-
Colorectal cancer (CRC) ranks third globally in prevalence and second as a major cause of cancer-related deaths[1]. Despite notable declines in colorectal cancer (CRC) incidence and mortality driven by early screening and therapeutic advances, there remains an urgent need to identify novel therapeutic targets and robust biomarkers to enable early diagnosis, precision treatment, and longitudinal disease monitoring, thereby improving patient outcomes[2]. CircRNAs have been a pivotal class of primarily non-coding RNAs, with many exerting critical influences on cancer development and progression through various mechanisms of action[3]. They exhibit specific and differential expression across diverse tissues and fluids, including CRC, under various pathological conditions. For instance, circPLCE1 is downregulated in CRC tissues and correlates with unfavorable survival outcomes. Mechanistically, it directly binds to the serine/arginine-rich splicing factor 2 (SRSF2) protein, inhibiting SRSF2-mediated splicing of PLCE1 pre-RNA, which contributes to CRC progression[4]. Conversely, circCCDC666 exhibited high expression in colon cancer tissues and was inversely associated with patient prognosis[5]. Collectively, accumulating evidence highlights the critical involvement of circular RNAs (circRNAs) in the tumorigenesis of CRC.
The long noncoding RNA (lncRNA) ANRIL, located at the 9p21 locus on human chromosome 9, has been implicated in tumor progression, cardiovascular disease, and diabetes. ANRIL is involved in the long-term repression of the tumor suppressor gene locus-CDKN2A/B and maintenance of chromatin silencing through interactions with polycomb protein complexes 1 and 2 (PRC1 and PRC2), thereby regulating cell proliferation and senescence, primarily through modulating the expression of pivotal cancer-related genes like the P53 axis[6]. In addition to its involvement in the regulation of tumor development, ANRIL is also tightly related to chemosensitivity. Knocking down the expression level of ANRIL increases the sensitivity of CRC cells to 5-FU and promotes apoptosis. Conversely, upregulation of ANRIL expression significantly enhances chemoresistance in these cells[7]. Therefore, ANRIL holds potential as a therapeutic target for inhibiting chemoresistance in CRC. The ANRIL gene consists of 21 exons, which undergo alternative splicing (AS), generating at least 28 distinct linear isoforms as well as 30 circular isoforms[8]. Notably, an analysis of circular-to-linear isoform ratio in cohorts of patients with cardiovascular disease (CVD) and melanoma revealed marked differences compared with control groups[9]. Maintaining a precise balance between circular and linear isoforms is crucial for proper gene regulation modulation, while any imbalance in this ratio can affect cellular physiology. Future research focused on thoroughly characterizing ANRIL isoforms across various cancer subtypes may provide valuable insights into ANRIL's role in tumor progression and metabolic disorders.
Inconsistent expression of circular isoforms was observed in colon cancer cells overexpressing the oncogenic linear isoform of ANRIL-P14AS[10], with down-regulation of hsa_circCDKN2B-AS_012 (circ86591) expression, which is one of the isoforms of circANRIL. However, whether and how the circ86591 functions in CRC remains unknown. Subsequently, in vitro and in vivo tests utilizing cell-coupled mouse models were performed to elucidate the biological role of circ86591 involved in tumorigenesis and metabolism of CRC.
-
HCT116, SW480, LOVO and HEK293FT cell lines were generously donated by Professor Dajun Deng from Peking University Cancer Hospital and were successfully identified using STR methods by Qingke Biotechnology Co., Ltd. HEK293T (AW-CNH086) were provided and identified by Abiowell Biotechnology Co., Ltd. All cells were cultured in DMEM (Corning, VA, USA) with 10% fetal bovine serum (Gibco, Australia) and 1% penicillin/streptomycin (Gibco, NY, USA), coupled with incubated at 37 °C in a 5% CO2 incubator. ASO (antisense oligonucleotide, ASO) and primers of circ86591 were synthesized by RiboBio (Guangzhou, China). Circ86591 sequences targeted by ASO are CAACACTCCAGTAGAGACGG. All siRNAs were synthesized by Tsingke (Beijing, China). Full-length of circ86591 was constructed in PLO5-ciR vector by Geneseed Biotech Co., Ltd (Guangzhou, China) and ADV vector (pADM-CMV-circRNA-mMCV-copGFP) by WZ Biosciences Inc. (Shandong, China). CRC cells were plated in 6 cm dishes and, upon reaching 70%−80% confluence, transfected with siRNA (50nM) or ASO (30, 50, or 100 nM) or vectors (2.5 µg/well) utilizing Lipofectamine 3000 (Invitrogen, CA, USA). Cells were then analyzed after 48 h, with all sequences detailed in Supplementary Table S1.
RNA extraction and quantitative real-time (qRT)-PCR
-
For RNA extraction. Total RNA was isolated according to the protocol of Trizol (TransGen Biotech, Beijing, China), and 1.5 µg of isolated RNA was reverse-transcribed utilizing TransScript First-Strand cDNA Synthesis SuperMix kit (Roche, IN, USA). QRT-PCR analysis was conducted on an ABI-7500 Fast system (Applied Biosystems) with SYBR green (TransGen Biotech, Beijing, China) to determine the RNA levels of target genes with the following settings: 94 °C for 30 s; 40 cycles of 94 °C for 5 s, 62 °C for 15 s, 72 °C for 34 s. Expression levels were normalized to GAPDH RNA levels. Primers utilized for this research are detailed in Supplementary Table S1.
Western blot
-
Cells were harvested by scraping and lysed in a lysis buffer composed of NP-40 buffer (Solarbio, Beijing, China) and protease inhibitor cocktail (LabLead, Beijing, China), Protein lysates were run in a 12.5% or 10% SDS-PAGE gel (Epizyme Biomedical Technology Co., Ltd, Shanghai, China) and transferred to polyvinylidene difluoride membranes (Merck Millipore). After a 1 h blocking step with 5% non-fat milk, the membrane was incubated overnight at 4 °C with primary antibodies (Supplementary Table S2) diluted in 1% non-fat milk before incubation with appropriate secondary antibodies—goat anti-rabbit or goat anti-mouse (ZSGB Biotech, Beijing, China)—at room temperature for 1 h. Following three washes with 1 × PBS, the signals were visualized using a chemiluminescence detection system (Cytiva, Amersham ImageQuant 800, Japan).
CCK-8 assay
-
Viability of CRC cells was evaluated employing CCK-8 (TransGen Biotech, Beijing, China) and measured at 450nm/630nm with the BioRad microplate reader. 100 μL of media containing 20,000 or 80,000 cells/mL (for proliferation as well as cytotoxicity assays) was added to each well of a 96-well plate. In the cytotoxicity assay, following enabling cells to adhere, the media were replaced with fresh media containing varying concentrations (0, 1, 2, 4, 8, 16 μM) of 5-FU, gemcitabine, or doxorubicin (Selleck, Shanghai, China). At designated time points, 10 μL of CCK-8 solution was added to each well before an additional 2−3 h of incubation at 37 °C. For the cell proliferation assay, transfected cells were evaluated daily over five consecutive days, and mean absorbance values (OD450/OD630) were recorded.
Flow cytometry
-
For the cell cycle assay, the transfected cells were harvested and centrifuged at 1,000 rpm for 5 min at 4 °C. After adding 70% ethanol to the cell precipitate overnight at 4 °C, the cells were collected at 1,000 rpm for 5 min at 4 °C and a single wash with 1 × PBS. Samples were added 500 μL 1 × PBS containing 50 μg/mL PI and 100 μg/mL RNase A. The reaction was incubated for 30 min at 4 °C away from light and detected by BD FACS Canto. The results were analyzed by Modfit LT 5.0.
RNA sequencing
-
For downregulated RNA-seq, LOVO cells were transfected with either a control ASO or a 30 nM ASO specifically targeting circ86591. Following 48 h, cells were harvested for RNA extraction, and RNA-seq was conducted by Sangon Biotech Co., Ltd (Shanghai, China). Significantly differentially expressed mRNAs resulting from circ86591 downregulation were identified by analyzing fold changes (FC) in gene TPM (transcripts per million) values between ASO-NC and ASO 30 nM transfected cells (q-value < 0.05 and |FC| > 2). For upregulated RNA-seq, HCT116 cells were infected with the supernatant of ADV Ctrl or circ86591. After 48 h, these cells were harvested for RNA extraction, and RNA-seq was carried out by Novogene Biotech Co., Ltd (Beijing, China). Significantly differentially expressed mRNAs upregulated by circ86591 were identified by assessing FC in gene FPKM (fragments per kilobase per million mapped reads) values between ADV Ctrl and ADV circ86591 groups (p-value < 0.05 and |log2FC| > 0).
RNA immunoprecipitation
-
Immunoprecipitation-targeting circ86591 assay was performed utilizing an RNA-binding protein immunoprecipitation kit (Cat# 17–701, EZ Magna, Millipore, USA) following the manufacturer's instructions. In brief, protein A/G beads pre-coated with 3−5 μg of anti-rabbit or anti-mouse immunoglobulin G, EBP1, or ACC1 were incubated with pre-frozen cell lysates overnight at 4 °C. RNA-protein complexes were then gathered, washed six times with 1 × PBS, and subjected to proteinase K digestion followed by RNA extraction employing Trizol. Purified RNA was subsequently analyzed by qRT-PCR with primers specific to the target RNA.
