroidetes Terrimonas soli 0.10 0.38 0.16 Aerobic soil organic matter decomposer [30] Chloroflexi Chloroflexi bacterium 0.04 0.12 0.06 Marine thermophilic heterotroph [19] Bacterium YC-LK-LKJ27 0.06 0.13 0.05 Uncharacterized moderate halophile [31] Gemmatimonadetes Gemmatimonas sp. 0.08 0.07 0.21 Aerobic anoxygenic phototroph [32] Planctomycetes Phycisphaera mikurensis 0.03 0.02 0.02 Marine source anaerobic bacteria [33] Planctomycete LX80 0.06 0.15 0.11 Slow-growing fastidious heterotroph [34] Proteobacteria Sphingomonas horti 0.06 0.04 0.18 Aerobic soil organic matter decomposer [35] Reyranella soli 0.05 0.02 0.10 Microaerobic soil organic matter decomposer [36] Kaistobacter sp. 0.06 0.09 0.21 Soil organic matter decomposer [37] Sulfuriferula plumbiphila 0.31 0.15 0.58 Chemolithoautotrophic sulfur-oxidizer [22] Steroidobacter sp. 0.07 0.12 0.07 Aerobic soil organic matter decomposer [38] Verrucomicrobia Bacterium Ellin518 0.04 0.03 0.07 Soil organic matter decomposer [39] Opitutus sp. 0.04 0.10 0.09 Obligately anaerobic decomposer [40] Lacunisphaera anatis 0.07 0.10 0.05 Organic matter decomposer [41] Fungi (total) 1.72 1.90 2.09 Ascomycota Aspergillus saccharolyticus 0.45 0.42 0.16 Organic matter decomposer [23] Aspergillus saccharolyticus 0.15 0.13 0.12 Organic matter decomposer [23] Aspergillus tubingensis 0.46 0.61 0.41 Organic matter decomposer [42] Aspergillus brunneoviolaceus 0.54 0.36 0.34 Organic matter decomposer with high P solubility [43] Curvularia guangxiensis 0.12 0.38 1.06 Organic matter decomposer [44] Litter inoculation induces a shift of microbial functional groups
-
Rehabilitated TLF-1A and Inoculated 1A soils shared broad taxonomic similarity with the reference site soils, yet PCoA of Bray–Curtis distances revealed clear separation of soil microbiomes between WR-based soils and the reference site (Supplementary Fig. S3), and the WR-based soil microbiomes remain compositionally heterogeneous compared to the reference site soils. Across all soils, archaeal communities were predominantly composed of Crenarchaeota (1.8%–4.4%) and Euryarchaeota (1.5%–8.4%), with a minor contribution from Parvarchaeota (Fig. 5a). Functionally, Ammonia-oxidizing archaea (AOA), primarily Candidatus Nitrososphaera (1.8%–4.4%), consistently outnumbered ammonia-oxidizing bacteria (AOB), including Nitrosospira (0.5%–1.2%), which occurred at low abundance in oligotrophic, N-limited soils[45], while common AOB, e.g., Nitrosomonas, were absent. The AOA:AOB ratio was highest in Inoculated 1A soils (12.2), intermediate in TLF-1A (4.2), and lowest in reference soils (2.0), highlighting the dominant role of AOA in N cycling under nutrient-limited conditions.
