-
Viola tricolor is a biennial or annual cool-season flowering plant extensively cultivated due to its varied coloration, diverse floral patterns and forms, strong cold tolerance, and broad adaptability[1−4]. V. tricolor cultivars often exhibit a loose growth habit, making them prone to lodging, which can compromise the uniformity and ornamental quality of the plants. In contrast, V. tricolor cultivars with shorter internodes are more conducive to intensive cultivation and can significantly reduce maintenance costs in horticultural practices. Therefore, the development of V. tricolor cultivars with compact plant architecture and shorter internodes has been a key objective in the breeding of V. tricolor.
Plant height is a critical trait that influences plant architecture, while internode development is a key morphological characteristic that influences overall plant height[5−7]. Research has shown that plant height is closely associated with endogenous hormones, such as gibberellins (GAs), brassinosteroids (BRs), indole-3-acetic acid (IAA), abscisic acid (ABA), cytokinins (CKs), and auxin, all of which play significant roles in regulating internode growth[8−13]. Among them, GA plays a crucial role in regulating stem elongation by promoting cell division and elongation. Crops and trees with shortened internodes typically display dwarf or semi-dwarf phenotypes, primarily attributed to impaired gibberellin biosynthesis or defective signal transduction[14]. For instance, reduced expression of MdGA20ox1 in apples results in lower levels of active GAs in the shoot apex, leading to a dwarf phenotype[15]. In bamboo, mutations in CmaGA2ox2 are associated with shortened vine lengths[16].
DELLA belongs to the plant-specific GRAS transcription factor family, which typically includes a variable N-terminal domain and a highly conserved C-terminal GRAS domain[17,18]. Current studies on DELLA protein function indicate that DELLA plays a role in multiple processes of plant growth and development[19]. Among them, DELLA plays a key regulatory role in GA signal transduction. DELLA acts as a repressor in the GA signal transduction pathway by interacting with growth-promoting transcription/regulatory factors, thereby inhibiting plant growth[20]. For example, DELLA protein GmRGAb stabilizes GmSTF1/2 protein, leading to an increase in transcription levels of GmGA2ox-7b and a subsequent decrease in plant height[21]. CsIREH1 interacts with DELLA protein and phosphorylates DELLA protein, specifically targeting CsGAIP and CsGAI2, thereby regulating plant height[22]. The overexpression of the DELLA subfamily genes MiSLR1 and MiSLR2 significantly reduced the plant height of transgenic Arabidopsis[23]. In addition, DELLA protein is also involved in plant response to various stresses[24]. For example, OsJAZ6 interacts with the DELLA protein OsSLR1, disrupting the stability of OsSLR1, thereby inhibiting rice tillering and reducing rice tolerance to drought stress[25]. FveDREB1B and FveSCL23 interact with FveDELLA to form FveDREB1B-FveSCL23-FveDELLA modules, thereby enhancing the cold tolerance of strawberry[26].
In ornamental plant breeding, compact plant architecture is a core target trait closely linked to cultivation efficiency and ornamental value, and DELLA proteins have emerged as key molecular targets for achieving this goal[27]. For instance, in petunia, a classic ornamental species with similar landscape application scenarios to V. tricolor, the regulation of DELLA-mediated GA signal transduction has been proven to restrict internode elongation effectively and produce dwarf cultivars with stable ornamental traits, thereby avoiding the environmental and quality risks associated with chemical plant growth regulators[28]. However, in comparison with model plants and major ornamental plants like petunia and chrysanthemum, there have been no reports regarding the breeding of compact plant height in the field of V. tricolor[29].
To investigate the molecular mechanisms underlying variation in plant height and internode length before and after jointing in V. tricolor, transcriptomic analyses were performed on two inbred lines, long internode (08H) and short internode (JP). The results revealed significant alterations in plant hormone signal transduction pathways during the jointing stage. Gene co-expression analysis identified VtRGL1 (Vtr06G053680) as a potential candidate gene for further functional studies. Moreover, overexpression of VtRGL1 was found to inhibit bolting in Arabidopsis. These findings contribute to a better understanding of the genetic basis of dwarfism and provide a foundation for molecular breeding in V. tricolor, enabling the cultivation of plants with specific morphologies.
-
The experimental materials used in this study were the long internode homozygous inbred line '08H' and the short internode homozygous inbred line 'JP', which were developed by the research group in recent years. All plants were cultivated in the rearing room of the School of Horticulture and Landscape Architecture, Henan Institute of Science and Technology, Xinxiang, China. To investigate the intrinsic factors influencing internode length in V. tricolor, a photoperiod regime of 16 h light/8 h dark was implemented. Stem tip samples were collected from each treatment group, both before and after jointing (with five to six seedlings per biological replicate and three biological replicates per treatment) for RNA sequencing. GA3 solution was used (50 and 100 mg/L) to treat after jointing V. tricolor plants.
Histological examination
-
The internode of JP and 08H were fixed in FAA solution and stored at 4 °C[30]. Sample sections were prepared using paraffin embedding, stained with safranin and fast green, and photographed under an Olympus IX73 microscope (Japan).
RNA-seq and quantitative real-time PCR (qRT-PCR)
-
Total RNA was extracted from the stem tips of JP and 08H inbred lines before (JPQ and 08HQ) and after (JPA and 08HA) jointing using the MiniBEST Plant RNA Extraction Kit (TAKARA, Japan). The cDNA was constructed using the PrimeScript™ RT reagent Kit (TAKARA, Japan). RNA integrity was accurately assessed using the Qsep400 Bioanalyzer. The cDNA libraries were sequenced on the Illumina sequencing platform by Metware Biotechnology Co., Ltd. (Wuhan, China)[31]. After filtering out adapter sequences and low-quality reads, clean reads were mapped to the reference genome of the V. tricolor (PRJCA045009). Differentially expressed genes (DEGs) were identified based on the criteria |log2Fold Change| ≥ 1 and False Discovery Rate < 0.05. Functional enrichment analysis of DEGs was conducted using Gene Ontology (GO) annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis[32,33]. Venn diagrams and heatmaps were generated using TBtools software for data visualization. To validate the RNA sequencing results, qRT-PCR analysis was performed on 11 randomly selected genes using the PrimeScript™ RT reagent Kit (TAKARA, Japan). The qRT-PCR was performed on a CFX96 Real-Time PCR Detection System (Bio-Rad, USA) with the TB Green® Premix Ex Taq™ Kit (TAKARA, Japan). Gene expression levels were calculated using the 2−ΔΔCᴛ method, and all reactions were carried out in triplicate biological replicates. All primer sequences used are listed in Supplementary Table S1.