Subcellular fraction
-
Cells were washed with cold 1 × PBS and gathered in Buffer I (10 mmol/L Hepes, pH 8.0; 1.5 mmol/L MgCl2; 10 mmol/L KCl; 1 mmol/L DTT) to induce swelling. Following a 10-min incubation, NP-40 was added at a final concentration of 1%. Supernatants were gathered as the cytoplasmic fraction following vortexing and centrifugation. Pellet was resuspended in ice-cold Buffer II (20 mmol/L Hepes, pH 8.0; 1.5 mmol/L MgCl2; 25% glycerol; 420 mmol/L NaCl; 0.2 mmol/L EDTA; 1 mmol/L DTT with protease and RNase inhibitors) and incubated with rotation at 4 °C for 30 min. After centrifugation at 12,000 rpm for 15 min, the nuclear fraction was collected. Cytosolic and nuclear RNAs were extracted for use. Then, detection of circ86591 in the cytoplasmic and nuclear RNA of cells was via qRT-PCR. Adoption of GAPDH was as a cytoplasmic control, while U6 expression served as the nuclear-specific internal control for RT-PCR analysis of nuclear RNA.
RNA fluorescence in situ hybridization (RNA-FISH)
-
RNA-FISH tests were carried out following the manufacturer's instructions from Beyotime (Shanghai, China). In brief, HEK293 cells were seeded onto coverslips and incubated overnight in complete medium. Cells were then fixed with 4% paraformaldehyde for 10 min at room temperature, permeabilized with a solution containing 5 μg/mL Proteinase K and 0.5% Triton X-100 for 10 min, and blocked with pre-hybridization buffer for 20 min. Then, cells were incubated with 1.5 μg/mL FISH probes of circ86591 overnight at room temperature in the dark before washing with hybridization washing buffer I, II, and III in order at room temperature in the dark. Nuclei were counterstained with DAPI (1 μg/mL), washed three times with PBS, and images were captured employing LAS_X_Core (Leica Microsystems, Germany). The circ86591 subcellular localization was indicated by the distribution of red fluorescent signals.
RNA pull-down assay
-
Biotinylated circ86591 RNA baits and control probes were synthesized by RiboBio (Guangzhou, China). RNA pull-down assay was conducted employing PierceTM Magnetic RNA-Protein Pull-Down Kit (Thermo ScientificTM, Rockford, USA). Cell lysates were prepared using standard lysis buffer, followed by the binding of labeled RNA to streptavidin magnetic beads, and then incubating RNA-protein complexes overnight at 4 °C with rotation. Washing and elution of RNA-binding protein complexes, finally, RNA-interactive proteins were detected by WB and Coomassie blue stain analysis.
ATP detection
-
Cellular energy was measured utilizing the adenosine triphosphate (ATP)-luminescence assay kit (Dojindo, Japan). Briefly, HCT116 as well as SW480 cells were transfected with the indicated ASO and diluted to 5 × 104/mL. 100 μL cell suspension was added to a white 96-well plate, which was then added to a 100 μL working solution and left for 10 min at 25 °C away from light. The RLU signals were detected by Cytation 7 (Biotek, USA).
Mitochondrial membrane potential detection
-
Mitochondrial membrane potential was evaluated employing the JC-1 MitoMP detection kit (Dojindo, Japan). All images were obtained with a fluorescence microscope (Biotek, USA), and fluorescence was measured with filter pairs of 469/525 nm (green) and 586/647 nm (red). The results were shown as a ratio of fluorescence measured at 586/647 nm to that measured at 469/525 nm.
NAD+/NADH assay
-
Total amount of nicotinamide adenine dinucleotide (NAD)+/nicotinamide adenine dinucleotide hydrogen (NADH) was determined by the NAD+/NADH assay kit with WST-8 (Beyotime, China). Briefly, 1 × 106 cells were added to 200 μL extraction solution and centrifuged at 12,000 g for 10 min at 4 °C. The collected supernatant was detected for absorbance value at 450 nm.
Co-immunoprecipitation (Co-IP)
-
The primary anti-FBXW7 or rabbit control IgG was added to the cell lysate, and the mixture was incubated at 4 °C overnight. Thereafter, 25 μL protein A/G beads (88,804, Thermo scientificTM, Rockford, USA) were added and incubated at room temperature for 1 h. Proteins were eluted and detected using a WB assay with the primary anti-EBP1 and anti-FBXW7 and the secondary antibody.
Mass spectrometry analysis
-
Central carbon metabolism was detected utilizing an ultra-high performance liquid chromatography-tandem mass spectrometry (UHPLC-MS/MS) system (ExionLC™ AD UHPLC-QTRAP® 6,500+, AB SCIEX Corp., Boston, MA, USA). Thirty four standards and five stable isotope-labeled standards were sourced from ZZ Standards Co., Ltd (Shanghai, China). Methanol, acetonitrile, formic acid, and ammonium acetate (all Optima LC-MS grade) were procured from Thermo-Fisher Scientific (FairLawn, NJ, USA), while imino-bis (methylphosphonic acid) was purchased from Sigma-Aldrich (St. Louis, MO, USA). Ultrapure water was supplied by Millipore (MA, USA). For sample preparation, 100 mg of each sample was homogenized in 500 μL of methanol/water (4:1) containing mixed internal standards and incubated for 5 min. The homogenates were centrifuged at 12,000 rpm for 10 min, and 2 μL of supernatant was injected into the LC-MS/MS system by Novogene Biotech Co., Ltd (Beijing, China).
Animal experiments
-
To investigate the impact of circ86591 on tumor growth, HCT116 cells (5 × 106) suspended in 100 μL of PBS were subcutaneously injected into female NVSG (NOD-Prkdc scid Il2rgtm201(−/−)/V from Beijing Weishang Lide Biotechnology Co., Ltd) mice. Once tumors reached approximately 100 mm3, mice were randomly assigned to two groups. ADV-NC or ADV-circ86591 (WZ Biosciences Inc) (5 × 108 Pfu/mice) were administered once weekly. Tumor volume and mice's body weight were measured twice weekly, with tumor volume (mm3) computed as [ (Length) × (Width)2]/2. Mice were euthanized on day 14 post-treatment initiation.
Statistical analysis
-
Statistical analyses were conducted employing GraphPad Prism 9.5 software. Differences between indicated groups were compared utilizing a two-tailed Student t test. GEPIA (Gene Expression Profiling Interactive Analysis) serves as a widely utilized and highly cited tool for analyzing gene expression, leveraging tumor and normal tissue data from TCGA and GTEx databases. According to database documentation, quantitative comparisons of cell type proportions or TCGA/GTEx sub-dataset distributions were performed using ANOVA, while Pearson Correlation Coefficient was calculated for additional quantitative analyses, with results visualized in graphs. Image J was employed for WB quantification. p value < 0.05 (*), < 0.01 (**), and < 0.001 (***) was defined as significant.
Data availability
-
All supporting data are available upon request, with raw RNA-seq data deposited in Gene Expression Omnibus under accession number GSE280633.
-
First, the levels of circANRIL isoforms were examined by qRT-PCR in HCT116 and SW480 cells overexpressing oncogenic lncRNA-P14AS. Results demonstrated that circ86591 expression was reduced (Fig. 1a). In contrast, upregulating circ86591 expression did not change P14AS expression levels (Supplementary Fig. S1), suggesting that circ86591 might exert its effects through pathways independent of P14AS. Circ86591 is formed from exon 8 of the ANRIL gene, 1,077 nucleotides in length according to the Circular RNA Interactome database. Divergent primers specific to circ86591 were designed and amplified reverse transcription polymerase chain reaction (RT-PCR) product was verified by Sanger sequencing, confirming predicted head-to-tail splicing junction (Fig. 1b). Actinomycin D (5 μg/mL) treatment of SW480 cells revealed that circ86591 exhibited greater stability than linear GAPDH (Fig. 1c). Total RNA from HEK293FT and HCT116 cells was treated with RNase R and amplified by RT-PCR, revealing that circ86591 but not linANRIL was resistant to RNase R digestion (Fig. 1d). Then, subcellular localization of circ86591 was detected by cellular fraction assay and RNA-FISH, revealing that circ86591 was located in nucleus (Fig. 1e, f).
