-
Floral scent compounds represent a crucial class of plant volatile secondary metabolites, primarily consisting of volatile organic compounds (VOCs) synthesized and released by floral organs[1]. These compounds play essential roles in plant–environment interactions, not only attracting pollinators but also defending against pathogens and herbivores, and facilitating adaptation to environmental stresses[2,3]. Owing to their low molecular weight and high vapor pressure, VOCs can permeate cellular membranes and diffuse into the extracellular matrix and surrounding environment, enabling rapid dissemination and release[4]. Floral VOCs are chemically diverse, primarily comprising terpenoids, phenylpropanoids/benzenoids, fatty acid derivatives, and nitrogen-containing compounds[5]. Based on biosynthetic pathway specificity, volatile metabolites are systematically classified into three major categories: terpenoids, phenylpropanoid/benzenoid compounds, and fatty acid derivatives[6]. The composition and abundance of these VOCs vary significantly among flowering plants, defining their distinctive scent profiles[7].
The biosynthesis of floral terpenoids in plants is initiated by the condensation of two C5 precursors, isopentenyl pyrophosphate (IPP) and dimethylallyl diphosphate (DMAPP)[8]. These central metabolic precursors are primarily synthesized via the mevalonic acid (MVA) pathway and the methylerythritol phosphate (MEP) pathway[9]. The MVA pathway, localized in the cytosol, is responsible for the production of volatile sesquiterpenes and triterpenes. In contrast, the MEP pathway operates in plastids and contributes to the formation of volatile monoterpenes and diterpenes, as well as carotenoids and chlorophylls[10−13]. Although these isoprenoid biosynthetic pathways are compartmentalized, they may be interconnected through metabolite transport mediated by specific transporters[14,15]. Such metabolic crosstalk enables the MEP pathway to support terpenoid biosynthesis in the cytosol[16]. The MVA pathway begins with the sequential condensation of three acetyl-CoA molecules and involves six enzymatic steps, whereas the MEP pathway starts from the condensation of D-glyceraldehyde 3-phosphate and pyruvate and requires seven enzymatic reactions[17,18]. The methylerythritol phosphate (MEP) pathway is central to the formation of volatile terpenoids and certain floral traits[19,20]. A key example is linalool, the most abundant monoterpene in tea, whose synthesis is governed by terpene synthase (TPS) enzymes within this pathway[21,22].
Lilium spp., a genus of significant ornamental value within the Liliaceae family, has attracted increasing research attention due to its floral fragrance. More than 100 wild species are known, distributed primarily across the Northern Hemisphere at latitudes between 10° and 60°, including regions of Asia, Europe, and North America[23]. The floral scent, a key aspect of lilies' ornamental appeal, is largely determined by monoterpenoids, with variation among cultivars attributed to differences in monoterpene emission levels[24]. Major terpenoid volatiles identified in lilies include (E)-β-ocimene, linalool, and myrcene[25]. Recent advances in molecular biology have greatly facilitated the study of fragrance biosynthesis in lilies. Terpene synthase genes, such as LhTPS, play essential roles in the production and emission of scent compounds and represent key genetic determinants underlying floral aroma in lilies[26].
In the Oriental lily (Lilium 'Sorbonne'), previous studies have shown that heterologous overexpression of the two monoterpene synthase genes, LiLIS and LiMYS, in Arabidopsis thaliana promotes monoterpene biosynthesis[27]. In addition, LiDXS2, which encodes the first key enzyme of the MEP pathway, has been implicated in the regulation of floral scent emission in 'Sorbonne'[28]. Despite these advances, the molecular mechanisms underlying floral fragrance synthesis in Lilium 'Sorbonne' remain largely elusive. Further investigation into the key genes and regulatory networks controlling scent production is essential to advance the understanding of floral aroma formation and facilitate lily breeding and biotechnological applications.
-
The Oriental lily (Lilium 'Sorbonne'), along with seeds of Nicotiana benthamiana, tobacco (Nicotiana tabacum L. '95'), and wild-type Arabidopsis thaliana Columbia-0 (Col-0), were used in this study. All plant materials were preserved and cultivated in the Laboratory of Ornamental Plant Genetics and Breeding at Northeast Agricultural University. Plants were grown under the following controlled conditions: during the vegetative growth stage, a 12-h-light/12-h-dark photoperiod was applied; during the reproductive growth stage, the photoperiod was adjusted to 14-h-light/10-h-dark. The temperature was maintained at 22–24 °C, and the relative humidity was controlled between 40% and 60%.
RNA extraction and gene cloning
-
Total RNA was extracted from petals of Lilium Oriental Hybrid 'Sorbonne' using the FastPure Plant Total RNA Isolation Kit (Vazyme Biotech Co., Ltd, Nanjing, China). RNA concentration and purity were assessed using a microvolume spectrophotometer (Kaiao Science and Technology Development Co., Ltd, Beijing, China), and integrity was verified by 1.2% agarose gel electrophoresis. First-strand cDNA was synthesized from the extracted RNA using the EasyScript® All-in-One First-Strand cDNA Synthesis SuperMix for qPCR (One-Step gDNA Removal) (TransGen Biotech Co., Ltd, Beijing, China). Specific primers were designed based on transcriptome data (primer sequences are provided in (Supplementary Table S1) to amplify the target fragment via PCR. The amplified product was purified by gel extraction, cloned into the pBZ plasmid vector, and sequenced by Sangon Biotech (Shanghai) Co., Ltd (China) to obtain the full-length sequence of LiCMK.
Multiple sequence alignment and phylogenetic analysis of LiCMK
-
To investigate the evolutionary relationships of LiCMK, a multiple sequence alignment of its amino acid sequence was performed using ClustalX. The alignment was further validated through domain prediction based on the SMART and Pfam databases. A maximum likelihood (ML) phylogenetic tree was constructed with MEGA5 software[29], and node support was assessed with 1,000 bootstrap replicates. These analyses provide insight into the evolutionary conservation and functional potential of LiCMK within the context of terpenoid biosynthesis in lily.
Analysis of LiCMK expression patterns
-
Total RNA was extracted from various tissues of Lilium Oriental Hybrid 'Sorbonne', including anthers, styles, stamens, inner petals, outer petals, and leaves, as well as from floral tissues at five developmental stages: green bud stage, pink bud stage, partially open stage, fully open stage, and senescent stage. The RNA was reverse-transcribed into cDNA. Quantitative real-time PCR (qRT-PCR)[30] was performed using ChamQ Universal SYBR qPCR Master Mix (Vazyme Biotech Co., Ltd, Nanjing, China) according to the manufacturer's instructions. The reaction mixture (20 µL total volume) contained 1 µL cDNA template, 1 µL each of forward and reverse primers (10 µM), 10 µL of 2 × ChamQ Universal SYBR qPCR Master Mix, and 8.2 µL nuclease-free water. The amplification protocol consisted of an initial denaturation at 95 °C for 30 s, followed by forty cycles of 95 °C for 10 s and 58 °C for 30 s. A melting curve analysis was performed from 65 to 95 °C with a ramp rate of 0.5 °C/s to verify amplification specificity. Three biological replicates (independent plants) and three technical replicates were included for each sample. Relative expression levels were calculated using the 2−ΔΔCT method.
Subcellular localization
-
The open reading frame (ORF) of LiCMK was cloned into the pCAMBIA1300-GFP vector using a homologous recombination method to generate the fusion construct pCAMBIA1300-GFP-LiCMK. Both the recombinant plasmid and the empty vector control (35S::GFP) were introduced into Agrobacterium tumefaciens strain GV3101 competent cells[31]. The transformed agrobacteria were resuspended in an infiltration buffer (10 mM MgCl2, 10 mM MES, 100 μM acetosyringone, pH 5.6). The bacterial suspensions were infiltrated into the leaves of Nicotiana benthamiana using a needleless syringe. At 48 to 72 h after infiltration, GFP fluorescence signals were observed and imaged using a FV3000 laser scanning confocal microscope.
