-
Plant transformation and genome editing are indispensable tools for crop improvement and functional genomics[1,2]. However, transgene expression is often unstable or silenced because of the vector's design, the host background, or insertion sites, which limits their functional validation, line selection, and breeding applications[1,3−16]. Reliable, broadly applicable, and growth-neutral visual markers are therefore critical to accelerate transgene screening and improve the efficiency of transformation.
Existing reporters, such as fluorescent proteins and β-glucuronidase (GUS), require specialized equipment, show restricted expression, or can affect plant growth[17−22]. The 2A-linked unit of CYP76AD1, DODA, and Glucosyltransferase (RUBY) system, a natural pigment-based reporter visible as red coloration without instruments, offers a promising alternative[23−33]. However, its cross-species applicability, dependence on the vector backbone, and potential for synchronized monitoring of genome editing events remain unexplored. In this study, we constructed and optimized a RUBY-based visual platform, assessed the effects of the vector backbone on the stability of expression, and demonstrated its robust performance in Arabidopsis thaliana, tobacco (Nicotiana tabacum), and tomato (Solanum lycopersicum). We also developed inducible and CRISPR-associated protein 9 (Cas9)-porcine teschovirus-1 2A peptide (P2A)-RUBY modules for temporal monitoring and real-time visualization of editing events[34], providing a versatile solution for stable transgene screening and genome editing workflows.
-
The overexpression vectors p35S::RUBY-A and p35S::RUBY-B constructed in this study were engineered on the basis of the backbone vectors, namely the pGC-4 generated in the present experiment and the DR5::RUBY plasmid kindly provided by Professor Yunde Zhao's laboratory. The full-length coding sequence of RUBY was amplified via polymerase chain reaction (PCR) using the DR5::RUBY plasmid as the template, and the amplified fragment was used to replace the green fluorescent protein (GFP) gene sequence in the pGC-4 backbone vector. Thereafter, the cauliflower mosaic virus (CaMV) 35S promoter was cloned into the predigested pGC-4 backbone vector, and the recombinant overexpression vector was designated p35S::RUBY-B. For the construction of p35S::RUBY-A, the CaMV 35S promoter was inserted into the predigested DR5::RUBY vector to substitute for the original DR5-responsive element sequence, and the resulting vector was named p35S::RUBY-A. In addition, the LexA-VP16-Estrogen receptor (XVE)-inducible regulatory sequence was integrated into the predigested DR5::RUBY vector, with the resulting inducible expression vector named pXVE::RUBY. For the gene knockout assay, the pCAMBIA 1300 vector, provided by Professor Hao Chen's research group, was used as the backbone for constructing the knockout vector. The full-length RUBY sequence was inserted into the predigested pCAMBIA 1300 vector, and the final clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9-mediated knockout vector was designated p35S::Cas9-P2A-RUBY.
Genetic transformation of tobacco, Arabidopsis, and tomato
-
Recombinant plasmids were introduced into Agrobacterium tumefaciens C58/GV3101 for Agrobacterium-mediated transformation of tobacco, Arabidopsis, and tomato. For transient transformation in tobacco, bacterial suspensions were injected into the underside of the leaf and incubated under dark conditions for 24 h, followed by normal light exposure for further incubation and sampling at 48 and 72 h.
For stable transformation, plant explants from tobacco and tomato were precultured, infected with Agrobacterium, and co-cultivated under sterile conditions. Tobacco and tomato explants were transferred to a bud induction medium after dark treatment, with subsequent subculture of the callus tissues. Buds were allowed to develop, then were transferred to the rooting medium. Rooted seedlings were acclimated and transplanted into nutrient soil. Transformation of Arabidopsis was carried out via the floral dipping method. Homozygous transgenic plants from all species were selected for observing the phenotype.
Phenotypic observation and statistics
-
The leaf tissues of plants were observed and imaged using a Hitachi TM3030Plus tabletop scanning electron microscope in this study. The quantification of seven trichome types in tomato was performed with ImageJ software, following the following specific procedures: Opening the software and directly dragging the target images into the operation interface; navigating to the menu bar; selecting the options Plugins → Analyze → Cell Counter → Cell Counter–Keep Original, then clicking Initialize; and sequentially marking the seven trichome types with the labels Type I to VII for manual counting. Finally, the counted data were exported to Microsoft Excel for subsequent statistical analysis.