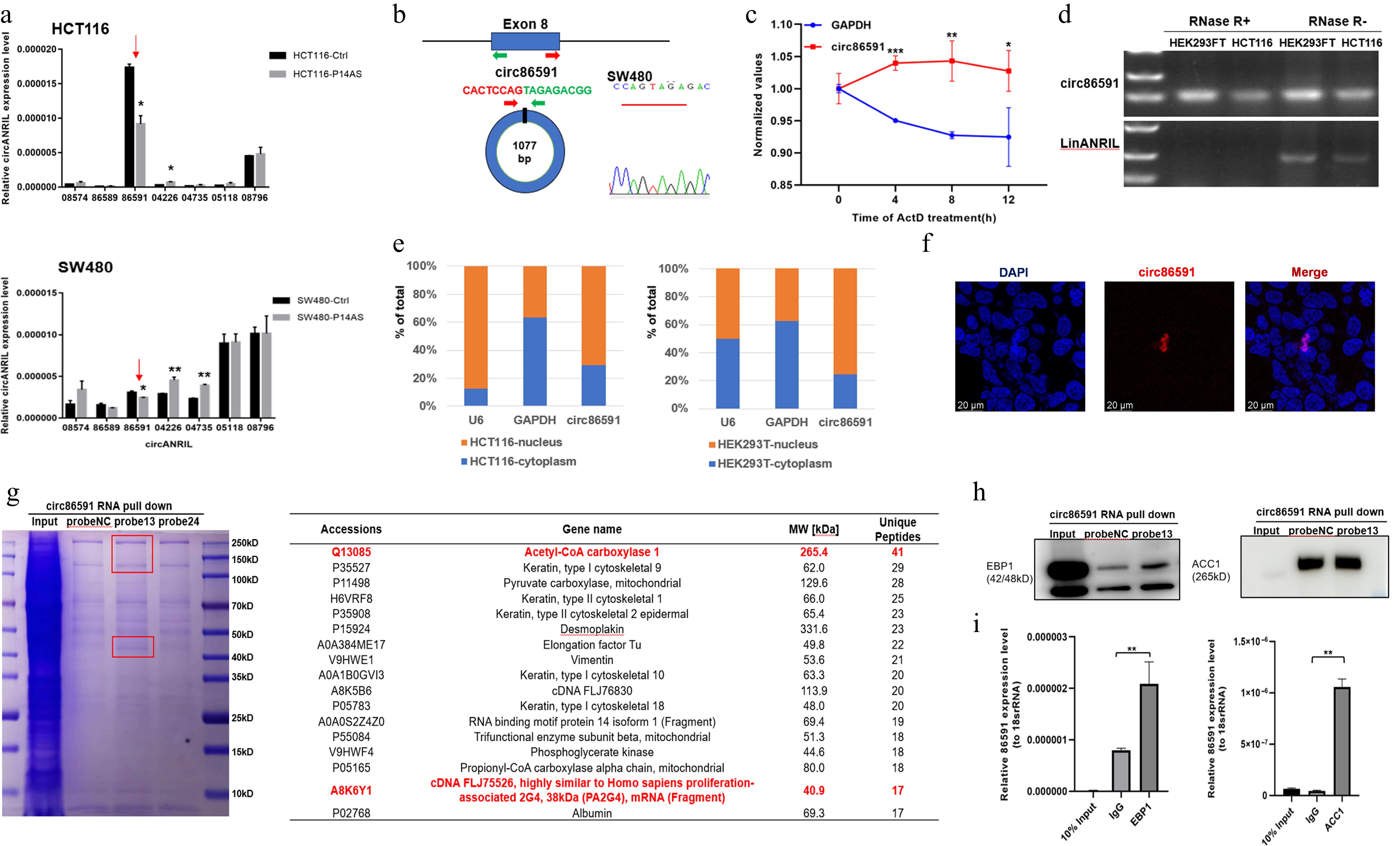
Figure 1.
The identification and characteristics of circ86591. (a) Circ86591 was down-regulated in CRC cells with P14AS OE group compared with control group by qRT-PCR. (b) Schematic diagram indicating the circularization of exon8 of CDKN2B-AS formed circ86591. Sanger sequencing demonstrated the presence of circ86591 and its circularization characteristics. (c) SW480 cells were treated with Actinomycin D (5 μg/mL) for duration as indicated, followed by qRT-PCR analysis to examine the expression of circ86591 or GAPDH. (d) Total RNAs extracted from cells were incubated with or without RNase R at 37 °C for duration as indicated, followed by RT-PCR analysis to examine the expression of circ86591 or linANRIL. (e) Circ86591 expression in cytoplasm (GAPDH used as cytoplasmic control) and nucleus (U6 used as nuclear control) of HCT116 and HEK293T cells was detected by qRT-PCR which indicating the circ86591 was enriched in the nuclear of cells. (f) The analysis of the distribution of circ86591 by RNA-FISH assay in HEK293T cells. (g) Biotin-labeled circ86591 pull-down complexes from cell lysate, following mass spectrometry. (h) Western blot analyses of circ86591-EBP1 and ACC1 complexes. (i) High levels of circ86591 were detected by RIP-PCR assay with an anti-EBP1 or anti-ACC1 antibody in HEK293T cells. IgG antibody acted as a negative control.
Furthermore, a biotin-labeled RNA pull-down assay combined with mass spectrometry was conducted to identify potential circ86591-binding proteins from HEK293FT cell lysates. The results showed that endogenous proliferation-associated 2G4 (PA2G4, also known as EBP1) as well as acetyl-CoA carboxylase 1 (ACC1) were identified as binding proteins for circ86591 (Fig. 1g). Western blot analysis validated the link of EBP1 or ACC1 with circ86591 (Fig. 1h). To assess whether circ86591 naturally interacts with EBP1 and ACC1 proteins in cells, RNA immunoprecipitation (RIP) assays were performed, revealing significant enrichment of circ86591 in EBP1 and ACC1 antibody complexes compared to IgG group (Fig. 1i), confirming the presence of endogenous circ86591-EBP1 and circ86591-ACC1 complexes in cells.
Down-regulated circ86591 regulated the nucleotide metabolism and cell cycle pathways
-
Next, mechanisms by which circ86591 exerts its functional role in CRC were elucidated. LOVO cells transfected with ASO-NC or ASO-circ86591, which targeted the junction region of circ86591, were collected for RNA-seq. QRT–PCR analysis confirmed a significant downregulation of circ86591 expression following ASO–circ86591 treatment. Differential expression gene analysis (DEG) uncovered that 195 genes were significantly downregulated in LOVO cells (q value < 0.05 and |Fold Change| > 2), which were shown by a heatmap (Fig. 2a). The results of eukaryotic orthologous group (KOG) enrichment analysis revealed that nucleotide transport and metabolism and cell cycle control pathways were the most enriched among genes positively modulated by circ86591 knockdown (Fig. 2b). The top pathway contained the phosphoribosyl pyrophosphate synthetase 2 (PRPS2), ribonucleotide reductase regulatory subunit M2 (RRM2), 3'(2'),5'-bisphosphate nucleotidase 1 (BPNT1), solute carrier family 25 member 24 (SLC25A24), adenosine deaminase tRNA specific 2 (ADAT2), deoxycytidine kinase (DCK), and phosphoribosyl pyrophosphate amidotransferase (PPAT) whereas the cyclin A2 (CCNA2) and cyclin E2 (CCNE2) genes were enriched in the cell cycle pathway (Supplementary Fig. S2a).
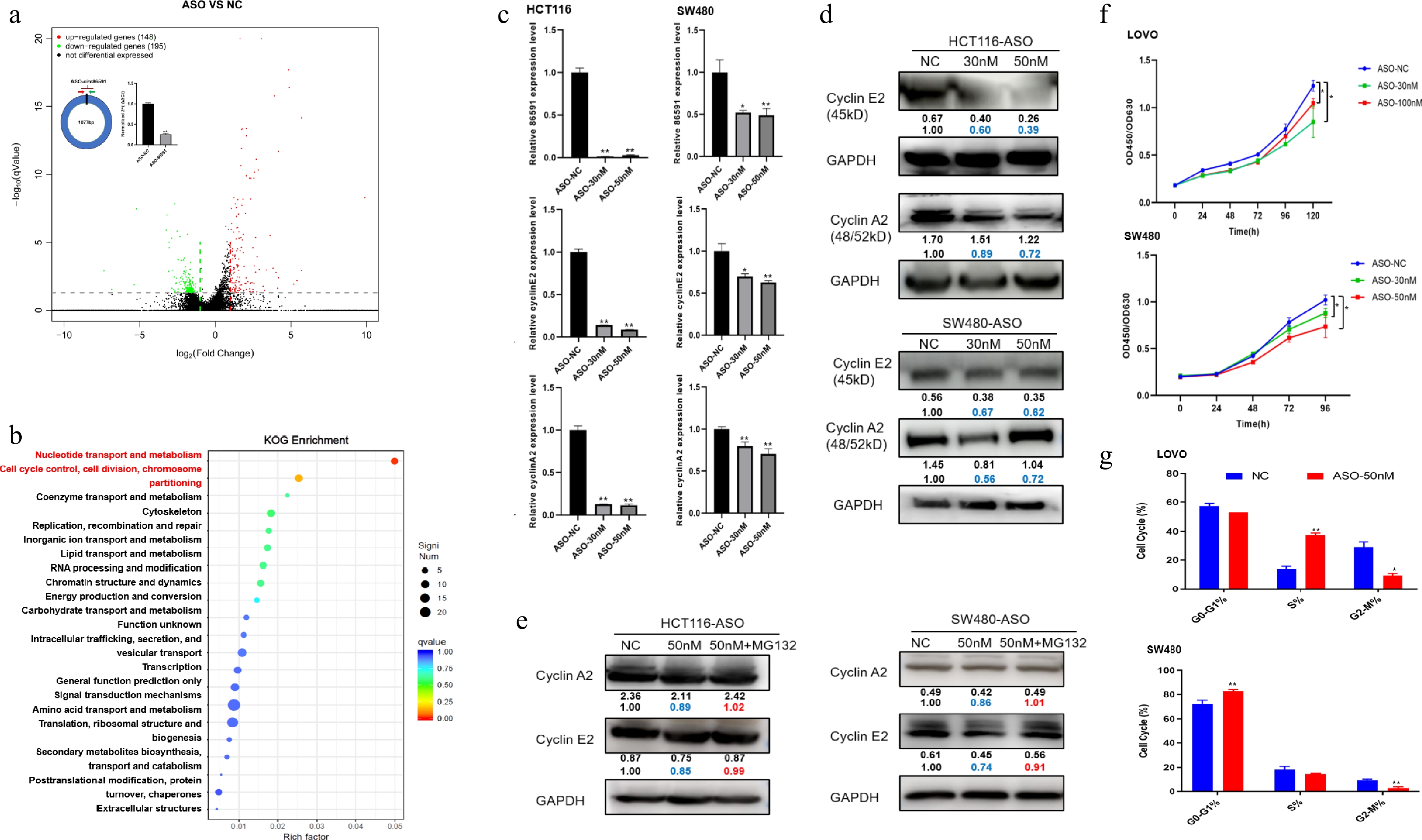
Figure 2.