Gene cloning and subcellular localization of VtRGL1
-
After being cloned from the JP of V. tricolor, the full-length coding sequence (CDS) of VtRGL1, except for the stop codon, was ligated into the plant expression vector GV1300-GFP[34]. To conduct a subcellular localization study, the VtRGL1-GFP construct was then converted into Agrobacterium tumefaciens. Subsequently, the obtained construct was co-transformed into tobacco plants along with the control nuclear cytoplasmic marker RFP. Following 24 h of dark culture and 48 h of normal growth conditions, the transformed tobacco tissues were examined using a confocal laser scanning microscope (LSM 710, Zeiss, Germany).
Phylogenetic analysis
-
The protein sequences of other species were obtained from the NCBI database for phylogenetic analysis of the VtRGL1 protein sequence with homologous proteins from other species. The GenBank accession numbers of protein sequences in the NCBI database were as follows: Arabidopsis thaliana OAP14146.1, NP_172945.1, NP_178266.1; Oryza sativa NP_001405437.1; Juglans regia XP 018811562.1; Pistacia vera XP 031264513.1; Carica papaya XP 021906703.1; Rosa chinensis XP 040368687.1; Prunus mume XP 008223345.1; Populus nigra XP 061974898.1. Multiple alignments of these protein sequences were conducted by Clustal W, and the neighbor-joining tree was constructed using MEGA 8 based on 1,000 bootstrap replications.
Overexpression of VtRGL1 in Arabidopsis thaliana
-
Using the dipping method, the VtRGL1-GFP plasmid was introduced into Arabidopsis thaliana. VtRGL1-OE lines were obtained by screening transgenic plants for resistance to hygromycin on Murashige and Skoog (MS) medium. A homozygous T3 generation line was obtained through continuous cultivation. Using the wild-type (WT) as a control, plants were cultivated for 30 d, after which the phenotypic differences and expression levels of VtRGL1 in both transgenic and control plants were analyzed.
Statistical analysis
-
All data were calculated using at least three replicate samples and are presented as mean values ± SD. The significance levels are defined as p < 0.05 (*), p < 0.01 (**), and Duncan's test (p < 0.05) was evaluated.
-
08H is an inbred line with long internodes, whereas JP is a short internode inbred line developed by this research team. The significant morphological differences began to appear between JP and 08H after plant jointing. These differences became more pronounced as the plants continued to grow (Fig. 1). After 70 d of differential emergence, the plant height of JP was markedly lower than that of 08H (Fig. 1a). Statistical analysis of key agronomic traits revealed that the plant height of JP was only 48.95% of that of 08H (Fig. 1b). Its plant width was also significantly reduced, reaching merely 71.21% of 08H (Fig. 1c). Further investigation of internode length showed that the internode length of JP was significantly shorter than that of 08H, amounting to just 58.94% of that in 08H (Fig. 1d). To investigate the cellular mechanisms underlying internode elongation, microscopic observations revealed that the epidermal cells in the internodes of the JP were significantly shorter than those in 08H (Fig. 1e, f). These findings suggest that the difference in plant architecture between the two cultivars is primarily attributable to variations in internode cell length.
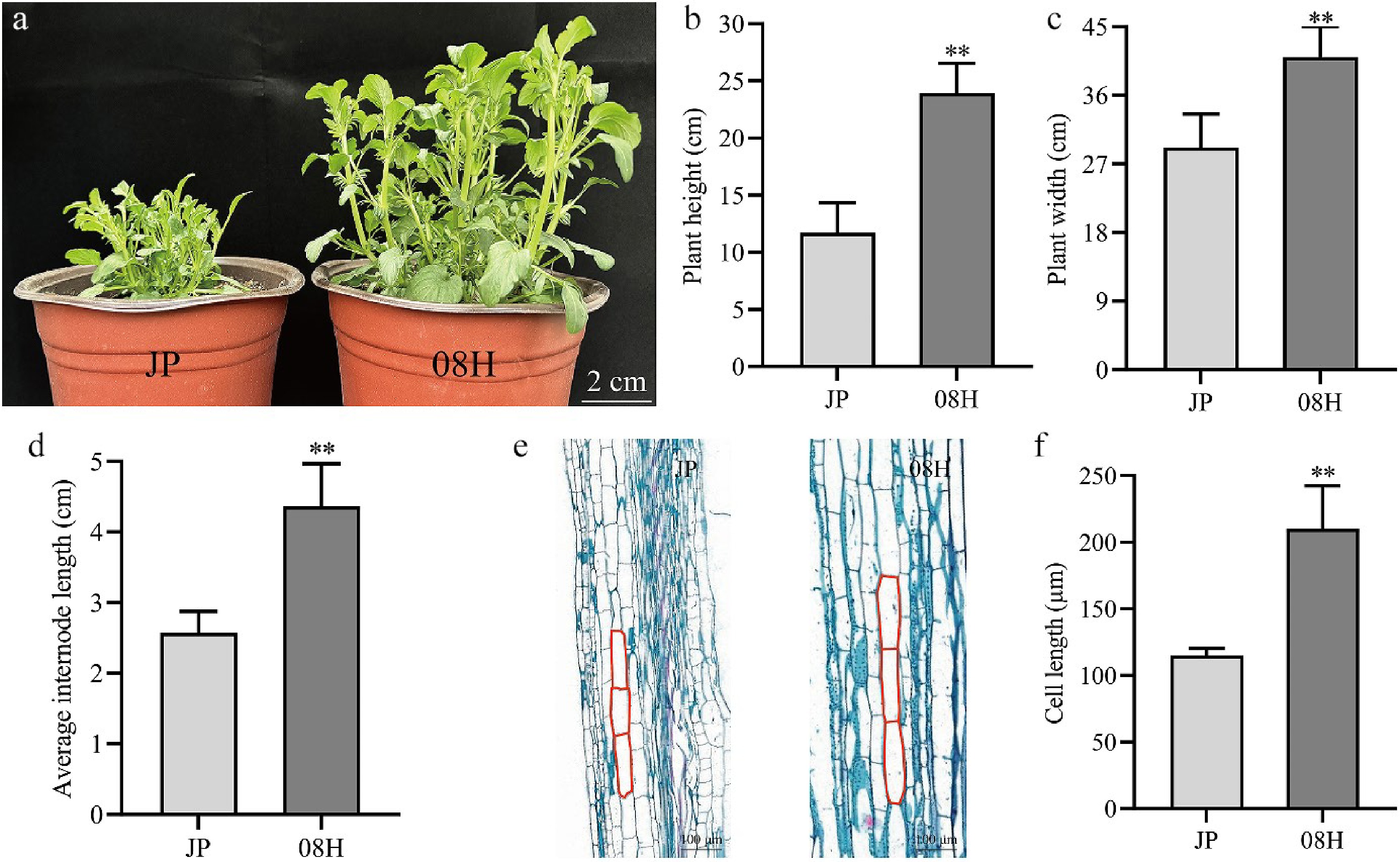
Figure 1.
