-
Jujube (Ziziphus jujuba Mill.), as the most important economic fruit tree species in the Rhamnaceae family, is renowned for its exceptionally high concentration of AsA, which can reach up to 1,000 mg·100 g−1 fresh weight (FW), significantly surpassing that of other fruits such as kiwifruit (Actinidia chinensis)[1,2]. With the growing trend toward healthy lifestyles, jujube has garnered increasing attention. Previous research on ascorbic acid (AsA) synthesis in jujubes has elucidated two primary pathways: the L-galactose pathway and the myo-inositol pathway[3,4]. However, the identification and characterization of the key genes and their regulatory mechanisms that influence AsA accumulation remain limited.
The L-galactose pathway is the primary route for AsA biosynthesis and has been demonstrated in various species[5−8]. GDP-D-mannose-3',5'-epimerase (GME) catalyzes a pivotal step in this pathway[9,10]. However, its role is not straightforward and exhibits considerable complexity, as evidenced by functional divergence among its homologs across species. For instance, in persimmon (Diospyros spp.), a GME gene correlates with fruit AsA[8], whereas in sweet orange (Citrus × sinensis), it functions in the leaves but not the fruits[11]. Further complexity is revealed in tomato (Solanum lycopersicum), where the expression of SlGMEs is upregulated under stress to enhance ascorbate accumulation and tolerance to oxidative stress, cold, and salt, whereas the individual homologs SlGME1 and SlGME2 also exhibit functional specialization through distinct roles in cell wall biosynthesis[12,13]. This context-dependent functionality underscores the necessity of characterizing GME within specific biological systems, particularly in species with unique AsA accumulation patterns like jujube.
Beyond the enzymatic genes themselves, transcription factors (TFs) have emerged as key regulators of AsA biosynthesis that are capable of orchestrating the expression of multiple pathway genes. Initial clues from large-scale studies, such as comparative transcriptomic and metabolomic analyses of the development of Rosa roxburghii fruit, suggest that a complex network of TFs (e.g., 168 TFs were identified) may co-regulate AsA synthesis[14]. Specific mechanisms are also being uncovered. In kiwifruit, the R1-MYB transcription factor AceMYBS1 binds to the promoter of AceGGP3 to activate its expression and enhance AsA biosynthesis[2,15]. However, in tomato, the HD-Zip I family TF SlHZ24 transcriptionally activates multiple structural genes, including SlGMP3, SlGME2, and SlGGP, thereby coordinately promoting AsA accumulation[16]. In addition to these families, members of the nuclear factor Y (NF-Y) family, known for their roles in diverse processes including stress responses and metabolism[17,18], have also been implicated. For instance, an NF-YA subunit (SlNFYA10) acts as a repressor of GME expression and AsA accumulation in tomato[19]. In contrast, the role of the NF-YC subunit in ascorbate biosynthesis remains completely unexplored. In particular, although kiwifruit and tomato have mapped regulators for moderate AsA levels[2,15,16], jujube—whose content is more than twofold higher—lacks any known upstream TF for GME paralogs; identifying which paralog drives the ultra-high AsA content and its transcriptional regulator is the gap we close here.
Here, we present a functional characterization of the GME gene family[20] in jujube. Phylogenetic and expression analyses of the two identified GME homologs pinpointed ZjGME1 as a putative key player in AsA biosynthesis. We confirmed its molecular function through subcellular localization and transient overexpression assays. Furthermore, via a yeast one-hybrid (Y1H) screen, we identified a transcription factor, ZjNF-YC9, that binds to the ZjGME1 promoter. Subsequent β-glucuronidase (GUS) staining and overexpression analyses demonstrated that ZjNF-YC9 activates ZjGME1 transcription, thereby promoting AsA accumulation.
-
Jujube fruits were obtained from the Jujube Germplasm Nursery of Shandong Fruit Tree Research Institute (36.15° N, 117.07° E). Fruits at distinct developmental stages were collected, encompassing young fruits (YF, 10 DAA), enlargement stage fruits (EN, 40 DAA), white ripe fruits (WR, 70 DAA), early red fruits (BR, 85 DAA), half red stage fruits (HR, 95 DAA), and fully red stage fruits (FR, 105 DAA). After harvesting, the fruits were cut into pieces, mixed thoroughly, rapidly frozen in liquid nitrogen and stored at −80 °C in a laboratory freezer.
Determination of AsA content
-
AsA was extracted and determined essentially as described in Zhang et al.[4]. Each fruit sample, weighing 1 g, was placed in a precooled mortar and extracted with 0.1% H3PO4. The resulting supernatant is obtained through centrifugation and diluted with an appropriate volume of a sufficient amount (up to 10 mL). The content of AsA was determined using the Shimadzu Nexera high-performance liquid chromatography (HPLC) system LC-30A, with the Nexera SPD-M30A photodiode array detector, and the Shodex KC-811 6E ion-exclusion column (6 × 250 mm). The mobile phase consisted of 2 mM HClO4 with a flow rate of 0.5 mL·min−1 at a column temperature of 50 °C. A sample injection volume of 20 µL was applied, and the detection wavelength for AsA was set at 243 nm. Each sample underwent analysis in triplicate to ensure biological robustness.
