-
With continuing urbanization in developing countries, the scale of construction activities continues to expand, resulting in increasing demands for building materials such as bricks and cement[1]. Globally, around 1,500 billion bricks are produced annually, among which, China accounts for more than 50% of bricks produced[1,2]. As an ancient technology, brick firing has made great contributions to the social-economic development of China. However, the traditional process of brick production may cause the enrichment of metal(loid) contaminants such as Pb, Cr, and As in the environment[3,4]. Among these, As is of primary concern due to its high environmental mobility and significant toxicity even at low concentrations, necessitating a targeted investigation into its impact on soil bacterial communities. These substances have strong toxicity and are non-biodegradable, persisting in the environment for many years and posing a threat to human health through invasion and runoff infiltration into soils[5]. Heavy metals enter the food chain through plants. They accumulate in the human body through the biological amplification effect, thus causing physical damage and physiological diseases[5,6]. To date, the environmental effects, health effects, and sustainable use of resources in the brick kiln industry have been widely discussed within the literature[7−9]. However, the specific metabolic pathways and detoxification mechanisms of bacterial communities in response to As stress within shut-down brick kiln soils remain to be fully elucidated.
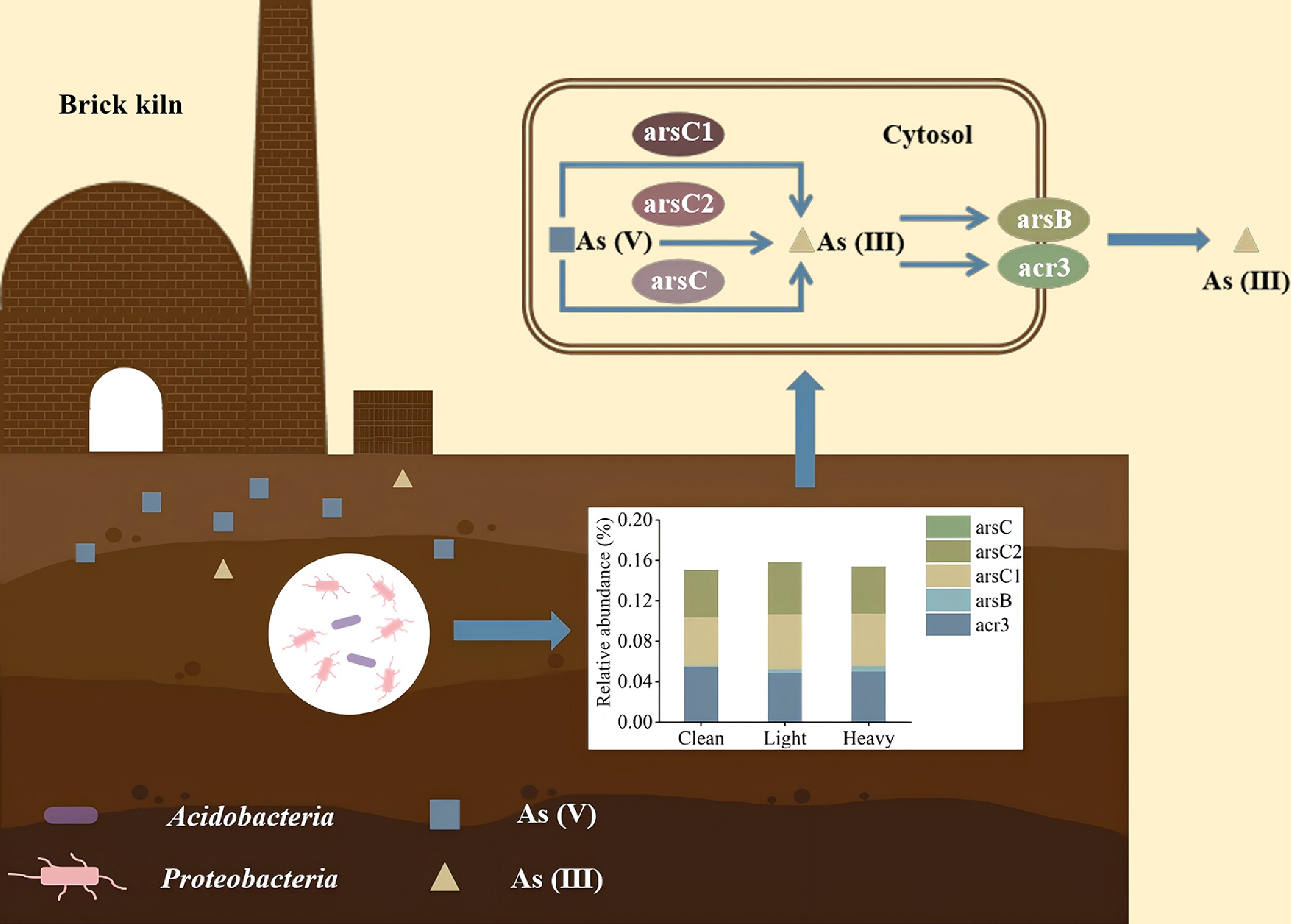
The structure, diversity, and functions of indigenous microorganisms can serve as biological indices to evaluate the health of the soil and have the potential to contribute to soil ecological restoration[10−12]. Microorganisms have been living in the environment with heavy metal stress for 3.5 billion years, during which time they have developed several defense mechanisms to relieve the toxic effects of heavy metals[13]. Heavy metal resistance genes (HMRGs) are able to regulate bacterial responses to heavy metals stresses, including detoxification and excretion[14]. The detoxification mechanism for As includes reduction (arsC and arsC1/arsC2) of arsenate As(V), excretion (arsB and acr3) of arsenite As(III), as well as methylation (arsM and arsH)[15]. In contaminated soils in the real world, As exists at gradient concentrations. Based on the Soil Environmental Quality Risk Control Standard (GB 36600-2018), the selected Clean, Light, and Heavy levels effectively represent the progression from background levels to concentrations exceeding regulatory screening thresholds, providing an ecologically relevant gradient for studying microbial stress responses[11]. Elevated levels of As adversely impact the structure of microbial communities, resulting in a reduction in both bacterial diversity and abundance[16]. Moreover, microbial metabolic processes, such as carbon, nitrogen, and sulfur metabolism, are also susceptible to arsenic stress[17]. This damage to community function can impair the normal operation and maintenance of soil ecosystems[16,18]. Therefore, exploring the structural and functional characteristics of soil microflora in brick kilns has important significance for subsequent ecological restoration and soil health management.