The phenotype of JP and 08H. (a) Phenotypic characteristics of long-internode and short-internode inbred lines at 70 d. Scale bar = 2 cm. (b)–(d) The plant height, plant width, and average internode length of the strains are shown in the diagram. (e) Longitudinal paraffin sections of JP and 08H. (f) Length of cell in internode from JP and 08H. Data are means (± SD). Error bars are the standard deviation. ** p < 0.01, Student's t-test was used to calculate the p-value.
Transcriptomics of the stem tips of V. tricolor
-
To investigate the molecular mechanisms underlying variations in internode length in V. tricolor, transcriptomic analysis was performed on two inbred lines (JP and 08H) at two stages: before and after the jointing stage. A total of 105.28 G of clean sequencing data was generated from 12 libraries, with individual sample sizes ranging from 7.77 to 9.79 G. The Q30 value exceeded 95%, and the GC content ranged from 46.13% to 46.86% (Supplementary Table S2). After removing low-quality reads, approximately 701.9 million clean reads were obtained, among which 80.92% to 86.34% of the clean reads were successfully located to the reference genome of V. tricolor. Principal Component Analysis (PCA) revealed a clear separation among the four experimental groups, with all biological replicates clustering closely together. PC1 and PC2 accounted for 45.27% and 20.25%, respectively, of the total variance, indicating high-quality sample preparation and reliable RNA-seq results (Fig. 2a).
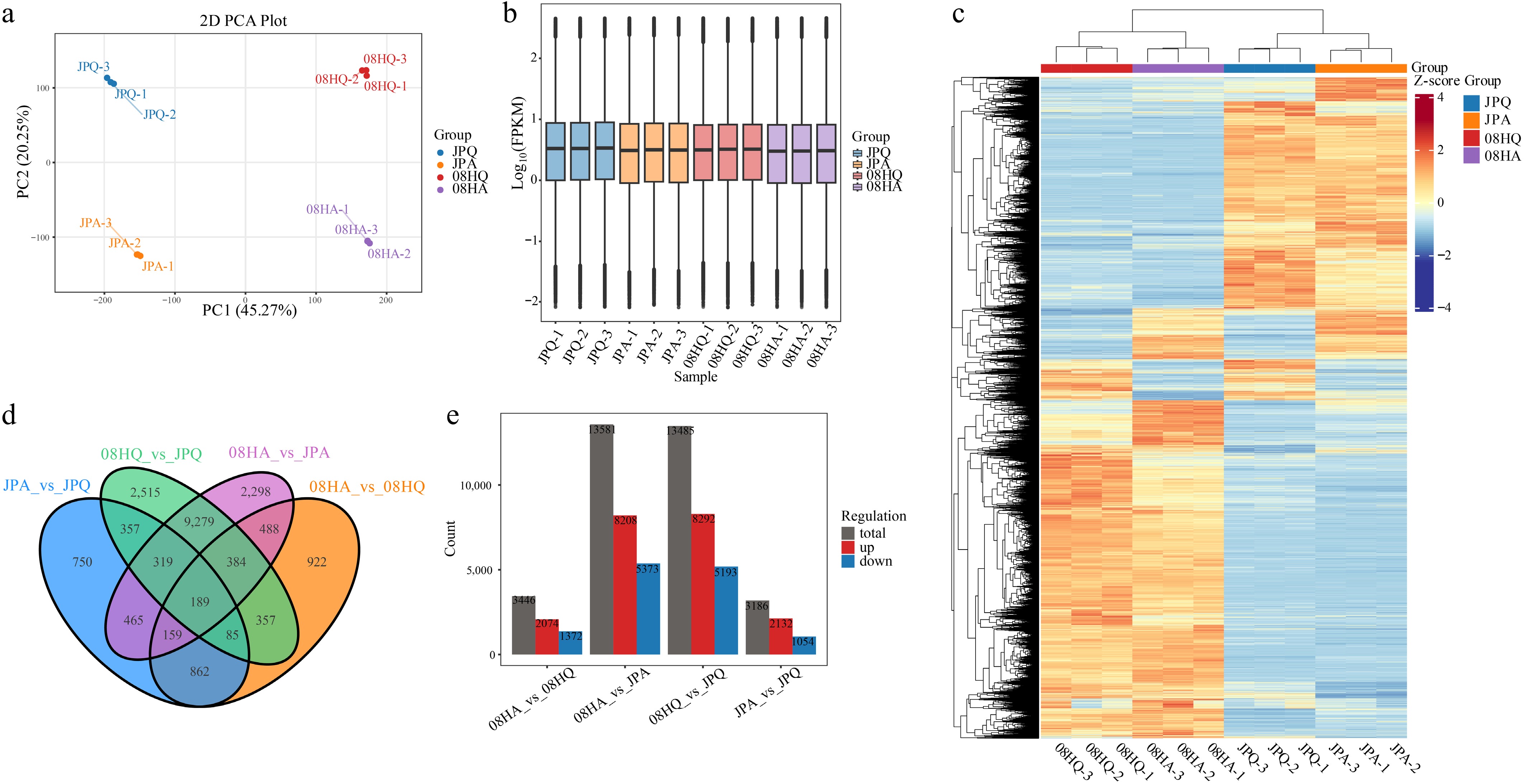
Figure 2.