Virus-induced gene silencing of LiCMK
-
The TRV2-LiCMK silencing vector was constructed by cloning a fragment of LiCMK into the TRV2 vector via homologous recombination and then introduced into Agrobacterium tumefaciens strain GV3101. Flower buds of Lilium 'Sorbonne' were infected using a vacuum infiltration method (0.07 kPa, 5 min) with a mixed agrobacterial suspension containing TRV1 and TRV2-LiCMK (1:1 ratio). A control group was infiltrated with a mixture of TRV1 and empty TRV2 (1:1 ratio). After infiltration, plants were kept in darkness for 24 h and then transferred to a growth chamber set at 22 °C under a light cycle until full blooming. Total RNA was extracted from petals of silenced lilies[32] and reverse-transcribed into cDNA[33]. The silencing efficiency was evaluated by qRT-PCR using primers designed with Primer3 software, with LrTIP41 (GenBank: KJ543466) as the internal reference gene. Expression changes of terpenoid biosynthesis-related genes were also analyzed.
Heterologous expression of LiCMK in Arabidopsis
-
The pCAMBIA1300-LiCMK expression vector was constructed using homologous recombination and introduced into Agrobacterium tumefaciens strain GV3101 via the freeze-thaw method[34]. Arabidopsis thaliana transformation was carried out using the floral dip method. Developing inflorescences were immersed in an infiltration solution containing 5% sucrose and 0.01%–0.05% (v/v) Silwet L-77. After infiltration, plants were grown under standard conditions until seed maturation. T0 seeds were surface-sown on MS medium containing 30 mg/L hygromycin for selection of transformants[35]. Putative positive transformants were identified by RT-PCR. Seeds from confirmed T1 plants (T2 generation) were collected. Total RNA was extracted from leaves of transgenic plants, reverse-transcribed into cDNA, and used as a template for PCR verification. Positive lines were advanced to the T3 generation (The Arabidopsis mutant used in the subsequent experiments) through repeated screening under the same selection conditions for subsequent experiments.
Analysis of gene expression and phenotype in transgenic Arabidopsis
-
Total RNA was extracted from homozygous T3 transgenic Arabidopsis plants and reverse-transcribed into cDNA. Three independent transgenic lines with high LiCMK expression levels were selected by qRT-PCR using AtActin2 (AT3G18780) as the internal reference gene[36]. Chlorophyll a, chlorophyll b, and carotenoid contents were measured in both wild-type and LiCMK-overexpressing plants. Phenotypic differences in seed development and leaf morphology were also analyzed. Furthermore, expression levels of key genes involved in terpenoid metabolism (DXS, DXR, TPS14, TPS21) were examined by qRT-PCR to elucidate the functional role of LiCMK.
Tobacco transformation and identification of LiCMK transgenics
-
Transgenic tobacco plants were generated via Agrobacterium-mediated leaf disk transformation[37]. After acclimatization, plants were transferred to a peat-vermiculite (1:1) substrate. One month later, they were repotted into a mixture of peat : vermiculite : perlite (1:1:1) and grown for three to four months until the flowering stage. Leaves were collected for RNA extraction, and PCR, along with qRT-PCR (using NtActin7; Gene ID: 107831145 as the internal reference) was performed to identify LiCMK-overexpressing lines for subsequent analysis of volatile compounds.
Phenotypic and gene expression analysis in transgenic tobacco
-
To investigate the effects of LiCMK overexpression on tobacco growth, key phenotypic traits—including leaf size, flower size, and flower color at full bloom—were compared between transgenic and wild-type plants. To confirm LiCMK overexpression, photosynthetic pigment contents were measured in leaves from one-month-old plants. Total RNA was extracted from pink petals of three independent transgenic lines with the highest LiCMK expression (as determined by RT-PCR) and wild-type controls. qRT-PCR was performed using established reaction conditions with gene-specific primers designed via NCBI for assessing the expression of terpenoid-related genes. Additionally, volatile compounds were analyzed in three selected high-expression transgenic lines and three wild-type plants using solid-phase microextraction (SPME)[38] coupled with gas chromatography–mass spectrometry (GC–MS)[39]. Automated high-throughput SPME was employed for extraction, and volatile identification was conducted by comparing mass spectra with the NIST 08 database, combined with manual interpretation.
Determination of phytohormone content in transgenic tobacco
-
Leaf samples (0.25 g) from one-month-old transgenic and wild-type tobacco plants were ground in liquid nitrogen and extracted with 1.5 mL of pre-cooled methanol (–20 °C) for 12 h at 4 °C. After centrifugation at 8,000 rpm for 10 min, the supernatant (A) was collected. The pellet was re-extracted with 0.5 mL of 80% methanol, and the resulting supernatant (B) was combined with A. The combined supernatant was dried under a nitrogen stream at 35 °C. The extract was subsequently processed as follows: (1) defatted three times with petroleum ether (aqueous phase retained); (2) acidified to pH 2.5 with citric acid and extracted twice with ethyl acetate (2 mL each); the organic phases were combined after centrifugation at 3,000 rpm for 3 min; (3) concentrated under nitrogen at 35–40 °C, reconstituted in 200 μL methanol, sonicated (40 kHz, 2 min), and filtered through a 0.22 μm PVDF membrane. Quantification of GA3 and ABA was performed using a Waters 2695 HPLC system equipped with a Compass C18(2) reversed-phase column.
Analysis of floral volatiles by GC–MS
-
Volatile organic compounds from lily and tobacco flowers were extracted using headspace solid-phase microextraction (HS-SPME). Following extraction, the SPME fiber was thermally desorbed in the injection port of a gas chromatograph–mass spectrometer (GC–MS). GC–MS analysis was performed under the following conditions: the initial oven temperature was set at 40 °C and held for 2 min, then increased to 250 °C at a rate of 5 °C/min, and held for 5 min. Helium was used as the carrier gas at a constant flow rate of 1.0 mL/min. Mass spectrometry detection was conducted in electron impact (EI) mode with an ionization energy of 70 eV. Ethyl decanoate (dissolved in methanol, 1:1,000, v/v) was used as an internal standard for quantification of target terpenoids, including (E)-β-ocimene, myrcene, and linalool[40,41].
-
Gene cloning and sequencing revealed that LiCMK has a full-length coding sequence of 1,191 bp, encoding a protein of 396 amino acids. Multiple sequence alignment of the deduced LiCMK protein with CMK homologs from other plant species—including Asparagus officinalis L. (AoCMK), Arabidopsis thaliana (L.)Heynh.(AtCMK), Artemisia annua L. (AaCMK), Canna indica L. (CiCMK), Nicotiana tabacum L. (NtCMK), Dioscorea zingiberensis C.H.Wright(DzCMK), Hevea brasiliensis (HbCMK), Pinus massoniana Lamb.(PmCMK), and Acorus calamus L. (AcCMK)—showed that LiCMK shares 77.19% sequence identity with AoCMK from Asparagus officinalis L. (Fig. 1a). Phylogenetic analysis further indicated that LiCMK is closely related to DzCMK from Dioscorea zingiberensis (Fig. 1b), a species within the order Liliales known to be involved in terpenoid biosynthesis. Notably, DzCMK has been functionally implicated in the synthesis of the terpenoid diosgenin. These findings suggest that LiCMK may play a role in terpenoid metabolism in lilies.