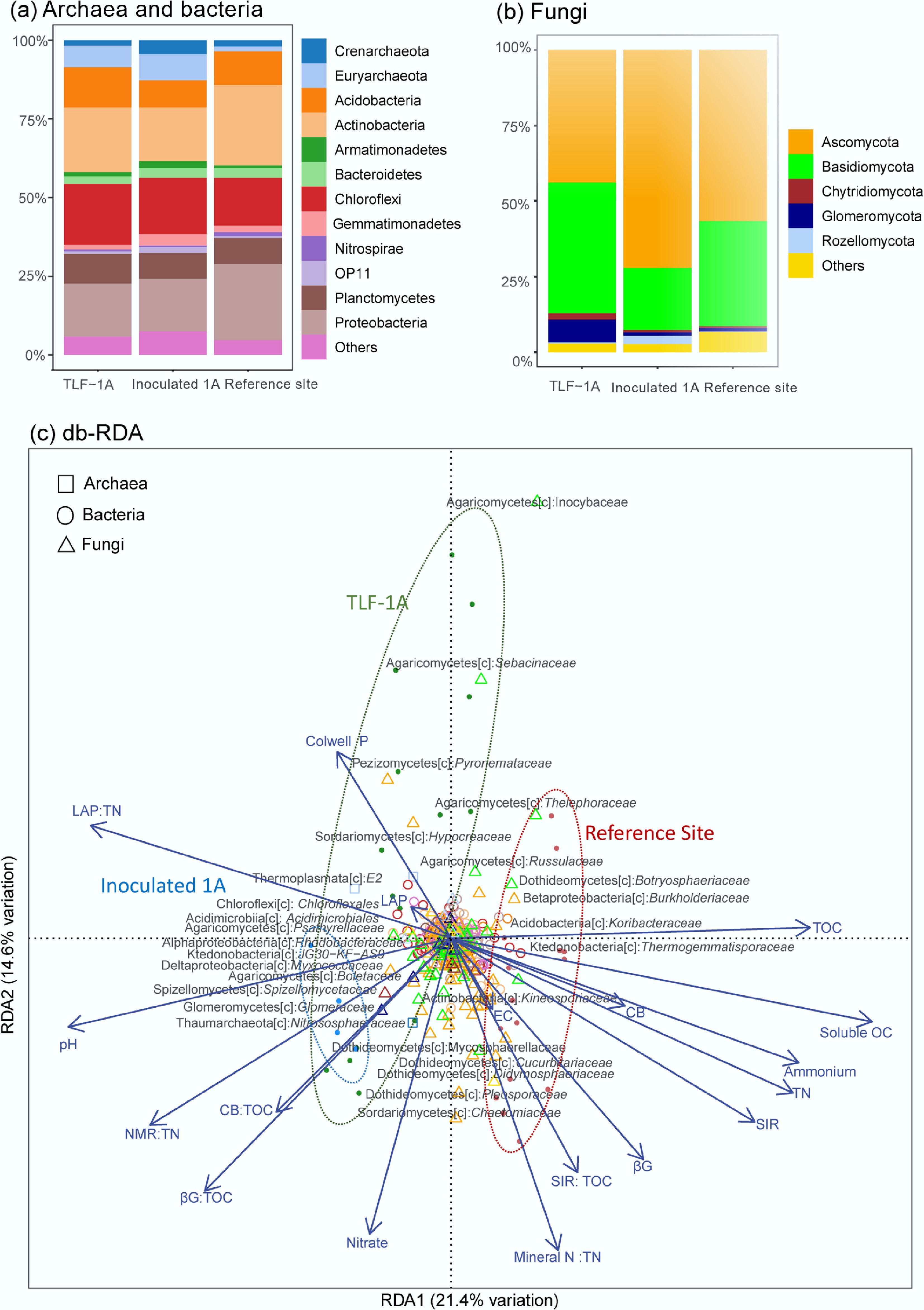
Figure 5.
Relative abundance of soil microbiomes among different sites: (a) Archaeal and bacterial phyla. (b) Fungal phyla. (c) Distance-based redundancy analysis (db-RDA) of soil microbiomes at the family levels and soil biochemistry and function, with archaeal families shown as squares, bacterial families shown as rings and fungal families shown as triangles. The color represents their respective phylum. Each sample is represented by a dot, annotated with its treatment within 95% confidence ellipses, and the environmental drivers are indicated by arrows. The taxonomy annotation is based on the BLAST results in the format of name of class[c]: family group. Abbreviations for soil biochemistry include TOC: total organic carbon; TN: total nitrogen; NMR: net N mineralization rate; SIR: sucrose-induced respiration; βG: β-1,4-glucosidase activity; CB: cellobiohydrolase activity; and LAP: L-leucine aminopeptidase activity.