We hypothesized that (i) increasing As concentrations would lead to a reduction in bacterial diversity and a shift in community composition toward arsenic-tolerant taxa; (ii) As stress would destabilize overall network cohesion while potentially intensifying specific survival-driven bacterial interactions; and (iii) bacterial communities would adapt through the upregulation of specific metabolic pathways and arsenic detoxification genes. Based on these hypotheses, soil samples with different As contamination gradients obtained from a typical shut-down brick kiln in Anhui Province, China, were analyzed. This study offers a new scientific understanding of the response mechanisms of microflora in As-contaminated soils surrounding shutdown brick kilns.
-
Soil samples were collected from a contaminated area of a shutdown brick kiln in Anhui Province, China. To address the issue of data reliability, a total of 1,975 original samples were collected in the early stage of this study (see Supplementary Table S1). Subsequently, we conducted preliminary analysis on the data, selecting representative sites that reflected a clear As gradient (Clean, Light, Heavy) after a systematic screening of the 1,975 original samples to ensure that the chosen sites were ecologically representative while minimizing the impact of environmental heterogeneity, such as pH variation. The sample number reflects a balance between analytical depth and cost-efficiency. Finally, 12 uniform representative soil samples were selected and stored in two parts. According to the Soil Environmental Quality Risk Control Standard for Soil Contamination of Development Land (GB 36600-2018), the screening value of As for Class-I land is 20.00 mg/kg[19]. Therefore, taking into account the average concentration and concentration range of As at each level, as well as funding and geographical location, soil samples were denoted as Clean (average concentration: 8.42 mg/kg), Light (20.87 mg/kg), and Heavy (54.46 mg/kg) according to their As concentrations. Soil samples (100 g) were collected at each site and mixed evenly, with four replicates collected at each site. The sample number reflects a balance between analytical depth and cost-efficiency. The first was used for the analysis of the physicochemical properties (stored at 4 °C) of the soil, and the second was used for the analysis of the molecular biology (stored at −80 °C) of the soil[20].
Test of soil physicochemical properties
-
Soil organic matter (OM) was determined by the potassium dichromate oxidation method, total nitrogen (TN) by the Kjeldahl method, and total phosphorus (TP) and total potassium (TK) were analyzed using spectrophotometry and flame photometry, respectively. Soil texture was determined using the hydrometer method, identifying the samples predominantly as silty clay loam. All physicochemical data are provided in Supplementary Table S2, and the results of these analyses are discussed in Section 'Correlations between the physicochemical properties of contaminated soils from a brick kiln and the microflora structure of the soil'[17,21,22]. Additionally, the higher clay proportion in the Heavy group might have led to a higher As concentration.
Extraction and sequencing of soil DNA
-
The Fast DNA SPIN Kit for soil (MP Biomedical, USA) was used to extract soil DNA. The DNA concentration and mass were tested with a NanoDrop One (Thermo Scientific, Singapore) and agarose gel electrophoresis (AGE) (Bio-Rad, Hercules, CA, USA)[23]. The primer pair 338F (5'-ACTCCTACGGGAGGCAGCA-3') and 806R (5'-GGACTACHVGGGTWTCTAAT-3') were used for analysis of the V3–V4 areas of the 16S rRNA of amplified bacteria[24]. The database was built by Guangzhou Meige Gene Technology Co., Ltd. The Illumina MiSeq platform was used for high-throughput sequencing[12]. The sequencing data from this study have been stored in the NCBI Short Read Archive Database (Accession Number: PRJNA1049149).
Species annotation and function annotation
-
Operational taxonomic unit (OTU) clustering of non-repetitive sequences with a similarity of 97% was carried out using UPARSE (v7.1). While ASV-based methods offer higher taxonomic resolution, the 97% OTU approach was adopted here to facilitate direct comparison with existing heavy metal contamination datasets, and its potential limitations in resolution are acknowledged in our functional interpretations. The chimera was eliminated, and sequences were optimized and mapped onto the OTU representative sequences, thus producing the OTU table[25]. The OTU table was compared with the SILVA database (QIIME2, 2019.4) for species taxonomic annotation (Kingdom, phylum, class, order, family, genus, species); the confidence threshold was 70%[26]. PICRUSt2 (v2.2.0) was chosen for function prediction. The abundance information of the KO and pathways of the samples were obtained. Given that the primers used in this study (338F and 806R) specifically target the bacterial 16S rRNA gene, the subsequent functional predictions were strictly limited to the bacterial community, excluding fungal gene profiles[27,28].
Statistical analysis
-
Origin 2022 software was used for graphic plotting. The vegan package in R language (v3.4.1) was used to calculate the α diversity of the soil microorganisms. Subsequently, the significance of microflora differences in the clean and As-contaminated soils (Clean, Light, and Heavy) was evaluated by the ANOSIM method[29]. Further, the edge files and point files of the bacterial co-occurrence network were acquired using OTUs with a relative abundance greater than 0.05% across all samples to ensure network robustness, and these were input to Gephi (0.9.2) for visualization[30]. Prior to analysis, relative abundance data were arcsine-square-root transformed to improve normality, and the assumptions of normality and homoscedasticity were verified using Shapiro-Wilk and Levene's tests, respectively. Variables that met these assumptions were subjected to one-way ANOVA to calculate significant differences among samples. For data that did not conform to a normal distribution, the non-parametric Kruskal-Wallis test was employed (α = 0.05, 0.01, and 0.001).
