-
Plastic debris smaller than 5 mm is classified as microplastics (MPs)[1], and these particles have been detected across diverse ecosystems worldwide[2−4]. Critically, MPs exhibit distinct physicochemical properties, including hydrophobicity, surface charge, and buoyancy that foster colonization by specialized microbial assemblages. These unique biofilm communities, collectively termed the plastisphere[5], demonstrate significant divergence from planktonic counterparts in taxonomic composition, community assembly patterns, biomarker species, and functional metabolic pathways. Such differentiation underscores the plastisphere's role as an ecological niche that selectively enriches microbial taxa with implications for biogeochemical cycling and contaminant dissemination[6].
Biofilms not only provide a substrate for the attachment of foundational microbial groups such as bacteria, fungi, and algae[7,8], but also selectively enrich antibiotic resistance genes (ARGs). Numerous studies have shown that MPs in aquatic environments can preferentially accumulate ARGs and pose a potential risk for their dissemination. For instance, Wu et al.[9] reported that the abundance of ARGs on MPs in rivers was significantly higher than in the surrounding water, approximately three times that in the river water. Similarly, Yang et al.[10] found that in the North Pacific, the abundance of ARGs on MPs was 5.7 times higher than in seawater. Similar observations have also been reported in other environments, such as coral reef areas and wastewater treatment plants[11−15], further confirming the widespread impact of MPs.
Biofilm carriers can undergo long-distance transport in aquatic environments, allowing the associated microorganisms to 'hitchhike', thereby posing a series of ecological risks[16−18]. Previous studies have reported that during the migration of biofilm carriers, intense exchanges of microorganisms and ARGs occur between the water column and the biofilm[19]. Zhou et al. further compared the effects of biodegradable and non-biodegradable plastic matrices on the dissemination of ARGs, finding that biodegradable polybutylene adipate terephthalate (PBAT) poses a higher dissemination risk[20]. In addition, a study by Metcalf et al. demonstrated that pathogens hitchhiking on plastic and glass substrates discharged from wastewater treatment plants can survive for at least 25 d after entering seawater[21].
Although existing studies have confirmed that MPs can act as vectors for the dissemination of ARGs, little is known about the dynamic responses of microbial communities upon MP introduction into receiving water bodies, or about the underlying mechanisms driving these changes. To address this knowledge gap, this study further investigated this emerging pathway of ARG dissemination. Taihu Lake and its outflowing river, the Liangxi River, were selected as the study system, conducting in situ MP incubation in Taihu Lake. Through microcosm experiments, the temporal dynamics of microbial communities and ARGs were tracked following the translocation of MPs into the Liangxi River. From a metagenomic perspective, this work systematically characterizes the changes and responses of microbial communities and ARGs in the receiving water body after MP introduction.
-
This study conducted a one-month in situ incubation of MPs in Taihu Lake (31.25° N, 120.13 °E) during August 2023. Considering the widespread occurrence of MPs in aquatic systems, four representative MP pellets were chosen for this study, including three non-biodegradable polymers—polypropylene (PP), polyethylene (PE), and polyvinyl chloride (PVC)—and one biodegradable polymer, polylactic acid (PLA), which were mixed at an equal mass ratio. All materials were supplied by Prospect New Material Ltd. (Dongguan, China). Sieved MPs (2–4 mm particle size) were placed in stainless steel incubation devices (0.4 L volume, 1.2 mm aperture), submerged approximately 30 cm below the water surface. After 1 month, samples were retrieved and rinsed three times with phosphate-buffered saline (PBS) to remove impurities before further processing.
The Liangxi River is one of the main outflow rivers of Taihu Lake. Approximately 3 L of surface water was collected from the Liangxi River for laboratory use. Sterilized 250 mL conical flasks were divided into three groups: Group A received 200 mL of sterile ultrapure water (G-CLONE, CB1005), to which 5 g of MP samples incubated in Taihu Lake were added. Group B received 200 mL of Liangxi River water with the same MP samples added. Group C received 200 mL of Liangxi River water without MPs.
All samples were incubated in triplicate in illuminated incubators at 25 °C under 2,000 lux light intensity with a 12-h:12-h light:dark cycle. Flasks were shaken daily to maintain dissolved oxygen. A nutrient solution containing mineral salts, trace elements, and growth factors was supplemented according to OECD Guideline 301 to sustain microbial growth. After 1–5 weeks of incubation, biofilm-adhered microorganisms and planktonic microbes underwent metagenomic sequencing to analyze the temporal dynamics of microbial communities and ARGs in both MP biofilms and recipient waters. The methods of DNA extraction and metagenomic sequencing were described in the Supplementary Text 1.
Data processing
-
Prior to analysis, raw sequencing data underwent quality control using fastp[22] to ensure the reliability of subsequent results. Sequences were then assembled using Megahit[23], followed by open reading frame (ORF) prediction performed on contigs with Prodigal[24]. The predicted gene sequences were clustered using CD-HIT[25] to construct a non-redundant gene set (identity ≥ 90%, coverage ≥ 90%). This gene set was subsequently aligned against relevant databases using BLASTp[26] with an e-value cutoff of 1e-5. ARGs were annotated against the Comprehensive Antibiotic Resistance Database (CARD), while other functional genes were annotated using both the Clusters of Orthologous Groups (COG) and UniProt KB databases. Beta diversity distance matrices were calculated using QIIME[27], with non-metric multidimensional scaling (NMDS) analysis and visualization performed using the vegan package[11] in R.