At finer resolution, db-RDA (Fig. 5c) revealed clear shifts in archaeal composition following litter inoculation. As shown in Fig. 5a, specifically, prior to inoculation, TLF-1A soils were enriched with Thermoplasmata (E2), thermophilic taxa associated with exposed, high-temperature microhabitats[46]. Their prevalence in TLF-1A likely reflects the harsher microclimatic conditions, such as exposed soil surfaces. Following litter inoculation, Thermoplasmata abundance declined, while Crenarchaeota increased, consistent with their known role in soil nitrification under acidic and N-limited conditions[47]. The enrichment of AOA in inoculated soils coincided with higher NO3-N concentrations and elevated N mineralization rates, likely contributing to the enhanced potential for microbial N cycling in these WR-based soils. Bacterial communities were dominated by five phyla—Acidobacteria (9%–13%), Actinobacteria (17%–26%), Chloroflexi (15%–19%), Planctomycetes (8% to 9%), and Proteobacteria (17%–24%)—accounting for 69%–84% of the total bacterial community across all sites (Fig. 5a). At the family level, litter inoculation increased community diversification but did not substantially alter the structure of these dominant groups in TLF-1A soils (Supplementary Fig. S4). Among 338 bacterial families, aerobic SOM-degrading taxa (e.g., Gaiellaceae, Gemmataceae, Hyphomicrobiaceae, and orders RB41/iii1-15) were prevalent, primarily consisting of oligotrophic genera such as Gaiella occulta and Gemmata, which thrive during late-stage litter decomposition when labile substrates are depleted[48]. Phototrophic Chloroflexales remained enriched in rehabilitated soils (Supplementary Table S3). Litter inoculation enriched some putative predatory bacteria Myxococcaceae (Deltaproteobacteria lineage)[49] and JG30-KF-AS9 (a Ktedonobacteria lineage known for its high carbohydrate degradation capacity)[50], while reducing the dominance of Burkholderiaceae (Betaproteobacteria) and Koribacteraceae (Acidobacteria), which are typically associated with plant-rich, acidic soils (e.g., rhizosphere, endosphere, peat soils)[51]. These shifts corresponded with enhanced SOM mineralization and mineral N availability following litter inoculation.
Fungal communities (Fig. 5b) were primarily composed of Ascomycota (43.9%–72.1%) and Basidiomycota (20.5%–34.9%), with smaller contributions from Glomeromycota (1.2%–7.5%), Chytridiomycota (0.5%–2.1%), and Rozellomycota (0.1%–2.8%). Ascomycota, well known for cellulolytic capacity[52], are often replaced by Basidiomycetes during litter decomposition[53]. Ascomycota were less abundant in TLF-1A soils than in the reference site, and litter inoculation significantly increased their relative abundance, reflecting stimulation of cellulose-degrading fungi by fresh organic inputs. In contrast, arbuscular mycorrhizal fungi (AMF, Glomeromycota) were more abundant in TLF-1A soils than in reference soils but declined following litter inoculation, likely due to reduced plant dependence on symbiotic fungi under improved soil conditions[10]. At finer taxonomic resolution, all soils supported abundant saprotrophic fungi. The most abundant group belonged to the Aspergillaceae (the class of Eurotiomycetes), comprising 5% to 6% of the fungal community. This included 48 ASVs closely related to Aspergillus, efficient decomposers of complex polymers and stress-tolerant taxa in fluctuating soils[54], and 24 ASVs closely related to Penicillium, often found in harsh soil environments characterised by low moisture and temperature fluctuations[55]. These taxa likely contribute to maintaining relatively high cellobiohydrolase (CB) functional capacity (Fig. 2). Unlike reference soils, which were enriched with symbiotrophic fungi such as Russulaceae (Pleosporaceae 3.06%), TLF-1A soils were particularly enriched with early-successional and stress-adapted fungi, including Thelephoraceae and Inocybaceae (averaging at 22.8%), Pezizomycetes (Pyronemataceae, 17.42%), and root-associated Glomeraceae (Glomeraceae 3.81%) (Supplementary Table S3). Following litter inoculation, fungal communities shifted toward increased saprotrophic capacity, reduced reliance on AMF, and enrichment of previously rare families.
-
Waste rock-based soils present challenging substrates for ecosystem development due to low SOM and fertility, and extreme physicochemical conditions. Previous studies have demonstrated that, over years of revegetation, WR-based soils can support diverse microbial communities that contribute to SOM decomposition and nutrient cycling. Overall, WR-based soils in this study have begun to approach the properties of native Eucalyptus woodland soils, but substantial differences remain. TOC, TN, and mineral N pools in WR-based soils were < 30% of reference values, and pH was slightly elevated, likely reflecting reduced proton production from root respiration, humification, and nitrification. These persistent differences highlight the challenges of fully restoring WR-based soil ecosystems and emphasize the need for field-based intervention strategies. The results from this study suggest that field-based litter inoculation can accelerate recovery by stimulating SOM transformation and promoting the reassembly of soil microbial communities.
