-
Antimicrobial resistance (AMR) has escalated into a defining global health crisis that threatens to dismantle the foundations of modern medicine[1]. In 2019, bacterial AMR was associated with an estimated 4.95 million deaths worldwide, and directly caused 1.27 million deaths, with a disproportionate burden in low and middle-income countries[2]. If left unaddressed, this toll is projected to rise substantially by 2050, potentially resulting in a cumulative global economic loss of US
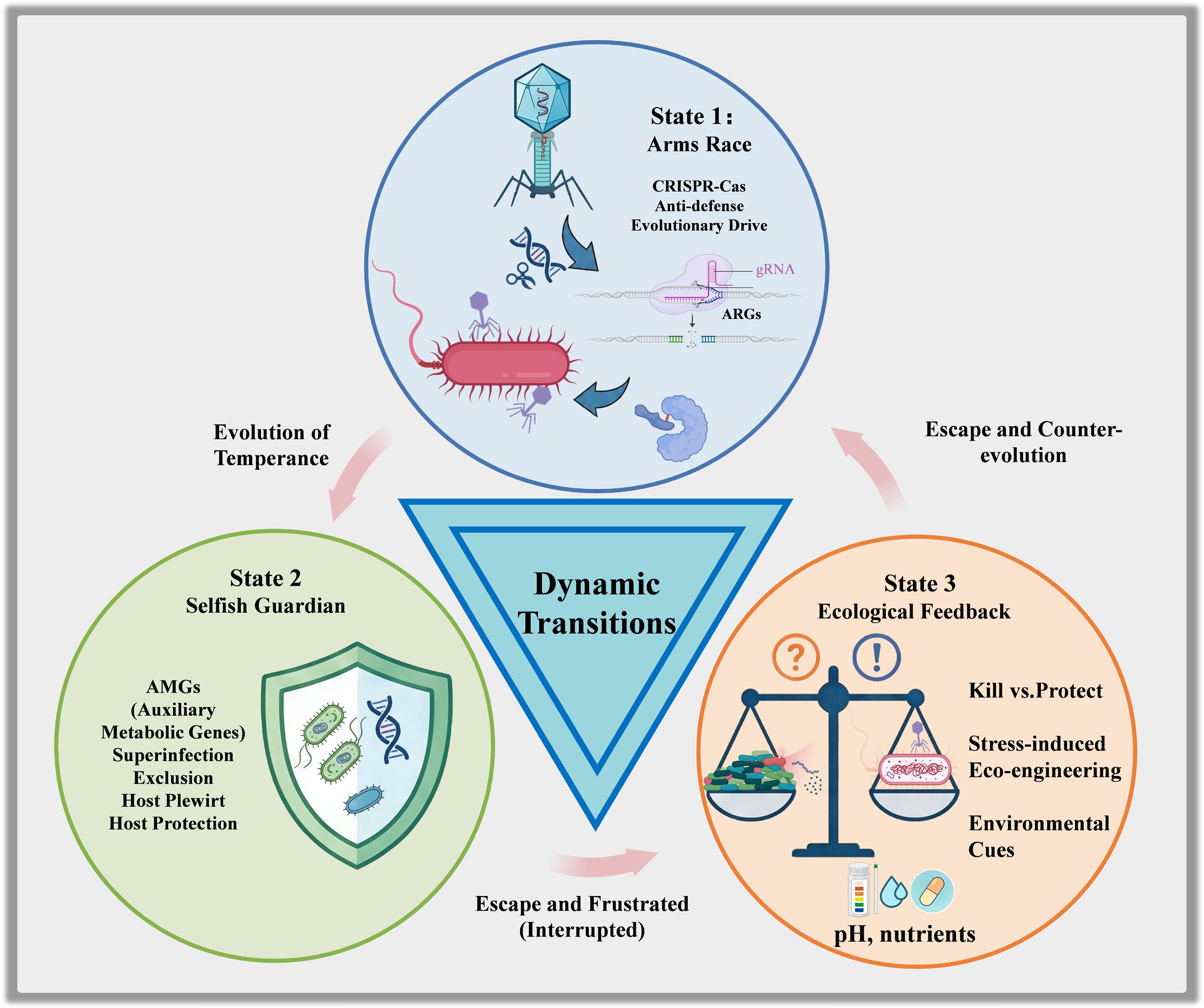
$\text{\$} $ In response to this crisis, bacteriophage (phage) therapy has re-emerged as a promising precision tool, historically prized for its lytic capability to infect and destroy specific antimicrobial-resistant bacteria (ARB) without disrupting the broader microbiome[5,6]. However, recent advances in viral metagenomics and molecular biology reveal that lysis is merely one outcome of a sophisticated evolutionary negotiation. To fully harness the potential of phages against AMR, a comprehensive framework has been proposed that describes how phage-host interactions progress through three distinct yet interconnected evolutionary states (Fig. 1). The first is an antagonistic arms race state that drives the diversification of defense and counter-defense systems. The second is a protective state, in which successfully infected phages function as selfish guardians that enhance host fitness. The third involves ecological feedback mechanisms, regulated by host density and environmental stress, which determine the transition between protection and lysis. Understanding this triad is essential for designing effective interventions. Manipulating these evolutionary states may significantly advance AMR control strategies.