-
Taxonomic annotation revealed Proteobacteria, Cyanobacteria, Bacteroidota, Planctomycetota, and Actinobacteria as the dominant phyla across all samples (Supplementary Fig. S1). Although microbial communities in MP biofilms exhibited similar compositions, planktonic communities diverged from biofilm assemblages, with Proteobacteria and Cyanobacteria consistently remaining predominant. Notably, Cyanobacteria showed significant enrichment in biofilms. This enrichment led to substantially higher proportions of Cyanobacteria in Group B compared to Group C (Supplementary Fig. S2). Conversely, Proteobacteria demonstrated markedly greater relative abundance in ultrapure water (Group A) than in Liangxi River water (Groups B/C). In this simplified environment, Proteobacteria exhibited enhanced resource-utilization and niche-occupancy capabilities, resulting in elevated dominance. By contrast, established ecological conditions in the Liangxi River water intensified microbial competition, enabling phyla like Chlorophyta and Planctomycetota to establish a significant presence.
At the family level, significant changes occurred in the microbial community structure of MP-amended river water (Group BW) compared to MP-free river water (Group CW). MPs notably increased microbial richness in recipient waters, with Chao1 indices demonstrating higher richness in BW (5,452.6) than in CW (4,886.6). The treatment groups contained 329 unique species. The significantly divergent families—Acaryochloridaceae, Thermosynechococcaceae, Calotrichaceae, Scytonemataceae, and Hapalosiphonaceae—all belonged to Cyanobacteria. As the dominant phylum in Taihu Lake, diverse cyanobacterial taxa were enriched on MP biofilms. During MP transport, these cyanobacteria may disperse into new aquatic environments, posing ecological risks. Notably, biofilm-associated families (Leptolyngbyaceae, Sphaerotilaceae, Acaryochloridaceae, Oculatellaceae, Thermosynechococcaceae, Nostocaceae, and Oscillatoriaceae) exhibited rapid proliferation in ultrapure water after MP incursion, particularly Acaryochloridaceae and Thermosynechococcaceae (Supplementary Fig. S3). Conversely, Pirellulaceae and Planctomycetaceae increased significantly in Liangxi River water but showed limited growth in ultrapure water. These environment-dependent responses suggest microbial adaptations linked to habitat conditions and niche competition.