Transcriptome sequencing and DEGs in V. tricolor inbred lines. (a) PCA of transcriptome sequencing data. (b) Gene expression of all samples, the boxplots with different colors indicate different samples analyzed. (c) Cluster analysis of DEGs based on gene expression of all samples. (d) Venn diagrams showing the number of DEGs among four comparisons. (e) DEGs of different components.
DEGs analysis and functional annotation
-
The expression levels of genes in each sample were evaluated based on the fragments per kilobase million (FPKM) values (Fig. 2b). Cluster analysis of differentially expressed genes was conducted to investigate the biological relationships among genes. The results revealed that the clustering patterns of highly or lowly expressed DEGs varied across sample groups, leading to distinct clustering outcomes. However, these DEGs were commonly located in the cross-sectional area, and similar expression patterns were observed within the same sample group (Fig. 2c). Through pairwise comparison of transcriptomic data before and after the jointing of two V. tricolor inbred lines, a total of 19,429 DEGs were identified, among which 189 DEGs exhibited differential expression across all comparisons (Fig. 2d). The highest number of DEGs was observed in the comparison between 08HA and JPA (13,581), including 8,208 upregulated and 5,373 downregulated genes. In contrast, the lowest number of DEGs was detected in the comparison between JPA and JPQ (3,186), with 2,132 upregulated and 1,054 downregulated DEGs (Fig. 2e).
Functional enrichment analysis of DEGs
-
GO functional annotation and enrichment analyses were conducted to investigate the potential biological functions of DEGs before and after jointing in two inbred lines of V. tricolor (Supplementary Fig. S1). These analyses revealed that DEGs exhibited significant variations in biological processes, cellular components, and molecular functions before and after jointing. The GO enrichment results indicated that genes related to JP and 08H were significantly enriched in biological processes such as cellular process and metabolic process. Then, binding and catalytic activities played a crucial role in molecular function. Regarding cellular components, they primarily pertain to cellular anatomical entities.
To delve deeper into the metabolic pathways of DEGs before and after jointing in two V. tricolor inbred lines, this study annotated and analyzed the KEGG metabolic pathways of these DEGs (Fig. 3). The KEGG pathway enrichment analysis revealed that 3,141 DEGs were annotated into 140 KEGG pathways in 08H between before and after jointing. These pathways were mainly significantly enriched in Metabolic pathways, Biosynthesis of secondary metabolites, and Plant hormone signal transduction. In JP, a total of 2,656 DEGs before and after joining were annotated to 133 KEGG pathways. Significantly enriched pathways primarily included Biosynthesis of secondary metabolites, Plant hormone signal transduction, and Starch and sucrose metabolism (Fig. 3). Based on these findings and prior research, it was speculated that the key metabolic pathways of 08H and JP, both before and after jointing, are related to plant hormone signal transduction.
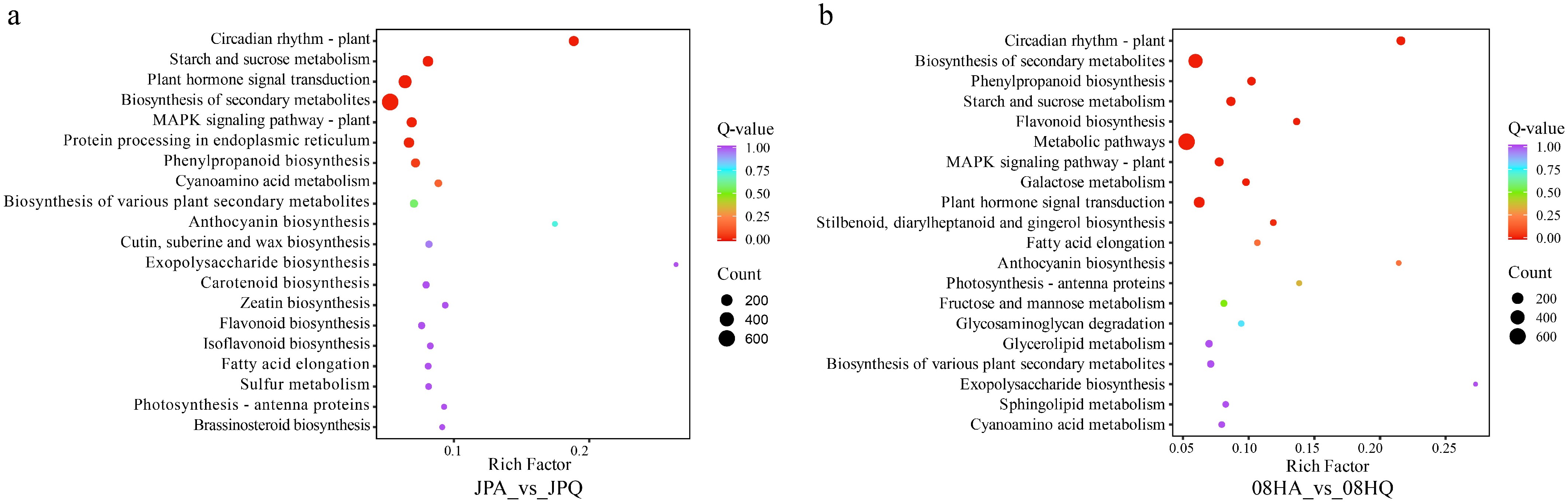
Figure 3.
KEGG analysis of DEGs identified in two different V. tricolor inbred lines. (a) KEGG enriched pathways for DEGs of 08H inbred line. (b) KEGG enriched pathways for DEGs of the JP inbred line.
Analysis of expression patterns of relevant DEGs in the GA signaling pathway
-
To gain a deeper understanding of the impact of plant hormones on the plant type of V. tricolor, a specific concentration of GA3 solution was sprayed on V. tricolor before the jointing stage. The findings revealed that GA treatment significantly increased the internode length of V. tricolor, particularly in the case of JP (Supplementary Fig. S2). This increase was observed in both plant height and width, while the number of internodes did not exhibit a significant change (Supplementary Table S3). Based on these results, it was proposed that the GA-related pathway plays a significant role in determining the plant architecture of V. tricolor. Hierarchical clustering analysis was conducted on DEGs within the GA signaling pathway using transcriptome data, and a corresponding heatmap was generated (Fig. 4a, Supplementary Table S4). The results indicated that several DEGs in this pathway exhibited significant differences in transcript accumulation before and after jointing in the two inbred lines. For instance, genes such as Vtr05G044690, Vtr02G010490, and Vtr02G097210 showed higher expression levels before jointing, whereas other DEGs, including Vtr02G086670 and Vtr11G049880, showed higher expression levels after jointing (Supplementary Fig. S3). Taken together, these findings suggest that the majority of DEGs identified in the GA signaling pathway are specifically associated with variations in plant type morphology in V. tricolor.
