-
Mammalian iron homeostasis depends on a network that coordinates systemic regulation via the hepcidin-ferroportin signaling pathway, along with intracellular processes of post-transcriptional control largely mediated by interactions of two iron regulatory proteins (IRPs), IRP1 and IRP2, with RNA stem-loops found in multiple iron metabolism transcripts. Intracellular ferritin stores iron, lysosomes recycle stored iron for use, and iron-sulfur (Fe-S) cluster biogenesis is required to maintain iron homeostasis through its role in regulating IRP function and turnover[1−4]. Although these canonical processes describe how cells detect the availability of iron to satisfy metabolic needs, it has been unclear how transcription programs, and chromatin states, contribute to the maintenance of iron availability in a variety of pathophysiological conditions[1]. Distinguishing cell populations that respond to changes in iron levels is a precondition for identifying upstream regulators of iron-responsive pathways. However, direct measurements of iron levels are difficult to identify the unsuspected contributors to iron homeostasis.
To overcome the limitations of traditional biochemistry, Martinelli et al. recently developed a sensitive and simplified tool by utilizing IRP2 protein levels as an indicator of iron status. In combination with CRISPR-based functional screening, they were able to perform functional screening of the iron metabolism network, enabling them to discover that cellular iron metabolism is subject to epigenetic regulation. They demonstrate that SETD2 regulates alterations of chromatin that modify accessibility for transcription of iron metabolism genes through alterations in histone methylation, closing a gap in our understanding of epigenetic regulation of iron uptake, distribution, and utilization[5]. Their work adds iron-dependent control of chromatin status to the known classical post-transcriptional and metabolic regulatory systems.
-
The IRP2 protein level serves as a metabolic sensor of iron loading. At high iron levels, IRP2 is degraded by FBXL5, while at low iron levels, high IRP2 expression is induced that binds to iron regulatory elements (IREs), thereby regulating iron homeostasis[6]. Accordingly, the authors of the present study employed IRP2 expression as a metabolic indicator of iron levels for screening upstream networks regulating iron homeostasis. Based on this rationale, the present authors designed and used a novel and elegant IRP2-fused fluorescent reporter with genome-wide CRISPR functional screening to uncover unknown players in the critical network of cellular iron regulation. In this screening system, the present authors demonstrated that iron levels are tightly controlled by several well-known regulators, such as transferrin receptor-mediated cellular iron uptake and recycling, NCOA4-dependent ferritinophagy, and Fe-S cluster assembly, which supports the validity of their functional screening approach. More importantly, the authors of the present study were able to use this detection tool to uncover new factors involved in endolysosomal trafficking, epigenetic regulation, and exocyst complex formation (Fig. 1a), all of which contribute to the regulation of intracellular iron flux[1,5,7]. Although alterations in IRP2 expression levels may not fully capture all iron homeostasis pathways, their screen maps the landscape of iron regulators correlated with IRP2.