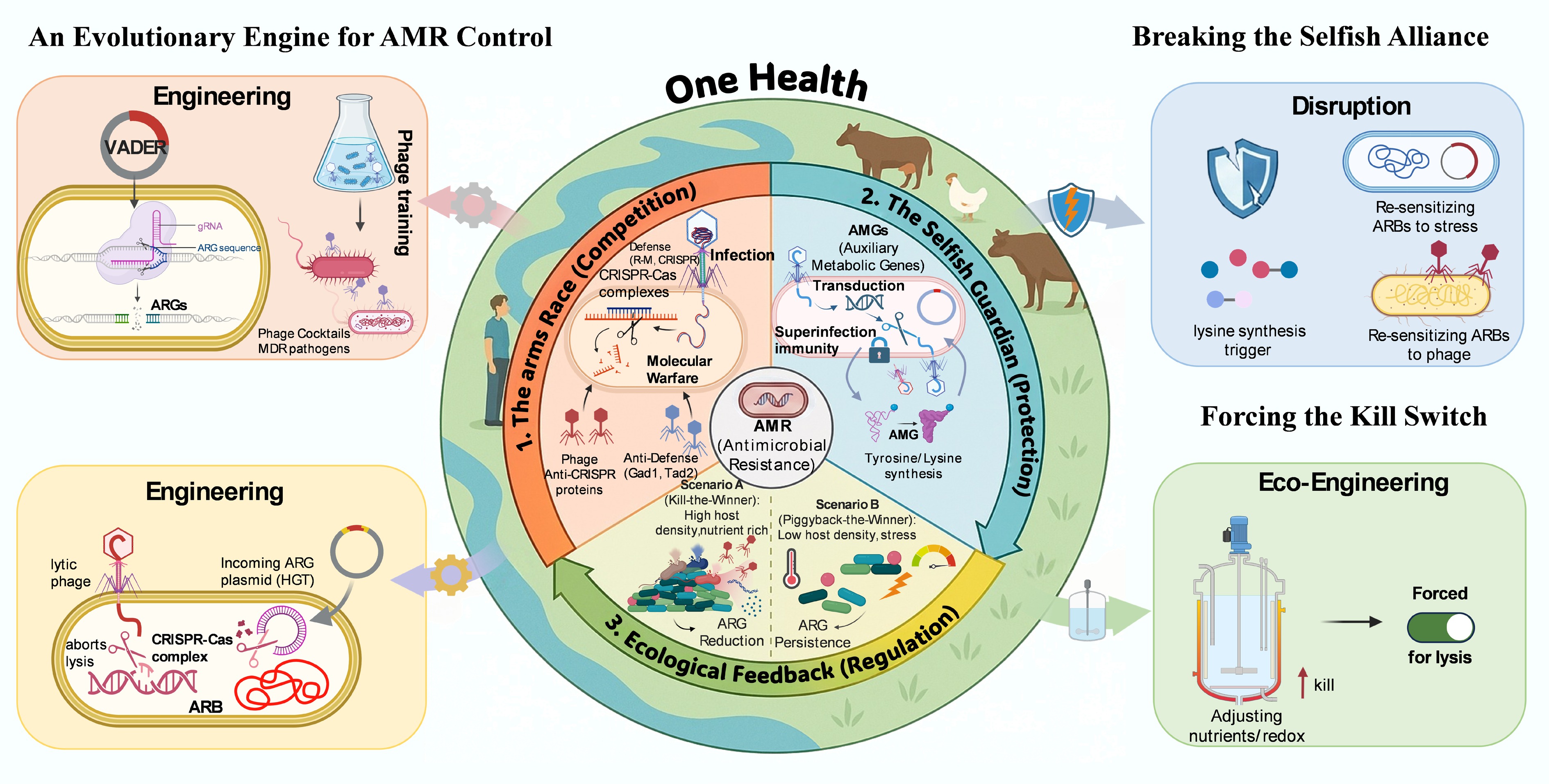
Figure 1.
Phage-host evolutionary triad and strategies for AMR control. The three evolutionary states of phage-host interactions within the One Health context are illustrated. State 1: Arms Race. Phages and hosts engage in molecular conflict (e.g., CRISPR-Cas vs anti-defense proteins). This state provides a genetic arsenal for engineering precision antimicrobials, such as VADER, and optimized phage cocktails. State 2: Selfish Guardian. Successful phages protect their hosts from biotic and abiotic stressors via Auxiliary Metabolic Genes (AMGs) and superinfection exclusion, thereby ensuring survival. Strategies here focus on disrupting this protection to sensitize antimicrobial-resistant bacteria. State 3: Ecological Feedback. Environmental cues (e.g., host density, nutrients) dictate the switch between kill-the-winner (lysis) and piggyback-the-winner (lysogeny). Ecological engineering aims to drive this switch towards lysis, maximizing AMR reduction while minimizing environmental transduction risks.
-
An eternal, high-stakes antagonism defines the first state of the phage-host interaction. For millions of years, phages have engaged in a perpetual molecular arms race to breach host defenses, while bacteria have evolved a sophisticated defensome to block infection[7,8]. This defense includes well-characterized mechanisms like receptor mutations, restriction-modification (R-M), and CRISPR-Cas, as well as recently discovered systems such as CBASS, Thoeris, and the RNA-based TIGR-Tas