Root length was also measured via ImageJ software, and the specific operational steps were as follows: Launching the software and importing the target images by direct dragging, then selecting the Segmented Line tool to preliminarily calibrate the image scale. Next, Analyze → Set Scale was selected, the actual known distance and the corresponding unit of length were input according to the image scale, with the other parameters kept to the default settings. After scale calibration, the Segmented Line tool was reselected to trace and measure the root length on the image, and the shortcut Ctrl + M was used to obtain accurate measurement results. All measured data were exported to Microsoft Excel for further statistical analysis.
Growth conditions and cultivation conditions for plant materials
-
All experimental materials in this study were grown in the greenhouse. The temperature during the day was 28 °C with 16 h of light; at night, it was 18 °C with 8 h of darkness. The humidity was 60%, and the light intensity in the greenhouse was 120−130 µmol·m2·s. Regular watering and fertilization were applied, and all plants were cultivated in 9 cm × 9 cm nutrient pots with PINDSTRUP as the growth medium.
-
The different binary vector backbones markedly influenced transgene expression. Using the RUBY system, we constructed two binary vectors, p35S::RUBY-A and p35S::RUBY-B (Supplementary Fig. S1a, S1b). Transient expression in tobacco readily produced visible red pigmentation, indicating that both constructs maintained functional integrity (Supplementary Fig. S2). In Arabidopsis, both vectors exhibited comparable expression with only minor differences (Fig. 1a). In contrast, in stably transformed tomato and tobacco plants, p35S::RUBY-A produced strong and stable red pigmentation in the flowers and fruits. In the leaves, tobacco displayed pronounced red coloration, whereas tomato leaves remained largely colorless (Supplementary Fig. S3); by comparison, p35S::RUBY-B showed minimal pigmentation across nearly all tissues (Fig. 1b-g). Notably, during tissue culture, the two species also displayed differential expression: Tobacco calli showed clear red coloration, whereas tomato calli exhibited almost no visible pigment. Together, these results indicate that the vector backbone not only affects the stability of transgene expression but also largely determines expression efficiency and tissue- and species-specificity. This property establishes a foundation for the application of RUBY in transgene screening.
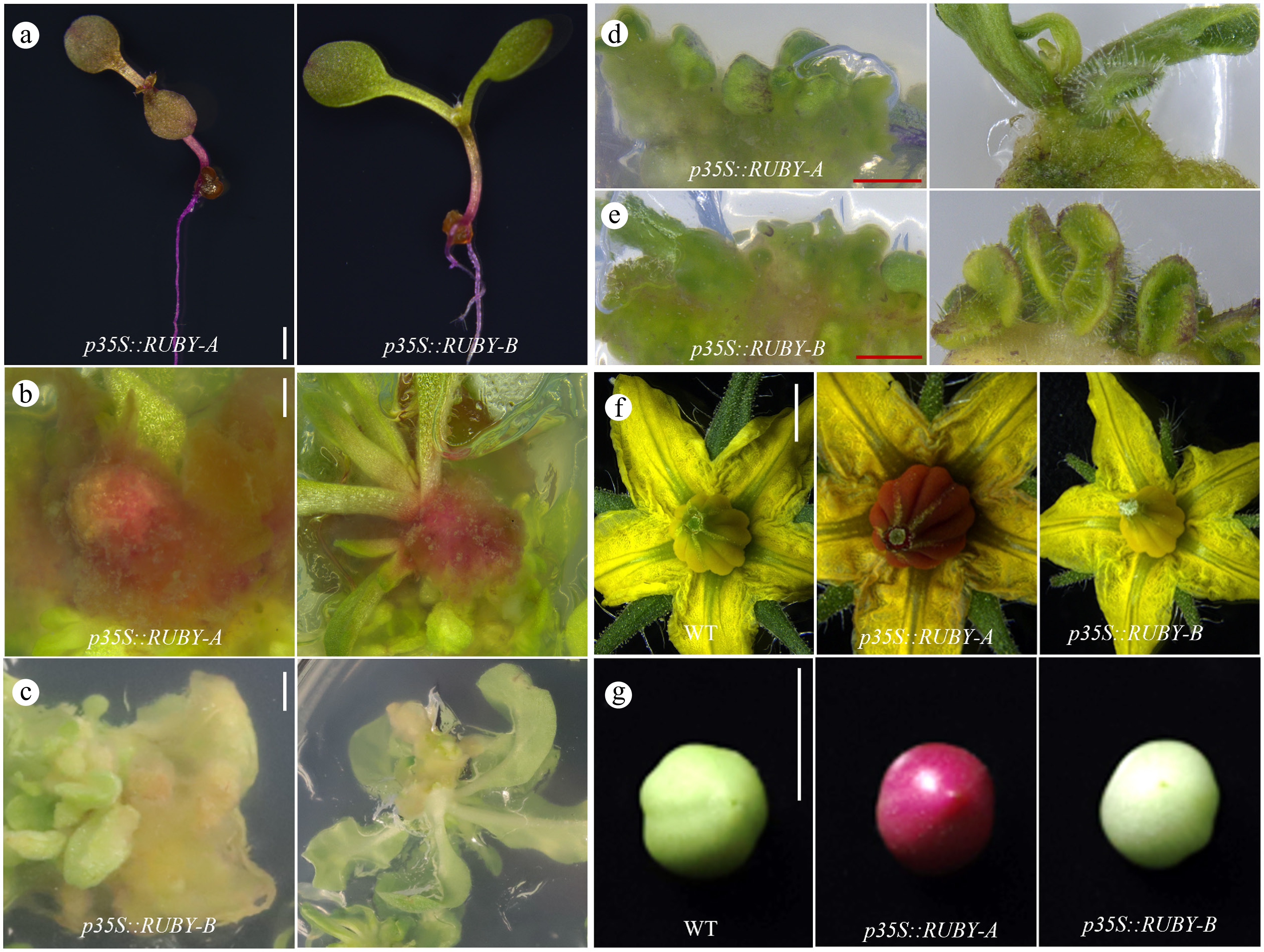

Figure 1.
The vector backbone affects the expression of RUBY system across species and tissues. (a) Different vector backbones affect the expression of RUBY system in Arabidopsis seedlings, scale bar = 1 mm. (b), (c) Different vector backbones affect the expression of RUBY in tobacco (Nicotiana tabacum) tissue culture, scale bar = 1 mm. (d), (e) Different vector backbones induced no obvious pigment deposition in tomato tissue culture, scale bar = 1 mm. (f), (g) The RUBY system's expression in tomato flowers and fruits. (f) Scale bar = 1 mm. (g) Scale bar = 1 cm.
RUBY enables rapid identification of single-copy transgenes
-
Early and accurate identification of single-copy transgenic lines is critical for subsequent genetic analyses and phenotypic evaluation. Given the stable and visually distinct expression of RUBY at the seedling stage, we assessed its utility for single-copy transgene screening. T1 seedlings of Arabidopsis lines carrying p35S::RUBY-A or p35S::RUBY-B were analyzed, and could be directly distinguished on the basis of red versus green coloration. For example, seedlings of the p35S::RUBY-A#3 line segregated approximately 3:1 for red versus green (Fig. 2a), consistent with single-copy Mendelian inheritance (χ2 test, p > 0.05) (Fig. 2b). These findings demonstrate that RUBY provides a rapid, intuitive, and instrument-free strategy for efficiently identifying single-copy transgenic lines at the seedling stage, offering a reliable tool for transgene screening in plants.