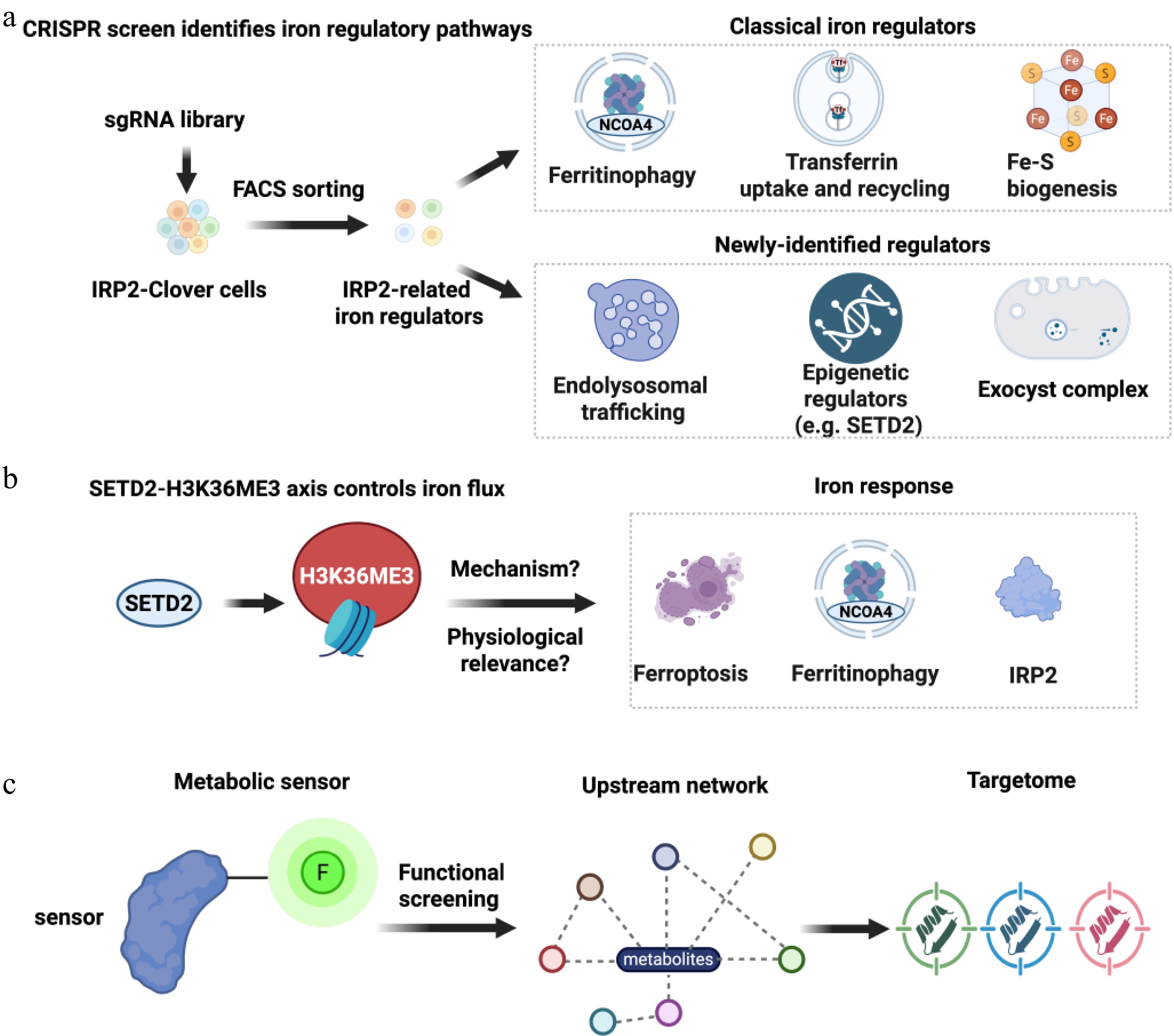
Figure 1.
Integrating genome-wide CRISPR screening and metabolic sensors to reveal cellular iron regulatory networks. (a) Genome-wide CRISPR screening with IRP2-Clover reporter cells revealed classical pathways and new regulators of cellular iron homeostasis. (b) A suggested paradigm that has the SETD2-H3K36me3 axis in the regulation of ferroptosis sensitivity, ferritinophagy, and IRP2. (c) Model for use of the fluorescent-fused metabolic sensor as a proxy for metabolite levels to screen and identify the upstream network and potential targetome. Note: figure was created using BioRender.com.
-
One of the most important discoveries of the original study is that SETD2, a histone H3K36 methyltransferase[8], is a new regulator of iron homeostasis[5]. Epigenetic modifications may have direct effects on the availability of iron, and the function of iron-sensitive metabolic pathways[9,10]. Since the post-transcriptional control of intracellular iron uptake, storage, and utilization has long been considered to be mainly regulated by the IRP/IRE system[1,3,4], the finding that SETD2 influences iron regulatory programs, through indirect regulation of IRP2 levels, elevates the role of the chromatin modification status in directly regulating the iron response. Depletion of SETD2 impairs ferritinophagy, and it correlates with decreased vulnerability to ferroptosis, suggesting a place for epigenetic control of iron-dependent cell death pathways[5,11,12] (Fig. 1b). This is especially applicable to cancer, in which the instances of SETD2 mutations are prevalent[13]. It is possible that SETD2-mutant tumors can thus be affected by therapies targeting ferroptosis.
-
Though specific regulatory pathways in iron metabolism have been studied extensively, a high-throughput way to more completely map the landscape of iron metabolism was previously lacking. By utilizing the novel screening system, the authors here developed a potential high-throughput method for detection of unknown iron metabolic pathway regulators, which may be applicable to related pathophysiological conditions such as anemia, cancer, and neurodegeneration, and may identify potential novel targets for therapy. Identification of SETD2 as an iron homeostasis regulator demonstrates that the methylation state of chromatin can affect therapeutic responses, due to having a potent effect on iron metabolism. Despite the fact that SETD2 is not a directly druggable target, due to its role in global chromatin regulation, some iron-handling mechanisms controlled by SETD2, such as ferritinophagy, IRP2-mediated regulation, the control of the labile iron pool, and ferroptosis, could be more accessible to pharmaceutical interventions. The reduced sensitivity of ferroptosis in SETD2-deficient tumors indicates that the tumors have disturbed iron recycling, which may restrict the efficacy of ferroptosis-based therapy[5]. Simultaneously, the changes can result in the development of new iron-dependencies which can be targeted in developing new therapies. Together, these findings indicate that the role of chromatin modifiers in developing iron-related metabolic vulnerability is possible, and that mapping the landscape of different iron-related diseases will be useful for finding novel therapeutic targets.