system[9,10]. Phages, in turn, evolve countermeasures that neutralize immunity, alter adsorption, and restore productive infection[11]. This evolutionary tug-of-war continually generates new molecular parts and design principles that can be translated into AMR control strategies.
One immediate opportunity is to repurpose defence mechanisms as precision tools. CRISPR-Cas systems, for example, can be engineered to target specific resistance determinants. The CRISPR platform, the VADER system, has been developed to specifically degrade plasmid-borne ARGs, offering a route to resensitize resistant populations without broad-spectrum killing[12]. Besides, the phage machinery itself offers a powerful delivery route. Recent advances have used engineered non-replicative phage capsids (phagemids) as cargo vehicles. By exploiting the viral entry mechanisms evolved during the arms race, these systems can inject CRISPR payloads to specifically target resistance genes. For instance, phagemid-delivered CRISPR-Cas13a has been successfully employed to silence mecA in MRSA[13] and reverse carbapenem resistance in P. aeruginosa[14], effectively resensitizing resistant populations without the broad-spectrum lysis associated with traditional phage therapy. In parallel, the trans-cleavage activity of Cas proteins has been incorporated into rapid detection platforms; a recent study used Cas12a in combination with recombinase polymerase amplification to sensitively monitor AMR markers in diverse aquatic environments[15]. These approaches exemplify how arms-race biology can be harnessed to develop programmable systems for identifying and disabling AMR.