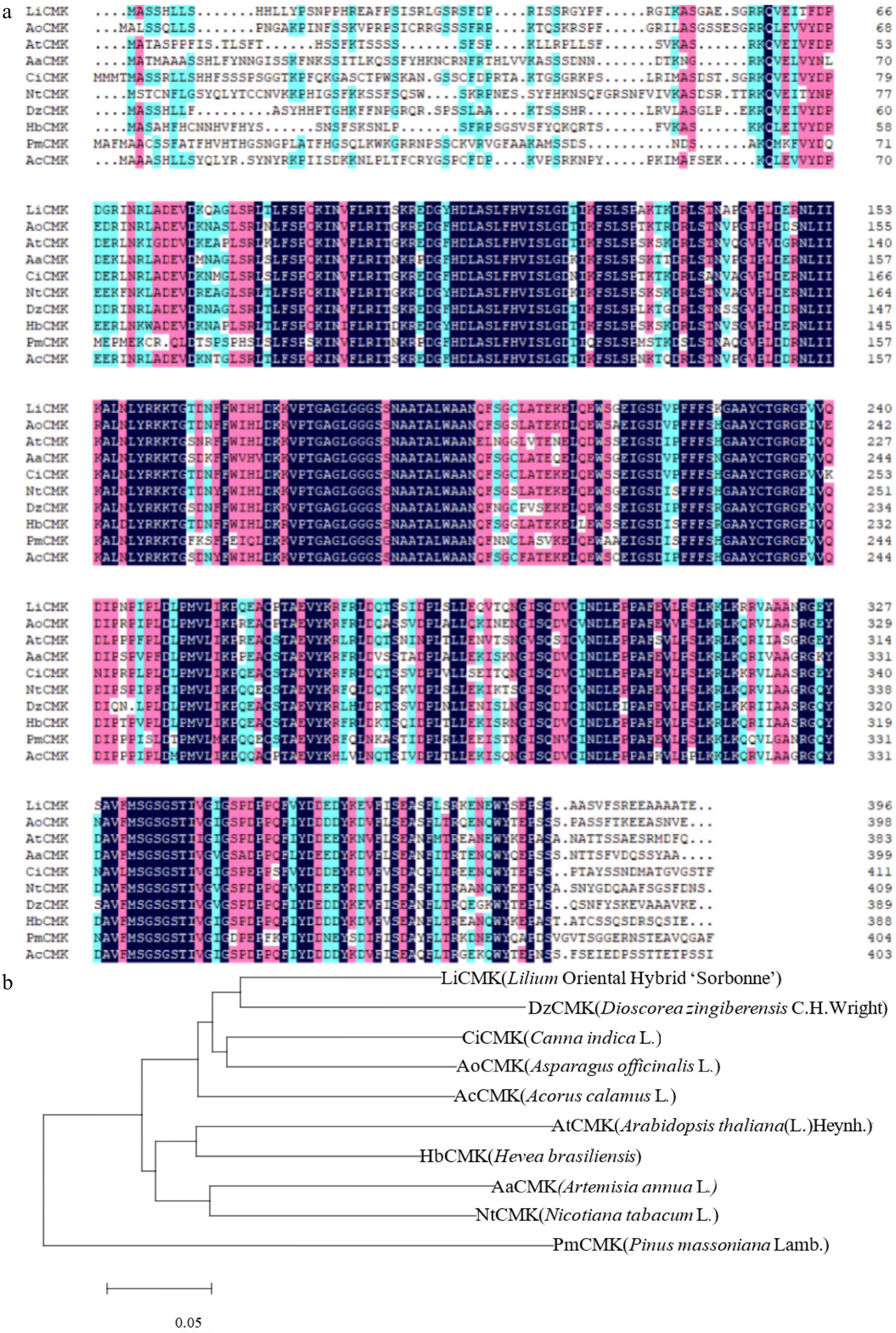

Figure 1.
Phylogenetic analysis and multiple sequence alignment of LiCMK with known homologs. (a) Multiple sequence alignment of the amino acid sequences of LiCMK and CMK proteins from other plant species. (b) Phylogenetic analysis of LiCMK from Lilium 'Sorbonne' and CMK homologs from other species.
Expression analysis of LiCMK
-
The expression dynamics of LiCMK were examined by qRT-PCR in petals of Lilium Oriental Hybrid 'Sorbonne' at five developmental stages (Fig. 2a). The results showed that LiCMK expression increased initially and then decreased, peaking at the full blooming stage. To further investigate the tissue-specific expression of LiCMK, total RNA was extracted from various organs—including filaments, stigma, anthers, leaves, inner petals, and outer petals—and analyzed by qRT-PCR. LiCMK expression was lowest in anthers and highest in outer petals, followed by inner petals; expression in outer petals was approximately 12-fold higher than that in anthers (Fig. 2b). Given its specific expression in petals—the main site of fragrance emission in 'Sorbonne'—it is proposed that LiCMK plays a critical role in floral scent formation.
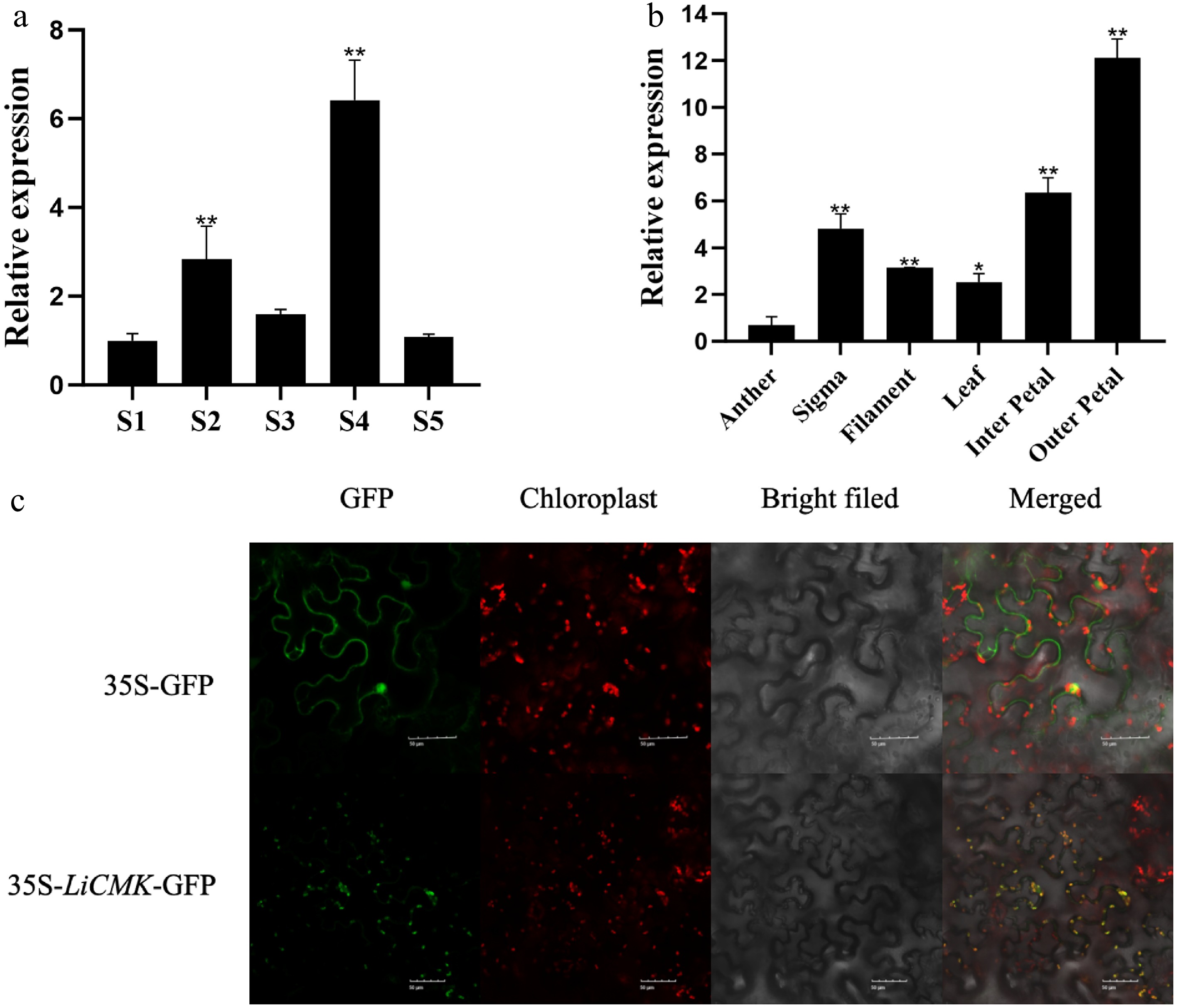
Figure 2.