Circ86591 affected cell cycle pathway through binding EBP1 protein. (a) The LOVO cells transfected with ASO-NC or ASO-circ86591 were subjected to qRT-PCR to examine the expression of circ86591. The volcano plot showing the expression profile of LOVO cells with ASO-NC and ASO-circ86591. (b) The KOG analysis for genes positively regulated by circ86591 knockdown was shown. (c) The qRT-PCR, and (d) WB assays confirmed the downregulation of CCNE2 and CCNA2 involved in cell cycle in circ86591 knockdown cells. (e) The WB assay was used to detect the protein levels of CCNE2 and CCNA2 after MG132 drug treatment in HCT116 and SW480 cells transfected with ASO-NC or ASO-circ86591. (f) The CCK8 assay exhibited changed in the growth of LOVO and SW480 cells with circ86591 knockdown compared with controls. (g) Flow cytometry indicated that the G0-G1 and S phase arrest in the cells with circ86591 knockdown compared with controls.
ErbB3 binding protein 1 (EBP1) produces two alternatively spliced isoforms, P48 and P42, which exhibit different roles in human cancers. P42 functions as a tumor suppressor and enhances tumor suppressor function of F-box and WD40 domain protein 7 (FBXW7) by promoting its interaction with target proteins, including c-MYC and Cyclin E, to the ubiquitination degradation pathway. Therefore, circ86591 was hypothesized to regulate the cell cycle by competitively interacting with EBP1. To confirm this hypothesis, the impact of circ86591 on CCNA2 and CCNE2 expression in SW480 and HCT116 cells transfected with ASO-NC and ASO-circ86591 was explored by qRT-PCR and WB assays. Results indicated that CCNE2 and CCNA2 expression were significantly decreased upon circ86591 knockdown (Fig. 2c, d). CCNE2 and CCNA2 expression, regulated by the knockdown of circ86591, were restored by MG132 (a proteasome inhibitor) treatment in HCT116 and SW480 cells (Fig. 2e). Furthermore, cells treated with MG132 for diverse durations were collected and cultured, and expression changes of CCNE2 and CCNA2 were analyzed by WB assay. The results demonstrated that CCNE2 and CCNA2 levels in the ASO-circ86591 group were lower than those in the control group at early time points (2 and 4 h). Conversely, this trend was reversed at later stages (6 and 24 h), with both proteins exhibiting higher expression in the ASO-circ86591 group. These findings indicate that although circ86591 knockdown initially suppresses CCNE2 and CCNA2, prolonged proteasome inhibition (≥ 6 h) leads to progressive accumulation of both proteins (Supplementary Fig. S2b). Through Co-IP experiments, increased binding of EBP1 to FBXW7 protein was observed in the ASO-circ86591 group compared with the control group, thereby promoting degradation of FBXW7-mediated target proteins (Supplementary Fig. S2c).
To explore the function of circ86591, the CCK8 assay was performed to test the proliferation ability of LOVO and SW480 cells, transfected with the ASO-NC or ASO-circ86591. The outcomes showed that knockdown of circ86591 inhibited cell proliferation in LOVO and SW480 cells (Fig. 2f). Then, the cell cycle of LOVO and SW480 cells transfected with ASO-NC or ASO-circ86591 was analyzed by flow cytometry. Circ86591 knockdown induced S-phase and G0/G1-phase arrest, accompanied by a reduced proportion of colorectal cancer (CRC) cells entering the G2/M phase of the cell cycle (Fig. 2g).
Circ86591 knockdown influences energy metabolism including nucleotide metabolism in CRC cells
-
For nucleotide transport and metabolism pathway, circ86591-related transcript levels of PRPS2, RRM2, BPNT1, SLC25A24, ADAT2, DCK, and PPAT were detected by qRT-PCR and WB assays, demonstrating that the expression levels of these genes decreased with circ86591 knockdown (Fig. 3a, b). Nucleotides are mainly involved in the composition of nucleic acids and single nucleotides like adenosine triphosphate (ATP), which are related to energy metabolism. ATP level and total amount of NAD+/NADH were detected in HCT116 and SW480 cells transfected with ASO-NC or ASO-circ86591. The results indicated that downregulation of circ86591 resulted in a decrease in the total amount of ATP and NAD+/NADH (Fig. 3c, d).
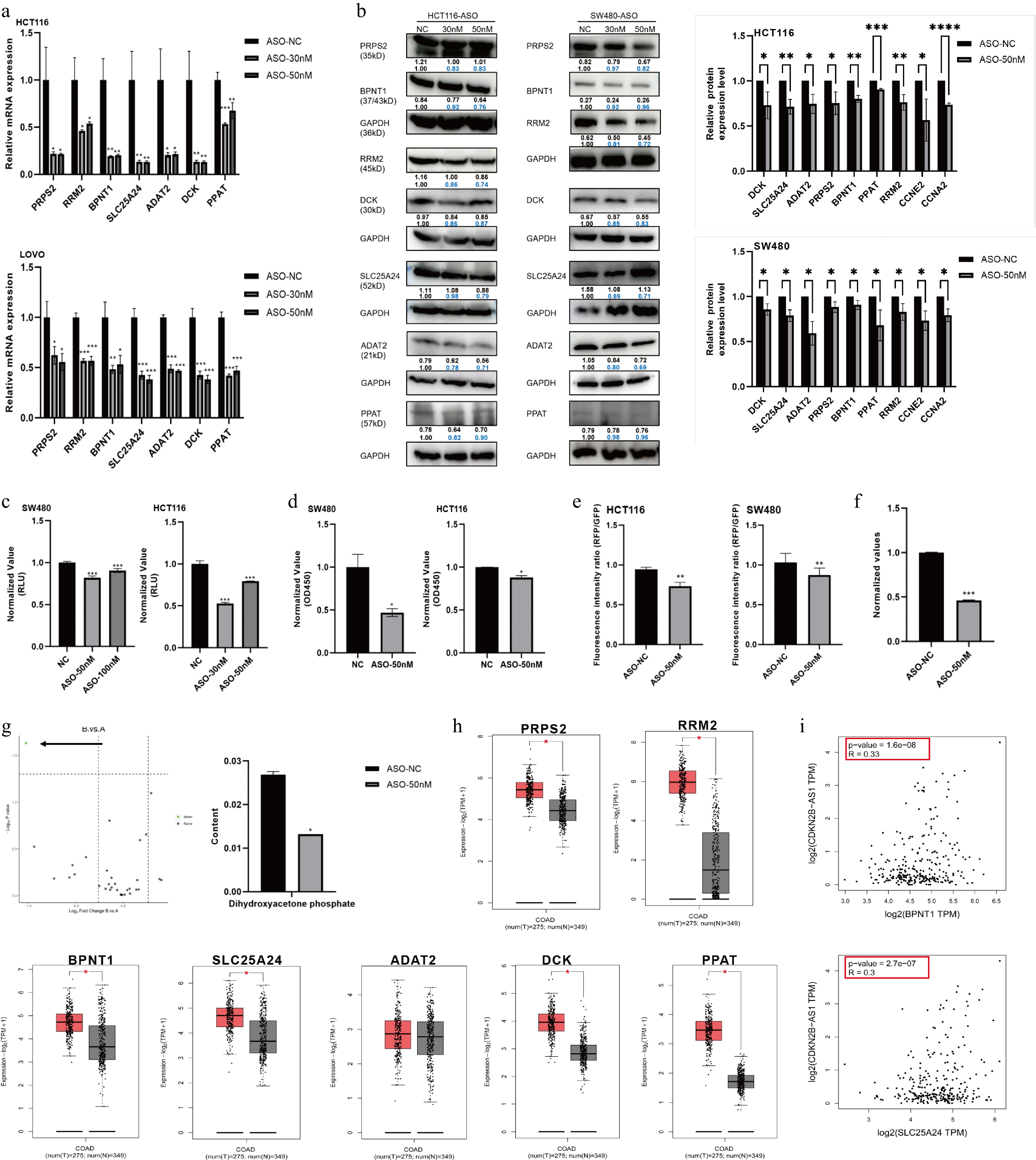
Figure 3.
circ86591 knockdown influence energy metabolism including nucleotide metabolism in CRC cells. The (a) qRT-PCR, and (b) WB assays confirmed the downregulation of genes involved in nucleotide metabolism in circ86591 knockdown cells. (c) The ATP assay exhibited changes in the ATP content of HCT116 and SW480 cells with circ86591 knockdown compared with controls. (d) The NAD+/NADH assay exhibited changes in the content of HCT116 and SW480 cells with circ86591 knockdown compared with controls. (e) The JC-1 MitoMP detection kit showed changes in the mitochondrial membrane potential abilities of HCT116 and SW480 cells with circ86591 knockdown compared with controls. (f) The L-Lactic Acid (LA) colorimetric assay kit exhibited changes in the lactate content of SW480 cells with circ86591 knockdown compared with controls. (g) The mass spectrometry analysis showed that the content of DHAP decreased in the ASO-circ86591 group compared with controls. (h) The genes regulated by circ86591 were up-regulated in COAD tissues compared with normal tissues. (i) The GEPIA database analysis of BPNT1 and SLC25A24 expression were positively correlated with CDKN2B-AS1.