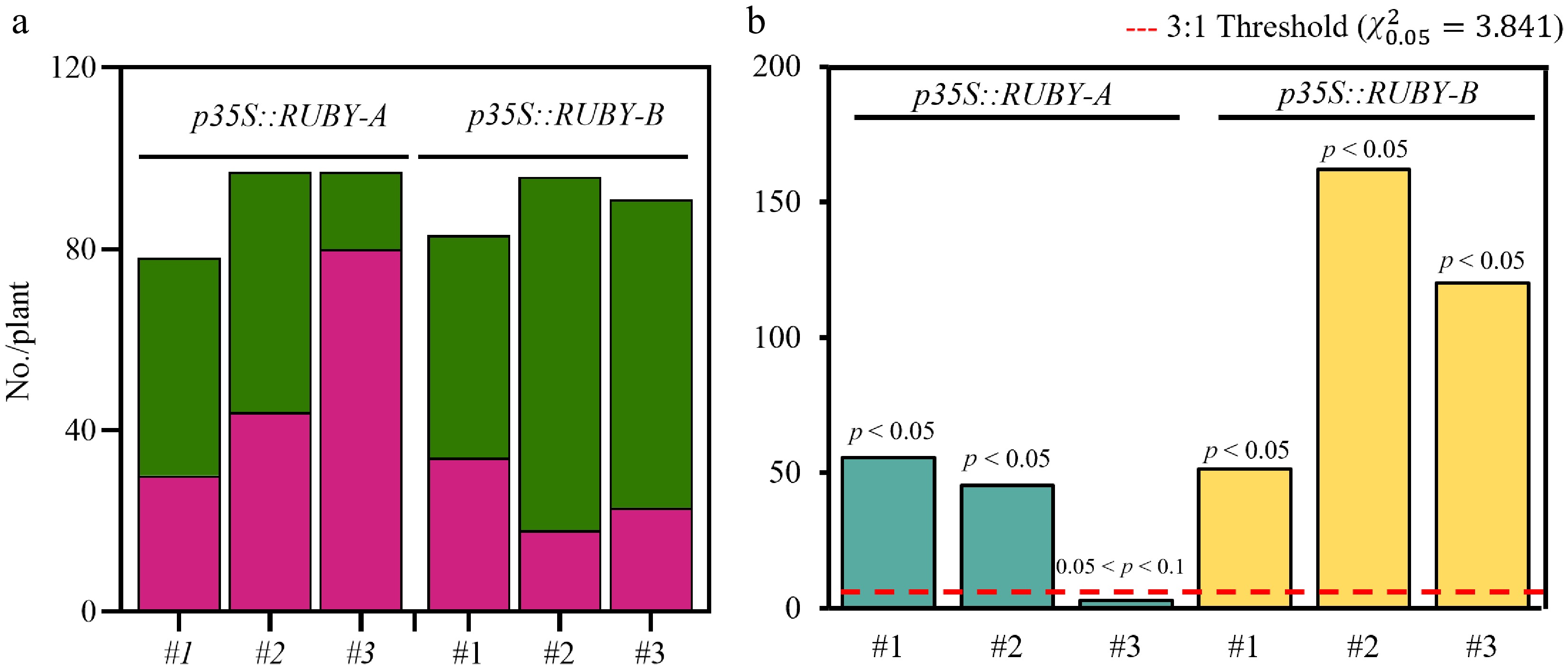
Figure 2.
Copy number detection of the RUBY system in Arabidopsis. (a) Quantification of RUBY's expression in p35S::RUBY-A and p35S::RUBY-B lines was performed to assess positive and false positive rates. Positive, red pigmentation; false positive, green pigmentation. (b) Chi-square analysis: The red dashed line = 3:1 segregation threshold (χ20.05 = 3.841); bars show the χ2 values; p < 0.05 = significant fit to single-copy segregation.
RUBY does not affect plant growth or epidermal traits
-
Despite the excellent performance of RUBY in expression monitoring and transgene copy identification, whether its constitutive expression may cause unintended physiological effects still required verification. Therefore, we conducted a systematic assessment of growth performance and epidermal characteristics in p35S::RUBY-A transgenic lines. Compared with the wild-type, no obvious differences were observed in root growth, plant height, leaf morphology, or trichome development (Fig. 3a-h). These results indicate that continuous RUBY expression does not impose a detectable growth penalty or alter epidermal traits, thereby supporting its applicability as a growth-neutral and reliable visual marker for plant biotechnology.