-
Despite the original study revealing that SETD2 is a chromatin level regulator of intracellular iron homeostasis, there are still some major questions left. To define how SETD2 and H3K36me3 control iron metabolism, a major step will be to determine what transcriptional and splicing targets allow SETD2 to control iron availability, especially those that affect lysosomal activity, vesicular trafficking, and ferritinophagy. To have a wider mechanistic perspective, more epigenetic or other regulation layers that coordinate iron with Fe-S cluster biogenesis, oxidative stress response, mitochondrial activity, and operation remain to be studied. Moreover, considering that iron metabolism is highly optimized to serve tissue-specific requirements, it will also be of interest to evaluate how the regulation of SETD2 activity occurs within physiologically relevant models, such as hepatocytes, renal epithelium, erythroid precursors, and tumor microenvironment cells. Lastly, the broader iron-related disease landscape, including anemia, cancer, and neurodegeneration, can be explored to find novel targets for potential therapy.
More broadly, this commentary presents a conceptual shift in studying the cellular homeostasis of iron. Contrary to more traditional methods, which usually aimed to identify an individual pathway or a single regulatory factor, CRISPR-based functional genomics, combined with a tool for sensing iron levels, can be used to more completely interrogate iron regulation in a systems biology approach. In this way, the authors discover a constellation of regulators that would be part of a putative iron-regulatory network, thereby advancing a more comprehensive and integrated insight into iron metabolism. Though causal and hierarchical connections in this network are yet to be completely addressed, the future combination of the functional screening with artificial intelligence-designed analytical models may be useful in determining causality and key driver nodes. These observations highlight the need for therapeutic modulation of iron homeostasis, which may reflect the activities of key drivers in an integrated regulatory network, constituting a targetome through which iron dysregulation may be manipulated. Use of a fluorescent tool as a high-throughput readout not only fits the sensing of iron levels by IRP2, but can also be utilized in other types of research investigating how cells sense and react to alterations in the various intracellular metabolite contents. This study employs a fluorescence-labeled metabolic sensor screen to map upstream networks and identify unsuspected components, revealing a potential targetome and establishing a generalizable sensor-based functional screening paradigm for systematically expanding target discovery across metabolic landscapes (Fig. 1c).
-
Not applicable.
-
The authors confirm their contributions to the commentary as follows: study conception, manuscript revision, and supervision: Zhang Y, Rouault TA; draft manuscript preparation and figure creation: Zhang Y. Both authors reviewed the results and approved the final version of the manuscript.
-
Data sharing is not applicable to this commentary as no datasets were generated or analyzed.
-
The authors thank the Eunice Kennedy Shriver National Institute of Child Health and Human Development for support. This work was funded by the Intramural Research Program of the National Institutes of Health (1ZIA HD001602-Regulation of Intracellular Iron Metabolism).
-
The authors declare no conflict of interest.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of China Pharmaceutical University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhang Y, Rouault TA. 2026. An iron sensor-based approach discovers a role for epigenetic regulation in iron homeostasis. Targetome 2(1): e007 doi: 10.48130/targetome-0026-0008
An iron sensor-based approach discovers a role for epigenetic regulation in iron homeostasis
- Received: 28 January 2026
- Revised: 01 February 2026
- Accepted: 02 February 2026
- Published online: 13 February 2026
Abstract: Iron homeostasis is tightly regulated by systemic and intracellular pathways, yet how transcriptional programs and chromatin states contribute to the maintenance of iron availability remains poorly understood. A recent study developed an elegant tool by leveraging iron regulatory protein 2 (IRP2) as a metabolic sensor, in combination with CRISPR-based functional screening to map the regulatory landscape of cellular iron metabolism. Using this strategy, the histone methyltransferase SETD2 is identified as a chromatin-based regulator of IRP2 levels, ferritinophagy, and ferroptosis sensitivity. These findings reveal a new epigenetic layer of iron regulation, and provide a broader model for the metabolite-responsive sensors, through functional screening, to identify upstream networks and potential targetome.
-
Key words:
- Iron homeostasis /
- Epigenetic regulation /
- SETD2 /
- Ferritinophagy /
- H3K36 methylation /
- IRP2