Vector construction and transient expression in jujube
-
The ZjGME1 coding sequence (CDS) was cloned into both the IL-60 and pCaMV2300-GFP vectors to generate overexpression constructs. Agrobacterium suspensions (optical density [OD600] = 0.6) were injected into the fruit peel (100 μL per fruit) as described by Zhang et al.[21]. Gene expression analysis and determination of AsA content were conducted 3–4 d after fruit injection.
Reverse transcription–quantitative polymerase chain reaction analysis
-
Reverse transcription–quantitative polymerase chain reaction (qRT-PCR) was used to analyze ZjGME1 expression levels during different stages of jujube's development, as well as in overexpressed fruit samples. The Plant Total RNA Isolation Kit Plus (Foregene) was used for total RNA extraction from each sample, followed by cDNA reverse transcription using All-in-One First-Strand Synthesis MasterMix (BestEnzymes Biotech). Fluorescence quantification experiments were performed using Taq SYBR Green qPCR Premix (BestEnzymes Biotech) on a fluorescent quantitative polymerase chain reaction (PCR) instrument (CFX 96 Bio-rad). Each 20-μL reaction mixture contained 10 μL of Taq SYBR Green qPCR Premix, 0.4 μM of each primer, and 2 μL of 1:10 diluted cDNA, with the remainder supplemented by nuclease-free water. The thermal cycling conditions were as follows: Initial denaturation at 95 °C for 30 s, followed by 40 cycles of denaturation at 95 °C for 10 s and annealing/extension at 60 °C for 30 s, and a final melting curve stage. The UBQ gene served as the internal reference[22], which demonstrated its stable expression across jujube fruits' developmental stages. The relative expression levels of each gene in each sample were determined with three biological replicates. The primers used are listed in Supplementary Table S1.
Subcellular localization of the proteins
-
A fusion expression plasmid was constructed by cloning the coding sequence of ZjGME1 downstream of the CaMV 35S promoter in the pCaMV2300-GFP vector, following the method of Liu et al.[23]. The resulting construct, 2300-35S-ZjGME1-GFP, was then introduced into the GV3101 strain of Agrobacterium tumefaciens. For transient expression, the transformed agrobacteria harboring this construct were co-infiltrated with a strain containing pRT101-AtCBL3–red fluorescent protein (RFP) into the leaves of Nicotiana tabacum. After infiltration, the plants were kept in the dark for 12 hours and subsequently transferred to a growth chamber under normal conditions for 48 h. Fluorescence was observed using a high-resolution confocal laser scanning microscope (ZEISS LSM-880).
Y1H assay
-
For the Y1H assay, the cis-acting elements in the promoter region of the ZjGME1 gene were predicted using PlantCARE, a database of plant promoters and cis-acting regulatory elements. Subsequently, the potential fragments were cloned into the pAbAi vector. The recombinant vectors were then transformed into GOLD1 yeast cells and cultured in SD/-Ura medium. To determine the minimum inhibitory concentration, the bait strains were cultured in SD/-Ura medium supplemented with various concentrations of aureobasidin A (ABA). Furthermore, yeast strains containing the pBait-AbAi vector were transformed with cDNA AD library vectors previously constructed and cultured in SD/-Leu/+ABA medium. Positive AD vectors underwent sequencing to identify binding transcripts. Finally, one-on-one bait-bait interactions were conducted to validate the effectiveness of positive recombinant bait vectors.
GUS staining assay
-
We integrated the synthesized ZjGME1 promoter fragment acquired during the preceding step into the P1300-GUS vector and fused the chosen transcription factor from the Y1H to the pGreenII 62SK vector. The treatment group was composed of a combination of P1300-proGME-GUS and pGreenII62SK–target gene at a ratio of 1:1, whereas the control group consisted of a combination of P1300-proGME-GUS and the empty pGreenII62SK vector. The aforementioned liquid was injected into both sides of a tobacco leaf. To provide an internal reference, the right half of each leaf was infiltrated with the treatment mixture, whereas the left half received the control mixture. All infiltrations were carried out on the same fully expanded leaf of 4-week-old N. benthamiana grown at 25 °C, 70% relative humidity, and 16 h light/8 h dark. GUS staining was performed after sampling 3 d later using both leaf disc and whole-leaf staining methods. Information on the plasmid vectors is shown in Supplementary Table S2.
-
Two genes encoding GME were identified within the jujube genome[3,24,25], located on chromosomes 4 and 5, and were designated as ZjGME1 and ZjGME2, respectively. To investigate their evolutionary relationships, a phylogenetic tree was constructed with GME protein sequences from diverse plant species (Fig. 1a). The phylogenetic analysis revealed that the GME proteins are primarily divided into two major clades. ZjGME1 clustered within one clade alongside orthologs such as CsGME from citrus and MnGME1 from mulberry (Morus notabilis), whereas ZjGME2 grouped into a distinct clade with another mulberry member. The GMEs from Arabidopsis thaliana (AtGME) and S. lycopersicum (SIGME) were positioned at the base of the tree. To further explore the origin of these two jujube genes, a collinearity analysis of the jujube genome was performed. The results demonstrated a significant collinear relationship between the chromosomal region harboring ZjGME1 and the region containing ZjGME2 (Fig. 1b), strongly suggesting that ZjGME1 and ZjGME2 likely originated from a jujube-specific segmental duplication event.