Temporal-spatial characterization and subcellular localization of LiCMK in Lilium 'Sorbonne'. (a) Expression analysis of LiCMK at different developmental stages in Lilium 'Sorbonne'. (b) Tissue-specific expression analysis of LiCMK in Lilium 'Sorbonne'. (c) Subcellular localization of LiCMK. Asterisks denote statistically significant difference, * p < 0.05, ** p < 0.01.
Subcellular localization of LiCMK protein
-
The constructed pCAMBIA1300-LiCMK-GFP fusion vector was transiently expressed in tobacco leaf epidermal cells via Agrobacterium tumefaciens-mediated transformation, with the empty GFP vector serving as a control. The 35S:LiCMK-GFP fusion protein produced green fluorescent signals that colocalized with chloroplasts (Fig. 2c), indicating that LiCMK is localized to the chloroplast. This result supports its potential role in plastidial terpenoid biosynthesis via the MEP pathway.
Analysis of terpenoid-related gene expression in LiCMK-silenced Lilium
-
To elucidate the biological function of the LiCMK gene in Lilium 'Sorbonne', this study employed virus-induced gene silencing (VIGS) for functional characterization. The results (Supplementary Fig. S1) showed that LiCMK expression was significantly reduced in the experimental groups compared with the pTRV2 control group, indicating successful silencing of LiCMK in Lilium 'Sorbonne'. Among the silenced plants, lines 1#, 2#, and 5# exhibited high silencing efficiencies of 85.1%, 84.9%, and 59.6%, respectively, and were therefore selected for subsequent experiments.
Petals from these high-efficiency silencing lines (1#, 2#, and 5#) were sampled and compared with those of the pTRV2 control. Changes in the levels of the three major volatile terpenes—(E)-β-ocimene, linalool, and myrcene—in Lilium 'Sorbonne' are shown in Supplementary Fig. S2. Silencing of LiCMK led to a 2.8-fold decrease in (E)-β-ocimene content (Fig. 3a), and a 2.6-fold reduction in linalool (Fig. 3b). These results demonstrate that suppression of LiCMK expression impairs the accumulation and synthesis of monoterpenes in Lilium 'Sorbonne'.
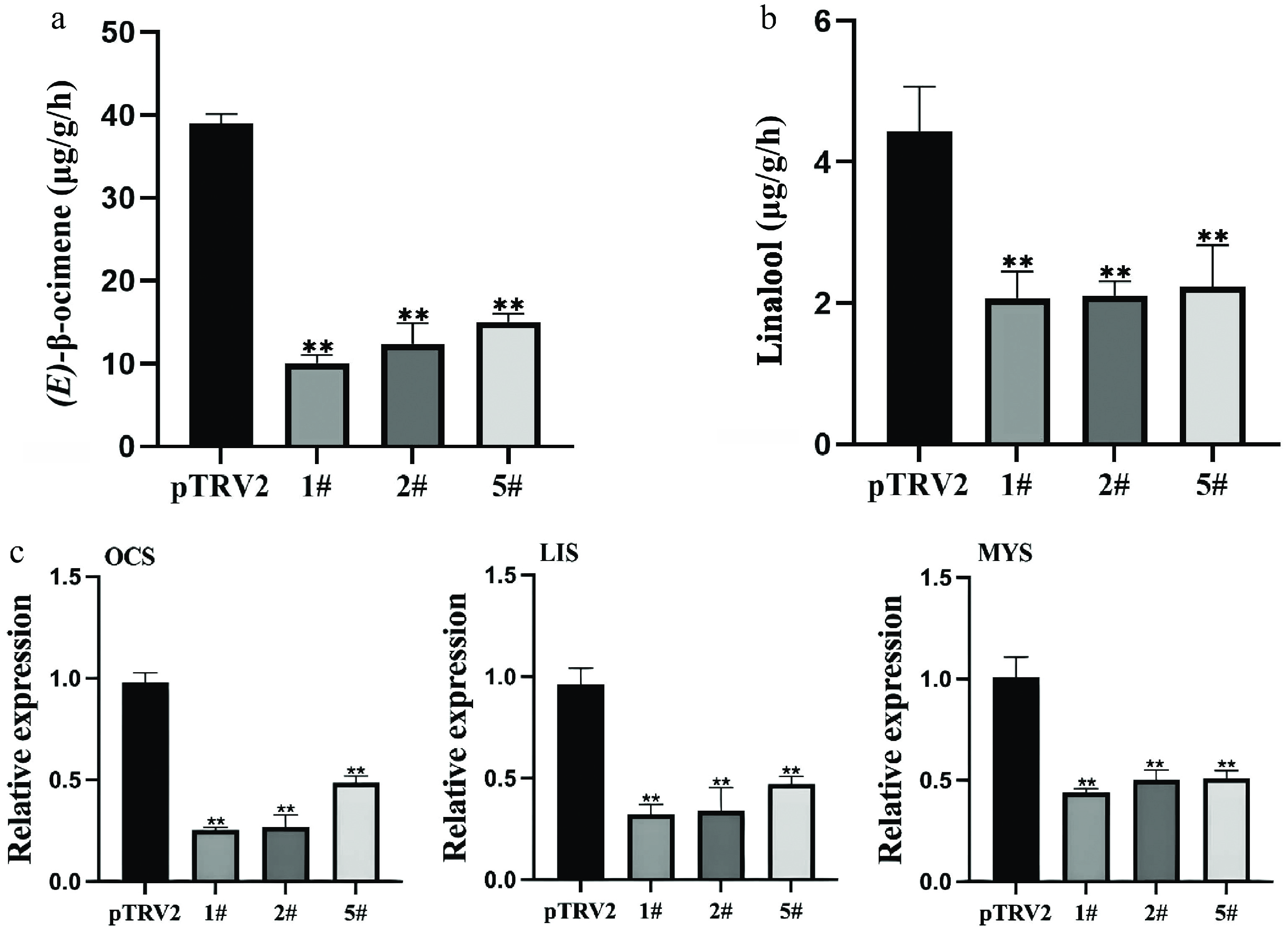
Figure 3.
Effects of LiCMK silencing on monoterpene accumulation and synthase gene expression in Lilium 'Sorbonne'. (a), (b) Contents of (E)-β-ocimene and linalool in petals after LiCMK silencing. (c) Relative expression levels of monoterpene synthase genes in response to LiCMK silencing. pTRV2: empty vector control; 1#, 2#, and 5#: LiCMK-silenced lines; OCS: (E)-β-ocimene synthase; LIS: linalool synthase; MYS: myrcene synthase. Data are presented as mean ± SD (n = 3). Asterisks indicate significant differences compared to the pTRV2 control group (one-way ANOVA followed by Duncan's test, * p < 0.05, ** p < 0.01).
Furthermore, silencing of LiCMK also influenced the expression of other terpenoid-related genes. qRT-PCR analysis revealed that the expression levels of key monoterpene synthase genes were significantly downregulated compared with the control. Specifically, (E)-β-ocimene synthase (OCS) expression decreased by approximately 66%, linalool synthase (LIS) by 62%, and myrcene synthase (MYS) by 52% (Fig. 3c). Thus, LiCMK plays a crucial role in regulating the accumulation and synthesis of major floral scent compounds in lilies.
Phenotypic comparison of transgenic and wild-type Arabidopsis
-
Seeds from T3 transgenic lines were noticeably larger than those of wild-type plants (Fig. 4a). Determination of thousand-seed weight further confirmed a significant increase in seed weight in transgenic lines (Fig. 4b). Biometric analysis revealed that the average seed weight of transgenic plants increased by approximately 34% compared to the wild-type, supporting a role for LiCMK in influencing seed development. In addition, rosette leaves of LiCMK-overexpressing transgenic plants (T3 generation) were significantly larger than those of wild-type plants under identical growth conditions (Fig. 4c).