NMDS analysis of microbial communities revealed a tight clustering of MP-associated samples from weeks 1 to 5, indicating greater temporal stability of biofilm communities (Fig. 1). This temporal stability may be attributed to the protective microenvironment provided by the biofilm matrix on MP surfaces, which buffers the microbial community against fluctuations in the surrounding water. In contrast, planktonic microbial communities were more widely dispersed and exhibited clear temporal shifts, reflecting the greater variability of environmental conditions in the water column. In Group A, the Bray–Curtis dissimilarity between planktonic microbial communities and biofilm communities was initially high in the first week, gradually declined, and reached its lowest point by week 3, after which it increased again (Fig. 2a). This temporal trend suggests the progressive release of microorganisms from MP biofilms into the water column following MP exposure, leading to a relatively equilibrated state between the two habitats by week 3. However, the distinct environmental conditions in biofilms and water subsequently favored the enrichment of different dominant taxa, resulting in the divergence of community structures in each habitat over time.
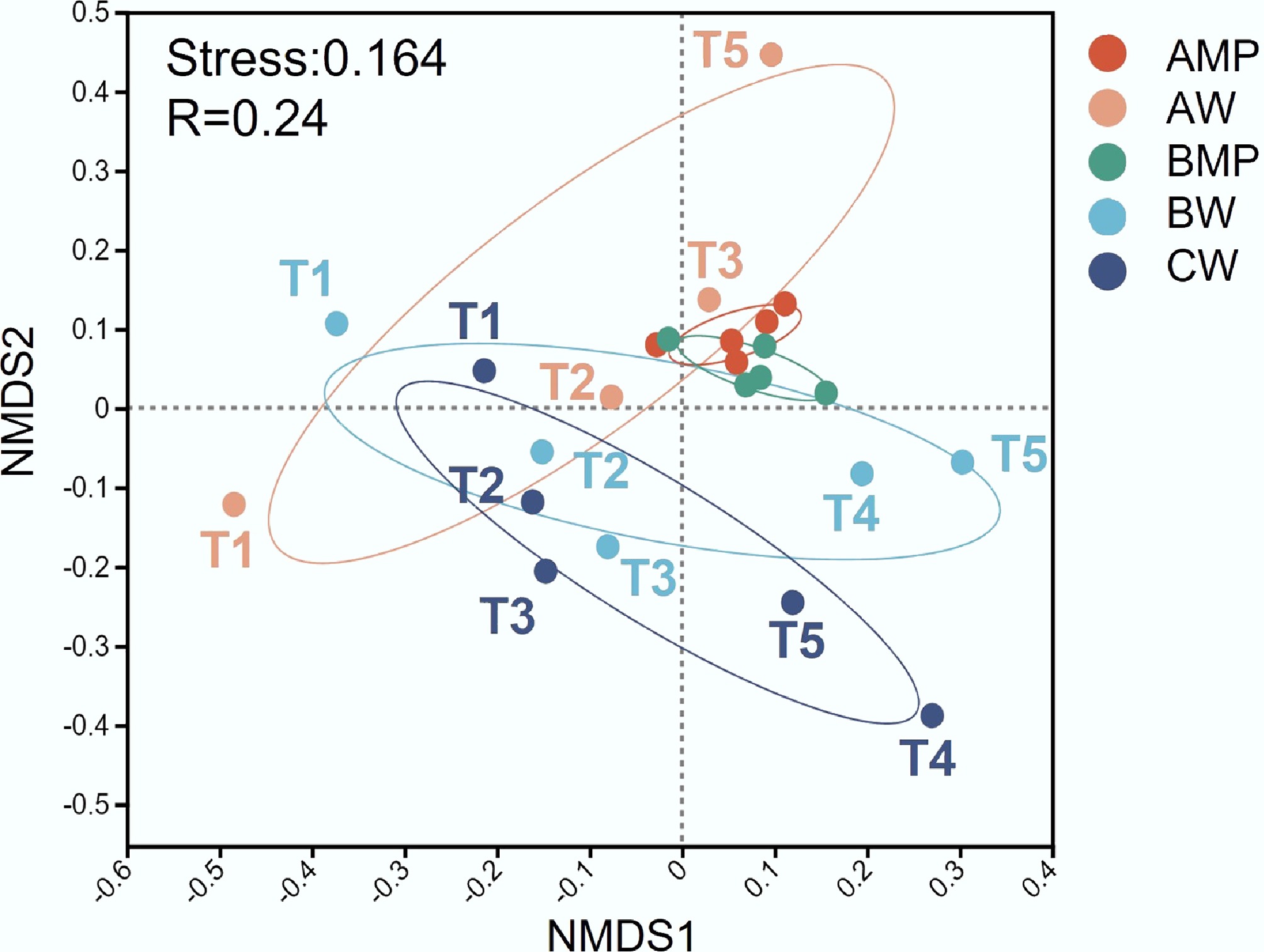
Figure 1.
NMDS analysis of microbial community beta-diversity across samples. AMP and AW represent the microplastic and water samples of Group A, respectively. BMP and BW represent the microplastic and water samples of Group B, respectively. CW represents the water sample of Group C.
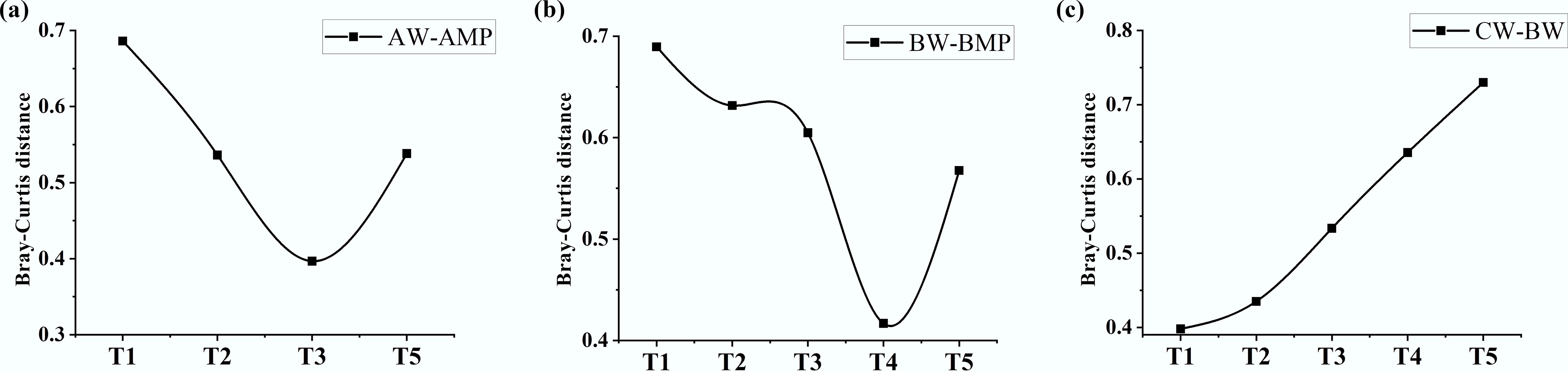
Figure 2.
Temporal variation in intergroup Bray-Curtis distance of microbial communities from week 1 to week 5. (a) Pure water group (Group A): plastisphere vs water. (b) River water group (Group B): plastisphere vs water. (c) River water of experimental group (Group B) vs control group (Group C).