Mitochondria act as the major locus of ATP production by aerobic respiration. Reduced mitochondrial activity was found in the ASO-circ86591 group compared with the ASO-NC group in HCT116 and SW480 cells (Fig. 3e). Furthermore, lactate content, another energy product of metabolism, was investigated, and lactate levels were inhibited in the ASO-circ86591 group (Fig. 3f). Differences in central carbon metabolism between the two groups were analyzed by targeted metabolomics. The central carbon metabolism, also known as energy metabolism, refers to the metabolic pathways related to maintaining the most basic life activities of living organisms, which mainly include the tricarboxylic acid cycle (TCA cycle), glycolysis, and the pentose phosphate pathway. The results indicated that the content of dihydroxyacetone phosphate (DHAP) decreased in the ASO-circ86591 group (Fig. 3g). During glycolysis, DHAP can be converted to glyceraldehyde-3-phosphate to generate energy and metabolic intermediates. In the process of lipid synthesis, DHAP can be converted to glycerol, which in turn synthesizes triglyceride with fatty acids. ACC1 is closely related to fatty acid synthesis, suggesting that circ86591 plays a critical role in lipid metabolism.
The expression of PRPS2, RRM2, BPNT1, SLC25A24, ADAT2, DCK, and PPAT in COAD tumor tissues and normal tissues was analyzed through GEPIA (http://gepia.cancer pku.cn/index.html) database, showing that, in comparison with normal tissues, PRPS2, RRM2, BPNT1, SLC25A24, DCK, and PPAT were highly expressed in tumor tissues (Fig. 3h), while BPNT1 and SLC25A24 were positively correlated with another ANRIL transcript, CDKN2B-AS1 (Fig. 3i). Moreover, the link between ACC1 with nucleotide metabolism-related proteins was examined through the GEPIA database, showing that ACC1 was positively related to PRPS2, RRM2, ADAT2, and PPAT (p value < 0.05) (Supplementary Fig. S3). Collectively, the above findings uncovered the metabolic process effects of circ86591 in vitro.
Overexpression of circ86591 regulated the level of tumor suppressor P53
-
On the contrary, circ86591 was overexpressed in tumor cells to explore the function of circ86591 under high-expression conditions. HCT116 and LOVO cells were transiently transfected with PLO5-circ86591 or control plasmids before RT-PCR measurement. Surprisingly, tumor cells with overexpressed circ86591 still exhibited lower proliferation ability compared to the control group (Fig. 4a). Furthermore, adenoviral supernatants for circ86591 (ADV circ86591) or control (ADV ctrl). were constructed and obtained. Subsequently, the supernatants were infected into HCT116 and SW480 cells, and green fluorescence was observed under a fluorescence microscope (Fig. 4b). QRT-PCR's results demonstrated that circ86591 expression was elevated in the ADV circ86591 group compared to the ADV ctrl group (Fig. 4c), suggesting that the packaging and infection process was successfully accomplished. Next, the expression of nucleotide metabolism- and cell proliferation–related proteins regulated by circ86591 knockdown was examined through qRT-PCR and WB assays, indicating that upregulation of circ86591 expression did not affect these levels of metabolism and proliferation-related proteins (Supplementary Fig. S4).
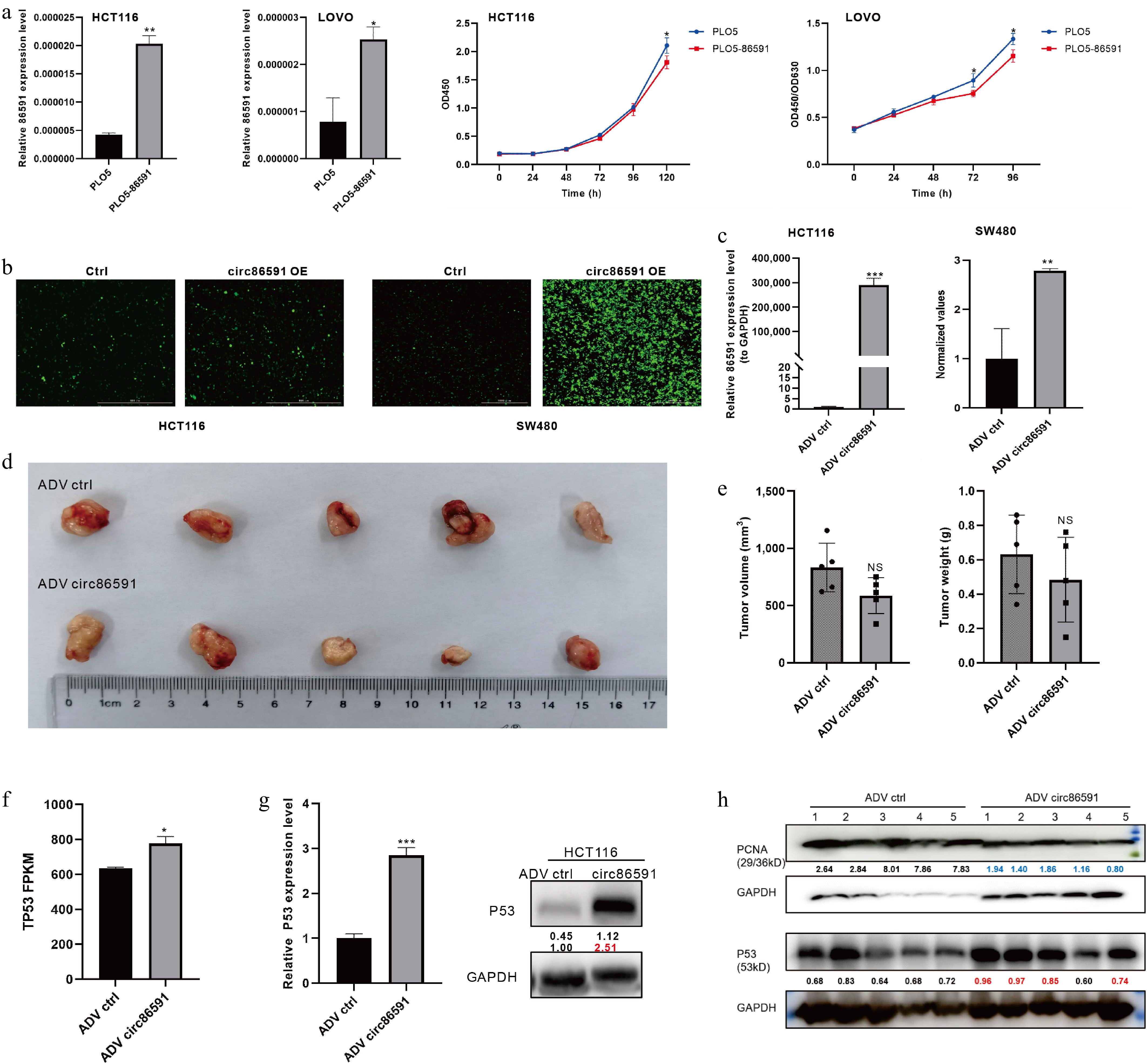
Figure 4.
The effect of circ86591 in vivo. (a) The up-regulated circ86591 in HCT116 and LOVO cells were detected by qRT-PCR and the CCK8 assay exhibiting changes in the growth of CRC cells with circ86591 overexpression compared with controls. (b) The green protein expression of HCT116 and SW480 cells infected with ADV ctrl or ADV circ86591 were showed by fluorescence microscopy. (c) The qRT-PCR assay confirmed the overexpression of circ86591 in ADV ctrl and ADV circ86591 group. (d) NVSG mice were subcutaneously implanted with HCT116 cells, and treated with ADV ctrl or ADV circ86591 for three weeks (once per week) and the tumors were collected and photographed. (e) The volume and weight of tumors as described in (d) was shown. (f) The RNA-seq analysis of P53 gene expression level between ADV ctrl and ADV circ86591 group. (g) The WB assay confirmed the elevation of P53 protein levels in HCT116 cells after infection with ADV circ86591. (h) The P53 and PCNA protein levels in tumor of (d) were detected by the WB assay.