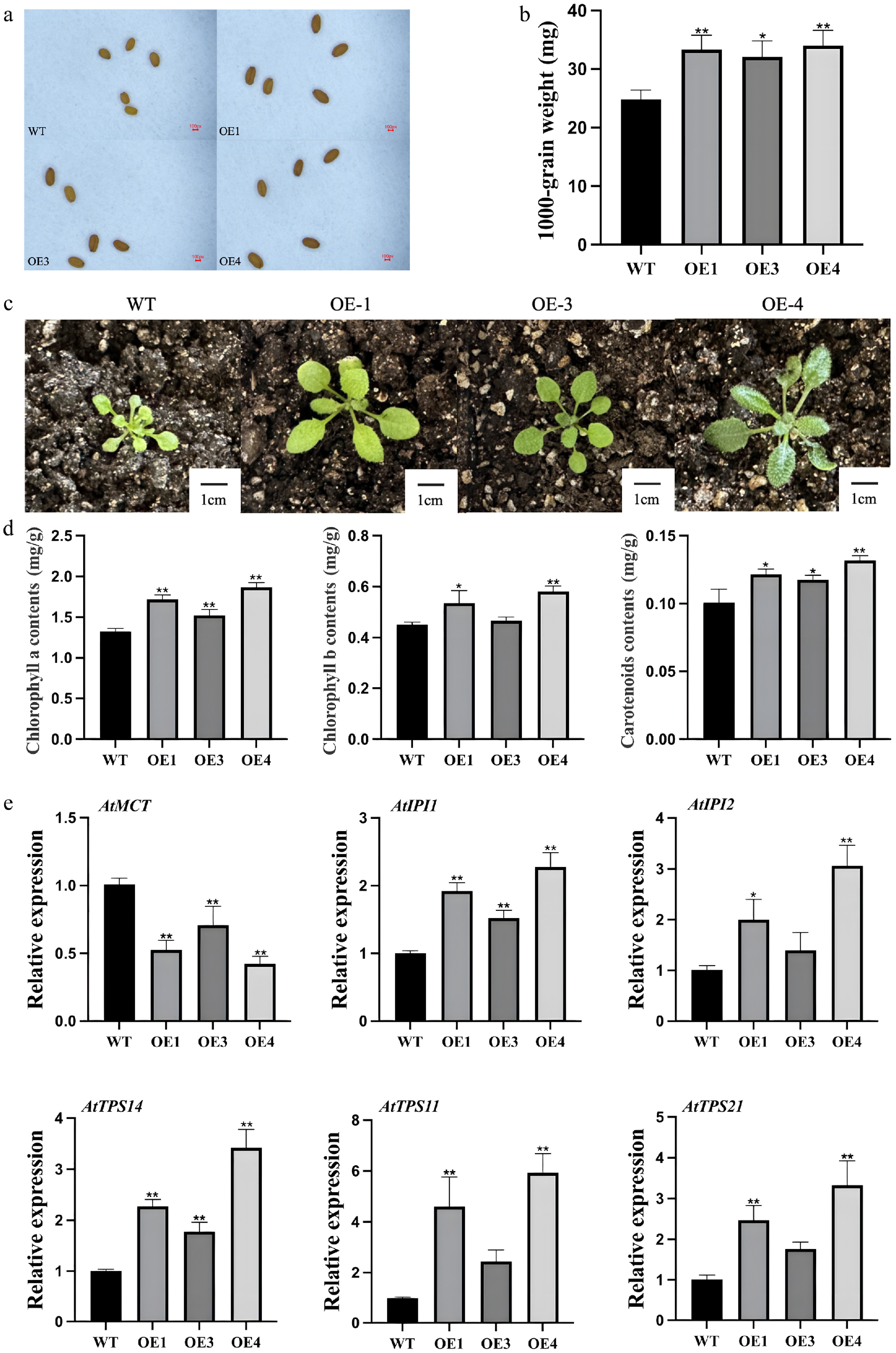
Figure 4.
Phenotypic and gene expression analysis of transgenic and mutant Arabidopsis thaliana. (a) Comparison of seed size between wild-type and transgenic Arabidopsis plants. (b) Thousand-seed weight analysis of wild-type and transgenic lines. (c) Leaf size comparison among different genotypes. (d) Photosynthetic pigment content in leaves of LiCMK-overexpressing Arabidopsis plants. (e) Expression levels of terpenoid biosynthesis-related genes in Arabidopsis. WT, wild-type; OE1, OE3, OE4, LiCMK-overexpression transgenic lines. Data are shown as mean ± SD (n = 3). Asterisks indicate significant differences between each transgenic line (OE), and the wild-type (WT) control (one-way ANOVA followed by Duncan's test, * p < 0.05, ** p < 0.01).
Analysis of photosynthetic pigments in transgenic Arabidopsis
-
Based on expression levels, two-week-old leaves from three independent T3 transgenic lines with high LiCMK expression and wild-type controls were subjected to spectrophotometric quantification of chlorophyll a, chlorophyll b, and carotenoid contents. The results indicated that the accumulation of all three photosynthetic pigments was significantly higher in transgenic lines compared with the wild-type (Fig. 4d), suggesting that overexpression of LiCMK promotes the biosynthesis of photosynthetic pigments.
Expression profiling of terpenoid biosynthetic genes in transgenic Arabidopsis
-
To investigate the regulatory effect of LiCMK on key genes involved in terpenoid metabolism, qRT-PCR was performed using flowers collected at full bloom from three independent overexpression lines (OE1, OE3, and OE4). As shown in Fig. 4e, within the MEP pathway, the expression of the upstream gene AtMCT was downregulated, whereas downstream genes AtIPI1and AtIPI2 were significantly upregulated. This expression pattern is consistent with the enzymatic position of CMK, which functions downstream of MCT and upstream of IPI in the MEP pathway. Moreover, among terpene synthase genes, expression of the monoterpene synthase gene AtTPS14 and the sesquiterpene synthase genes AtTPS11 and AtTPS21 was upregulated. In summary, these findings demonstrate that overexpression of LiCMK, a key enzyme catalyzing the fourth step of the MEP pathway, markedly alters the expression of downstream terpenoid biosynthetic genes.
Transformation and identification of LiCMK in tobacco
-
qRT-PCR analysis was performed to detect the expression level of LiCMK in transgenic tobacco lines (Supplementary Fig. S3). Three lines with high expression (3#, 6#, and 9#) were selected for further experiments. Phenotypic comparison between wild-type (WT) and transgenic plants from the seedling to flowering stages revealed no significant differences in floral organ morphology between the transgenic and WT plants (Fig. 5e, a). At one month old, leaf length (Fig. 5b) and fresh weight (Fig. 5c) were measured. The results showed that WT leaves were significantly larger than those of the transgenic lines. The average leaf length and fresh weight of transgenic plants were approximately 75% and 55% of the WT values, respectively. It was also observed that the WT controls exhibited precocious flowering compared with the LiCMK transgenic lines, and the latter exhibited a slower growth rate (Fig. 5d).