In Group B, the temporal dynamics of Bray–Curtis dissimilarity between planktonic and biofilm microbial communities followed a pattern similar to that observed in Group A: a marked dissimilarity in the early stage, followed by an overall decrease, and a subsequent increase after week 4 (Fig. 2b). Compared to Group A, the changes in Group B occurred with a slight delay, suggesting that the Liangxi River ecosystem exhibits a certain degree of resistance to MP-induced disturbances. Moreover, the Bray–Curtis dissimilarity between BW and CW increased progressively over time (Fig. 2c), indicating that the influence of MPs on the recipient water intensified throughout the 5-week incubation. This cumulative impact is closely associated with microbial community succession, underscoring the potential long-term ecological consequences of MP transport in aquatic environments.
Specifically, within 1 week after the transfer of MPs into sterile water, Burkholderiales, Nitrosomonadales, and Chitinophagales were quickly released from the MPs and grew rapidly in the surrounding water (Supplementary Fig. S4). These bacteria have strong movement abilities, such as flagellar movement, allowing them to disperse in aquatic environments. In particular, Nitrosomonadales exhibit strong chemotaxis, enabling them to move toward favorable conditions by sensing gradients of ammonia and oxygen. Compared to biofilms, the water column provides a more nutrient-rich and oxygenated environment, which allows Nitrosomonadales to rapidly locate optimal habitats.
Following the introduction of MPs into the Liangxi River water, the relative abundance of Oscillatoriales, Chitinophagales, and Nostocales significantly increased in sample BW during the first week (Supplementary Fig. S5). In contrast, within the sterile water system, Nitrosomonadales, Burkholderiales, and Chitinophagales showed significant increases over the same period. Although Nitrosomonadales and Burkholderiales were also relatively abundant in the BW samples, they did not exhibit a significant increase there. This is likely because these orders are commonly found in the Liangxi River under natural conditions. Overall, Chitinophagales proliferated rapidly during the early phase of the experiment, while Burkholderiales, Nitrosomonadales, Synechococcales, and Chlamydomonadales exhibited significant growth later in the experiment. The early proliferation of Chitinophagales may be attributed to its chemotactic response to environmental changes[28]. However, with nutrients gradually depleting, the microbial community in the water underwent succession and favored the growth of bacteria possessing greater environmental adaptability. For instance, Burkholderiales possesses diverse metabolic capabilities, enabling it to degrade some organic compounds; Nitrosomonadales obtains energy via chemoautotrophic ammonia oxidation to nitrite; and Synechococcales and Chlamydomonadales perform photosynthesis.
Microplastic-mediated spread of ARGs
-
After MPs entered the Liangxi River water, 102 new subtypes of ARGs emerged (Supplementary Fig. S6), among which ARO:3002635, ARO:3002567, and ARO:3002319 were identified as the dominant subtypes. ARO:3002635 and ARO:3002567 confer resistance to aminoglycoside antibiotics, while ARO:3002319 encodes an enzyme hydrolyzing β-lactam antibiotics. Both antibiotic classes are clinically essential, and the spread of these resistance genes could challenge clinical treatment. Furthermore, MP transfer significantly increased ARG richness: the Chao1 index rose from 606.2 in the control group to 654.6 in the MP-exposed group (Supplementary Fig. S7).
Multiple ARGs also proliferated in Group A, with the most prominent being ARO:3000833, ARO:3000838, ARO:3003067, ARO:3000829, ARO:3003841, ARO:3003066, ARO:3002931, and ARO:3002937 (Supplementary Fig. S8). Notably, ARO:3000838 and ARO:3002937 likewise showed significant proliferation in the river water of Group B. This spread and proliferation of ARGs demonstrates that environmental MPs act as vectors for disseminating resistance genes. Specific ARGs, particularly ARO:3000838 (conferring tetracycline resistance), and ARO:3002937 (conferring carbapenem resistance), proliferated substantially across different water bodies, indicating their adaptive capacity and dissemination potential in diverse ecosystems.
NMDS analysis of ARGs revealed closer clustering between groups AWP and BWP (Fig. 3). Biofilm-associated ARGs exhibited higher temporal stability, while aquatic ARGs showed greater temporal dynamics—a pattern similar to that observed in the microbial community (Fig. 4). Under sterile conditions, microorganisms and ARGs released into the water column are primarily derived from the biofilm, resulting in a waterborne ARG profile that closely mirrors that of the plastisphere. In contrast, the ARG composition in the Liangxi River is largely governed by the established native microbial community and its long-term environmental selection, which buffers the influence of microplastic-derived inputs.