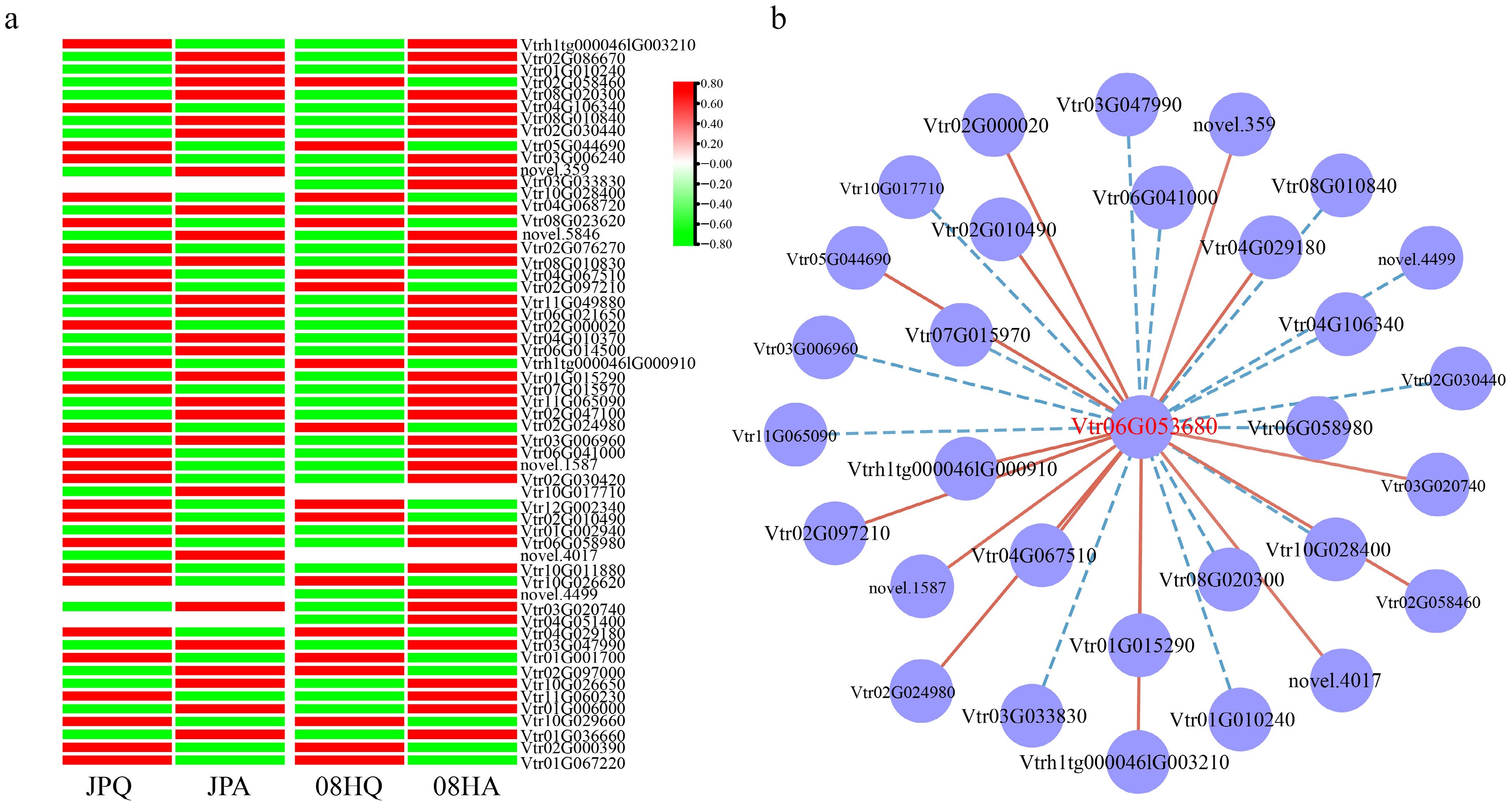
Figure 4.
The expression pattern of GA-related DEGs. (a) Cluster heat map of GA-related DEGs. (b) Co-expression network related to GAs. The red line represents positive correlation, and the blue line represents negative correlation.
The DELLA family has been demonstrated to play a critical role in the GA signaling pathway[35]. Therefore, a co-expression network involving VtDELLAs and DEGs was constructed within the GA signaling pathway. The results revealed that Vtr06G053680, encoding a DELLA transcription factor, exhibited strong correlations with 30 GA-related genes (Fig. 4b, Supplementary Fig. S4). Specifically, Vtr06G053680 showed significant positive correlations with 15 GA-related genes and significant negative correlations with the remaining 15 genes. These findings suggest that Vtr06G053680 may play a significant role in the signal transduction of GA.
Characterization of the VtRGL1 gene
-
To better characterize the function of Vtr06G053680 in V. tricolor, a 1,593 bp CDS encoding 530 amino acids was cloned from JP (Supplementary Fig. S5). Phylogenetic analysis revealed that Vtr06G053680 is closely related to RGL1 proteins from other plant species; therefore, it was designated as VtRGL1 (Fig. 5a, Supplementary Fig. S6). To investigate the subcellular localization of VtRGL1, recombinant plasmids containing VtRGL1-GFP and a GFP empty vector, along with a nuclear and cytoplasmic marker (RFP), were transiently expressed in tobacco epidermal cells. Subcellular localization was determined by observing GFP signal via confocal microscopy. In the control group, GFP fluorescence was distributed throughout the entire cell, whereas VtRGL1-GFP fluorescence was specifically localized to the nucleus and cytoplasm (Fig. 5b).