Litter inoculation triggers SOM and enhances functional potential
-
Plant litter composition strongly influences decomposition dynamics, nutrient release, and soil fertility. Previous studies have shown that litter from Eucalyptus-dominated TLF-1A soils contains only ~50% of the N and P found in native woodland litter[11]. In contrast, litter collected for inoculation from a native Eucalyptus woodland—co-dominated by leguminous Acacia and native grasses—provided a substrate rich in cellulose. As Acacia and understory grasses increase in cover, this litter is expected to be relatively low in polyphenols and to contain more readily available N compared with TLF-1A plant litter[56]. Accordingly, wet-season litter inoculation likely supplied some degradable organic compounds and nutrients while introducing a diverse assemblage of native soil microbiota.
Compared with reference woodland soils, WR-based soils have lower canopy cover, which exposes surface litter to stronger UV radiation and photodegradation, which could accelerate litter decomposition[57]. The timing of inoculation coincided with heavy rainfall (1,125 mm, 64% of annual total) and warm temperatures (23.7–37.6 °C) conditions that could enhance leaching of organic inputs, thereby stimulating microbial activity and promoting SOM turnover[58]. In this system, these factors likely triggered mineralization of existing SOM, contributing to the observed decline in TOC following litter inoculation (Fig. 2). This mechanism aligns with the known activity of extracellular enzymes produced by broad-spectrum or specialized soil microorganisms under warm and moist conditions[59]. Supporting this, WR-based soils showed elevated SIR: TOC and βG: TOC ratios after litter inoculation. While these shifts indicate enhanced potential for microbial SOM decomposition, the shrinking TOC pool and subsequent substrate depletion likely triggered stress-induced respiration or energy demands. Without measuring microbial biomass C, it is challenging to distinguish definitively between a surge in population-level functional enhancement and a stress-driven increase in metabolic rate per cell. Expressing enzyme activity and respiration per unit of microbial biomass C would provide a more precise indicator of physiological efficiency. Future research should incorporate biomass data to further decouple the effects of microbial community size from metabolic stress during priming events. This is because bacteria are key drivers of both soil respiration and β-glucosidase production[60]. These shifts suggest that bacterial communities responded strongly to fresh litter inputs. These two elevated ratios likely reflect a combination of rapid bacterial turnover and metabolic costs associated with the depletion of C pools. In contrast, CB, a key enzyme in cellulose degradation predominantly derived from fungal communities[61], remained relatively stable, indicating that the fungal component maintained a steady potential for cellulose degradation. This observation aligns with previous studies suggesting that bacteria play a more prominent role than fungi in early-stage SOM mineralization[5,62].
Substantial evidence indicates that nutrient availability is a major constraint on natural Eucalyptus woodland development in tropical Australia, with improved nutrient supply strongly promoting tree establishment and productivity[63,64]. WR-based soils contained relatively high P concentrations (0.07%), approximately threefold greater than reference soils (0.02%), suggesting alleviation of P limitation (Supplementary Table S1). In contrast, N availability remained low, with ammonium reaching only 17% of reference levels (21 mg/kg; Supplementary Table S1). This imbalance highlights N as a key limiting factor in the Eucalyptus woodland development on WR-based soils, where supply depends heavily on microbial SOM mineralization. During this process, large organic N polymers (e.g., proteins, chitin) must be cleaved into smaller monomers (e.g., amino acids), ultimately determining both the rate and extent of mineral N production[65].