-
A total of 796,917 optimized bacterial sequences were obtained from high-throughput sequencing of 12 soil samples, with read counts ranging from 54,231 to 72,145 per sample. Samples were rarefied to the minimum library size to ensure even sampling depth, which was sufficient to capture the representative bacterial diversity as indicated by the plateauing rarefaction curves (Fig. 1a). A total of 1,124 core OTUs were common in the Clean, Light, and Heavy groups. Specifically, the Clean group contained 225 specific OTUs, while the Light and Heavy groups contained 80 and 29 specific OTUs, respectively. There were 227 common OTUs between the Clean and Light groups, and there were 52 common OTUs between the Clean group and the Heavy group. It can be seen from Fig. 1b that Acidobacteria, Proteobacteria, and Bacteroidetes were the dominant bacteria at the phylum level in the Clean, Light, and Heavy groups. This is consistent with previous research results, indicating that soil microorganisms can adapt to different degrees of heavy metal contamination by adjusting their microflora structures (Supplementary Table S3)[31,32]. In the Clean group, the relative abundances of Acidobacteria, Proteobacteria, and Bacteroidetes were 37.85%, 27.37%, and 15.43%, respectively, accounting for 80.65% of the bacteria in the group. In the Light group, the relative abundance of Acidobacteria decreased to 35.73%, while the relative abundance of Proteobacteria increased to 28.68%, and the relative abundance of Bacteroidetes increased to 18.75%, accounting for 83.16% of the bacteria in the group. In the Heavy group, the relative abundance of Acidobacteria decreased to 31.47%, the relative abundance of Proteobacteria increased to 38.54%, and the relative abundance of Bacteroidetes decreased to 14.81%, accounting for 84.83% of the bacteria in the group. Generally speaking, the relative abundance of Acidobacteria decreased with increases in the degree of contamination, while Proteobacteria gradually began to dominate, and were able to tolerate environments with high concentrations of heavy metals[6,33].
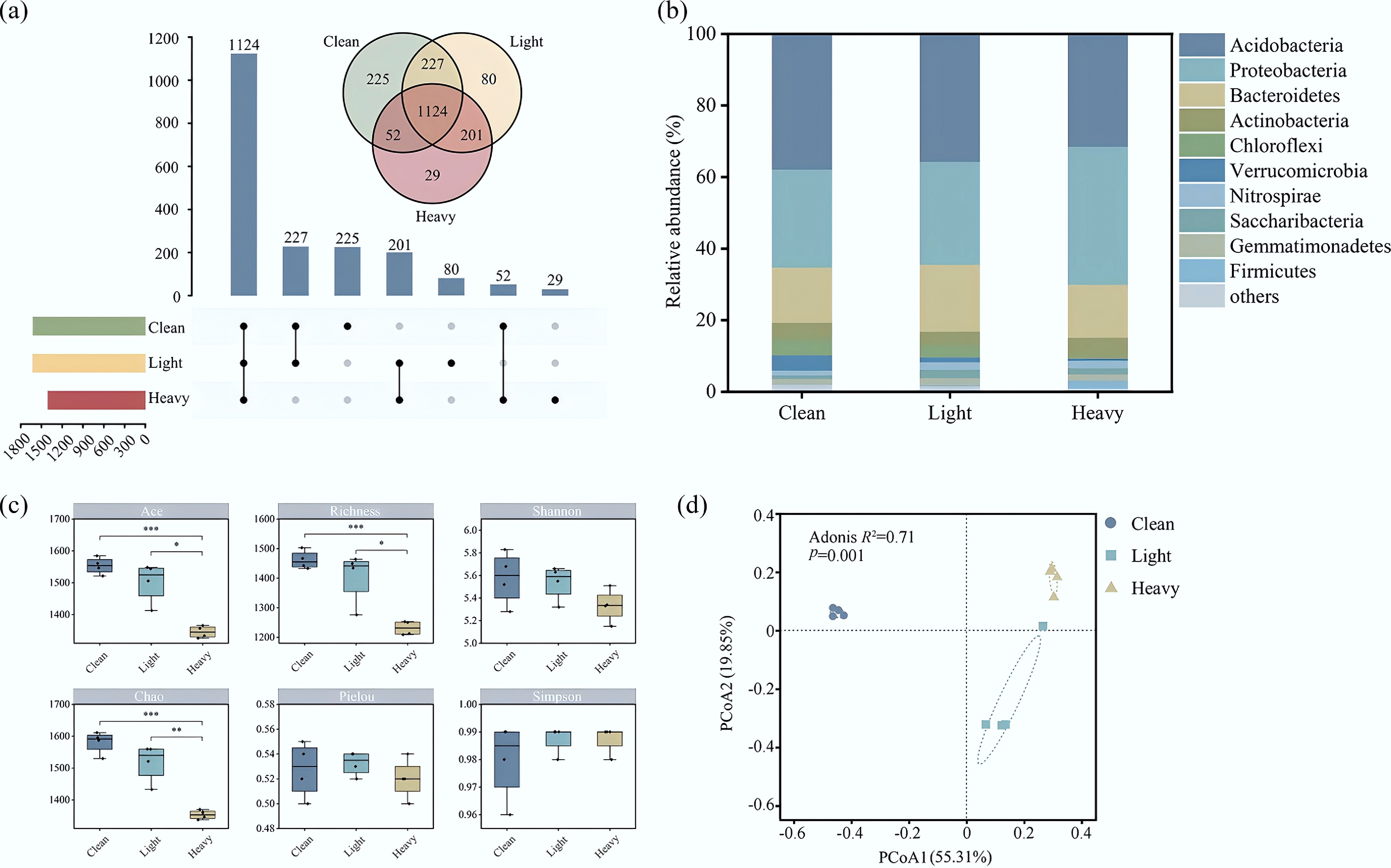
Figure 1.
Composition of the bacterial communities in contaminated soils obtained from a shut-down brick kiln. (a) Differences in the OTU quantity among different sites; (b) relative abundances of the top 10 bacteria at the phylum level at different sites; (c) α diversity of the bacterial communities; (d) β diversity of the bacterial communities.
The influence of soil As on the α diversity of bacterial communities is shown in Fig. 1c. With respect to the Shannon index and Simpson index, the following trend was observed: Clean > Light > Heavy. This indicates that with increases in the As concentration of the soil, the diversity of indigenous microorganisms decreased. Similarly, the Ace index and Chao index exhibited decreasing trends, indicating that the abundance of the bacterial communities was significantly associated with arsenic gradients (p < 0.001, Supplementary Table S4). The decreasing Pielou index indicates that species uniformity decreased in parallel with the increasing concentrations of pollutants (PCoA was used to compare the microbial communities among different contaminated areas, Fig. 1d). The bacterial community structure between contaminated soil samples (Light, Heavy) was similar, but differed from that of Clean soil samples. The ANOSIM analysis further demonstrated significant differences in the bacterial communities of the clean and contaminated soils (R2 = 0.71, p = 0.001). These results indicate that the ecological niche of clean soil and contaminated soil changes with the variation of arsenic pollution, causing changes in the bacterial community structure[34]. Although the highest As concentration measured in this study may not be classified as extremely high, similar concentrations have been reported to significantly impact microbial diversity, especially in soils with low buffering capacity or previously low background As levels[35]. This supports our observation of a notable decrease in both richness and evenness of bacterial communities in the Heavy group.