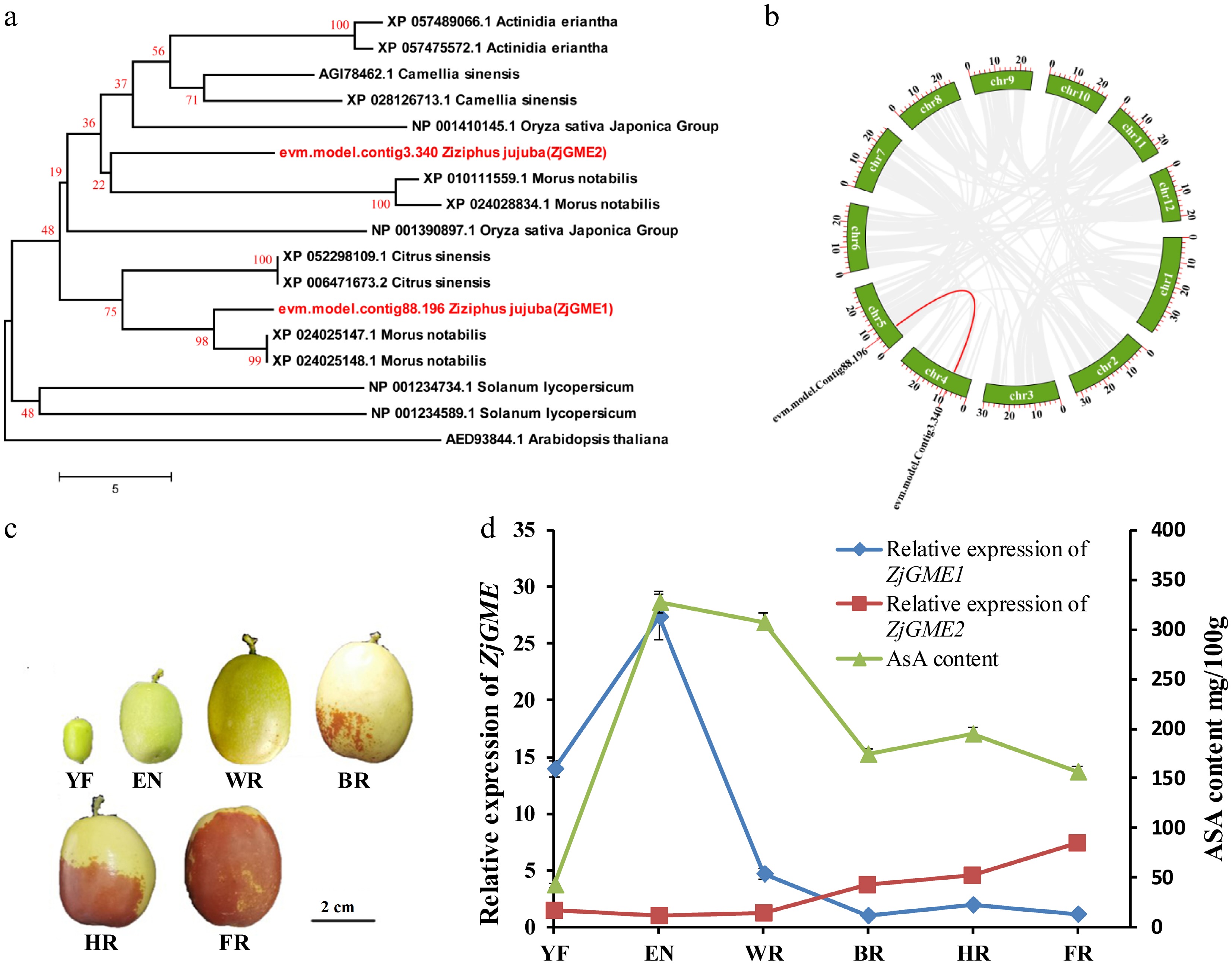
Figure 1.
Phylogenetic evolution and expression of GME genes in jujube. (a) Phylogenetic analysis of ZjGMEs and homologous genes from Morus notabilis, Citrus sinesis, Oryza sativa, Camellia sinensis, Actinidia eriantha, Solanum lycopersicum, and Arabidopsis thaliana. (b) Collinearity analysis of the two jujube GMEs within the jujube genome. (c) Phenotypic characteristics of the jujube fruits during different development stages. Scale bar = 2 cm. (d) ZjGMEs (ZjGME1 and ZjGME2) mRNA abundance and AsA content at six fruit developmental stages. YF: young stage; EN: enlargement stage; WR: white mature stage; BR: early red stage; HR: half red stage; FR: full red stage.
To investigate the genetic regulation of AsA accumulation, we analyzed the expression patterns of the two GME genes alongside AsA levels during fruit development. As shown in Fig. 1c, d, the transcript level of ZjGME1 was relatively high in young fruit, increased significantly to a peak during the enlargement stage (a 1.96-fold increase), and subsequently declined upon ripening. The overall expression trend of ZjGME1 was broadly consistent with the dynamic pattern of AsA accumulation throughout these developmental stages. In contrast, the expression of ZjGME2 exhibited nearly the opposite pattern, showing a steady increase as the fruit matured. This suggests a potential role for ZjGME1 in AsA biosynthesis during fruit development.