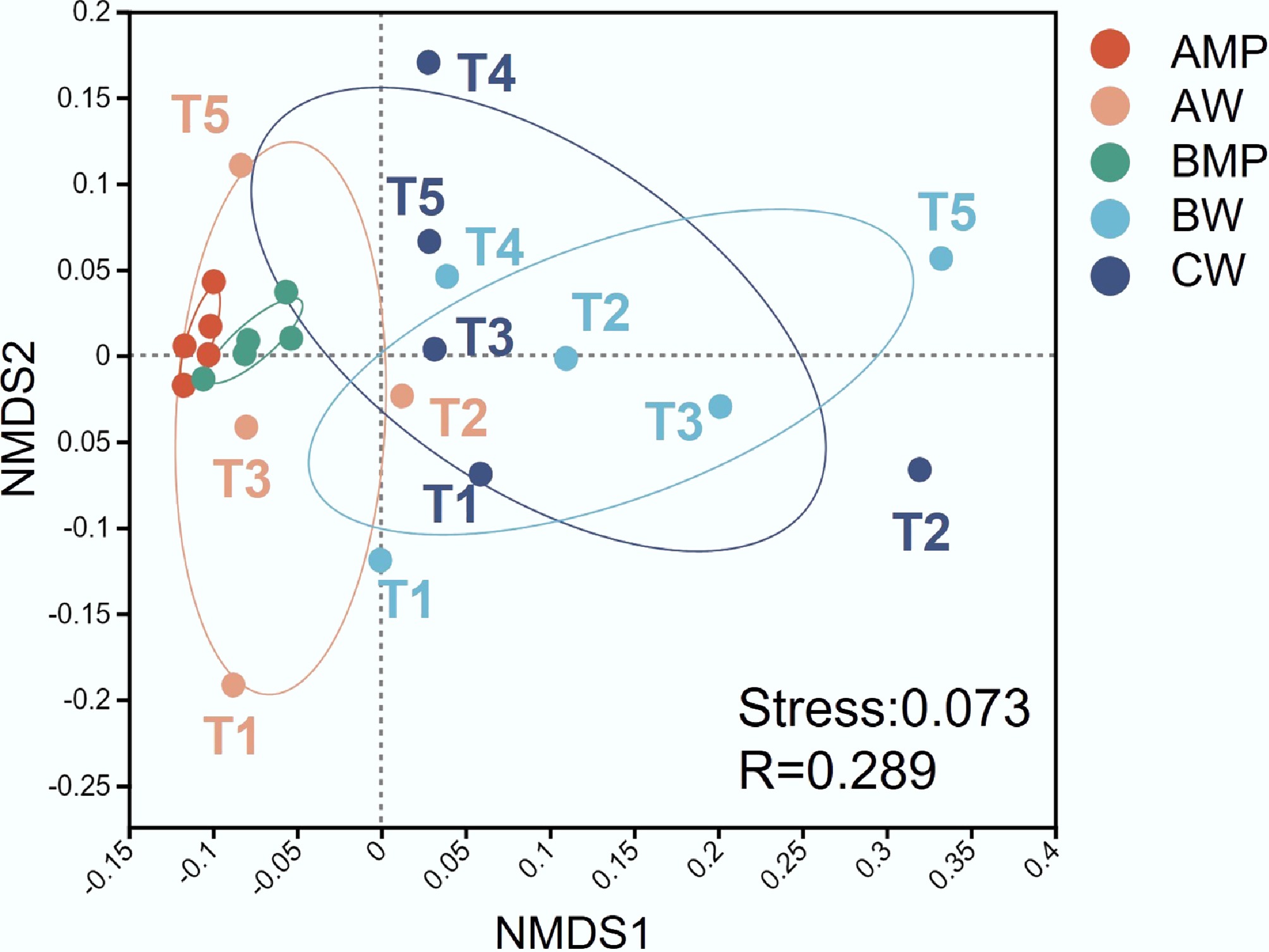
Figure 3.
NMDS analysis of ARGs across samples. AMP and AW represent the microplastic and water samples of Group A, respectively. BMP and BW represent the microplastic and water samples of Group B, respectively. CW represents the water sample of Group C.
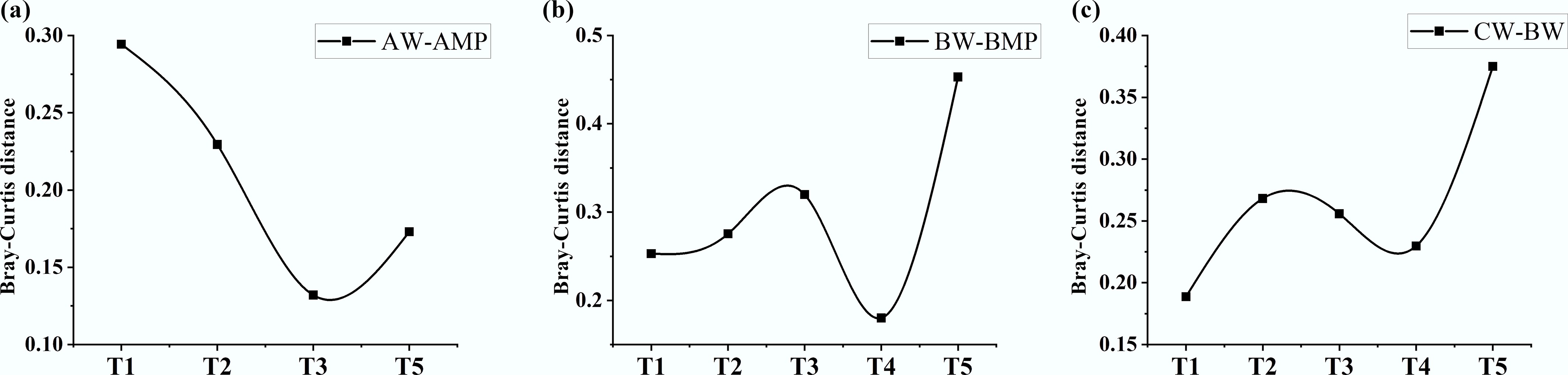
Figure 4.