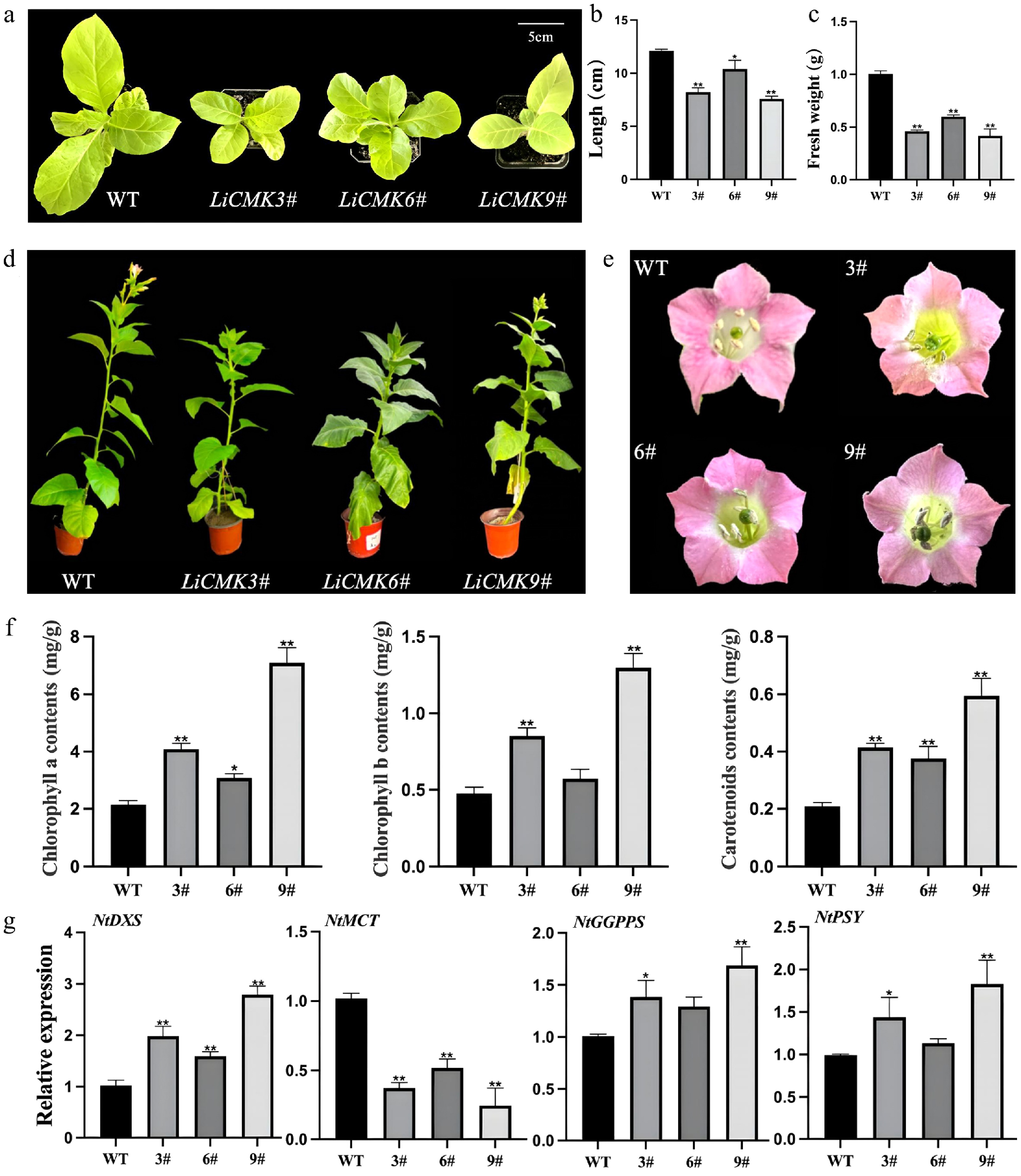
Figure 5.
Phenotypic and gene expression analysis of transgenic tobacco. Comparison of (a) one-month-old seedling phenotype, (b) leaf length, and (c) fresh weight between wild-type and transgenic lines. (d) Comparison of the phenotypes of wild type (WT) and LiCMK transgenic lines of tobacco. (e) Phenotypic comparison of flowers between wild-type and transgenic tobacco plants. (f) Analysis of photosynthetic pigments in leaves of transgenic tobacco plants. (g) Relative expression levels of genes involved in the terpenoid biosynthesis pathway in transgenic tobacco. WT, wild-type; 3#, 6#, 9#, transgenic tobacco lines. Data are presented as mean ± SD (n = 3). Asterisks indicate significant differences between the wild-type (WT) and transgenic lines (one-way ANOVA followed by Duncan's test, * p < 0.05, ** p < 0.01).
Detection of photosynthetic pigments in transgenic tobacco
-
The contents of photosynthetic pigments in leaves of wild-type and three transgenic tobacco lines were measured, with WT serving as the control. The results indicated that the levels of chlorophyll a, chlorophyll b, and carotenoids in transgenic plants were higher than those in the control (Fig. 5f). Specifically, line 9# exhibited the highest chlorophyll a content, which was 3.30-fold that of the control, while line 6# showed the lowest level at 1.43-fold. For chlorophyll b, line 9# had the highest content (2.72-fold of the control), and line 6# had the lowest (1.20-fold). Regarding carotenoids, line 9# accumulated the highest level (2.84-fold of the control), and line 6# showed the lowest (1.79-fold). These results suggest that the introduction of the LiCMK gene promotes the synthesis and accumulation of photosynthetic pigments, which are downstream products of the MEP pathway.
Analysis of terpenoid biosynthesis gene expression in transgenic tobacco
-
To investigate the effect of LiCMK overexpression on genes involved in terpenoid biosynthesis in tobacco, RNA was extracted from flowers of WT and overexpression lines (3#, 6#, and 9#), followed by qRT-PCR analysis. Candidate genes included NtDXS and NtMCT from the MEP pathway, NtGGPPS from the GGPP biosynthesis branch, and NtPSY from the carotenoid biosynthesis pathway. As shown in Fig. 5g, compared with the WT control, the expression levels of NtDXS, NtGGPPS, and NtPSY were significantly up-regulated in the overexpression lines, whereas the expression of NtMCT, a gene in the MEP pathway, was significantly down-regulated. These findings indicate that overexpression of LiCMK in tobacco suppresses the expression of its upstream gene MCT, while promoting the expression of genes related to terpenoid biosynthesis and the MEP pathway.
Analysis of volatile compounds in transgenic tobacco
-
To evaluate the effect of LiCMK gene overexpression on volatile profiles, floral volatiles from a mixture of high-expression transgenic lines (3#, 6#, 9#) and wild-type (WT) tobacco were analyzed using solid-phase microextraction (SPME) (Fig. 6a, b). The results showed significantly higher levels of (E)-β-ocimene and linalool in transgenic plants compared to WT, while the caryophyllene content did not change significantly (Fig. 7a). Specifically, linalool content in transgenic tobacco was approximately 3.2-fold that of WT, indicating that LiCMK is a key gene in terpenoid biosynthesis.