Moreover, the immunodeficient mice were inoculated subcutaneously with HCT116 cells. When the volume of tumors was approximately 100 mm3, the mice were classified into two groups and treated with ADV ctrl or ADV circ86591 once weekly. As compared with ADV control, the volume and weight of tumors treated with ADV circ86591 were inhibited (Fig. 4d, e). To discover the mechanism of inhibition of cell proliferation by overexpression of circ86591, RNA-seq analysis of HCT116 cells treated with ADV ctrl or ADV circ86591 was performed. Circ86591 was observed to upregulate the expression level of the well-known tumor suppressor gene P53 by analyzing circ86591-related DEGs (Fig. 4f). The results of WB verified that P53 protein levels were significantly increased in the HCT116 cells infected with ADV circ86591 compared to ADV control and P53 protein levels were higher in most ADV circ86591-injected tumors in mouse tissues than in ADV ctrl group, but proliferating cell nuclear antigen (PCNA) expression was suppressed (Fig. 4g, h).
Circ86591 was associated with the chemoresistance of 5-FU, doxorubicin and gemcitabine
-
5-Fluorouracil (5-FU) is an anti-metabolite with substitution of fluorine for hydrogen at the C-5 position of uracil. Since the 1950s, 5-FU-based chemotherapy has been the main treatment regimen for patients with CRC[11]. The anticancer role of Doxorubicin (Dox) is already recognized due to its intercalation between deoxyribonucleic acid (DNA) base pairs, inhibition of topoisomerase II enzyme activity, interference with mitochondrial function, along with enhancement of free-radical production and oxidative stress[12]. DCK catalyzes phosphorylation of the drug-Gemcitabine (Gem), which exerts antitumor effects by inhibiting DNA replication and repair in tumor cells[13]. The above drugs were chosen for the next experiments. The cellular activity of HCT116 and SW480 cells after knockdown of circ86591 against the drugs 5-FU, Dox, and Gem was examined by CCK-8 assay, revealing that downregulation of circ86591 expression increased resistance of tumor cells to these drugs (Fig. 5a), which might be caused by a reduction in the level of cellular metabolism resulting in elevated resistance to chemotherapeutic drugs. Aside from the observation that downregulation of circ86591 leads to reduced DCK protein expression, contributing to Gem resistance in tumor cells, the overexpression of circ86591 did not result in any significant difference in drug sensitivity after treatment with Gem (Fig. 5b), suggesting its potential as a biomarker for predicting chemotherapy response for Gem treatment.
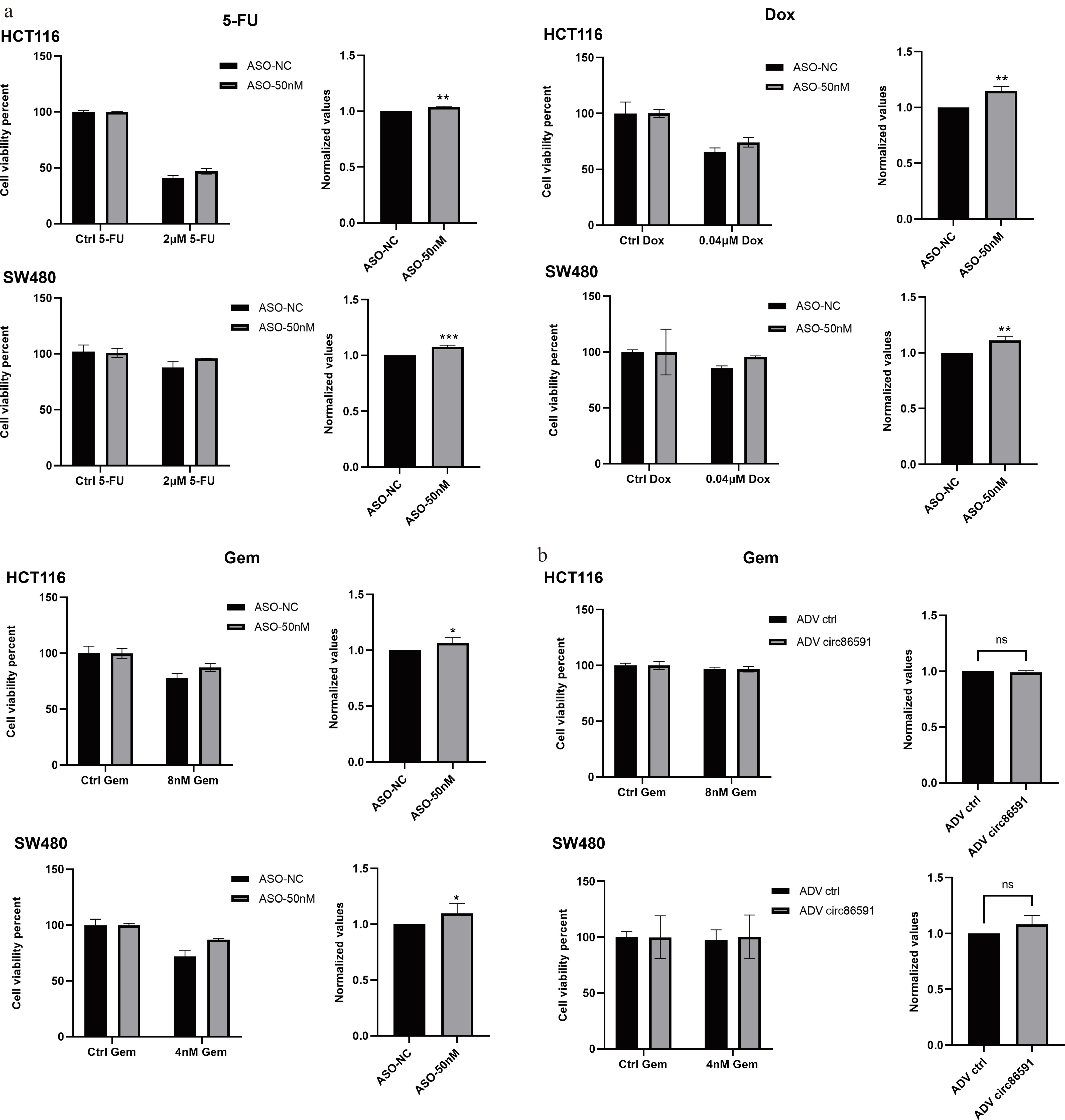
Figure 5.
circ86591 knockdown could increase drug resistance in CRC cells. (a) The CCK8 assay was performed for cell survival after drug treatment in ASO-NC and ASO- circ86591 groups. (b) The CCK8 assay was performed for cell survival after GEM treatment in ADV ctrl and ADV circ86591 groups.
Circ86591 was related to lipid metabolic pathway via analysis of RNA-seq
-
The analysis of Kyoto Encyclopedia of Genes and Genomes (KEGG) in DEG, which were regulated by circ86591 knockdown, revealed that its effects on sulfur metabolism, lipid metabolism pathway and cholesterol metabolism, in addition to cell cycle and nucleotide metabolism pathway (Fig. 6a). Based on KEGG analysis, the sortilin 1 (SORT1), CD36, and oxysterol binding protein like 8 (ORP8) genes were found to be enriched in the cholesterol metabolism pathway, and 3-hydroxyacyl-CoA dehydratase 3 (HACD3) and ELOVL fatty acid elongase 5 (ELOVL5) were enriched in the unsaturated fatty acid biosynthesis and fatty acid elongation metabolism pathway. ORP8 is localized in the endoplasmic reticulum and acts as a negative regulator for macrophage cholesterol efflux[14,15]. The elongation of very long-chain fatty acids (ELOVL) family catalyzes fatty acid elongation, with ELOVL5 playing an indispensable role in the elongation of polyunsaturated fatty acids (PUFAs), which is crucial for lipid metabolism[16]. Hence, the expression level of ORP8 and ELOVL5 was detected to determine the status of the cholesterol metabolism and fatty acid elongation metabolism pathway. WB assay results indicated that circ86591 downregulation inhibited ELOVL5 and ORP8 expression levels (Supplementary Fig. S5). Lipid droplet synthesis was examined in HCT116 and SW480 cells with circ86591 knockdown and control group, and Lipi probe with red fluorescence showed weaker fluorescence intensity in the ASO-circ86591 group compared to ASO-NC group under fluorescence microscopy after incubation with a red-fluorescent Lipi probe (object mean RFP/object mean bright) (Fig. 6b).