Co-occurrence network analysis was performed to examine the complicated interaction modes and ecological laws among the microorganisms[36,37]. The bacterial co-occurrence network (weighted degree: 10.608) of all samples was composed of 46 nodes and 301 sides, covering seven modules (Fig. 2). The nodes in the bacterial co-occurrence network were mainly composed of Proteobacteria (21 nodes), Bacteroidetes (eight nodes), and Acidobacteria (five nodes). According to the betweenness Centrality value, g_unclassified_c_OPB35_soil_group (66.628), g_Paucimonas (48.442), g_Flavisolibacter (45.926), and g_Ferruginibacter (44.527) served as bridges in the bacterial co-occurrence network. Further, the average weighted degrees of g_Paucimonas (21.923) and g_unclassified_c_OPB35_soil_group (21.524) were the highest. Hence, the role of g_Paucimonas and g_unclassified_c_OPB35_soil_group in the whole bacterial community cannot be ignored. Some studies suggest that g_Paucimonas can take a dominant role in an oligotrophic environment, and is related to the oxidation of Fe(II)[38]. g_unclassified_c_OPB35_soil_group belongs to Verrucomicrobia, and the abundance of Verrucomicrobia is related to the C, N, and P contents of soils[39]. Ferruginibacter is related to the N cycle of soil[12]. These taxa exhibited the highest weighted degrees, suggesting their pivotal role in maintaining community structure under As stress.
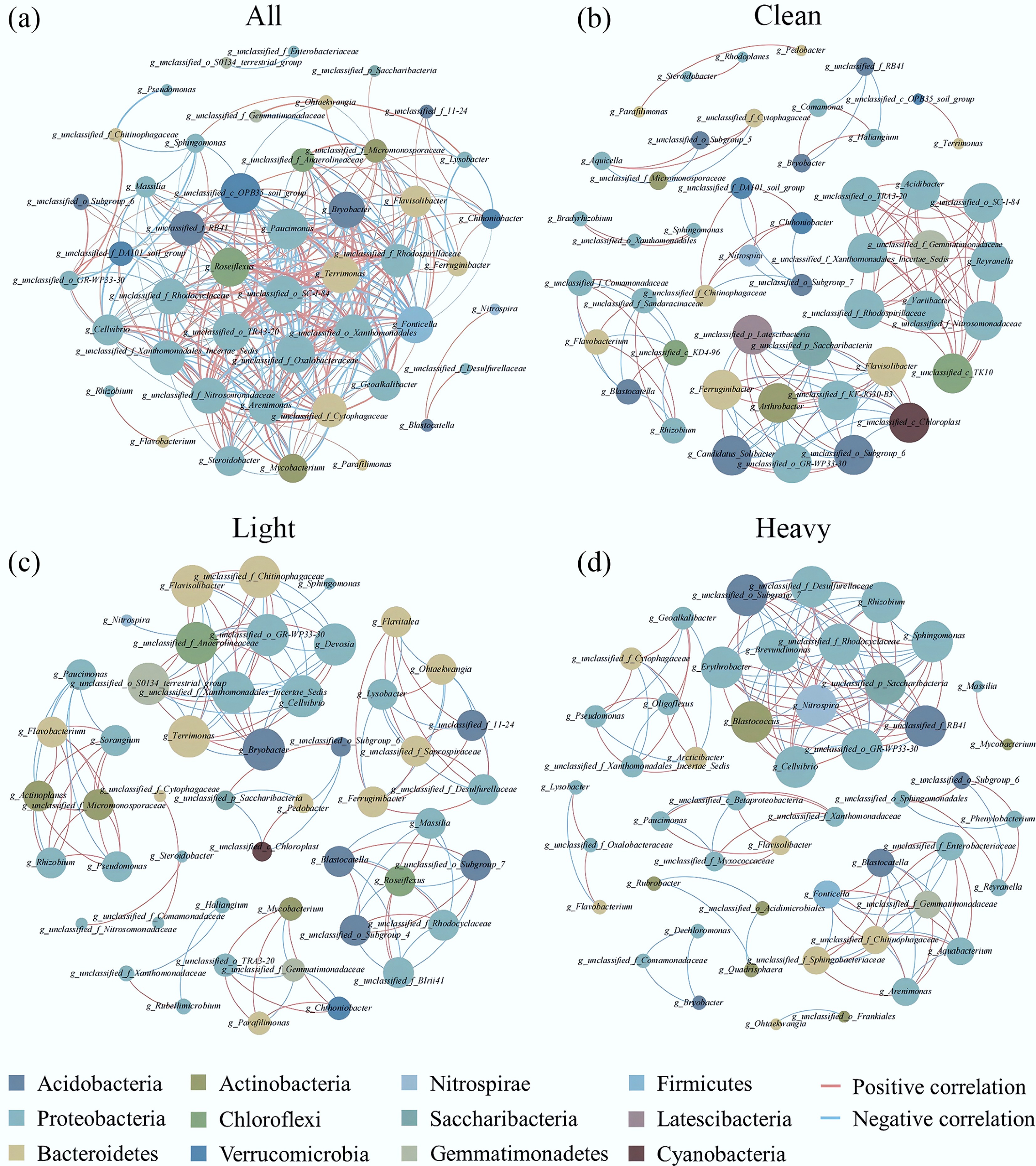
Figure 2.
Co-occurrence network of the top 50 bacteria in terms of their relative abundance at the genus level (Spearman correlation coefficient: r > 0.8, p < 0.01). (a) All samples (Clean, Light, and Heavy); (b) Clean, (c) Light, (d) Heavy. Red indicates a positive correlation and blue indicates a negative correlation. Please refer to the web version of this paper for more detail about the interpretation of the colors.