Temporal variation in intergroup Bray-Curtis distance of ARGs from week 1 to week 5. (a) Pure water group (Group A): plastisphere vs water. (b) River water group (Group B): plastisphere vs water. (c) River water of experimental group (Group B) vs control group (Group C).
In Group A, the Bray-Curtis distance of ARGs between water and biofilm peaked in week 1, declined to a minimum by week 3, and then rebounded—mirroring microbial community dynamics. This progression reflects the dissemination of ARGs into the water column in the early phase. Subsequently, as gene exchange approached saturation, community succession became the dominant driver, ultimately increasing the Bray-Curtis distance.
In Group B, the Bray-Curtis distance of ARGs between water and biofilm increased gradually during the first 3 weeks, decreased subsequently, and then rebounded after week 4. The ARG dynamics diverged from microbial trends, implying that factors beyond host microorganisms influence ARG distribution. Moreover, the distance between sample BW and CW showed an overall increasing trend throughout the experiment, indicating progressively intensifying impacts on ARGs in receiving water over the 5 weeks—though with dynamics differing from those of the microbial community.
Although the MP concentration used in this study exceeds typical environmental levels, it serves to enhance the detectability of MP-associated biofilm processes under laboratory conditions. MP concentration is expected to regulate biofilm development primarily through changes in available attachment surface area and particle–microbe encounter frequency. At lower, environmentally relevant MP concentrations, MP-associated biofilms may still function as ARG reservoirs and vectors, but their influence on waterborne ARG pools is likely to be modulated by background microbial communities and hydrodynamic dilution. Therefore, our results highlight the potential of MP-associated biofilms to facilitate ARG dissemination, rather than providing a direct quantitative prediction of environmental risk.
Functional genomic responses to MP transport
-
Functional annotation was performed on samples from each group using COG, successfully assigning functions across 25 of the 26 categories. Notably, genes associated with general function prediction, signal transduction, and the biosynthesis, transport, and breakdown of secondary metabolites were significantly more abundant in biofilms compared to the water column. This enrichment likely reflects key ecological demands within biofilms: coordinating microbial activities via complex signaling networks requires a higher abundance of signal transduction genes. Additionally, microbial competition within biofilms drives substantial production of secondary metabolites (e.g., antibiotics, toxins, enzyme inhibitors); furthermore, these metabolites can also serve as signaling molecules or mediators, enhancing interactions and synergistic effects during cooperative microbial processes.
Further analysis revealed that microbial communities in receiving water adjusted their gene expression in response to MP disturbance (Fig. 5a; Supplementary Fig. S9), with significantly higher abundance of genes COG0515, COG1008, and COG1943 in the experimental group (BW) vs the control (CW). The enrichment of COG0515—involved in RNA synthesis and modification—suggests enhanced RNA processing for rapid, precise gene regulation during environmental adaptation. Concurrently, elevated COG1008 levels, which facilitate membrane protein functions, likely assist responses to membrane perturbations (e.g., osmotic stress, toxins), thereby preserving cellular integrity. Similarly, increased COG1943 expression, governing sugar metabolism and transport, ensures efficient carbon source utilization to sustain energy production and microbial growth.
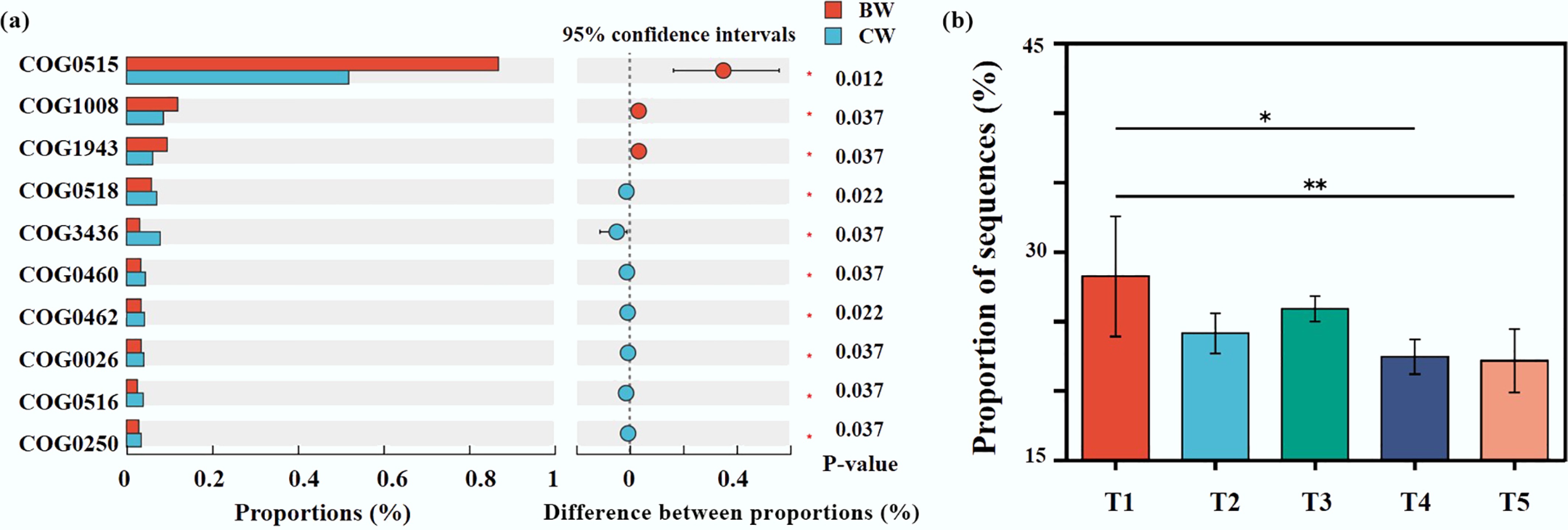
Figure 5.