The original litter possessed a wide C:N ratio (54–67), and the litter-inoculated WR-based soils exhibited a significantly lower ratio (14.6 ± 1.7), indicating rapid C mineralization and net N immobilization or concentration following litter inoculation. This study found that when normalized to total N, the soils exhibited disproportionately high potential for NMR and LAP, particularly following wet-season litter inoculation (Fig. 2). According to ecological stoichiometry theory, soil microbiomes maintain a relatively constrained internal C:N ratio (7 to 8) comparing to surrounding soil matrix (17 to 18) in the terrestrial system[66,67]. This creates a 'stoichiometric imbalance' when compared to the WR-based soils, with the C:N ratio lowered following litter inoculation (14.6 ± 1.7). Thus, the soil microbial community remained focused on N acquisition to bridge the gap and could accelerate the breakdown of organic matter, thereby maintaining stoichiometric homeostasis. Consequently, the reduction in soil C:N alongside elevated LAP activity provides strong evidence for enhanced net N mobility and a transition toward a more nutrient-active state in rehabilitated woodlands. Given weaker stabilization mechanisms in WR-based soils—where SOM is less effectively protected within aggregates or bound to clay minerals[68], this makes SOM and associated N more susceptible to microbial decomposition. Consequently, despite low total N pools, WR-based soils demonstrated high potential for rapid N release and dynamic cycling when supplemented with fresh litter, conditions that can temporarily boost fertility, stimulate plant growth, and foster positive plant–microbe feedback during the wet season[11,12].
From an applied perspective, the findings suggest that fresh litter inoculation is a low-input, field-scale strategy to stimulate microbial activity, trigger biological SOM turnover, and enhance short-term C and N cycling potential. These functional gains are particularly relevant for supporting vegetation during the wet season. However, the effect may represent a transient disturbance, with limited long-term benefits unless complemented by continuous organic inputs from litter and root turnover. Therefore, sustained monitoring of SOM pools, microbial networks, and nutrient cycling processes is essential to determine whether litter inoculation can provide lasting improvements and facilitate convergence with mature woodland soils.
Litter inoculation boosts microbial diversity and function recovery
-
Abundant evidence shows that soil functions such as SOM transformation, nutrient mineralization, and structural development are closely linked to microbial diversity, composition, and ecological interactions[69,70]. This linkage is particularly critical for WR-soil development and the long-term sustainability of rehabilitated Eucalyptus woodlands[11]. Rehabilitation programs frequently use reference ecosystems as benchmarks[1], yet it is generally unrealistic to expect WR-based soils microbiomes to exactly replicate the taxonomic composition of reference soils. In this study, known functional microbial groups were integrated with measured metabolic activities, providing a robust biochemical indicator of soil function. Although functional uncertainty remains when assigning specific metabolic roles based on taxonomic identity, the results suggest that litter inoculation promotes a degree of functional equivalence between WR-based soils and reference woodland soils at a broad community scale. These findings provide a practical framework for guiding field interventions aimed at restoring essential biogeochemical processes.
Microbial diversity reflects both environmental selective pressures and the breadth of metabolic capabilities present. In this study, TLF-1A soils exhibited the lowest diversity, indicative of strong selection for stress-adapted taxa (Fig. 3). Litter inoculation significantly increased diversity in WR-based soils, promoting a more balanced and versatile microbiome. Higher richness enhances functional redundancy, wherein multiple taxa contribute overlapping roles in biogeochemical processes—a feature critical for maintaining soil function under environmental stress such as extreme temperatures or drought[69]. Thus, these diversity gains represent progress toward functional recovery, rather than taxonomic convergence. Beyond diversity, the presence of core microbiota plays a pivotal role in maintaining functional stability across soil profiles[71]. In this study, genomic and functional attributes of the core microbial taxa provide further evidence of the functional recovery in the WR-based soils. Although often low in abundance, these groups—spanning C and N cyclers, stress-adapted specialists, and mineral-weathering bacteria—play disproportionate roles in establishing soil chemical and structural properties[72]. Litter inoculation accelerated the reassembly of these core taxa in WR-based soils, increasing overlap with the reference site and suggesting improved persistence of key biogeochemical functions.