The bacterial co-occurrence network of the Clean group (weighted degree: 5.542) was composed of 48 nodes and 133 sides. The bacterial co-occurrence network of the Light group (weighted degree: 5.306) was composed of 49 nodes and 130 sides. The bacterial co-occurrence network of the Heavy group (weighted degree: 6.041) was composed of 49 nodes and 148 sides (Fig. 2). The increasing density of the bacterial networks might be related to increases in the As concentration, thus increasing the interaction density of the bacterial community (Clean: 0.118, Light: 0.111, and Heavy: 0.126). Further, the decreasing degree of modularization implies rebuilding of the network structure, and this might make the bacterial communities more susceptible to external disturbance (Clean: 0.748, Light: 0.793, and Heavy: 0.669)[37]. While environmental stress has been broadly shown to destabilize overall microbial network cohesion, the concurrent increase in interaction density observed here suggests that severe As stress may selectively drive surviving tolerant taxa to strengthen specific cooperative or competitive linkages as an adaptive strategy. Under contamination stress, the positive correlations among the abundant dominant bacteria (e.g., Proteobacteria) in the Clean group decreased, indicating that these bacteria are able to change from a cooperative relationship to a competitive relationship[40]. Hence, As pollution might decrease the soil ecosystem stability in shut-down brick kilns.
Correlations between the physicochemical properties of contaminated soils from a brick kiln and the microflora structure of the soil
-
Soil physicochemical properties shifted across the As gradient. The TP and TK contents of the clean soil were significantly higher than those in contaminated soils (p < 0.01), and the accumulation of As(V) was more pronounced than that of As(III) (Fig. 3a). Soil constitutes a complex system, with its physical and chemical properties (comprising organic carbon, total nitrogen, particle size distribution, etc.) playing a significant role in shaping the microbial community structure within it[41]. Metal contaminants pose a potential threat to soil microbes, particularly in the context of industrial activities like brick kilns[42]. Nevertheless, soil microbial communities exhibit a degree of adaptability, and can acclimate to metal pollution[43].
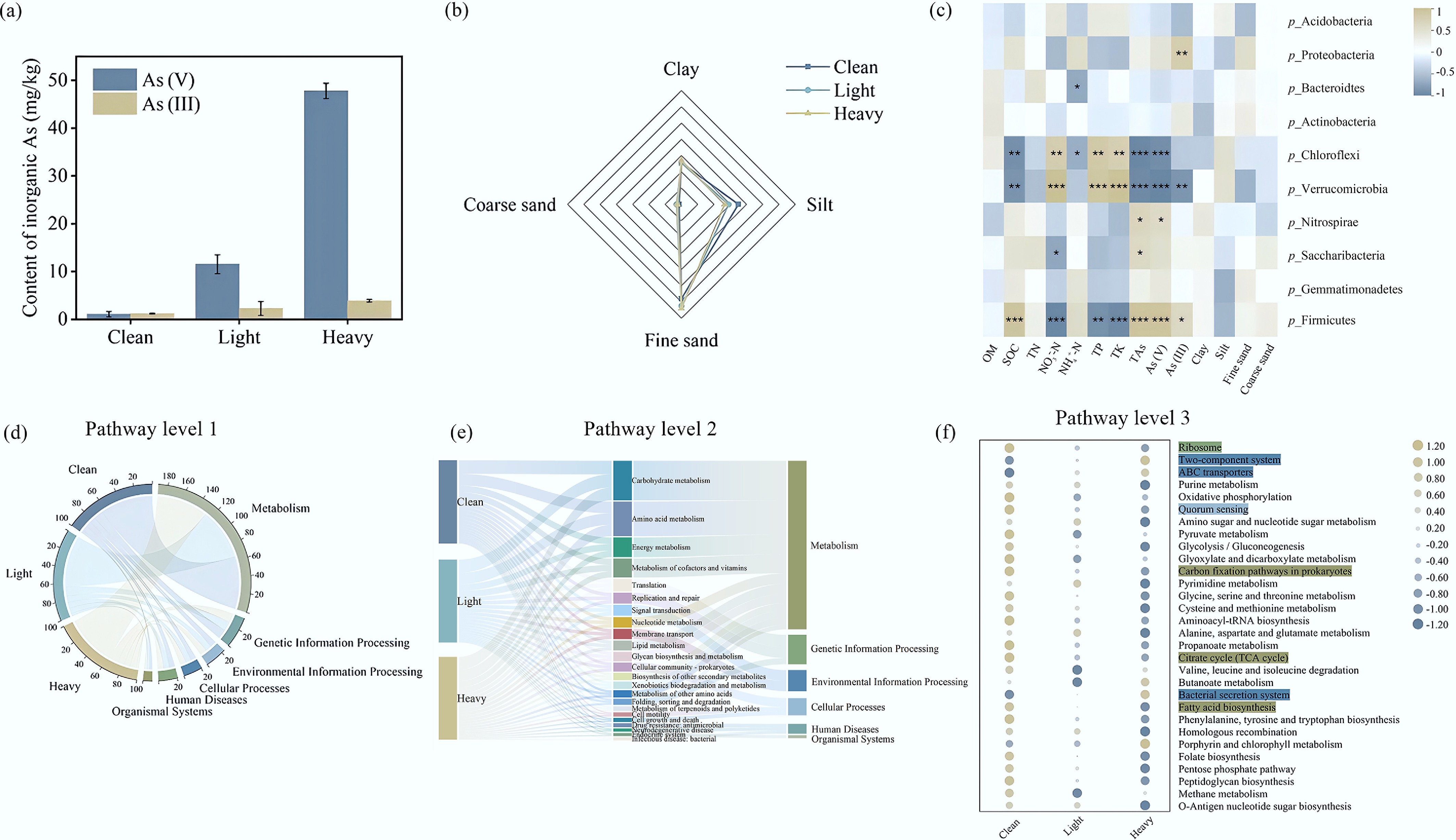
Figure 3.
(a) Inorganic As content; (b) mechanical composition of soils; (c) heat maps of the correlations between the bacterial community (phylum level), the soil physicochemical properties, and heavy metals contamination. OM: organic matter; SOC: soil organic carbon; TN: total nitrogen; TP: total phosphorus; TK: total potassium; TAs: total arsenic; (d) Circos diagram illustrating the inter-group distribution of KEGG Level 1 pathways; (e) Sankey diagram showing the hierarchical mapping and contribution of Level 1 categories to the 23 most abundant Level 2 metabolic pathways; (f) the bubble diagram is composed of the top 30 level-three metabolic pathways in terms of relative abundance.