Functional and metabolic responses of the receiving water to MP introduction. (a) Comparative analysis of COG functions. (b) Temporal dynamics of the autoinducer from week 1 to week 5.
Quorum-sensing responses to MP transport
-
Quorum sensing (QS) is a regulatory mechanism for bacterial group behavior. Many bacterial species synthesize and release autoinducers as signaling molecules to facilitate intercellular communication. By detecting these molecules, bacteria monitor population density and environmental shifts, subsequently activating relevant gene expression to coordinate collective behaviors, including growth and biofilm formation, among others.
QS annotation analysis was performed on all samples (Supplementary Fig. S10). The results revealed a predominance of autoinducer, transporter, regulator, and decomposer categories. Notably, autoinducer producers exhibited significant variation among sample groups, with biofilm samples showing higher abundance. This pattern likely reflects increased demands for autoinducer synthesis driven by complex microbial interactions and competition within biofilms. Further analysis identified Actinoplanes, Komagataeibacter, and Roseovarius as the primary autoinducer producers in biofilms, which generate AHLs and AI-2 autoinducers, playing key roles in biofilm development and maintenance.
Temporally, autoinducer levels exhibited pronounced fluctuations (Fig. 5b). During the initial phase of MP intrusion into receiving waters, both biofilm-associated and planktonic microbial communities markedly increased autoinducer production to coordinate collective behaviors and address environmental alterations. Subsequently, microbial communities underwent adaptive adjustments to these changing conditions, resulting in declining autoinducer proportions. Following this initial stress response, communities reached a relatively stable equilibrium state, with autoinducer levels stabilizing accordingly.
-
When MPs migrate into new aquatic environments, the microbial communities and ARGs within their biofilms demonstrate high stability, whereas significant changes occur in the receiving water (AW and BW). This migration can alter the structure and diversity of microbial communities in the receiving water and, by releasing specific microorganisms, pose potential ecological risks. Simultaneously, acting as carriers, MPs spread ARGs, increasing their diversity in the receiving water, with specific ARGs proliferating significantly in the new environment. Time-series analysis revealed that during the initial migration phase, microbes and ARGs from the biofilm gradually diffuse into the water column, leading to increasingly similar distributions between the biofilm and water. Subsequently, however, due to environmental differences, the microbial communities and ARGs in the biofilm and water begin to evolve independently. Notably, changes in microbial communities and ARG dynamics are not synchronized, with ARGs exhibiting higher stability over time. COG functional analysis indicates that microbial communities in the receiving water adapt to MP stress by regulating the expression of genes involved in RNA synthesis, cell membrane function, and sugar metabolism. Furthermore, quorum-sensing studies demonstrate that these communities maintain ecological balance by dynamically adjusting the levels of autoinducers, which follow a pattern of initial increase, subsequent decrease, and eventual stabilization. Overall, this study demonstrates that MPs act as stable reservoirs and effective vectors for microbial communities and ARGs in freshwater systems, thereby reshaping microbial ecology in receiving river environments. These findings highlight the potential ecological and public health risks associated with MP transport.
-
It accompanies this paper at: https://doi.org/10.48130/newcontam-0026-0007.
-
The authors confirm their contributions to the paper as follows: Qiji Zhang: conceptualization, methodology, data curation, and writing − original draft preparation. Xin Qian: writing − reviewing and editing, resources, and supervision. Both authors reviewed the results and approved the final version of the manuscript.
-
The data that support the findings of this study are available from the corresponding author upon reasonable request.
-
This study was supported by the National Key R&D Program of China (Grant No. 2022YFC3204400).
-
The authors declare no conflicts of interest.
-
Full list of author information is available at the end of the article.