A second opportunity is to use arms-race logic to improve the phage candidate itself. A major barrier to durable phage efficacy is the speed and diversity of bacterial anti-phage immunity. Experimental evolution 'phage training', where phages are repeatedly passaged on resistant hosts, can select for counter-defence mutations that expand host range and delay resistance. Together with rational cocktail design, this expands the host range and suppresses the evolution of antiviral defenses in multidrug-resistant Klebsiella pneumoniae and Enterobacter cloacae[16,17]. By maximizing lytic efficiency through directed evolution, the kill-the-winner dynamic can be enhanced, which potentially reduces the burden of AMR.
Finally, the arms race influences the dissemination of AMR in complex ways. Bacterial defense systems play a dual role: they protect the host from lethal phage infection, but they can also act as barriers to the uptake of foreign DNA, including AMR plasmids[18,19]. However, this relationship is nuanced. Our analysis of the ICE (Integrative and Conjugative Elements) database reveals that approximately 25% of ARGs are flanked by antiviral defense genes[20], and a recent Science study by Kieffer et al. found that mobile integrons act as cost-efficient and highly mobile defense islands at the crossroads of antibiotic and bacteriophage resistance[21]. This suggests that while antiviral defense systems can block horizontal gene transfer (HGT) of AMR plasmids, they are also co-selected with ARGs, potentially stabilizing AMR lineages by protecting them from viral predation. Therefore, AMR control strategies must navigate this duality: leveraging bacterial immunity mechanisms (such as CRISPR) to block plasmid uptake while simultaneously employing evolved phages to breach antiviral defense mechanisms.
-
Once a phage successfully breaches host defenses, it faces a branching point: kill the host immediately or farm it for future gain? If environmental conditions are unfavorable for lysis (e.g., low host density or high stress), the phage enters the lysogenic cycle as a phase of selfish protection, and chronic or pseudo-lysogenic states can also occur in some systems. Phage genomes may persist and alter bacterial fitness in ways that can support the survival and competitiveness of antimicrobial-resistant bacteria (ARB). Here, the phage acts as a guardian, enhancing the fitness of its bacterial host, not out of altruism, but to ensure a robust vessel for its own future replication.
While transduction is frequently cited as a mechanism for AMR dissemination, recent metagenomic surveys in anaerobic digestion and urban rivers have shown that viral communities contribute significantly more to the lysis of ARB (reducing ARG loads) than they do to the transduction of ARGs[22,23]. Besides, recent planetary-scale virome analysis indicated that phages rarely encode ARGs[24]. This finding challenges the fear that phage application will inevitably spread AMR, supporting the ecological safety of utilizing phages for AMR control. However, these observations do not eliminate the risks of ARGs transduction, which has been widely validated. For instance, in high-density clinical environments, the potential for generalized transduction, in which bacterial DNA is accidentally packaged into phage capsids, cannot be entirely ruled out[25], and transduction has also been observed in anaerobic digestion of livestock manure and residual sludge[22]. Consequently, stringent genomic screening of therapeutic phage products is mandatory to ensure they are free of ARGs, virulence factors, and integrases, thereby satisfying both safety requirements and regulatory mandates. The ecological role of phages in AMR should not be reduced to the simplistic narrative that phages merely spread antibiotic resistance genes (ARGs). A more nuanced risk is that phages can enhance the persistence of antibiotic-resistant bacteria (ARB) even under conditions of weak antibiotic selection. Phages offer a multifaceted defense against both biotic and abiotic stressors. Specifically, prophages confer superinfection immunity, preventing secondary infections by other lytic phages and thereby protecting ARB from viral predation[26,27]. They may also encode toxin systems or other virulence factors that eliminate bacterial competitors[18]. Abiotically, phages carry auxiliary metabolic genes (AMGs) that bolster host resilience against environmental pressures such as temperature, pH, and even antibiotics[28]. The role of AMGs in this selfish guardianship is profound. In soil ecosystems, viral AMGs modulate carbon and nutrient cycling, enhancing host survival in nutrient-poor niches[29,30]. Viral manipulation of methane metabolism can optimize host energetics[31]. In the human gut, a striking example involves AMGs for the synthesis of energetically costly amino acids, such as tyrosine (e.g., prephenate dehydrogenase) and lysine (e.g., diaminopimelate decarboxylase). Since these amino acids are scarce in the gut and absorbed by the host epithelium, phages that gift their bacterial hosts the ability to synthesize them provide a decisive competitive advantage[32]. This metabolic subsidy promotes host growth and ecological dominance, aligning with the piggyback-the-winner dynamic, in which lysogeny is favored to maintain a stable host population.