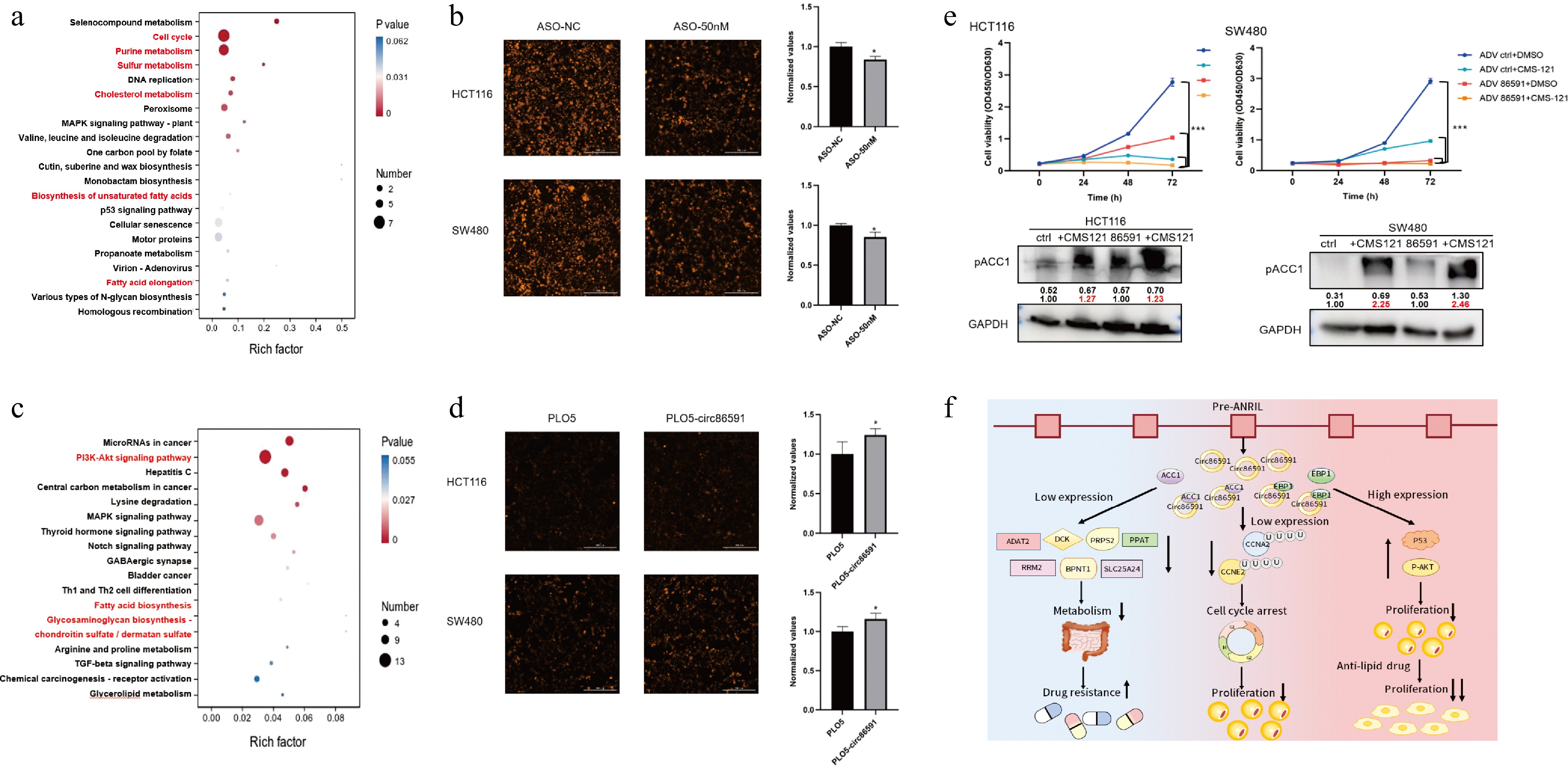
Figure 6.
circ86591 was related to lipid metabolic pathway via analysis of RNA-seq. (a) The KEGG analysis for genes positively regulated by circ86591 knockdown was shown. (b) Fluorescence microscopy was used to observe lipid droplet synthesis in the ASO-circ86591 or ASO-NC group, and three fields of view were randomly selected for fluorescence intensity analysis. (c)The KEGG analysis for genes positively regulated by circ86591 overexpression was shown. (d) Fluorescence microscopy was used to observe lipid droplet synthesis in the ADV circ86591 or ADV ctrl group, and three fields of view were randomly selected for fluorescence intensity analysis. (e) The HCT116 and SW480 cells infected with ADV ctrl or ADV circ86591 combined with DMSO or CMS121 were subjected to cell proliferation assay (up) and WB assay (below). (f) Schematic representation of the functional role of circ86591.
Besides, the analysis of KEGG in DEGs, which were regulated by circ86591 overexpression, revealed that its effects also on fatty acid biosynthesis (Fig. 6c). The results of lipid droplet synthesis demonstrated that Lipi probe with red fluorescence showed stronger fluorescence intensity in the ADV circ86591 group compared to ADV ctrl group (Fig. 6d). Furthermore, the inhibitor of ACC1, which acts as the first rate-limiting enzyme in fatty acid synthesis, CMS121 (10 μM), was combined with ADV circ86591 or ctrl supernatant to treat tumor cells. CCK-8 assay results indicated that combined treatment with CMS121 and ADV-circ86591 significantly inhibited cell proliferation compared with CMS121 alone, ADV-circ86591 alone, or control treatment. WB assay results indicated that CMS121 treatment increased the phosphorylation of ACC1 at serine 79 (Fig. 6e). Collectively, the above findings indicated that circ86591 is a critical regulator of CRC lipid metabolism and tumorigenesis. ADV-circ86591 demonstrated potent inhibitory effects on CRC growth and significantly improved the therapeutic efficacy of lipid metabolism–targeted therapy.
-
CircRNAs show a unique class of non-coding RNAs, marked by a covalently closed loop structure that lacks typical 5' and 3' ends. While circRNAs were initially considered byproducts of RNA splicing with no functional significance, accumulating evidence demonstrates that circRNAs play vital roles in tumorigenesis and progression by regulating gene expression, functioning as microRNA (miRNA) sponges, and mediating protein interactions. Despite typically being divided as non-coding, some circRNAs have been observed to have translational potential, encoding small peptides that may play distinct roles in tumor progression[17]. The antisense noncoding RNA in INK4 locus (ANRIL), also known as CDKN2B-AS, is an antisense long non-coding RNA (lncRNA) situated at the CDKN2A/B gene locus on human chromosome 9p21.3. In tumor cells, ANRIL upregulation promotes cell proliferation and metastasis coupled with epithelial-mesenchymal transition (EMT), whereas ANRIL downregulation inhibits tumor growth and metastasis, as well as promotes apoptosis and senescence. In metabolic diseases, polymorphisms in the ANRIL gene are associated with diabetes, atherosclerosis, and obesity[6].
However, research on the various ANRIL transcripts in CRC is limited and lacks a unified conclusion. In a previous study, a novel linANRIL P14AS (GenBank Accession No. MK574077) was discovered. P14AS expression level in colon cancer tissues was markedly higher than that in the margin tissues and normal tissues. P14AS could promote the growth ability of tumor cells by competitively binding heterogeneous nuclear ribonucleoprotein D (HNRNPD) and thereby increasing the stability of linANRIL RNA[10]. In addition to cis-regulating neighboring gene expression levels, P14AS could also regulate the protein level of ubiquitin conjugating enzyme E2 D3 (UBE2D3) by sponging miR-23a-5p, which affected the proteasomal degradation pathway of NF-κB inhibitory protein-IκBa and ultimately activated NF-κB-dependent inflammatory signaling pathway[18]. Drawing on this, inconsistent expression of circANRIL transcripts was found in colon cancer cells overexpressing linANRIL-P14AS, and the expression of circ86591 was down-regulated, suggesting that linANRIL and circANRIL play different roles in tumor development. Further investigation of the function of circ86591 showed that the proliferative ability of tumor cells was impaired either by upregulation or downregulation of circ86591, leading to the hypothesis that circ86591 exerts suppressive effects through a specific mechanism. Through experiments such as cytosolic/nuclear fractionation and RNA-FISH, circ86591 was confirmed to be present in the cytosolic and nucleus, albeit mainly in the nucleus. EBP1 and ACC1 proteins were shown to interact with circ86591, as analyzed by the results of experiments such as RNA pull-down and RIP.
ErbB3-binding protein 1 (EBP1) is a member of the PA2G4 family of proteins that regulate proliferation. The two isoforms of EBP1, P48 as well as P42, have vital roles in tumors because the difference in N-terminus between P48 and P42 directs unique function and links with various binding proteins[19]. In various cancer cell types, like glioblastoma multiforme and lung cancer cells, P48 exhibits notably elevated expression due to its oncogenic function. On the contrary, P42 functions as a tumor suppressor[20]. In CRC cells, P48 and P42 bind to F-box and WD40 domain protein 7 (FBXW7), a substrate recognition subunit of SCF E3 ligase complex, and P48 is ubiquitinated and degraded in a GSK-3b-dependent way, while P42 EBP1 remains resistant to degradation. The tumor suppressor function of FBXW7 is accelerated by P42 to improve the link of FBXW7 and its oncogenic targets, like Cyclin E. P48 contributes to re-localization of FBXW7 from nucleus to cytoplasm, where it fails to bind to its oncogenic nuclear targets[20]. In this study, the expression of nuclear-localized circ86591 was knocked down using ASO reagents. The results showed that CCNE2 and CCNA2 expression were decreased in CRC cells with transfected ASO-circ86591, which was restored by the treatment of the proteasome inhibitor MG132. Circ86591 is hypothesized to regulate the cell cycle by competitively binding to EBP1. Downregulation of circ86591 increased the binding of EBP1 to FBXW7 and CCNE2 and CCNA2, resulting in the ubiquitination-dependent degradation of CCNE2 and CCNA2. Consequently, CRC cells were arrested in the G0−G1 and S phase of the cell cycle and were unable to proliferate.