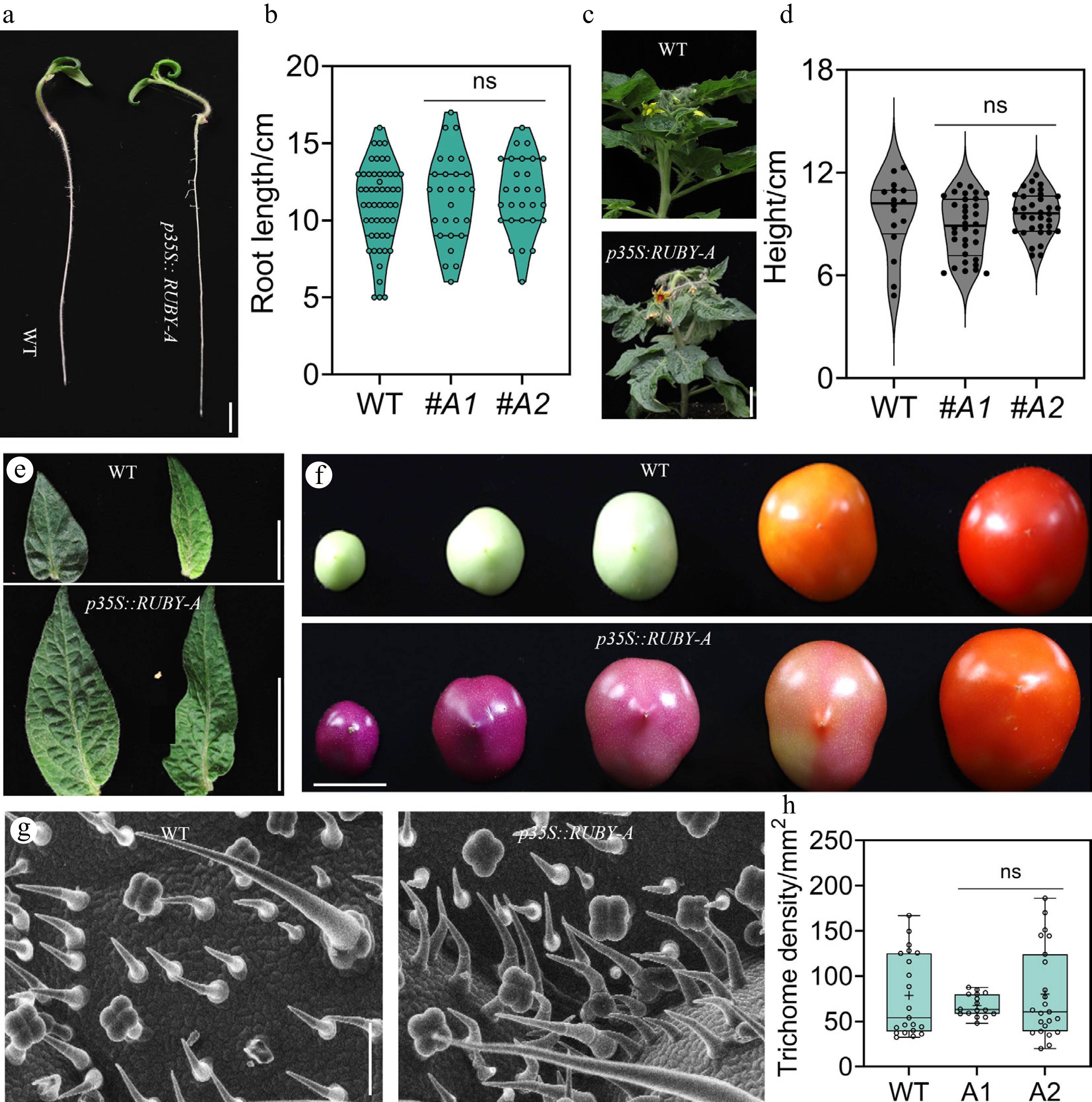
Figure 3.