This protective state fundamentally alters the approach to AMR control. If phages are actively helping ARBs cope with environmental stress (e.g., heavy metals in wastewater or nutrient scarcity in the gut), then current strategies that focus solely on blocking transduction are insufficient. We must recognize that phage-mediated protection can stabilize AMR populations even in the absence of antibiotic selection. However, this selfish guardian role also presents a therapeutic opportunity. By deciphering the specific metabolic boosts phages confer (e.g., lysine synthesis), interventions can be designed to disrupt these pathways and strip ARBs of their viral advantage. Furthermore, the selfish nature of this protection creates a vulnerability: the phage waits for the right moment to kill. A large-scale analysis of over 267,000 Escherichia coli genome assemblies unexpectedly revealed 373 integrated virulent phage genomes that persist in a chronic infection state within the host chromosome without causing significant cell death[33]. Understanding the triggers that end this protective state is key to forcing the phage to betray its host.
-
The transition from the selfish guardian phase (Stage 2) back to the arms race (State 1) is not random; a sophisticated ecological feedback loop governs it. This state determines the fate of the host population: will the phage continue to protect its host to ensure survival, or will it trigger lysis or prophage excise to capitalize on host abundance? This branching point is dictated by population-level dynamics, most notably the tension between kill-the-winner and piggyback-the-winner strategies. Theoretical frameworks, such as extended Lotka-Volterra models, demonstrate that these dynamics are sensitive to host density and metabolic activity[34,35]. For example, nutrient pulses or redox potential fluctuations can act as critical environmental cues that shift the equilibrium, forcing temperate phages from a stable piggyback lysogenic state into an active kill-the-winner lytic cycle[35]. Understanding these mathematically predictable transitions is essential for developing ecological engineering strategies that can precisely trigger host lysis in high-density AMR populations.
Ecological theory suggests that in nutrient-rich (eutrophic) environments with high host densities, phages adopt the kill-the-winner strategy. Here, the abundance of hosts favors lytic replication, causing phages to aggressively infect and lyse the dominant bacterial populations (the winners). This dynamic results in a positive correlation between phage and host abundances and drives rapid turnover of biomass[36]. Conversely, in nutrient-poor (oligotrophic) or stressful environments, the piggyback-the-winner dynamic prevails[37]. When host densities are low or the host is under stress, phages switch to lysogeny (State 2), integrating into the host genome to piggyback on its survival rather than risking lysis in a host-scarce environment.
This switching mechanism is crucial for ecosystem stability. For instance, the recent study observed that in methanogenic anaerobic digestion systems, a delicate balance exists between volatile fatty acid (VFA) producers and VFA-consuming methanogens. Under stable conditions, temperate phages likely piggyback on them, protecting them and maintaining system function. However, if volatile fatty acids (VFAs) accumulate, indicating ecosystem stress and overgrowth of VFA producers, phages may shift to a kill-the-winner strategy, lysing the fast-growing acidogens to restore equilibrium. This homeostatic function demonstrates that phages act not only as killers, but also as regulators of ecosystem health. Such shifts in phage lifestyle have been documented in various ecosystems[36].