Therefore, a thorough investigation into the interplay between microbial communities and the soil's physical and chemical properties in areas tainted by brick kiln pollution is crucial for providing a solid scientific foundation and theoretical framework to support soil remediation efforts and ecological environmental protection initiatives. In this study, the relationships between the top 10 microflora (phylum level) in terms of relative abundance, the soil physicochemical properties, and the level of heavy metals contamination were analyzed (Fig. 3b, c). p_Proteobacteria was significantly positively correlated with As(III) (p < 0.01), which may reflect their active involvement in Arsenate reduction and Arsenite tolerance, consistent with the increased abundance of arsC2 and arsB genes in contaminated soils[6]. Soils with a higher NH4+–N content might be disadvantageous for the survival and reproduction of Bacteroidetes (p < 0.05). High concentrations of NO3−–N (p < 0.01), TP (p < 0.01), and TK (p < 0.01) in soils might be good for the survival of Chloroflexi, while high concentrations of SOC (p < 0.01), NH4+–N (p < 0.05), TAs (p < 0.001), and As(V) (p < 0.001) might be disadvantageous for the survival of Chloroflexi. Similar to the Chloroflexi results, soils with high concentrations of NO3−–N (p < 0.001), TP (p < 0.001), and TK (p < 0.001) appeared to be beneficial for the survival and reproduction of Verrucomicrobia, but soils with high concentrations of TAs (p < 0.001), As(V) (p < 0.001), As(III) (p < 0.01), and SOC (p < 0.01) appeared to be disadvantageous[39]. Soils with high concentrations of TAs (p < 0.001), As(V) (p < 0.001), As(III) (p < 0.05), and SOC (p < 0.001) appeared to be beneficial for the survival of Firmicutes, but soils with high concentrations of TP (p < 0.05), NO3−–N (p < 0.001), and TK (p < 0.001) appeared to be disadvantageous for the survival of Firmicutes. Although Acidobacteria and Bacteroidetes were among the dominant phyla in terms of relative abundance, no statistically significant correlation with As species was observed for these groups. This suggests that their variation may be driven by other soil factors, such as organic matter or nutrient content, rather than directly by As concentrations. In contrast, Firmicutes showed positive correlations with TAs, As(V), and As(III) (p < 0.05), implying their potential resistance and adaptability under arsenic stress. However, they were negatively correlated with NO3––N, TP, and TK, which may reflect a trade-off between heavy metal tolerance and nutrient dependency. Acidobacteria, although often considered oligotrophic, showed weak or no significant correlations, suggesting their overall sensitivity to As accumulation. These patterns highlight how nutrient dynamics and metal contamination jointly influence microbial community assembly. For instance, the positive correlation of Firmicutes with As concentration may be due to their robust spore-forming ability and diverse resistance genes, while the negative correlation of Verrucomicrobia with TAs suggests high sensitivity to metal toxicity despite their role in nutrient cycling. These patterns highlight how nutrient dynamics and metal contamination jointly influence microbial community assembly through selective pressures and stress adaptation mechanisms. It is important to acknowledge that because As concentrations frequently co-vary with soil nutrient dynamics in such industrial legacy sites, the observed shifts likely reflect the integrated response of the soil microenvironment rather than the isolated toxicity of As.
Microflora metabolic pathway features of contaminated soils from a brick kiln
-
The potential ecological functions of the bacterial communities in contaminated soils from a brick kiln were examined by analyzing the differences in the pathway levels among the bacterial communities in the clean soil and contaminated soils[12]. Six level-one metabolic pathways of bacterial communities were obtained from the KEGG database (Fig. 3d). With increases in the As concentration, the relative abundances of the metabolism (64.44%–63.58%–62.51%) and genetic information processing (11.54%–11.21%–11.02%) pathways decreased gradually. Moreover, the relative abundances of environmental information processing (7.53%–7.86%–8.31%), cellular processes (6.65%–6.90%–7.50%), human diseases (6.57%–7.11%–7.28%), and organic systems (3.28%–3.35%–3.38%) pathways increased gradually.
Next, 23 level-two metabolic pathways with relative abundances higher than 1% were analyzed (Fig. 3e). Specifically, the relative abundances of level-two metabolic pathways (carbohydrate metabolism: 15.26%–14.46%; amino acid metabolism: 13.18%–12.78%, and energy metabolism: 7.54%–7.32%) and level-two metabolic pathways of genetic information processing (translation: 4.89%–4.52%) were significantly decreased (p < 0.05). On the contrary, the relative abundances of level-two metabolic pathways (signal transduction: 3.78%–4.38%) of environmental information processing and level-two metabolic pathways (cell motility: 1.31%–1.97%) of cellular processes were significantly increased (p < 0.05). The relative abundances of most level-two metabolic pathways exhibited no significant changes (p > 0.05).
Additionally, analysis of the top 30 KEGG level-three metabolic pathways in terms of relative abundance (Fig. 3f) demonstrated that with increases in the As concentration, the relative abundances of level-three metabolic pathways (ABC transporters, two-component system, and bacterial secretory system) of environmental information processing increased gradually. In contrast, the relative abundances of level-three metabolic pathways (ribosome) of genetic information processing and level-three metabolic pathways (fatty acid biosynthesis, citrate cycle, and carbon fixation pathways in prokaryotes) of metabolism decreased gradually.
Generally speaking, As contamination inhibits the metabolism and genetic information processing of bacterial communities, resulting in changes to the metabolic pathways of indigenous bacterial communities. Bacterial communities adapt to the environment by adjusting the relative abundances of metabolic pathways at different levels[44]. The increase in As concentration affects pathways such as energy, carbon, nitrogen, and sulfur metabolism[45,46]. In this study, significant decreases were observed in carbohydrate metabolism, amino acid metabolism, and energy metabolism (p < 0.05), which is consistent with findings from previous research. Furthermore, a significant rise in the relative abundance of signal transduction and cell motility was observed, suggesting that microbial communities may respond to As stress by augmenting these mechanisms.
The stress caused by pollutants may cause changes in the structures and metabolic pathways of indigenous microflora, thus influencing the abundance of functional genes[47]. Specifically, pollutants mainly influence the nutrient metabolism function of microflora. With increases in As concentration, the relative abundances of methane metabolism and carbon sequestration gradually reduced, while the relative abundances of nitrogen metabolism and sulfur metabolism gradually increased (Fig. 4a). The correlations between functional genetic groups (carbon metabolism, nitrogen metabolism, sulfur metabolism, xenobiotics biodegradation and metabolism, HMRGs, and As detoxification) in soils with different pollution gradients, environmental factors (OM, SOC, NO3––N, NH4+–N, TN, TP, TK, As, As[V], As[III]), and microflora (Fig. 4b) were analyzed by the Spearman correlation coefficient. While formal multiple testing corrections were not applied, the potential for false discoveries was mitigated by the use of a highly stringent significance threshold (r > 0.8 and p < 0.01) to ensure the identification of only the most robust ecological associations. Methane metabolism and carbon sequestration were significantly positively correlated with the Shannon index (p < 0.05). This might be because, with increases in the degree of pollution, the bacteria with methane metabolism and carbon sequestration functions decreased or disappeared. Heavy metal detoxification (p < 0.05), nitrogen metabolism (p < 0.01), and sulfur metabolism (p < 0.001) were positively correlated with Proteobacteria. This indicates that the increasing abundance of Proteobacteria plays an important role in detoxification, as well as nitrogen and sulfur metabolism, in contaminated soils[46].