Fatty acid synthesis controls the balance between carbon source storage and energy expenditure. This process influences various metabolic pathways, including amino acid metabolism as well as glucose metabolism. Consequently, fatty acid synthesis is often alternated to harness the intracellular metabolism network to satisfy demands for materials and energy during the progression of cancer. ACC1 plays a critical role in CRC progression. ACC1 is an enzyme in fatty acid synthesis, which is responsible for the conversion of acetyl-coenzyme A to malonyl-coenzyme A[21]. ACC1 overexpression is tightly associated with tumor progression and poorer prognosis in CRC. During CRC progression, several factors have been reported to regulate fatty acid synthesis in tumor cells via ACC1. Silencing of tumor suppressor protein-protein tyrosine phosphatase receptor type O (PTPRO) can activate the AKT/mTOR signaling axis to increase sterol regulatory element-binding protein 1 (SREBP1) expression and its target protein ACC1, thereby promoting de novo fatty acid production[22]. In addition to protein-protein interactions, many ncRNAs can also regulate ACC1 expression to participate in the fatty acid synthesis process. CircACC1 from ACC1 pre-mRNA plays a vital role in the cellular response to metabolic stress. Transcription factor c-Jun is elevated, and circACC1 is preferentially produced over ACC1 mRNA in a serum-deprived environment. It stabilizes and enhances adenosine monophosphate-activated protein kinase (AMPK) holoenzyme activity by generating a ternary complex with its regulatory β and γ subunits. circACC1 cellular levels regulate fatty acid β-oxidation and glycolysis, leading to significant alterations in cellular lipid storage. Silencing or overexpressing circACC1 inhibits or enhances tumor growth, respectively, in vivo[23]. Therefore, ACC1 is a therapeutic target for cancer treatment and metabolic diseases. Although the molecular mechanism by which circ86591 binds ACC1 to play a role is not yet clear, downregulation of circ86591 decreased the metabolic level of the cells, including nucleotide metabolism and lipid metabolism, leading to impairment of the cells' proliferative capacity.
Many available chemotherapeutic drugs target rapidly dividing cells, whereas tumor cells can evade the effects of drugs by modulating the cell cycle or entering a resting state, which in turn increases drug resistance. CCK-8 assay was used to detect cell survival of colon cancer cells after knockdown of circ86591 against the antimetabolites and antibiotic analogs 5-FU, Gem, and Dox. The results indicated that downregulation of circ86591 expression increased the resistance of tumor cells to 5-FU, Gem, and Dox. In spite of the fact that knockdown of circ86591 induced lower metabolism and slower proliferation of tumor cells, it caused an elevation of the tumor cells' resistance to chemotherapy, balancing the relationship between cell growth and drug resistance is a challenge that circ86591 must overcome as a therapeutic method.
Beyond exploring the molecular mechanism of downregulation of circ86591, the function of upregulation of circ86591 was also investigated. Strikingly, contrary to the prediction that high expression of circ86591 would promote tumor cell growth, the proliferative capacity of tumor cells remained inhibited after raising the expression level of circ86591. Overexpression of circ86591 in HCT116 and LOVO cells led to a significant decrease in cell proliferation, determined by CCK-8 assay. The HCT116 cells were injected subcutaneously into immunodeficient mice, and when tumors grew, they were divided into two groups and injected with ADV circ86591 or ADV ctrl (n = 5 per group). Tumors derived from the ADV circ86591 group exhibited smaller volumes compared to those from the ADV ctrl group. RNA-seq data between the ADV circ86591 and ADV ctrl groups of HCT116 cells demonstrated that the expression level of tumor suppressor gene P53 was elevated. In addition, the Notch and AKT signaling pathways were abnormally activated by the Reactome enrichment analysis.
In the present study, upregulation of circ86591 might shift its location from mainly in the nucleus to the cytoplasm, resulting in its exertion as a tumor suppressor. High expression of circ86591 in the cytoplasm activated negative regulation of AKT signaling pathways, which may have led to elevated P53 and AKT activation, and continued AKT activation in turn led to negative feedback regulation, which inhibited cell growth. Tumor cell proliferation was significantly impaired by the combined treatment of circ86591 adenovirus supernatant and ACC1 inhibitor-CMS121, indicating that circ86591 could be a potential RNA to improve the anti-tumor efficiency of lipid metabolism inhibitors. Following ASO-mediated knockdown of nuclear circ86591, cell proliferation remained suppressed. This finding suggested that circ86591 downregulation might influence tumorigenesis through distinct pathways. RNA-seq analysis revealed that nuclear circ86591 modulated nucleotide metabolism and cell cycle pathways, thereby affecting tumor cell sensitivity to anticancer drugs (Fig. 6f). Contrary to most studies, both upregulation and downregulation of circ86591 expression induced tumor suppression. This is attributed to the critical role of circRNA localization in determining its function, as different subcellular sites or translocations may lead to distinct outcomes. Additionally, the dynamic equilibrium and complex regulation of RNA within tumor cells might also contribute to this phenomenon.
Despite the valuable findings, several limitations should be acknowledged. First, precise molecular consequences of the circ86591-EBP1/ACC1 interactions (like on EBP1-FBXW7 binding or ACC1 enzymatic activity) are not fully elucidated. The therapeutic potential of targeting circ86591 is not robustly validated in vivo using a treatment-oriented model. Second, the clinical relevance of the findings would be strengthened by correlating circ86591 expression with patient outcomes or drug responses in clinical cohorts. This study is limited by the lack of analysis in human clinical samples. While the findings offer compelling evidence for the role of circ86591 in anti-cancer effects in preclinical models, future validation in patient-derived tissues is essential to confirm its clinical relevance. Ongoing efforts are focused on establishing the necessary collaborations and resources to address this critical aspect in subsequent work. These limitations underscore the necessity for more rigorous experimental validation to improve the reliability and clinical applicability of this study.
-
In summary, circ86591 demonstrated a unique effect on tumor growth, and its binding proteins, including EBP1 and ACC1, played a critical role in cell cycle and lipid metabolism disorders. Either knockdown of circ86591 in the nucleus or elevation of circ86591 in the cytoplasm inhibited cell proliferation. Therefore, circ86591/EBP1/ACC1 may be a promising therapeutic target for CRC treatment.
This study was supported by the National Natural Science Foundation of China (Grant No. 82103289). We express our gratitude to the dedicated staff and participants involved in this study for their valuable contributions.
-
All mice experiments were carried out in accordance with the Guidelines for the Care and Use of Laboratory Animals published by Ministry of Health, People's Republic of China, and approved by the Ethics Review Committee for Animal Research-Beijing Viewsolid Biotechnology Co., Ltd (Approval No. VS2126A01218).
-
The authors confirm contribution to the paper as follows: study conception and design, original draft preparation, review & editing: Ma W; investigation, supervision: Hu J. Both authors reviewed the results and approved the final version of the manuscript.
-
The original contributions presented in the study are included in the article and its supplementary material. Further inquiries can be directed to the corresponding author on reasonable request.
-
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this article.
- Supplementary Table S1 Primer sequence information used in the study.
- Supplementary Table S2 Antibodies information used in the study.
- Supplementary Fig. S1 The relationship between circ86591 and linANRIL.
- Supplementary Fig. S2 Down-regulated circ86591 regulated the nucleotide metabolism and cell cycle pathways.
- Supplementary Fig. S3 The correlation between ACC1 and nucleotide metabolism-related proteins through the GEPIA database.
- Supplementary Fig. S4 Upregulation of circ86591 in cells altered the expression of proteins involved in nucleoside metabolism and cell proliferation.
- Supplementary Fig. S5 The WB assay confirmed the decline of ELOVL5 and ORP8 protein levels in HCT116 and SW480 cells transfected with ASO-circ86591 or ASO-NC.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Ma W, Hu J. 2026. circ86591's suppression of colorectal cancer cell proliferation via cell cycle and metabolic regulation. Epigenetics Insights 19: e004 doi: 10.48130/epi-0026-0001
circ86591's suppression of colorectal cancer cell proliferation via cell cycle and metabolic regulation
- Received: 17 July 2025
- Revised: 18 November 2025
- Accepted: 25 December 2025
- Published online: 27 February 2026
Abstract: Circular ribonucleicacids (circRNAs), marked by their covalently closed-loop structures, serve as crucial regulators in tumor development and progression. This study aimed to explore how circ86591, one of the isoforms of circular ANRIL, affects the progression of colorectal cancer (CRC). Real-time quantitative polymerase chain reaction (qRT-PCR) was utilized to examine circ86591 expression in cells. RNA pull-down, mass spectrometry, RNA immunoprecipitation, and western blotting (WB) were applied to elucidate molecular mechanisms underlying circ86591 and its binding proteins. RNA-sequencing, along with gain- and loss-of function assays, was conducted to uncover the tumor-suppressive effect and the relevant signaling pathways of circ86591. Circ86591 is lowly expressed in CRC cells overexpressing oncogenic linANRIL. Mechanistically, circ86591 directly interacts with ErbB3-binding protein (EBP1) and acetyl coenzyme A carboxylase (ACC1) proteins to modulate cell cycle and lipid metabolism. The knockdown of circ86591 by antisense oligonucleotide (ASO) increases the drug resistance of CRC cells to 5-FU, gemcitabine, or doxorubicin. Overexpression of circ86591 suppresses CRC growth both in vitro and in vivo via upregulation of P53 and negative feedback of the protein kinase B (AKT) pathway. Combining circ86591 adenovirus (ADV) with the anti-lipid metabolism drug CMS121 yields synergistic anti-tumor effects. In conclusion, the circ86591 inhibits CRC progression and holds potential as a therapeutic target for those tumors with active lipid metabolism.
-
Key words:
- circ86591 /
- EBP1 /
- ACC1 /
- Lipid Metabolism /
- Cell cycle