In the context of AMR control, a key challenge is to identify and manipulate environmental triggers that direct phage-host systems toward outcomes that reduce the abundance of antibiotic-resistant bacteria (ARB), while limiting opportunities for gene mobilization. It is essential to determine the specific environmental factors, such as nutrient ratios, temperature, or particular stress signals, that induce phages to transition from the protective piggyback-the-winner mode to the lytic kill-the-winner mode, specifically targeting AMR populations. Engineering the environment to promote kill-the-winner dynamics for ARBs, while preserving piggyback-the-winner stability for beneficial commensals, may maximize the reduction of AMR pathogens and minimize the risk of antimicrobial resistance gene (ARG) dissemination via transduction. This ecological engineering approach resembles the final, integrative step in the phage-host evolutionary triad. However, the feasibility of this approach varies significantly across ecosystems. In engineered systems like wastewater treatment plants, adjusting organic loading or redox potential to trigger prophage induction in ARBs is relatively feasible. Empirical studies and ecological models suggest that nutrient-rich conditions and environmental stressors, such as redox shifts, can drive a transition from lysogeny to lysis[37]. However, it was acknowledged that this level of control is limited in natural environments. In soil, spatial heterogeneity and micro-niche governance constrain viral activity, making large-scale environmental manipulation to force the kill switch significantly more challenging[30].
-
Effective control of AMR requires translating knowledge of the phage-host evolutionary triad into actionable biotechnological and ecological strategies. These interventions should not be applied uniformly. Rather, they must be tailored to specific environmental contexts. Recent global surveys indicate that soil viral diversity is shaped by factors such as moisture and texture, resulting in micro-niches that spatially constrain viral activity[38]. Similarly, the vast, unexplored viral diversity uncovered in the Global Soil Virus Atlas suggests that the potential for biogeochemical impact via AMGs is immense but unevenly distributed[30]. Recognizing these spatial and environmental patterns is the first step toward predicting hotspots of ARG dissemination (via transduction) or reduction (via lysis), allowing for targeted management strategies that bridge environmental reservoirs with clinical and livestock health outcomes—a cornerstone of the One Health approach.
Building on this ecological foundation, a three-pronged approach to AMR control was proposed (Fig. 1). (1) Engineering the Phage-Host Arms Race: From Killing to Disarming. We should continue to expand the genetic editing toolkit derived from the phage-host conflict. While traditional phage therapy focuses on lysis, the arms race offers tools for precision genetic engineering. The development of standardized purification protocols[39] and the rational design of phage cocktails based on host-range matrices[17] are prerequisites for scalable application. Future research should pivot toward disarming strategies: utilizing conjugation-based systems (such as VADER) to deliver CRISPR-Cas payloads that specifically degrade resistance plasmids[12] and inducing antibiotic resensitization of ARB through the development of phage resistance[17]. This approach could revert multidrug-resistant populations to antibiotic sensitivity without the ecological disruption caused by broad-spectrum killing. (2) Disrupting Viral Protection: Breaking the Selfish Alliance. A critical frontier involves targeting the selfish guardianship phages that ARBs receive. If phages stabilize AMR populations through metabolic subsidies (e.g., AMG-mediated amino acid synthesis) or superinfection exclusion, strategies can be developed to break this alliance. For instance, counter-defense strategies can be employed to overcome prophage-conferred immunity. Recent discoveries of diverse phage-encoded anti-defense proteins suggest that therapeutic phages can be engineered to neutralize these protective systems, effectively breaching the viral shield[11]. Furthermore, by identifying specific metabolic boosts from AMGs, inhibitors can be designed to disrupt these viral-mediated pathways, stripping ARBs of their fitness advantage and turning viral protection into a liability for the host. (3) Ecological Engineering: Forcing the Kill Switch. The ultimate goal of environmental AMR control is to manipulate the ecological feedback loop to favor kill-the-winner dynamics against ARB. In engineered systems such as wastewater treatment plants, this could involve precise adjustments to organic loading rates, redox potential, or nutrient ratios to trigger prophage induction (lysis) specifically within ARB populations. In natural environments such as soils and rivers, understanding how moisture and nutrient pulses drive viral activity can help us predict ARG blooms and implement timing-based interventions. This may move the field from reactive therapy to proactive ecosystem management.