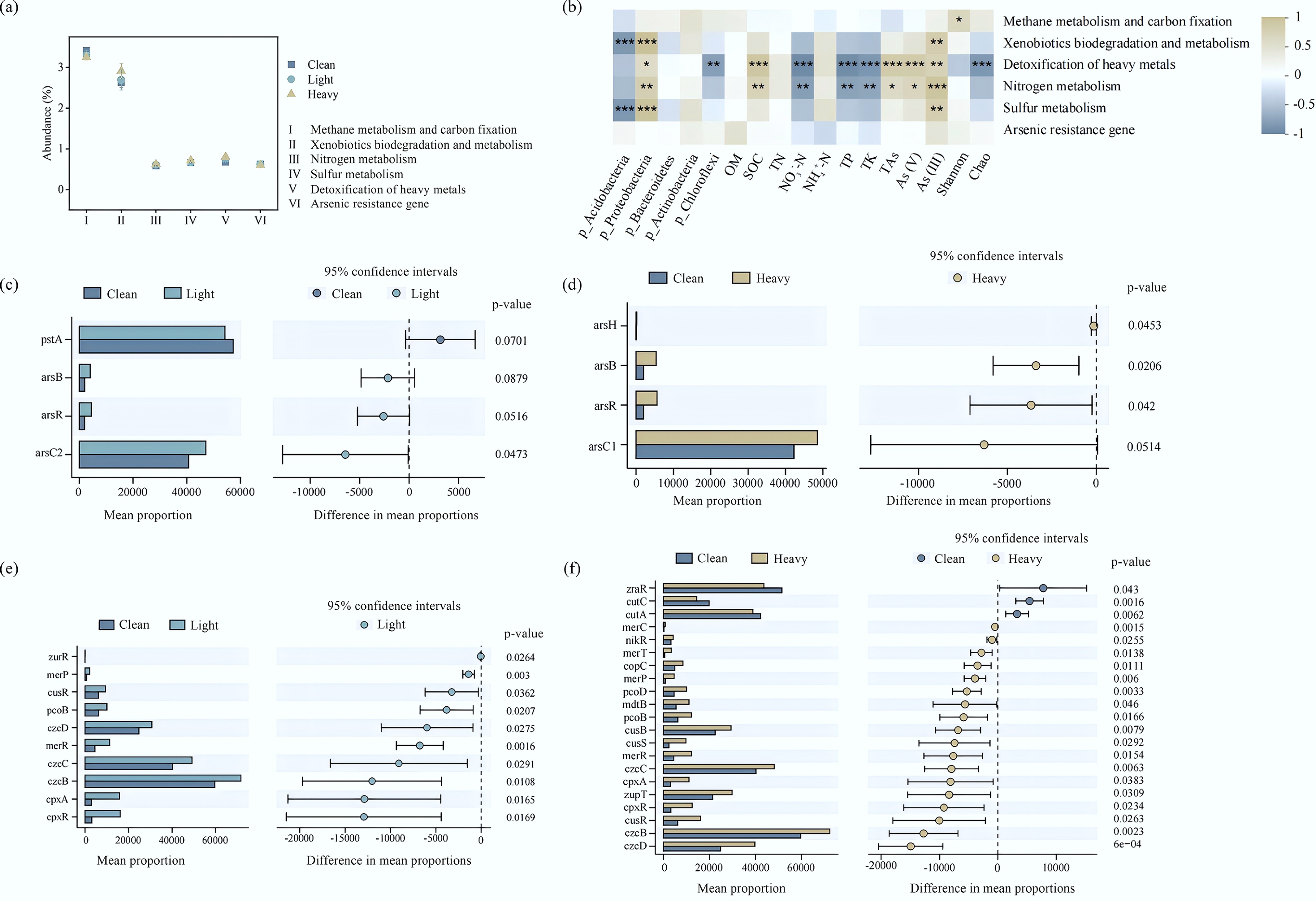
Figure 4.
(a) Changes in the relative abundances of carbon metabolism, nitrogen metabolism, sulfur metabolism, xenobiotics biodegradation and metabolism, and HMRGs of bacterial communities. (b) Correlations between functional gene clusters (carbon metabolism, nitrogen metabolism, sulfur metabolism, xenobiotics biodegradation and metabolism, HMRGs and As detoxification), and environmental factors of bacterial communities. (c) Stamp analysis of As resistance genes (Clean and Light). (d) Stamp analysis of As resistance genes (Clean and Heavy). (e) Stamp analysis of HMRGs (Clean and Light). (f) Stamp analysis of HMRGs (Clean and Heavy).
Indigenous bacteria cope with different degrees of heavy metal stress by upregulating the expression of HMRGs[12]. It can be seen from Fig. 4c that the abundance of As-resistance genes differed significantly between the clean and contaminated soils (Light and Heavy). Compared to the Clean group, the Light group exhibited upregulated abundances of arsB (p > 0.05), arsC2 (p < 0.05), and arsR (p > 0.05). The Heavy group exhibited significantly upregulated abundances of arsB, arsR, and arsH (p < 0.05). Transcription inhibits the regulatory effects of arsR on the expression of arsBC, while arsC2 reduces As(V) into As(III), which is then discharged out of cells through arsB. Additionally, arsH is able to oxidize MAs(III) into MAs(V) with lower toxicity[15,23,48]. Hence, although the concentration variation in As(III) was small (1.19–3.87 mg/kg), it was still far lower than the concentration changes (1.08–47.78 mg/kg) in As(V) (Fig. 3). The upregulated relative abundances of arsC2, arsB, and arsR demonstrate that indigenous bacteria achieve As detoxification through the reduction of As(V) and excretion of As(III). This might be the reason for the increasing As(III) concentration in contaminated soils.