Plants' development and epidermal differentiation with the RUBY system. (a), (b) Phenotypic observation and quantification of wild-type (WT) and p35S::RUBY plants, scale bar = 1 cm. (c), (d) Observation and quantification of plant height in WT and p35S::RUBY plants, scale bar = 1 cm. (e) Analysis of leaf morphology in WT and p35S::RUBY plants, scale bar = 1 cm. (f) Analysis of fruit morphology and size in WT and p35S::RUBY plants, scale bar = 1 cm. (g), (h) Observation and quantification of trichome morphology in WT and p35S::RUBY plants, scale bar = 1 mm.
Design of inducible and genome editing-compatible RUBY systems
-
To enable RUBY to function not only as a stably expressed visual selection marker but also as a versatile tool adaptable to more sophisticated regulatory and application scenarios, we also incorporated it into temporal control and functional monitoring systems. In many basic research and applied contexts, precise activation of transgene expression at specific time points or under defined conditions is critically required[30,35−39]; however, most existing visual markers lack the capability to intuitively monitor this process. To address this limitation, we constructed a pXVE::RUBY module driven by the well-established and widely applied estrogen-inducible system. This design allowed RUBY signals to be precisely activated upon exogenous induction, thereby enabling on-demand initiation of the red pigmentation signal and providing a continuous, controllable, and time-resolved visual readout of the induction process (Fig. 4a, b). This strategy substantially expands the applicability of RUBY in gene regulation studies and synthetic biology.
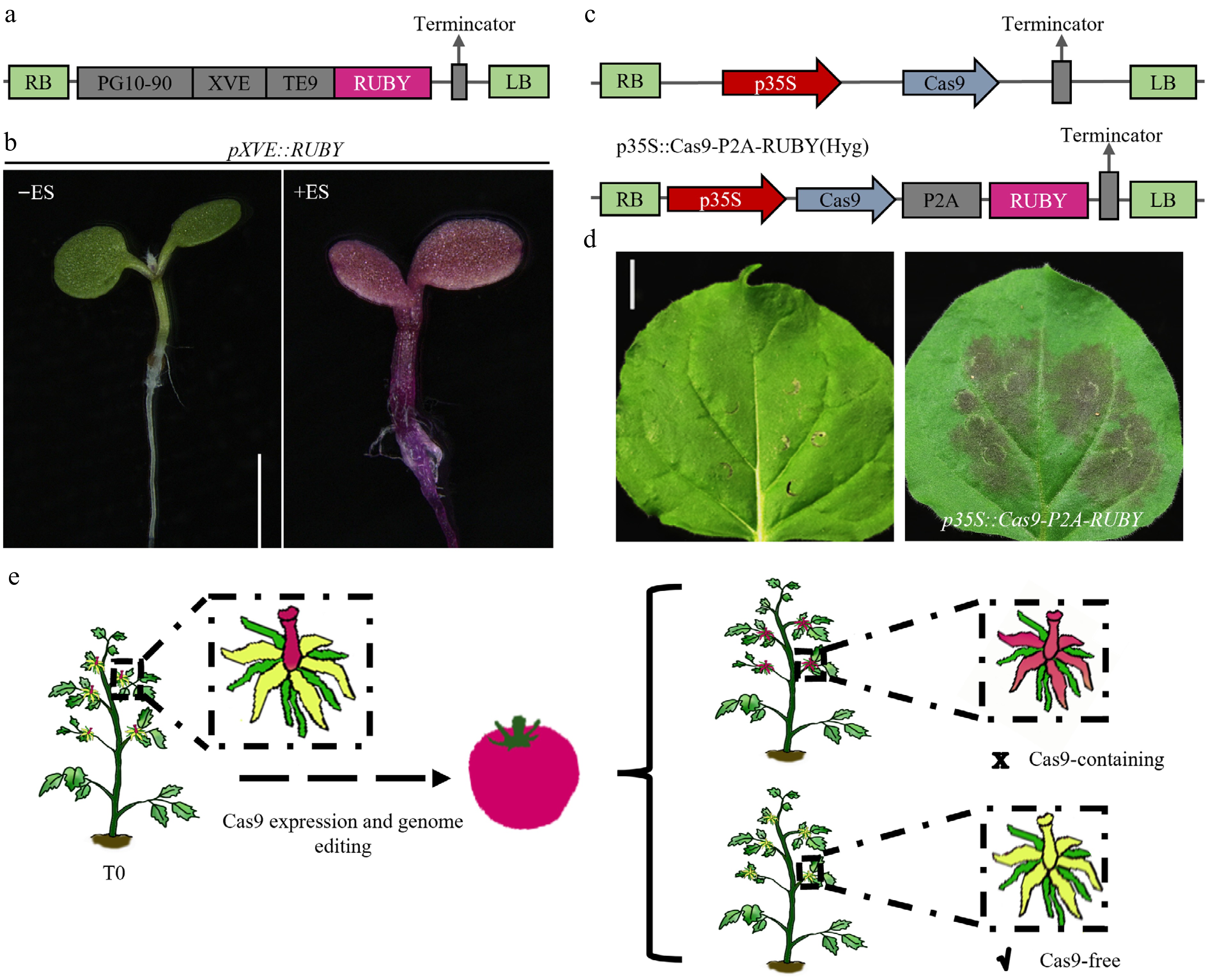
Figure 4.