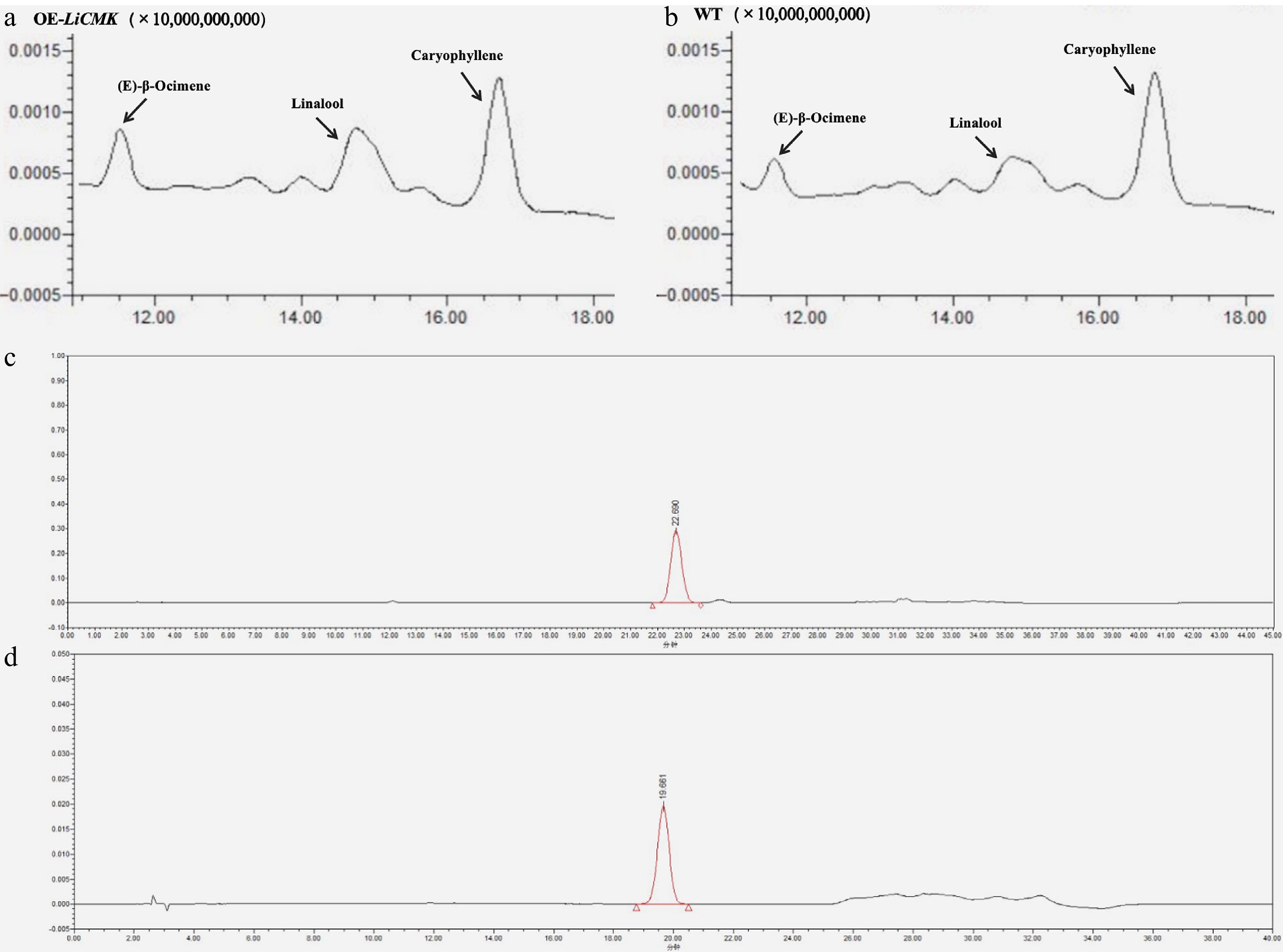
Figure 6.
Total ion chromatograms (TIC). (a) TIC of major volatile compounds in transgenic tobacco hybrid samples. (b) TIC of major volatile compounds in wild-type (WT) tobacco. (c) Chromatogram of GA3 standard. (d) Chromatogram of ABA standard.
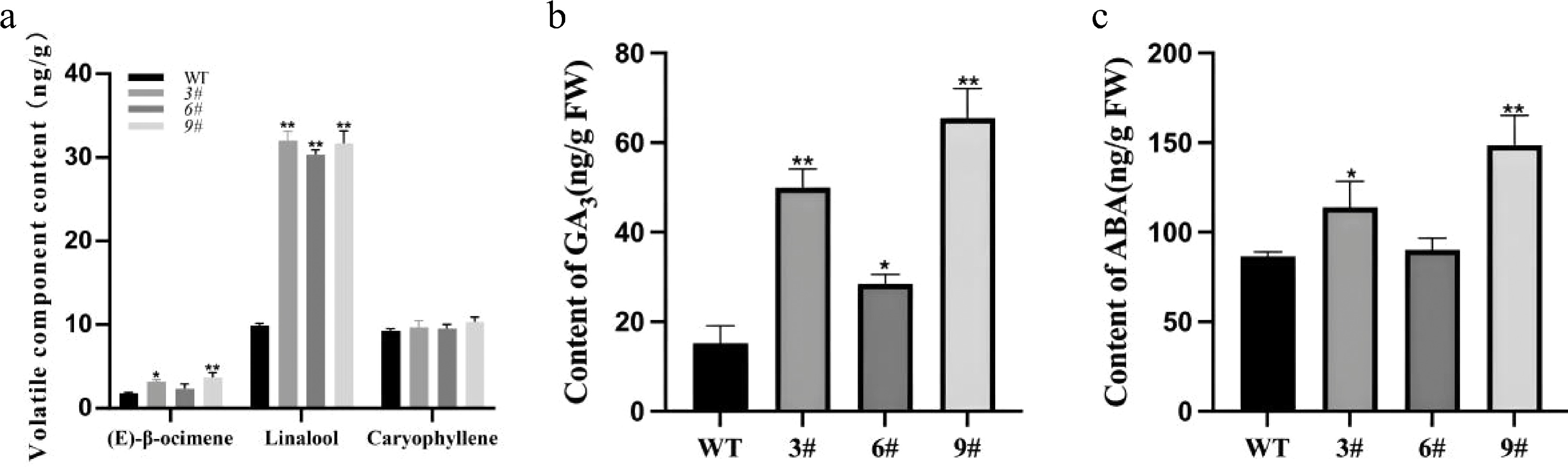
Figure 7.
Analysis of volatile compounds and phytohormones in transgenic tobacco. (a) Contents of major terpenes in flowers of wild-type and transgenic tobacco plants. (b),(c) Endogenous GA3 and ABA contents in LiCMK-overexpressing transgenic tobacco lines. WT, wild-type; 3#, 6#, 9#, independent transgenic lines. Data are shown as mean ± SD (n = 3). Asterisks indicate significant differences between the wild-type (WT) and transgenic lines (one-way ANOVA followed by Duncan's test, * p < 0.05, ** p < 0.01).
Plant hormone analysis in transgenic tobacco
-
Analysis of hormone levels in WT and transgenic lines (3#, 6#, 9#) was also conducted (Fig. 6c, d). Since LiCMK-overexpressing lines exhibited a late-flowering phenotype and flowering time is hormonally regulated, the results measured the contents of gibberellin (GA3) and abscisic acid (ABA) in one-month-old leaves. Both GA3 and ABA levels were significantly elevated in the transgenic lines relative to WT (Fig. 7b, c).
-
The main aromatic components of lily fragrance are terpenoid volatiles. Their biosynthetic precursors, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP), are generated via both the cytosolic mevalonate (MVA) pathway and the plastidial methylerythritol phosphate (MEP) pathway. The MEP pathway primarily contributes to the synthesis of monoterpenes, diterpenes, and polyterpenes. In its fourth enzymatic step, 4-(cytidine 5′-diphospho)-2-C-methyl-D-erythritol kinase (CMK) catalyzes the conversion of CDP-ME to CDP-ME2P, representing a key rate-limiting reaction toward the formation of IPP and DMAPP[42]. In this study, the full-length open reading frame (ORF) of LiCMK was isolated. Bioinformatics analysis indicated that it encodes a hydrophilic protein lacking both transmembrane domains and a signal peptide. Subcellular localization predictions suggested a chloroplast-specific localization for LiCMK. As the chloroplast is a central site for the synthesis of various floral scent volatiles, such as linalool[43], these results imply a potential role for LiCMK in floral fragrance metabolism. Tissue-specific expression profiling demonstrated that LiCMK expression was highest in the outer petals of Lilium 'Sorbonne', correlating with terpenoid emission patterns. Throughout floral development, LiCMK transcript levels peaked at the full-blooming stage (S4), consistent with the highest emission of monoterpenes at this phase. In contrast, the expression of LiDXR, another gene within the MEP pathway, reached its maximum at the half-opening stage (S3). This distinct temporal expression pattern suggests that LiCMK may specifically regulate monoterpene synthesis during the full-blooming stage in lilies.