ZjGME1 overexpression enhances AsA accumulation and subcellular localization analysis
-
To further elucidate the role of ZjGME1 in AsA biosynthesis in jujube, transient overexpression assays were performed using 'Jinsi 4' and 'Dongzao' fruits. For 'Jinsi 4', the IL60-ZjGME1 vector was constructed and injected, with the empty IL60-2 vector serving as the control. After 4 d, qPCR and HPLC analysis revealed that ZjGME1 expression was upregulated 2.0-fold in the overexpression group, accompanied by an 11% increase in AsA content (453.87 vs 407.83 mg·100 g−1 FW; Fig. 2a, b). In a parallel experiment, a 35S::ZjGME1 construct was introduced into 'Jinsi 4' fruits via Agrobacterium-mediated transformation, with the empty GFP vector as the control. After 4 d, ZjGME1 expression increased 2.79-fold, and AsA content rose from 339.57 to 355.26 mg·100 g−1 FW (Fig. 2a, b). Furthermore, in fresh on-tree 'Dongzao' jujube fruits injected with 35S::ZjGME1, a 2.57-fold increase in ZjGME1 expression was observed, along with a 56.8% elevation in AsA content (127.16 vs 81.11 mg·100 g−1 FW; Supplementary Fig. S1). Collectively, these results clearly demonstrate that ZjGME1 overexpression enhances AsA accumulation in jujube fruits.
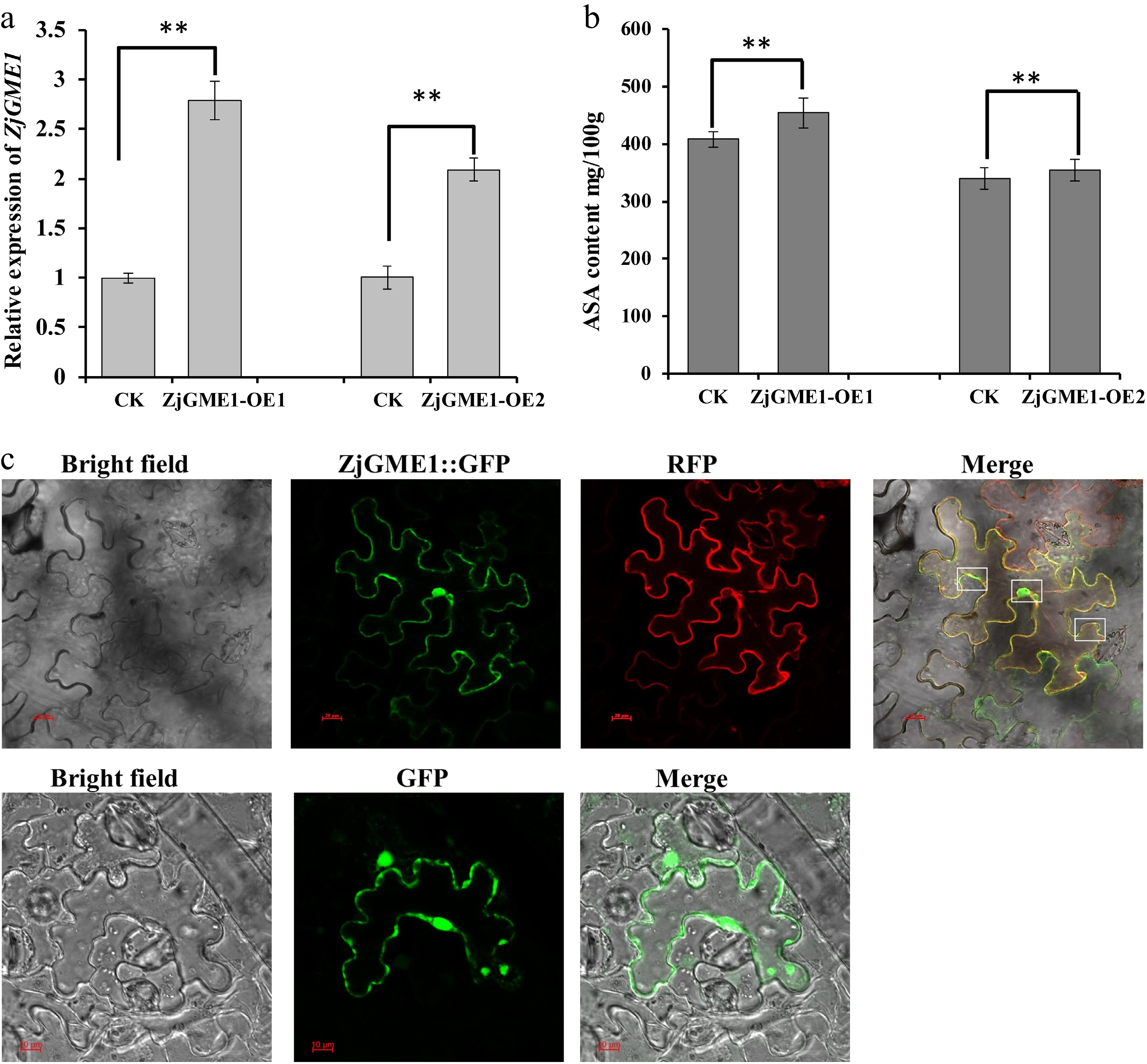
Figure 2.