Design and application of the RUBY system for inducible expression and visualizing genome editing. (a), (c) Schematic of the RUBY-labeled inducible expression and genome editing visualization system. (b) Validation of inducible expression using the RUBY system, scale bar = 1 mm. (d) Transient expression analysis of the RUBY-labeled genome editing visualization system in tobacco (Nicotiana benthamiana), scale bar = 1 cm. (e) A dual-phase CRISPR workflow enabling simultaneous gene editing and subsequent Cas9 elimination in tomato.
Meanwhile, in the field of genome editing, real-time and visual monitoring of editing events as well as the expression and activity status of Cas9 has been a longstanding technical challenge. Through a systematic examination of existing Cas9 editing lines (including the Slspx1 mutant background)[40], we found that the prolonged persistence of Cas9 protein after completion of the intended editing event may trigger unintended subsequent edits (Supplementary Fig. S4), thereby increasing the potential risk of genome instability. This observation underscores the necessity of implementing dynamic regulation and visual surveillance of Cas9 expression and the progress of editing in plant genome editing systems. Although strategies such as Cas9-P2A-GFP fusion constructs can report Cas9 expression, these approaches rely on specialized fluorescence-detection equipment, which increases operational complexity and limits the method's applicability in field evaluations and high-throughput screening. To overcome these constraints, we developed a Cas9-P2A-RUBY co-expression system that tightly couples Cas9's expression and editing activity with a readily visible RUBY pigmentation signal (Fig. 4c, d). This system enables an intuitive judgment of Cas9's functional status and efficient identification of edited materials without the need for sophisticated instrumentation. In tomato, this approach successfully achieved real-time visual tracking of genome editing events and markedly improved both screening efficiency and the controllability of the entire editing process (Fig. 4e).
-
Unstable or silenced transgene expression has long limited the efficiency of plant transformation and genome editing[4,6,15,41−48]. Here, we present a RUBY-based visual strategy that provides an integrated solution for monitoring transgene expression, identifying single-copy lines, and visualizing genome editing events. Unlike conventional fluorescent or GUS reporters, RUBY enables a direct observation of red pigmentation without specialized equipment[23−33], exerts minimal impact on growth or epidermal development, and demonstrates robust applicability across Arabidopsis, tobacco, and tomato[23,24,26,27,30,31,49,50].
The vector backbone substantially influences the stability and tissue specificity of expression, guiding the optimization of transgene design[5,7,8,12−14,16,50−55]. In this study, p35S::RUBY-A consistently yielded strong and stable pigmentation in tomato, whereas p35S::RUBY-B showed minimal expression, highlighting the importance of backbone selection in achieving robust, species-, and tissue-specific expression. The inducible pXVE::RUBY module and the Cas9-P2A-RUBY system further expand the utility of RUBY by enabling temporal control of transgene expression and real-time visualization of genome editing events. This dual functionality provides a powerful tool for monitoring induction kinetics, evaluating Cas9 activity, and improving the workflow's efficiency without reliance on specialized detection equipment. Compared with previous reports[23−33], our study further expands the understanding of RUBY's application potential. We not only validated the practical utility of RUBY in tomato and tobacco tissue culture systems and in stable transgenic lines, but also demonstrated that constitutive RUBY expression does not affect plants' growth or epidermal traits under routine expression levels. In addition, by integrating RUBY into a Cas9-linked system, we achieved a visual confirmation of genome editing events. Collectively, these results reinforce the versatility and growth-neutrality of RUBY as a visual marker.