Several limitations of this evolutionary triad warrant consideration. First, although enforcing the lytic kill-the-winner switch effectively reduces the abundance of ARB, the consequent release of intracellular DNA may increase the local reservoir of free-floating ARGs, potentially facilitating secondary dissemination through natural transformation. Second, although engineered phage capsids for CRISPR delivery offer precision, strict monitoring is required to prevent unintended transduction of host genomic fragments. Third, aggressive ecological engineering to promote lysis poses a risk of collateral damage, as the removal of AMR-carrying lineages may inadvertently disrupt essential biogeochemical cycles or symbiotic networks within the broader microbial community. Future strategies should incorporate real-time ecological monitoring to balance AMR reduction with ecosystem stability.
In summary, phages should not be regarded solely as bacterial killers but also function as genetic engineers, metabolic guardians, and ecological regulators. The evolutionary relationship between phages and their hosts transitions from intense competition for entry to a mutually beneficial alliance for survival, and ultimately to a population-level divergence shaped by ecological feedback. Recognizing these three states enables a shift from viewing phages merely as alternative antimicrobials to adopting a comprehensive approach to microbial ecosystem management. By deliberately steering the phage-host dynamic from protection toward competition and lysis, it is possible to leverage the virosphere to address the ongoing pandemic of AMR, thereby advancing the environmental objectives of the One Health framework.
-
The authors confirm their contributions to the paper as follows: Junya Zhang: conceptualization, writing – original draft, visualization, formal analysis; Jingxian Yang: conceptualization, validation, writing – review & editing; Yuansong Wei: writing – review & editing; Hans Hermann Richnow: conceptualization, writing – review & editing, validation. All authors reviewed the results and approved the final version of the manuscript.
-
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
-
This work is supported by the National Natural Science Foundation of China (Grant No. 42477403). Dr. Junya Zhang is also grateful for the financial support from the Alexander von Humboldt Foundation.
-
Authors declare there are no competing financial interests in relation to the work described.
-
Full list of author information is available at the end of the article.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhang J, Yang J, Wei Y, Richnow HH. 2026. Reconsidering phage-host evolutionary triad for antimicrobial resistance control. Biocontaminant 2: e006 doi: 10.48130/biocontam-0026-0003
Reconsidering phage-host evolutionary triad for antimicrobial resistance control
- Received: 13 January 2026
- Revised: 11 February 2026
- Accepted: 30 March 2026
- Published online: 16 April 2026
Abstract: The escalating crisis of antimicrobial resistance (AMR) requires innovative interventions beyond conventional antibiotics. Bacteriophages, as natural bacterial predators, represent a promising alternative. However, their influence is governed by a dynamic evolutionary continuum that determines whether they eradicate or maintain AMR populations. This work introduces a tripartite evolutionary framework to elucidate phage-host interactions: (1) an antagonistic arms race state that drives the diversification of molecular defenses; (2) a selfish protection state in which phages enhance host fitness to secure their own persistence; and (3) ecological feedback mechanisms that regulate transitions between these states. The potential for manipulating these evolutionary states to transform AMR control strategies is examined. Recent metagenomic and mechanistic evidence indicates that the arms race state provides a genetic toolkit for targeting AMR pathogens, whereas the protective state, mediated by auxiliary metabolic genes (AMGs) and superinfection exclusion, may inadvertently stabilize AMR lineages. Notably, environmental context is shown to govern the shift between kill-the-winner and piggyback-the-winner dynamics, ultimately influencing the efficiency of AMR transfer. This analysis highlights how targeted manipulation of this evolutionary triad could facilitate the development of advanced phage therapeutics and ecological engineering approaches to mitigate global AMR dissemination within the One Health framework.