Transient overexpression of ZjGME1. (a) Overexpressed ZjGME1 improved AsA content. (b) ZjGMEOE1 represents the IL60-ZjGME vector; ZjGMEOE2 represents the 35S::ZjGME1 vector. (c) Subcellular colocalization of the ZjGME1-GFP fusion protein with a AtCBL3 tonoplast marker. Boxes highlight the position of green fluorescence localized partially in the nucleus and cytoplasmic (outside the red vacuole member). Values are presented as the mean ± standard deviation (SD) on the basis of three repetitions. ** p < 0.01.
For subcellular localization, a ZjGME1::eGFP fusion construct was co-expressed with a vacuolar membrane marker (RFP) in Nicotiana benthamiana leaves via Agrobacterium infiltration. Confocal imaging (Fig. 2c, Supplementary Fig. S2) revealed that ZjGME1-eGFP was localized partially in the nucleus, as indicated by green fluorescence overlapping with the nuclear regions. Additionally, co-localization with the RFP-labeled tonoplast marker indicated cytoplasmic distribution. These results indicate that ZjGME1 localizes to both the nucleus and the cytoplasm.
Transcriptional regulation of ZjGME1
-
To investigate the transcriptional regulation of ZjGME1, a 3,000-bp promoter region upstream of the start codon was analyzed using PlantCARE. The analysis revealed that the CCAAT-box, MYB, and MYC binding sites were among of the most abundant cis-elements in this region (Supplementary Table S3). A fragment containing theses predicted cis-elements (Fig. 3a) was cloned into the pAbAi vector to generate a Y1H bait strain. After confirming the absence of self-activation, a cDNA library from jujube fruits was screened. Under AbA selection, yeast colonies showing positive growth were subjected to analysis with Matchmaker® Insert Check PCR Mix 2 (TaKaRa) and sequencing, leading to the identification of a putative transcription factor, ZjNF-YC9 (Zj.jz037921106, nuclear transcription factor Y [NF-Y] subunit C-9). NF-Y proteins are known to bind CCAAT-box elements[26], suggesting that ZjNF-YC9 may regulate ZjGME1 through this mechanism.
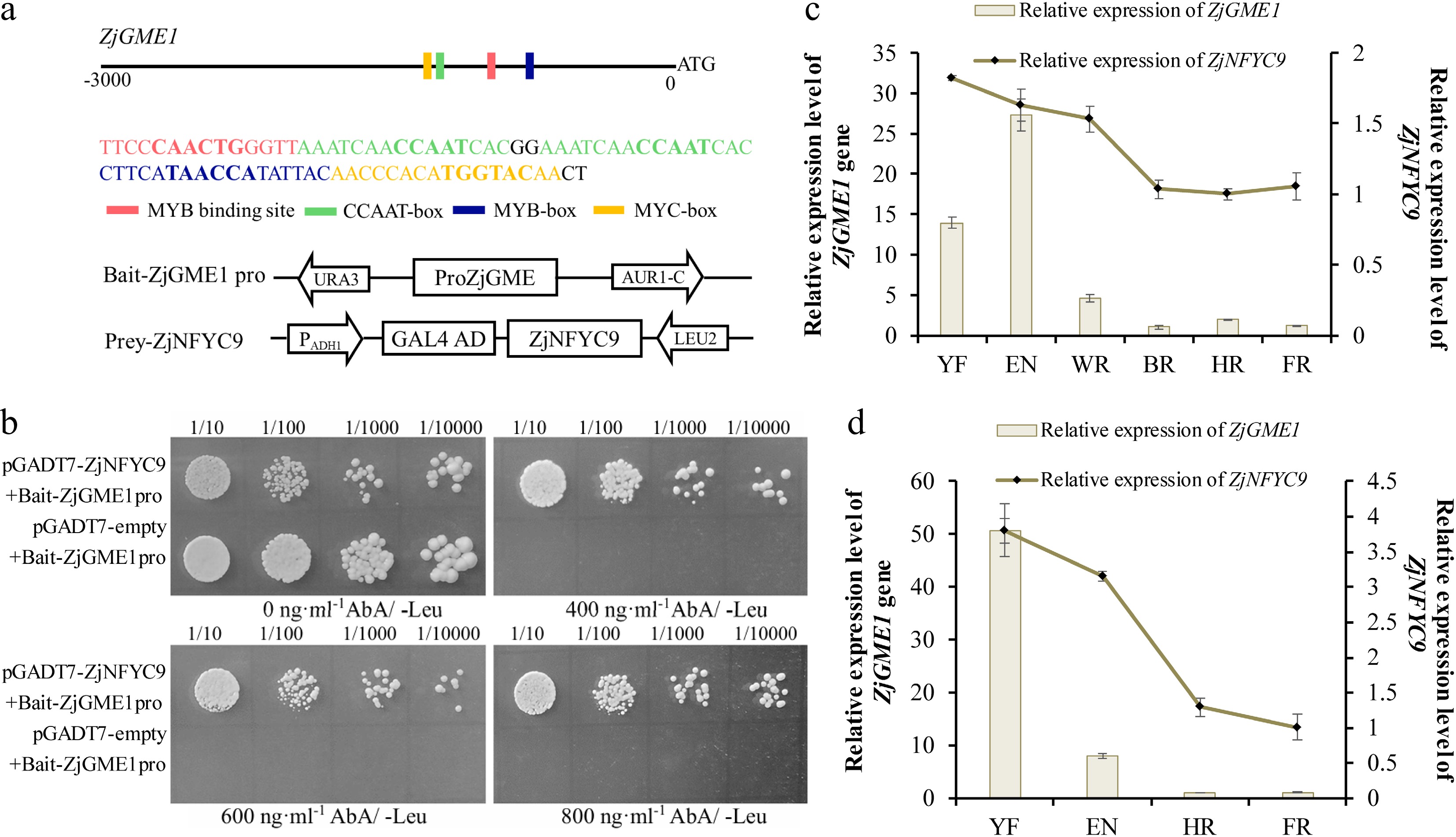
Figure 3.