Environmental implications
-
Arsenic is a toxic metalloid that poses a potential threat to the environment and human health. Direct evidence of the effects of As pollution on soil microbial communities can be provided by examining the structure and diversity of indigenous bacterial communities in As-contaminated soils from a shutdown brick kiln. The relative abundance of Acidobacteria gradually decreased with increasing As concentration. The observed decreasing trend in the relative abundance of Acidobacteria with increasing As concentration suggests that this phylum may serve as a potential indicator of As stress in soil ecosystems. However, further targeted studies are needed to verify its indicator reliability under varying environmental conditions. Exploring the metabolic pathways and detoxification potential of bacterial communities in soils with different As pollution gradients uncovered the response mechanisms of bacterial communities to As pollution stress. As affected the nutrient metabolism of indigenous bacteria, which was detrimental to the growth and reproduction of microbial communities[16,45]. Although the current findings are based on predictive bioinformatics tools (PICRUSt2) using 16S rRNA data, this approach has been widely validated for inferring functional potentials. In this study, the mean NSTI values were below 0.15, indicating high prediction accuracy for soil samples. However, we acknowledge the inherent uncertainty in functional inference compared to shotgun metagenomics. The predictive reliability is further supported by consistent changes in microbial community composition, network structure, and enrichment patterns of known detoxification-related taxa and genes. These multi-layered correlations lend credibility to the functional predictions despite the absence of direct biochemical validation. Bacterial communities detoxified by upregulating As resistance genes, which reduced their own intoxication[15,48]. Investigating the mechanisms regulating the expression of these As resistance genes can help develop more effective bioremediation technologies to mitigate the potential threat of As pollution to the environment and human health. Future studies will incorporate chemical quantification, gene cloning, and functional assays to directly verify the roles of genes such as arsC2 and arsB in As detoxification pathways.
-
In this study, the structural and functional changes and the diversity of indigenous microflora in heavy metal-contaminated soils from a typical brick kiln were comprehensively analyzed by combining high-throughput sequencing with an analysis of the soil physicochemical properties. With increases in the As concentration, the dominant microorganism in the bacterial communities changed from Acidobacteria to Proteobacteria. According to the analysis of the metabolism pathways, bacterial communities cope with heavy metal stress by increasing signal transduction and cell motility, and carbon metabolism-related functions decline. In polluted soils, the upregulated abundances of arsC2 and arsB indicate that the increased concentration of As(III) in contaminated soils might be related to the reduction of As(V) and the excretion of As(III). In summary, this study provides scientific insights into the ecological risks of legacy brick kiln sites by characterizing bacterial community changes along an As contamination gradient. The integration of co-occurrence network analysis with functional prediction (PICRUSt2) reveals shifts in detoxification potential and metabolic function, which offers a preliminary framework for understanding microbially mediated As attenuation in historically contaminated soils.
-
It accompanies this paper at: https://doi.org/10.48130/aee-0026-0009.
-
The authors confirm contribution to the paper as follows: Zhenchang Zhang: study conception and design, data collection, analysis and interpretation of results; Shuyue Liu: data collection, draft manuscript preparation; Hengxiang Cao: data collection; Qi Liu: data collection; Yiwu Wang: analysis and interpretation of results, draft manuscript preparation; Ningsha Feng: data collection; Mao Ye: study conception and design. All authors reviewed the results and approved the final version of the manuscript.
-
The data that support the findings of this study are openly available in the NCBI Short Read Archive (SRA) repository under accession number PRJNA1049149.
-
This work was financially supported by the National Natural Science Foundation of China (Grant No. 42477131), and the State Key Laboratory of Soil and Sustainable Agriculture (SKLSSA2508), Carbon Peak & Carbon Neutral Science and Technology Innovation Project of Jiangsu Province (BT2025015) .
-
All authors declare that they have no competing interests.
-
# Authors contributed equally: Zhenchang Zhang, Shuyue Liu
Full list of author information is available at the end of the article. - The supplementary files can be downloaded from here.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhang Z, Liu S, Cao H, Liu Q, Wang Y, et al. 2026. Characteristics and metabolic potentials of bacterial communities in arsenic-contaminated soils from a typical brick kiln in China. Agricultural Ecology and Environment 2: e011 doi: 10.48130/aee-0026-0009
Characteristics and metabolic potentials of bacterial communities in arsenic-contaminated soils from a typical brick kiln in China
- Received: 16 January 2026
- Revised: 21 March 2026
- Accepted: 10 April 2026
- Published online: 21 April 2026
Abstract: Production activities in brick kilns lead to arsenic (As) enrichment in surrounding soils. This poses serious threats to the ecological safety of the soil and human health. However, the metabolic ability and detoxification genes in As-contaminated soils in brick kilns remain to be fully elucidated. In this study, the microflora characteristics and metabolic potentials of soils with different degrees of As contamination obtained from typical brick kilns in Anhui, China were examined through high-throughput sequencing. The Ace index and Chao index exhibited decreasing trends, indicating that the abundance of the bacterial communities was impacted by pollutants (p < 0.001). The tightness of the bacterial co-occurrence network increased from 0.118 to 0.126, and the degree of modularization decreased from 0.748 to 0.669. This indicates that interactions between bacterial communities intensified, and the stability of community interactions declined with increasing As concentrations. A total of 376 metabolic pathways were detected. The relative abundances of the signal transduction (Clean: 3.78%, Light: 4.01%, Heavy: 4.38%) and cell motility (Clean: 1.31%, Light: 1.49%, Heavy: 1.97%) pathways were significantly increased (p < 0.05). Further, the abundances of the arsenate reduction gene (arsC2) and arsenite efflux penetrase gene (arsB) in contaminated soils were upregulated, indicating that the increase in the As (III) concentration of contaminated soils might involve the reduction of As(V) and the excretion of As(III). These results provide a basis for the further exploration of microflora response features and the formulation of effective microbial restoration strategies for As-contaminated soils in typical shut-down brick kilns. In conclusion, arsenic contamination significantly alters the diversity, network structure, and metabolic pathways of soil bacterial communities, driving an adaptive shift towards arsenic-tolerant taxa and the upregulation of specific detoxification genes. These findings provide a basis for the formulation of effective microbial restoration strategies for As-contaminated soils in typical shut-down brick kilns.
-
Key words:
- Brick kiln /
- Arsenic /
- Bacterial community /
- Metabolic potential