Soil microbiomes are inherently complex, multi-trophic systems characterized by diverse substrate preferences, spatial heterogeneity, and intricate interaction networks involving cooperation, competition, and resource partitioning[73]. In WR-based soils, litter inoculation restructured archaeal, bacterial, and fungal networks. Archaeal and bacterial networks exhibited higher modularity (Fig. 4a). It is likely that following the litter inoculation, there is the formation of sub-communities specialized in exploiting distinct substrate pools[74]. This shift likely reflects the influx of labile organic compounds from inoculated litter, which facilitated more efficient resource partitioning, potentially reduced interspecific competition, and enhanced metabolic efficiency and adaptability[75]. However, this structural change represents a critical ecological trade-off: while high modularity promotes specialized metabolic efficiency, it can simultaneously lead to community fragmentation and a lack of module connection, which could reduce overall community resilience. It is probable that the observed high modularity represents a transient successional stage rather than a final equilibrium. Tracking the shift from modular networks toward integrated, resilient communities over several growing seasons will be essential to confirm that the initial benefits of inoculation translate into sustained soil health and successful forest succession. Fungal networks, in turn, displayed increased connectivity and clustering, indicative of cooperative processes such as cross-feeding and metabolite exchange[76]. Because fungi dominate the decomposition of recalcitrant substrates like lignin and cellulose, this restructuring likely produces metabolites such as simple sugars and phenolics that subsequently support bacterial metabolism. The complementarity between bacterial and fungal networks may therefore underpin the sustained SOM turnover and nutrient mobilization observed in WR-based soils.
Litter inoculation also selectively enriched taxa associated with SOM decomposition and N cycling processes, including AOA (Candidatus Nitrososphaera), diverse organotrophic bacteria, and cellulolytic Ascomycota. While functional contributions of these groups require further validation, their enrichment aligns closely with observed improvements in C and N turnover following litter addition (Fig. 5). Metagenomic sequencing and qPCR validation of key functional genes—such as amoA for nitrification, nirK for denitrification—would provide a high-resolution map of the community's metabolic potential. An additional metatranscriptomics study would distinguish between the presence of functional genes and their active expression. Incorporating these molecular tools in future monitoring will be essential to validate whether stable, long-term functional recovery could be achieved with litter inoculation practice. Concurrently, the relative abundance of stress-adapted taxa, such as thermophilic archaea and AMF Glomeraceae, declined. This shift reflects a transition from stress-tolerant to resource-acquisitive microbial strategies, a pattern commonly associated with advancing soil development[25]. Mechanistically, this transition may result from improved soil conditions induced by the litter layer, including enhanced temperature stability, water retention, and UV shading[77], which favor metabolically versatile, functionally active microbes.
However, it is important to note that micro-scale variations in soil moisture, solar radiation, vegetation cover, and root density were not measured concurrently with soil sampling. In field-based systems, these factors represent potential uncontrolled variables that may contribute to the observed spatial heterogeneity in microbial community and enzyme profiles. While this study's findings capture the overall community response to litter inoculation, future studies incorporating high-resolution microclimate sensors and root biomass measurements would be beneficial. Such data would further decouple the direct biochemical effects of litter inputs from the indirect, stabilizing influences of the surrounding plant community and micro-topographical variation.
Overall, litter inoculation provides multiple synergistic benefits for WR-based soils' functional recovery: it increases microbial diversity, promotes core functional soil microbiomes, restructures microbial interaction networks, and drives a compositional shift toward resource-acquisitive strategies. By acting as both a substrate source and ecological stimulus, litter inoculation bridges immediate nutrient deficits while potentially fostering the emergence of a resilient, functionally versatile soil microbiome. While this study captures a relatively short-term successional phase, the observed microbial shifts represent a functional foundation for soil recovery. The narrowed soil C:N ratio likely reduces the nutrient limitation and contributes to the establishment of nutrient feedback loops between the soil and the aboveground ecosystem, especially during the wet season, potentially supporting future plant productivity and canopy development. These findings underscore the importance of microbial-based monitoring in rehabilitation programs. Integration with long-term direct measurements of biogeochemical processes and plant community development will further validate broader ecosystem-level outcomes and guide effective soil and Eucalyptus woodland restoration practices.
-
This study demonstrates that field-based litter inoculation can accelerate soil development on WR-based soils under tropical conditions by enhancing microbial structure and functional potential characteristic of Eucalyptus woodlands. Comparison with adjacent reference woodland soils indicates that a combination of prolonged Eucalyptus dominance and short-term native litter inoculation can stimulate microbiome reassembly and strengthen key biogeochemical processes.