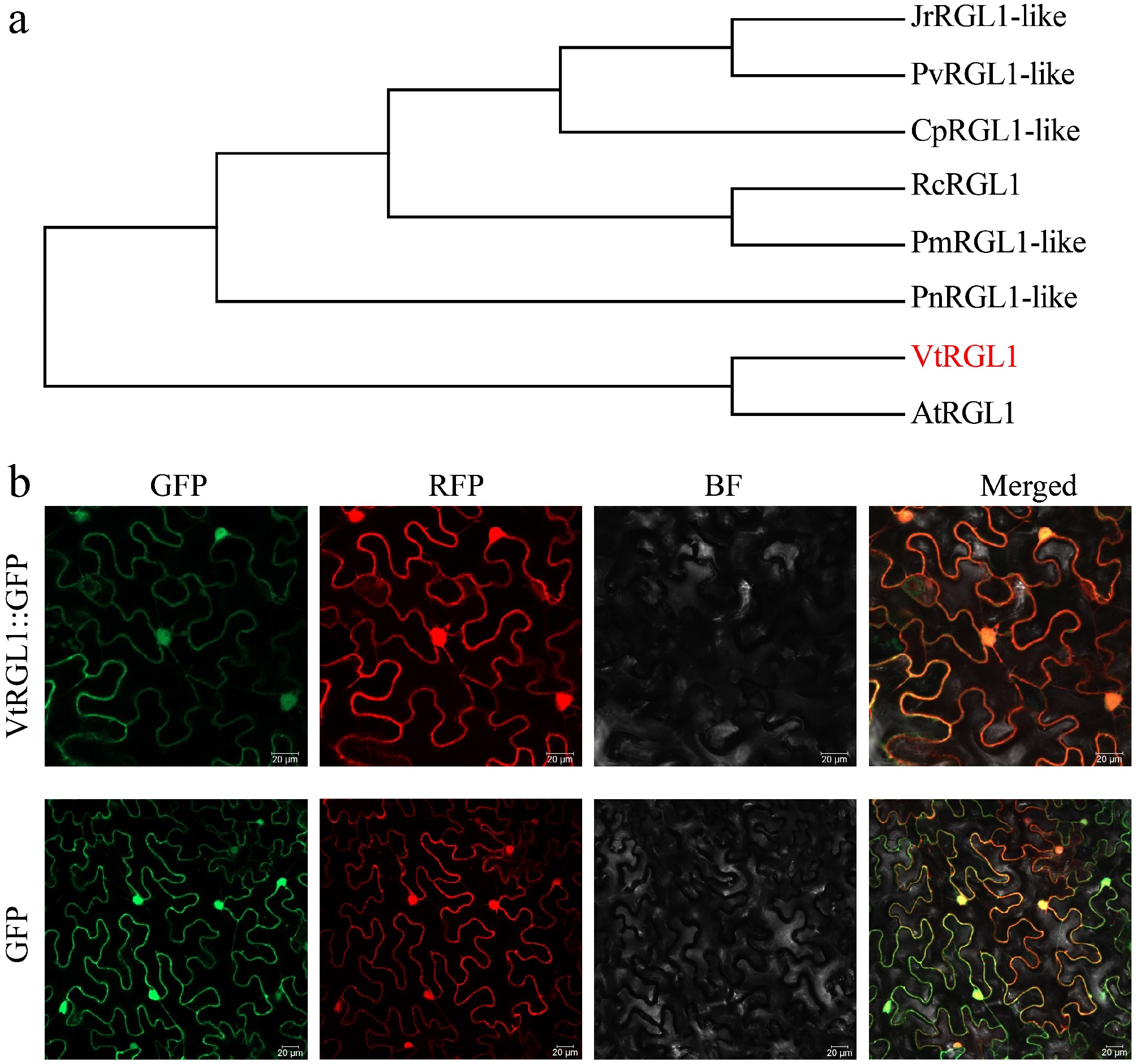
Figure 5.
Sequence analysis and subcellular localization of VtRGL1. (a) Phylogenetic tree analysis of VtRGL1. (b) Subcellular localization of VtRGL1. RFP, nuclear and cytoplasmic marker. Scale bar = 20 µm.
VtRGL1 overexpression results in a postponed bolting time in Arabidopsis
-
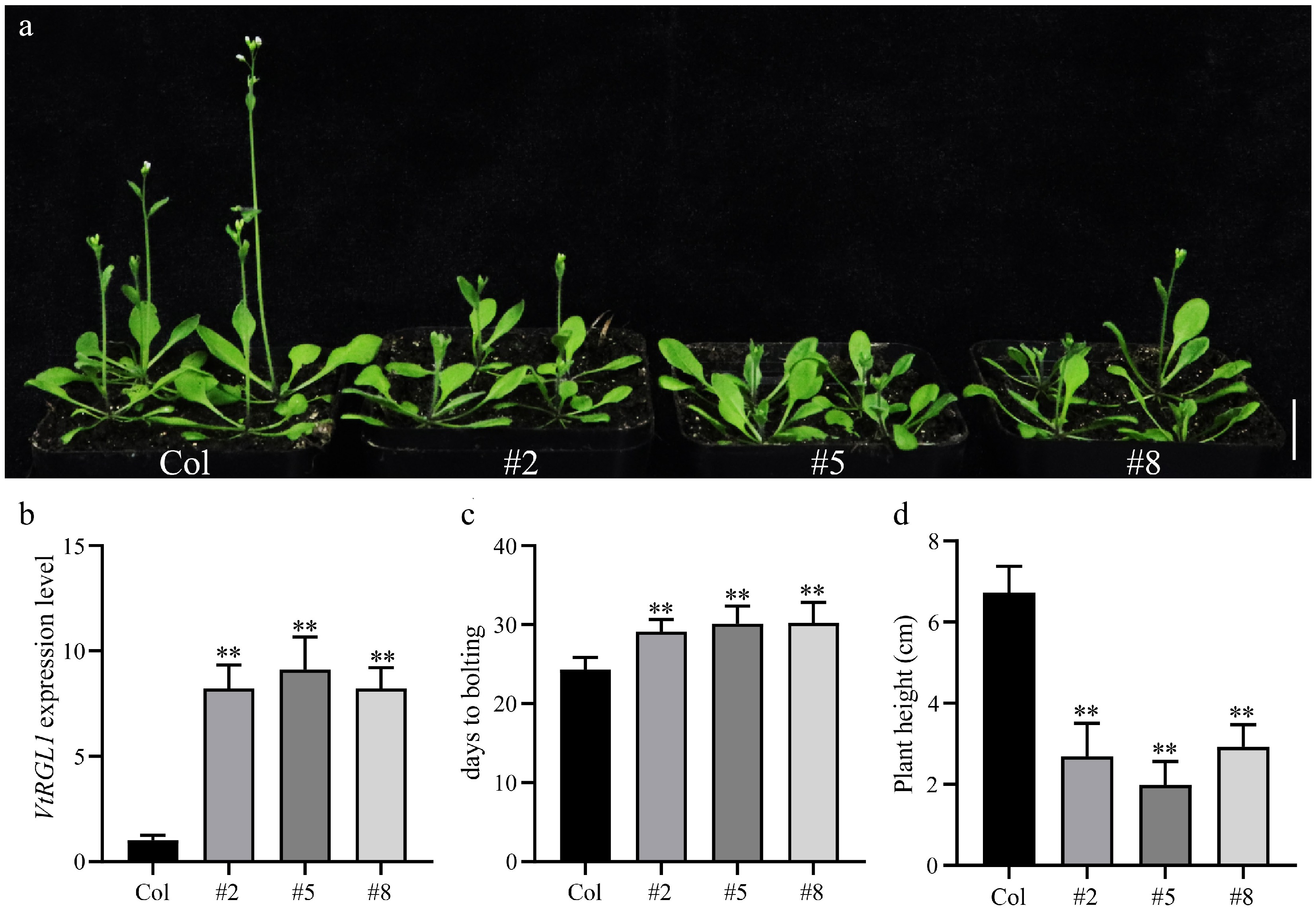
To investigate the functional role of VtRGL1, transgenic Arabidopsis thaliana overexpressing VtRGL1 was generated using an Agrobacterium-mediated transformation approach (GV3101) (Fig. 6). Three independent overexpression lines (lines two, five, and eight) were selected for further analysis (Fig. 6a). Both WT and VtRGL1 overexpressing plants were cultivated under controlled conditions (Fig. 6). The flowering time and plant height were systematically observed. The results revealed that the bolting time of VtRGL1 overexpression lines was significantly delayed compared to the WT (Fig. 6). These findings suggest that overexpression of VtRGL1 delays bolting, thereby affecting plant height (Supplementary Fig. S7).
-
Plant height, as a key component of plant architecture, is a fundamental agronomic and economic trait influencing the structural morphology and functional performance of ornamental species[12,36,37]. By precisely adjusting plant height to achieve optimal architectural form, the aesthetic value and environmental adaptability of ornamental plants can be significantly improved in diverse applications, including landscape design and container gardening[38]. Centuries of breeding efforts have yielded commercial horticultural flower cultivars with a diverse range of plant types[39]. Among these, cultivars exhibiting compact growth, optimal height, and consistent ornamental performance remain the most sought-after, directly aligning with market demands and consumer preferences[40]. In current horticultural practice, compact growth in flowering plants largely depends on synthetic plant growth regulators[41,42]. However, prolonged and extensive use of these chemicals has raised serious concerns about their sustainability, including issues such as environmental residue accumulation, ecological imbalance, and potential human health risk[40]. Consequently, there is growing interest in developing molecular biology-based strategies for compact plant development as a sustainable, eco-friendly alternative to chemical regulation. Accumulating evidence indicates that plant height is primarily governed by the coordinated regulation of two key physiological processes: cell division and cell elongation[43,44].
V. tricolor, as a typical species in cooling-season landscapes, is a flower plant that possesses significant ornamental and commercial value[45]. However, systematic studies on the regulatory mechanisms underlying plant height remain limited, with existing reports being fragmented and insufficient to support breeding efforts aimed at specific plant architectures. In this study, two inbred lines of 08H and JP were used as experimental materials. Quantitative measurements and comparative analyses revealed that JP exhibited significantly shorter internode lengths compared to 08H (Fig. 1). Further histological observations demonstrated that epidermal cell lengths in the internodal regions of JP were markedly reduced relative to those of 08H. These findings indicate that differential cell elongation is the primary cellular mechanism driving internode length variation and ultimately determining plant height differences between the two inbred lines. This conclusion aligns well with previous studies in diverse plant species such as maize and Chrysanthemum, which have established cell elongation as a key physiological process governing internode development and overall plant stature, thereby providing new empirical support for the conservation of plant height regulation mechanisms across species[46,47].