Binding of ZjNF-YC9 to the promoter of ZjGME1 from jujube. (a) Schematic diagrams of the bait and prey vectors that were used for the Y1H analysis. (b) Y1H assay of ZjNF-YC9 binding to the ZjGME1 promoter fragments of jujube. Yeast cells grown on SD/-Leu medium with different ABA concentrations (0, 400, 600, or 800 ng·mL–1). (c) and (d) Abundance of ZjNF-YC9 mRNA at different fruit development stages. YF: young stage; EN: enlargement stage; WR: white mature stage; HR: half red stage, FR: full red stage.
For validation, a pGADT7-NFYC9 vector was constructed and separately transformed into the bait strains along with the pGADT7 plasmid for pairwise verification. Both strains grew normally on SD/-Leu medium without AbA. However, under AbA selection (400–800 ng·mL−1), only the strain carrying pGADT7-NFYC9 survived (Fig. 3b), confirming a specific interaction. To correlate ZjNF-YC9 expression with its potential regulatory role, we examined its transcript levels during different fruit development stages in the cultivated jujube 'No. 4 Jinsi' and the sour jujube 'No. 3 Suanzao'. ZjNF-YC9 transcript levels gradually decreased in both varieties, showing a strong positive correlation with ZjGME1 expression (r = 0.74 in cultivated jujube; r = 0.81 in sour jujube; Fig. 3c, d).
To further verify the binding of ZjNF-YC9 to the ZjGME1 promoter, we performed a transactivation assay in Nicotiana benthamiana using co-transformed pGreenII-62SK-ZjNFYC9 and P1300-proGME-GUS. GUS staining after 3 d revealed stronger blue staining in leaves expressing ZjNF-YC9 compared with the empty vector control (Fig. 4a). Quantitative analysis confirmed significantly higher GUS expression in the experimental group (Fig. 4b), supporting ZjNF-YC9-dependent transactivation. Finally, transient overexpression of ZjNF-YC9 in jujube fruit led to a 1.3-fold increase in ZjNF-YC9 expression, a concomitant 0.2-fold up-regulation of ZjGME1, and a 24% rise in AsA content relative to the control (Fig. 4c–e). These results demonstrate that ZjNF-YC9 positively regulates ZjGME1 expression and enhances AsA accumulation.
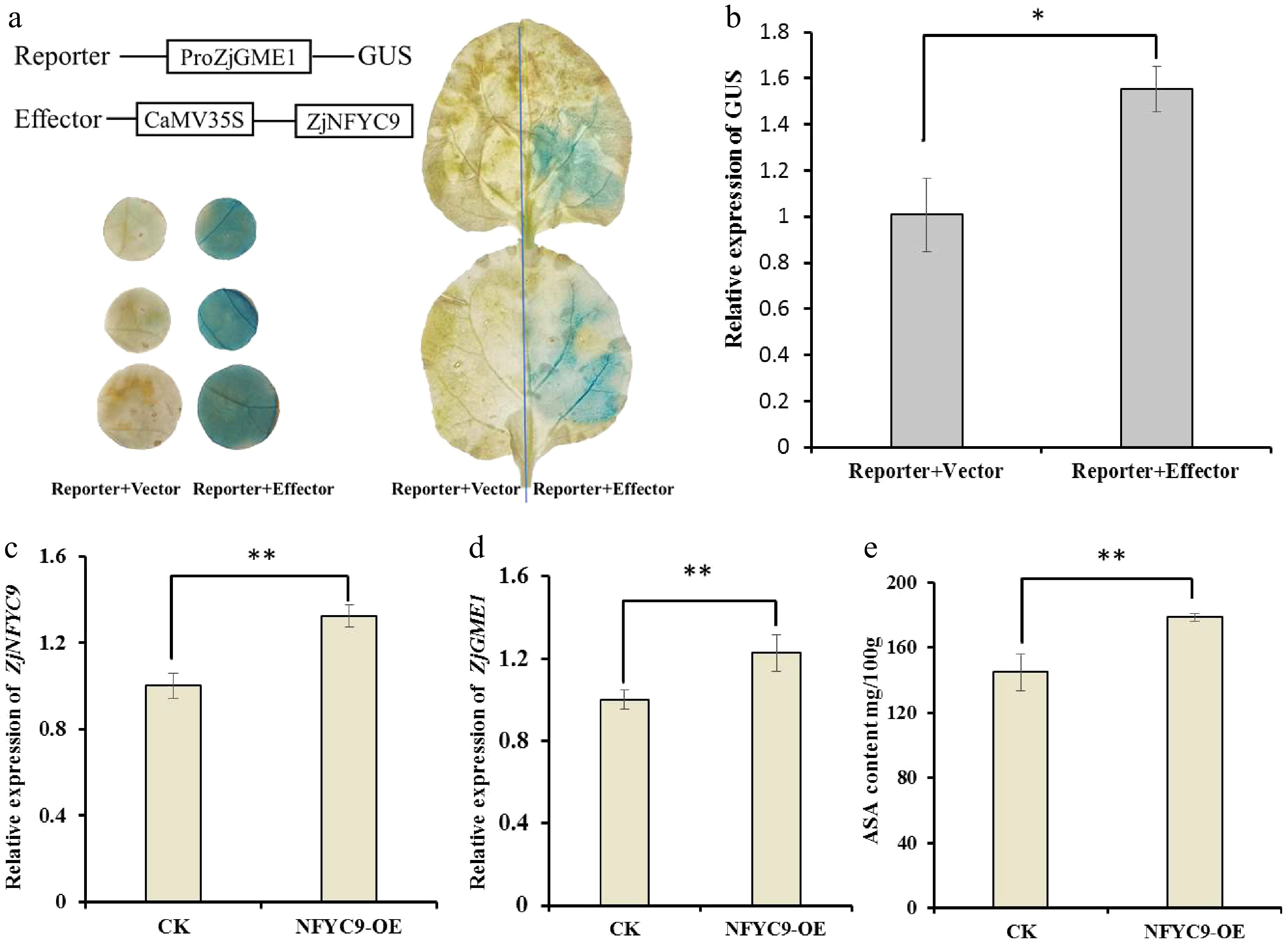
Figure 4.
ZjNF-YC9 activates ZjGME1 expression and promotes AsA accumulation. (a) Schematic of the effector and reporter constructs used in the GUS staining assay. (b) Relative expression analysis of the GUS reporter gene demonstrated that that ZjNF-YC9 significantly activates the promoter of ZjGME1. (c) ZjNF-YC9, (d)ZjGME1, and (e) AsA content around the injection sites in jujube fruits via ZjbNF-YC9 (NFYC9-OE) overexpression vector-based transformation. The empty vector (CK) was used as a control. Data are shown as the mean ± SD of three repetitions. Statistical significance was detected by Student's t-test. ** p < 0.01, * p < 0.05.