Although our study confirms the efficacy and broad applicability of RUBY, further validation under multiplexed editing, diverse genetic backgrounds, or field conditions will be valuable. Nevertheless, this system provides a simple, reliable, and instrument-free platform for high-throughput transgene screening, precise regulation of expression, and visual monitoring of genome editing events, supporting both basic research and applied crop improvement.
-
The authors confirm their contributions to the paper as follows: study conception and design: Wu S, Sun C; data collection: Yang X, Wu K, Zhou X, Wang J, Wu S, Wu S, Liu C, Yi W, Fu Y, Zhang J, Zhu H; analysis and interpretation of results: Sun C, An N, Wei J, Chen S; draft manuscript preparation: Sun C. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article and its supplementary information files.
-
This research was funded by the National Natural Science Foundation of China (32370354) and (32561143028) to Wu S, and Supported by the Joint Research Program of State Key Laboratory of Agricultural and Forestry Biosecurity (No. SKLJRP2506). We thank Professor Yunde Zhao for generously providing the DR5::RUBY plasmid used in this study[30]; we also thank Professor Haodong Chen for generously providing the pCambia 1300 plasmidp[34].
-
The authors declare that they have no conflict of interest.
-
accompanies this paper online at: https://doi.org/10.48130/vegres-0026-0007.
-
# Authors contributed equally: Ningkai An, Jinbo Wei, Shiliang Chen
- Supplementary Fig. S1 Circular map of different vectors.
- Supplementary Fig. S2 Construction and validation of RUBY system with different vector backbones.
- Supplementary Fig. S3 Phenotypic observation of the RUBY system in different species and vector backbones.
- Supplementary Fig. S4 Detection of gene-editing persistence in the progeny of Cas9-positive plants.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
An N, Wei J, Chen S, Yang X, Wu K, et al. 2026. A visual reporter system for transgene screening and monitoring genome editing in tomato. Vegetable Research 6: e013 doi: 10.48130/vegres-0026-0007
A visual reporter system for transgene screening and monitoring genome editing in tomato
- Received: 15 January 2026
- Revised: 16 February 2026
- Accepted: 25 February 2026
- Published online: 31 March 2026
Abstract: The stable expression of integrated transgenes remains a major restriction in plant transformation, as different vector backbones and host contexts often lead to variable or silenced expression. Systematic evaluations of the binary vector architecture and broadly applicable visual markers for rapid line selection are still lacking. In this study, we developed a streamlined 2A-linked unit of CYP76AD1, DODA, and Glucosyltransferase (RUBY)-based chromogenic reporter system and validated its robustness in the transformation in Arabidopsis thaliana, tobacco (Nicotiana tabacum), and tomato (Solanum lycopersicum). A comparative analysis of two binary vector backbones revealed strong structural effects on the stability of expression, with one backbone enabling consistently high pigmentation in tomato. RUBY's intensity provides an immediate readout of transgene activity and allows the rapid identification of single-copy, stably expressing lines through seedling segregation. RUBY expression does not influence plants' growth or epidermal development, demonstrating excellent biocompatibility. We also established an inducible pXVE::RUBY module and a CRISPR-associated protein 9 (Cas9)-porcine teschovirus-1 2A peptide (P2A)-RUBY system for real-time monitoring of transgene induction and visualizing genome editing events. This platform offers an efficient and readily implementable strategy to overcome the disconnect between transgenes' integration and expression, while substantially improving the overall performance of plant transformation and genome editing pipelines.