- The supplementary files can be downloaded from here.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhang Q, Qian X. 2026. Microplastics as vectors for the spread of antibiotic resistance genes across aquatic ecosystems: microbial adaptation and temporal dynamics. New Contaminants 2: e010 doi: 10.48130/newcontam-0026-0007
Microplastics as vectors for the spread of antibiotic resistance genes across aquatic ecosystems: microbial adaptation and temporal dynamics
- Received: 26 November 2025
- Revised: 24 January 2026
- Accepted: 30 January 2026
- Published online: 13 March 2026
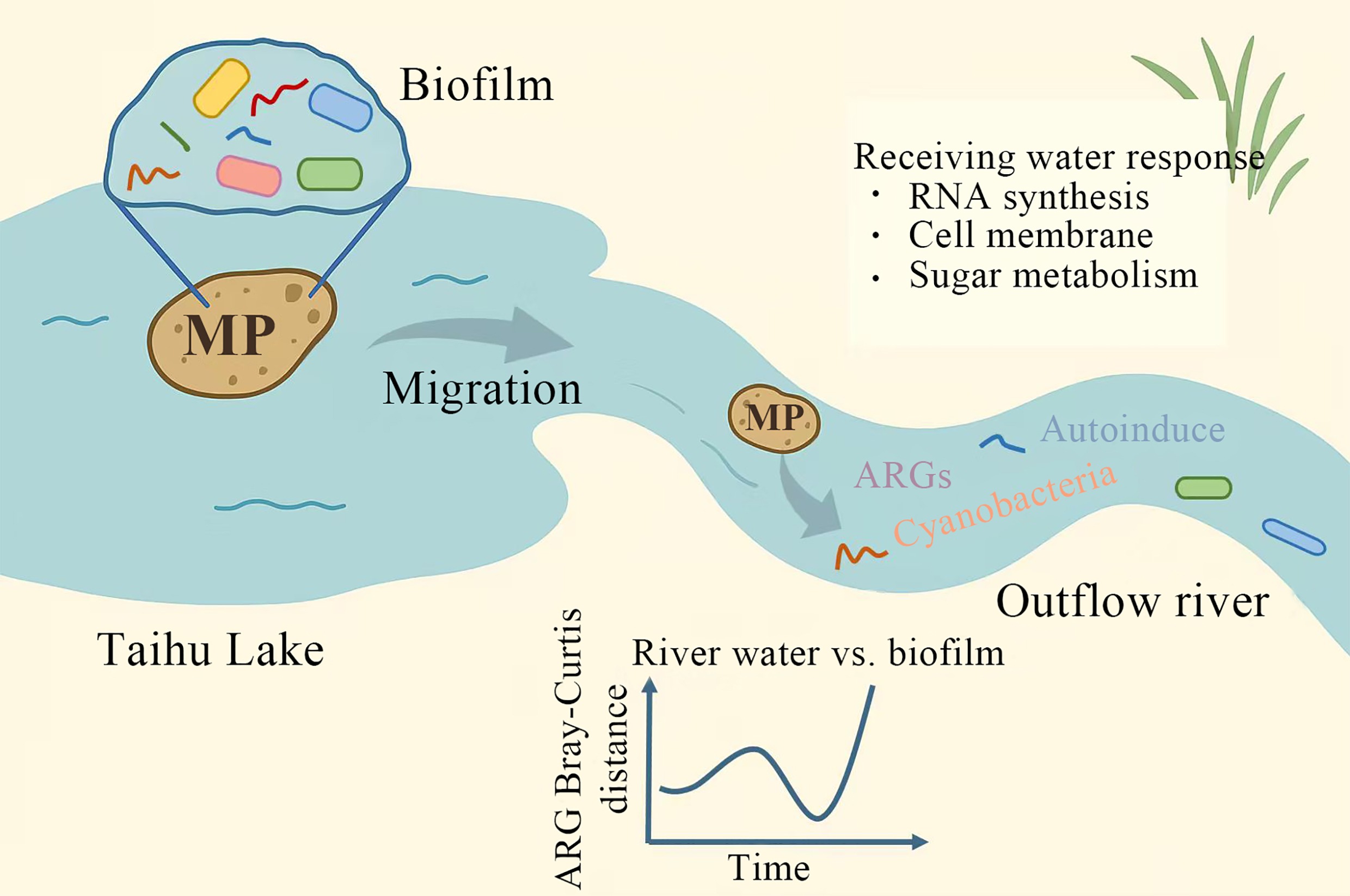
Abstract: Microplastics (MPs) carrying biofilms can travel across different aquatic ecosystems, representing an emerging route for the spread of antibiotic resistance genes (ARGs). However, this process remains poorly understood. In this study, the temporal dynamics of microbial communities and ARGs were examined in both biofilms and the surrounding water after MPs migrated from Taihu Lake into the Liangxi River, its outflow river, over a period of 1 to 5 weeks, during which 102 new ARG subtypes were detected. During the initial phase after migration, microbes and ARGs gradually diffused from the biofilm into the water, resulting in increasingly similar compositional profiles between the two habitats. Subsequently, however, microbial communities and ARGs in the biofilm and water began to diverge due to differing environmental conditions. Notably, shifts in microbial composition and ARG abundance were not synchronized—ARGs demonstrated greater temporal stability. Microbial communities in the receiving water adapted to MP stress by regulating genes related to RNA synthesis, cell membrane function, and sugar metabolism. Furthermore, quorum-sensing analysis indicated that these communities maintained ecological balance through dynamic adjustment of autoinducer levels, which followed a pattern of initial increase, subsequent decline, and eventual stabilization. These findings underscore the environmental significance of MP transport in shaping microbial and ARG dynamics across interconnected freshwater ecosystems.
-
Key words:
- Microplastic /
- Antibiotic resistance genes /
- Microbial /
- Plastisphere /
- Quorum sensing