Litter inoculation triggered a pronounced effect on biological SOM decomposition and elevated N cycling capacity. Microbial responses included increased diversity, enrichment of core taxa—particularly ammonium-oxidizing archaea (Nitrososphaeraceae) and diverse decomposers (Bacteroidetes, Ascomycota)—and a reduction of stress-adapted groups such as thermophilic archaea and root-associated AMF. Network analyses indicate improved modularity and connectivity, likely improving both resource partitioning and cooperative interactions. These microbial shifts corresponded with accelerated SOM decomposition and enhanced N cycling potential, supporting early-stage recovery of soil functions necessary for establishing Eucalyptus woodlands with higher productivity and coverage. While total SOM and microbial activity remain lower than in reference soils, litter inoculation provides a scalable, low-input strategy to jumpstart soil functional recovery. Sustained monitoring and repeated litter inputs are critical to consolidate these short-term gains into long-term stabilization of SOM pools and resilient ecosystem functioning.
-
It accompanies this paper at: https://doi.org/10.48130/een-0026-0003.
-
The authors confirm their contributions to the paper as follows: Longbin Huang led the conceptualization, design and fund acquisition. David Parry contributed to funding acquisition, supervised field operations, and assisted with manuscript editing and review. Fang You and Merinda Hall performed the experiments and collected the data. Fang You analyzed the data, interpreted the results, and wrote the manuscript with input from all authors. All authors reviewed, edited and approved the final version of the manuscript.
-
The data that support the findings of this study are available in the NCBI SRA depository. These data were derived from the following resources available in the public domains: www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA675659.
-
The work is financially supported by Advanced Queensland Industrial Research Fellow (Grant No. AQIRF021-2019RD2). This filed work was supported by Energy Resources of Australia (ERA). We gratefully acknowledge Dr. Ping Lu for assistance with establishing the field project and for valuable discussions, as well as field support from Aidan Wright and Megan Parry (ERA). We also thank the Australian Centre for Ecogenomics at the University of Queensland for conducting the Illumina sequencing analyses.
-
The authors declare that they have no conflict of interest.
-
Litter inoculation triggered biological soil C and N cycling potential.
Microbial diversity and core microbiome dominance rose with litter inoculation.
Litter inoculation enhanced microbial network modularity and clustering efficacy.
C and N cycling microbes rose, while stress-adaptive microbes declined with litter inoculation.
-
Full list of author information is available at the end of the article.
- The supplementary files can be downloaded from here.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
You F, Parry D, Hall M, Huang L. 2026. Biological triggering waste rock-based soil system with native plant litter establishes soil microbiome and biochemical functional potential typical of Eucalyptus woodland. Energy & Environment Nexus 2: e008 doi: 10.48130/een-0026-0003
Biological triggering waste rock-based soil system with native plant litter establishes soil microbiome and biochemical functional potential typical of Eucalyptus woodland
- Received: 31 October 2025
- Revised: 07 January 2026
- Accepted: 16 January 2026
- Published online: 27 February 2026
Abstract: Soil microbiomes native to Eucalyptus woodlands are critical to decomposing low-quality litter, characterized by high lignin content and wide carbon: nitrogen ratios (~60:1), supporting sustainable nutrient cycling in nutrient-poor forest soils. This study evaluated the effectiveness of native litter inoculation as a field-based strategy to establish key microbial groups in eco-engineered soils constructed from non-hazardous gravel waste rocks, aimed at facilitating Eucalyptus woodland restoration. Inoculation of native litter significantly increased microbial diversity and reshaped community composition in surface soils, enriching core taxa shared with adjacent native woodlands. Following inoculation, microbial communities exhibited increased abundance of C- and N-cycling taxa, including ammonia-oxidizing archaea (Nitrososphaeraceae) and diverse organic matter decomposers (Bacteroidetes, Ascomycota), alongside a decline in stress-adapted groups such as thermophilic Thermoplasmata and Glomeromycota. Network analyses revealed increased archaeal and bacterial modularity and enhanced fungal connectivity and clustering, reflecting improved resource partitioning and cooperative interactions potential. These microbial shifts coincided with the biological triggering of soil organic matter decomposition and enhanced N cycling during the wet season. The results demonstrate that native litter inoculation is a practical, low-input approach to accelerate microbial functional recovery in waste rock-based soils, providing a mechanistic basis for its integration into field-based Eucalyptus woodland rehabilitation.
-
Key words:
- Litter inoculation /
- Soil microbiomes /
- Network /
- Soil function /
- Eucalyptus woodland