Through transcriptomic analysis of the two V. tricolor inbred lines 08H and JP, before and after the jointing stage, it was found that DEGs were significantly enriched in two major pathways: biosynthesis of secondary metabolites and plant hormone signal transduction (Fig. 3). These findings suggest that the biosynthesis of secondary metabolites and plant hormone signaling may jointly constitute a key molecular mechanism regulating internode elongation. Previous studies have established that GAs, as key plant hormones, regulate diverse growth and developmental processes. The core function is to promote cell elongation, thereby influencing plant height and internode elongation. To test the hypothesis that the GA signaling pathway regulates internode elongation in these materials, exogenous GA3 was applied to both 08H and JP via stem tip spray (Supplementary Fig. S2). The results showed that exogenous GA3 treatment significantly promoted internode elongation in JP, effectively alleviating its dwarf phenotype. These results indicate that JP is a GA-responsive inbred line, and its dwarfism may result from either insufficient endogenous GA biosynthesis or impaired signal transduction. This observation aligns with findings in elephant grass; however, it differs from reports in grape and rice[48−50]. Endogenous hormones exhibit species-specific regulatory roles in internode elongation. The molecular basis underlying this specificity warrants further investigation.
The molecular mechanism of the GA signaling pathway has been systematically elucidated[51]. The core regulatory logic can be summarized as follows: GA molecules bind to the receptor protein GID1, inducing the formation of the GA-GID1-DELLA ternary complex. Conformational changes in this complex facilitate the recruitment of ubiquitin ligase complexes, leading to the ubiquitination of DELLA protein and its subsequent degradation via the 26S proteasome[52]. This degradation relieves the transcriptional repression exerted by DELLA proteins on downstream GA-responsive genes, thereby activating the GA signaling pathway and modulating plant growth and development. In this study, correlation analysis revealed that VtRGL1, a member of the DELLA protein family, exhibits significant co-expression relationships with 30 key genes involved in the GA signaling pathway (Fig. 4). This result indicates that VtRGL1 may play a role in regulating internode growth in V. tricolor. To test this hypothesis, a VtRGL1 overexpression vector was constructed and transformed into Arabidopsis. Phenotypic analysis reveals that transgenic Arabidopsis lines display a significantly delayed bolting time compared to WT plants (Fig. 6, Supplementary Fig. S6). This functional characteristic aligns with the reported regulatory roles of RGL1 in other species, such as Brassica rapa and lettuce, where RGL1 acts as a negative regulator of stem elongation by suppressing the GA signaling pathway[53,54]. Despite the conserved GA pathway regulatory function, VtRGL1 exhibits distinct functional differentiation from its homologs in Arabidopsis and other species. For instance, AtRGL1 in Arabidopsis mainly regulates stamen and anther development during the reproductive growth stage[55]. Taken together, these findings suggest that VtRGL1 may function as a negative regulator of internode elongation in V. tricolor. However, it is important to note that heterologous overexpression in Arabidopsis thaliana may not fully recapitulate the native expression patterns or protein interaction networks of VtRGL1 within its natural genetic context in V. tricolor. Therefore, the observed phenotype provides only preliminary functional insights. To precisely determine the biological role of VtRGL1 in internode development, further studies should incorporate a comprehensive physiological and molecular analysis alongside a V. tricolor-specific genetic transformation system.