-
Gene duplication is a key driver of functional innovation in plant genomes. Our identification of two GME genes in jujube, originating from a segmental duplication event, provides a genetic basis for potential functional specialization. Although a single GME copy exists in Arabidopsis, species like tomato and citrus possess two homologs; notably, four members are found in the closely related mulberry[12,27]. Phylogenetic analysis revealed that ZjGME1 and ZjGME2 cluster distinctly with different mulberry GME genes, suggesting an ancient duplication event prior to the divergence of the jujube and mulberry lineages. More importantly, we observed a clear functional divergence between these paralogs. The expression pattern of ZjGME1, peaking during fruit expansion in parallel with AsA accumulation, strongly implicates it as the primary contributor to AsA biosynthesis[28,29]. This role was functionally confirmed by transient overexpression of ZjGME1, which significantly enhanced ascorbate accumulation in jujube fruits. In contrast, the steadily increasing expression of ZjGME2 during ripening suggests a divergent physiological role, potentially linked to maturation processes or stress responses, as seen in our previous study[1].
ZjNF-YC9: a novel transcriptional activator elucidating the upstream regulation of AsA biosynthesis
-
NF-Y is a ubiquitous class of TFs found in eukaryotes[26,30], known for their roles in regulating plants' growth, development, and stress responses[17,31]. NF-Y comprises three distinct subfamilies: NF-YA, NF-YB, and NF-YC[32]. Members of this family exhibit diverse functional activities. For instance, some NF-Y members are involved in responses to salt and drought stress across various plant species[33−35], whereas certain members play a pivotal role in regulating fruit quality. A key example of the latter is provided by Wang et al.[18], who discovered that NF-YB family members can form complexes with NF-YC and NF-YA to regulate flavonoid biosynthesis in tomatoes. Chen et al.[19] identified SlNFYA10 as a negative regulator of AsA accumulation by binding to the promoter region of the SlGME1 gene and downregulating its expression. However, a role for NF-YC subunits in the transcriptional regulation of AsA biosynthesis has remained unexplored until now.
In the present study, we identified ZjNF-YC9 as a direct transcriptional activator of ZjGME1 in jujube fruit. Through comprehensive approaches including Y1H screening, GUS assays, and transactivation experiments, we demonstrated that ZjNF-YC9 specifically binds to the ZjGME1 promoter and enhances its transcriptional activity. To our knowledge, this represents the first experimental evidence that an NF-YC family protein positively regulates GME expression and promotes AsA accumulation. This finding expands the functional diversity of the NF-YC subfamily, whose members are known to participate in a range of processes beyond abiotic stress and flavonoid synthesis, such as photomorphogenesis and seed development[36,37]. The contrasting regulatory modes between ZjNF-YC9 (as an activator) and SlNF-YA10 (as a repressor)[19] suggest subunit- and species-specific functions of NF-Y transcription factors in AsA metabolism. This functional divergence highlights that AsA-regulatory modules are not universally conserved but have evolved distinct mechanisms in different plant species. Furthermore, the role of such a module in shaping natural variation of a complex quantitative trait like AsA content is likely nuanced. In our germplasm, ZjNF-YC9 and ZjGME1 expression showed a correlation (Supplementary Fig. S3), yet no simple linear relationship with the final AsA content was observed across varieties. This underscores that although the ZjNF-YC9-ZjGME1 axis is a key regulatory determinant, its contribution to the phenotype is integrated within a broader metabolic and genetic network, which is a common feature in the regulation of specialized plant metabolites[38]. An open question arising from our work is whether ZjNF-YC9 exerts its transcriptional activation independently or as part of a canonical NF-Y heterotrimeric complex with its NF-YA and NF-YB partners. Future studies using protein–protein interaction assays could elucidate this mechanism and provide further directions for understanding the regulatory network governing AsA accumulation in jujube.
-
Our study provides new insights into the molecular basis of the high AsA accumulation in jujube fruit. We demonstrate that the two GME paralogs in jujube have undergone functional divergence, with ZjGME1 emerging as the key regulator of AsA biosynthesis during fruit development. Significantly, we identified ZjNF-YC9 as a novel transcriptional activator that directly targets the ZjGME1 promoter, establishing a previously unrecognized ZjNF-YC9-ZjGME1 regulatory module. These findings substantially advance our understanding of the complex regulatory network controlling AsA accumulation in fruit and open new avenues for improving nutritional quality through targeted metabolic engineering, for instance, by modulating the expression of ZjNF-YC9 or ZjGME1 via gene editing to breed jujube varieties with enhanced AsA content.
-
The authors confirm contribution to the paper as follows: study conception and design: Bi J, Zhang C; data collection: Bi J, Xing Y; analysis and interpretation of results: Xing Y, Mao X, Zhao X; draft manuscript preparation: Bi J, Zhang C. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article.
-
We are grateful for the assistance provided by Professor Qiong Zhang and Associate Professor Zhongtang Wang from the Shandong Institute of Pomology during sample collection. This work was founded by the Key R&D Program of Shandong Province, China (2023LZGC016), the National Natural Science Foundation of Shandong Province (ZR2023MC028), and the Chinese National Natural Science Foundation (Grant No. 32371923).
-
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
-
accompanies this paper online at: https://doi.org/10.48130/frures-0026-0003.
-
# Authors contributed equally: Jingxin Bi, Yesheng Xing
- Supplementary Table S1 Sequence of primers used in this study (5'- 3').
- Supplementary Table S2 Information of plasmid vectors used in this study.
- Supplementary Table S3 Positions and sequences of zjGME1 promoter elements.
- Supplementary Fig. S1 Overexpress ZjGME1 improved AsA content in 'Dongzao' fruit.
- Supplementary Fig. S2 Subcellular colocalization of the ZjGME1-GFP fusion protein with DAPI.
- Supplementary Fig. S3 Correlation between the expression levels of ZjNF-YC9 and ZjGME1.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Bi J, Xing Y, Mao X, Zhao X, Zhang C. 2026. Transcription factor ZjNF-YC9 regulates ZjGME1 to promote ascorbic acid accumulation in jujube fruits. Fruit Research 6: e013 doi: 10.48130/frures-0026-0003
Transcription factor ZjNF-YC9 regulates ZjGME1 to promote ascorbic acid accumulation in jujube fruits
- Received: 07 November 2025
- Revised: 09 February 2026
- Accepted: 14 February 2026
- Published online: 03 April 2026
Abstract: Jujube (Ziziphus jujuba Mill.) fruit is an exceptional source of ascorbic acid (AsA), but the transcriptional mechanisms governing its extreme accumulation are unclear. Although GDP-D-mannose-3',5'-epimerase (GME) is a key enzyme for AsA biosynthesis, its functional divergence and regulatory context in jujube are unknown. Here, we identified two GME paralogs, ZjGME1 and ZjGME2, in the jujube genome. Despite their common origin, expression profiling revealed that only ZjGME1 displayed a positive correlation with AsA content during fruit development, suggesting functional divergence. Transient overexpression of ZjGME1 confirmed its specific role in significantly enhancing AsA accumulation. Furthermore, to elucidate the upstream regulation, we identified a transcription factor, ZjNF-YC9, which directly binds to the ZjGME1 promoter and activates its expression to promote AsA accumulation. Our study thus uncovers a complete ZjNF-YC9-ZjGME1 transcriptional regulatory module, providing fundamental insights for the metabolic engineering of nutritional quality in fruits.
-
Key words:
- Jujube /
- L-ascorbic acid (AsA) /
- ZjGME1 /
- ZjNF-YC9 /
- Transcriptional regulation