-
This study employed two Viola tricolor inbred lines (JP and 08H) as experimental materials. Through systematic agronomic trait evaluation and anatomical structure observation, it was identified that differences in stem cell length constitute the core anatomical factor underlying the significant divergence in plant height between the two inbred lines. To further elucidate the molecular regulatory mechanisms, transcriptome sequencing was conducted on both inbred lines, revealing that DEGs were significantly enriched in plant hormone signaling pathways. Gene co-expression network analysis demonstrated that the core candidate gene VtRGL1 exhibited strong co-expression with 30 key genes in the GA signaling pathway. To validate its functional role, a VtRGL1 overexpression vector was constructed and transformed into Arabidopsis. Phenotypic characterization revealed that transgenic Arabidopsis lines displayed a delayed bolting phenotype. In summary, this study provides the first evidence of the negative regulatory role of VtRGL1 in regulating the dwarfing trait in V. tricolor. These findings not only offer crucial genetic insights into the molecular regulatory network governing height development in V. tricolor but also supply a theoretical foundation and valuable candidate gene resources for the molecular breeding of dwarf V. tricolor.
-
The authors confirm their contributions to the paper as follows: study conception and design: Ji X, Du X, Liu H; experimental work: Ji X, Xie Y, Ye X, Li Y, Hu Y; data analysis: Ji X, Zhu X, Mu J; resources and materials: Du X, Liu H; draft manuscript preparation: Ji X, Du X. All authors reviewed the results and approved the final version of the manuscript.
-
All raw data have been submitted to the NCBI BioProject database (BioProject ID: PRJNA1337801).
-
This work was supported by the Training Program of Innovation and Entrepreneurship for Undergraduates (Grant No. S202510467041), the Science and Technology Program of Henan Province (Grant No. 252102110335), and the Key Research and Development Project of Henan Province (Grant No. 241111113100).
-
The authors declare that they have no conflict of interest.
- Supplementary Table S1 Primers used for PCR.
- Supplementary Table S2 Illumina sequencing data filtering statistics and RNA-Seq Map statistics of two V. tricolor inbred lines.
- Supplementary Table S3 The effect of external GA application on the plant types of 08H and JP.
- Supplementary Table S4 The expression profiles analysis of GA-related genes.
- Supplementary Fig. S1 Gene ontology (GO) enrichment of DEGs in two inbred lines of V. tricolor. (A) GO analysis of DEGs in 08H varieties. (B) GO analysis of DEGs in JP varieties.
- Supplementary Fig. S2 Response of 08H and JP to GA.
- Supplementary Fig. S3 . Transcriptional profiles of GA-related genes in JP (A) and 08H (B) before and after jointing. The asterisk indicates significant differences between the same strain before and after jointing (* p < 0.05, ** p < 0.01).
- Supplementary Fig. S4 Expression of VtDELLA genes in V. tricolor.
- Supplementary Fig. S5 Molecular characteristics of VtRGL1. (A) Cloning of VtRGL1. M: DNA 2000 marker, line 1: VtRGL1. (B) Nucleotide sequence and the deduced amino acid sequence of VtRGL1.
- Supplementary Fig. S6 Phylogenetic tree analysis of DELLA proteins across various species.
- Supplementary Fig. S7 Plant height phenotype of VtRGL1 transgenic Arabidopsis thaliana. (A) Plants of WT and VtRGL1 lines. Scale bar = 5 cm. (B) Plant height phenotype statistics of VtRGL1 transgenic Arabidopsis thaliana (** p < 0.01).
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Ji X, Xie Y, Ye X, Li Y, Hu Y, et al. 2026. VtRGL1, a DELLA transcription factor, negatively regulates internode elongation by modulating the gibberellin signaling pathway in Viola tricolor. Ornamental Plant Research 6: e015 doi: 10.48130/opr-0026-0004
VtRGL1, a DELLA transcription factor, negatively regulates internode elongation by modulating the gibberellin signaling pathway in Viola tricolor
- Received: 19 November 2025
- Revised: 15 December 2025
- Accepted: 06 January 2026
- Published online: 15 April 2026
Abstract: Plant height is a critical trait for ornamental plants, directly affecting their aesthetic value and commercial applications. Viola tricolor exhibits a compact growth habit due to its shortened internodes, which grant it significant advantages in landscaping. However, the molecular mechanisms regulating internode elongation in this species remain unclear. This study conducted a comparative transcriptomic analysis of the long-internode inbred line (08H) and the short-internode inbred line (JP) of V. tricolor before and after jointing. The results revealed significant differences in gibberellin (GA) signaling pathways, with exogenous GA3 application promoting internode elongation, particularly in JP. Among the differentially expressed genes, the DELLA family transcription factor VtRGL1 (Vtr06G053680) was identified as a key candidate. Subcellular localization confirmed its nuclear and cytoplasmic distribution, and overexpression of VtRGL1 in Arabidopsis thaliana resulted in delayed bolting and reduced plant height, consistent with its inhibitory effect on GA-mediated growth. Co-expression network analysis further supports its regulatory function in the GA signaling pathway. These findings provide new insights into the genetic regulation of plant morphology in V. tricolor, and offer a promising target for molecular breeding of compact ornamental varieties.
-
Key words:
- Viola tricolor /
- Internode length /
- DELLA protein /
- VtRGL1 /
- Transcriptomics