Virus-induced gene silencing (VIGS) is a powerful and versatile reverse genetic tool that enables targeted suppression of gene expression without the need for stable genetic transformation. This technique has been widely employed to investigate the function of fragrance-related genes; for instance, silencing of PsuLIS in tree peony led to reduced linalool synthesis[44], and suppression of LcTPS1 and LcTPS14 expression in crape myrtle resulted in decreased volatile production[45]. In this study, VIGS was applied to Lilium 'Sorbonne' by vacuum infiltrating stem segments with a virus-derived construct targeting LiCMK, achieving efficient gene silencing. qRT-PCR analysis confirmed a significant downregulation of LiCMK transcripts in silenced plants, accompanied by reduced expression of downstream terpenoid synthase genes (LiOCS, LiLIS, and LiMYS). GC-MS profiling of floral volatiles further revealed markedly decreased emissions of key monoterpenes, including (E)-β-ocimene, linalool, and myrcene. These findings provide direct in planta evidence that LiCMK positively regulates terpenoid biosynthesis in lilies. While the infiltration-based method used differs from the needle injection approach used by Yang et al. to silence LiSRM1 in Lilium 'Siberia'[46], both strategies effectively achieved silencing, demonstrating the adaptability of VIGS for functional genetic studies across lily cultivars.
To further investigate the function of LiCMK, heterologous overexpression assays were conducted in both Arabidopsis and tobacco. In Arabidopsis, overexpression of LiCMK significantly increased the contents of chlorophyll a/b and carotenoids and complemented the albino phenotype of the IspE mutant[47], indicating a role in promoting photosynthetic pigment and terpenoid production[48,49]. Transgenic lines also produced larger and heavier seeds, a phenotype consistent with previous reports concerning ZmHAK1 overexpression[50]. At the molecular level, LiCMK overexpression up-regulated key terpenoid biosynthetic genes, including AtTPS14, AtTPS11/21, and AtIPI1/2, but suppressed the expression of the upstream gene AtMCT, suggesting the presence of a feedback regulatory mechanism. Similarly, in tobacco, transgenic lines (3#, 6#, and 9#) overexpressing LiCMK exhibited notably elevated photosynthetic pigment levels, consistent with phenotypes reported in PmCMK transgenics[51]. qRT-PCR analysis revealed up-regulation of NtDXS and suppression of NtMCT, further supporting feedback inhibition at the transcriptional level[52]. The transgenic plants displayed developmental delay and late flowering, accompanied by elevated endogenous levels of GA3 and ABA. Exogenous application of ABA or GA3 promoted terpenoid accumulation[53−55]. Volatile profiling confirmed significantly increased emissions of (E)-β-ocimene and linalool, resembling the effects of LiDXS overexpression. These results demonstrate that LiCMK contributes to monoterpene biosynthesis, likely through phytohormone-mediated pathways.
This study reveals that heterologous overexpression of the lily LiCMK gene leads to contrasting growth phenotypes in Arabidopsis and tobacco, underscoring the significant impact of species-specific metabolic regulatory networks. Research in Arabidopsis and tomato reveals that species-specific architectures of the interconnected polyamine-ethylene network underlie their distinct stress responses[56]. In this present study, heterologous overexpression of the lily LiCMK gene in Arabidopsis enhanced the metabolic flux of the MEP pathway, promoting the accumulation of photosynthetic pigments and the expression of terpenoid biosynthesis-related genes, thereby increasing plant biomass. In tobacco, although photosynthetic pigment synthesis was similarly enhanced, carbon flux was reprogrammed toward terminal terpenoid products, accompanied by abnormal accumulation of GA3 and ABA, collectively leading to the suppression of vegetative growth. These findings indicate that while the catalytic function of LiCMK is conserved, the system-level responses triggered by its overexpression are strongly dependent on the host's inherent metabolic allocation and regulatory mechanisms[57]. In alignment with findings from studies on the lily LiMCT gene[58], this research further confirms that while core enzymatic functions of MEP pathway genes are maintained in heterologous systems, the ultimate phenotypic outcomes are governed by the distinct metabolic and regulatory contexts of the host species.
-
This study aimed to characterize the role of the CMK gene in terpenoid biosynthesis in lily, identified LiCMK (1,191 bp, encoding 396 amino acids), and confirmed its subcellular localization to the chloroplast. qRT-PCR analysis revealed that LiCMK is preferentially highly expressed in outer petals at full bloom, consistent with the emission pattern of monoterpenes. Silencing of endogenous LiCMK via a TRV2-based VIGS system led to downregulation of monoterpene synthase genes (LiOCS, LiLIS, and LiMYS) and significantly reduced petal contents of (E)-β-ocimene, linalool, and myrcene, as determined by SPME-GC-MS. Furthermore, heterologous overexpression of LiCMK in Arabidopsis and tobacco enhanced photosynthetic pigment accumulation, upregulating terpenoid biosynthetic genes and increasing the emission of linalool and (E)-β-ocimene in tobacco. These results demonstrate that LiCMK participates in floral scent formation by modulating terpenoid biosynthesis and influences plant development.
This research was supported by the Key Research and Development of the Ministry of Science and Technology (Grant No. 2016YFC0500306-02) and the project of the Science and Technology Department of Heilongjiang Province (Grant No. C2017030).
-
The authors confirm contribution to the paper as follows: study conception and design, methodology: Dong J; investigation, draft manuscript preparation: Zhang W, Chen C; software: Zhang W, Chen C, Qin Y; formal analysis: Zhang W, Chen C, Yu J; data curation: Zhang W, Chen C, Yu J, Dong J, Zhang J; visualization: Yang T; resources: Qin Y; supervision: Fan J. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article and its supplementary information files.
-
The authors declare that they have no conflict of interest.
-
# Authors contributed equally: Wuhua Zhang, Chen Chen
- Supplementary Table S1 The primers used in the experiment.
- Supplementary Fig. S1 Relative expression levels after LiCMK silencing (self-drawn).
- Supplementary Fig. S2 Total ion chromatogram (TIC) of major volatile compounds after LiCMK silencing.
- Supplementary Fig. S3 The expression level of LiCMK in different transgenic lines.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhang W, Chen C, Qin Y, Yu J, Zhang J, et al. 2026. Cloning and functional characterization of 4-Diphosphocytidyl-2-C-Methyl-D-Erythritol kinase (LiCMK) gene in oriental lily (Lilium 'Sorbonne') Func. Char. of CMK in Lilium 'Sorbonne'. Ornamental Plant Research 6: e012 doi: 10.48130/opr-0026-0002
Cloning and functional characterization of 4-Diphosphocytidyl-2-C-Methyl-D-Erythritol kinase (LiCMK) gene in oriental lily (Lilium 'Sorbonne') Func. Char. of CMK in Lilium 'Sorbonne'
- Received: 26 September 2025
- Revised: 26 November 2025
- Accepted: 04 January 2026
- Published online: 18 March 2026
Abstract: Oriental lily (Lilium spp.) has emerged as a model system for floral fragrance research due to its distinctive scent profile. Terpenoid compounds, major constituents of its floral volatiles, are primarily synthesized through the methylerythritol phosphate (MEP) and mevalonic acid (MVA) pathways. 4-Diphosphocytidyl-2-C-Methyl-D-Erythritol Kinase (CMK), a key enzyme in the MEP pathway, is involved in monoterpene biosynthesis. However, the function of CMK in lily remains uncharacterized. In this study, the role of LiCMK in Lilium 'Sorbonne' is investigated. Gene cloning revealed that LiCMK, previously identified in the present transcriptome analysis, has a full-length coding sequence of 1,191 bp encoding 396 amino acids, and the protein localizes to the chloroplast. Tissue-specific expression analysis showed that LiCMK is highly expressed in outer whorl petals during the full-blooming stage. Virus-induced gene silencing (VIGS) of LiCMK led to a significant reduction in volatile terpenoid emissions and downregulation of downstream genes (LiLIS, LiMYS, LiOCS), confirming LiCMK as a key rate-limiting enzyme in floral fragrance biosynthesis. Functional validation in model plants showed that overexpression of LiCMK in Arabidopsis led to enlarged leaves, increased photosynthetic pigments, and upregulation of terpenoid pathway genes. In transgenic tobacco, LiCMK overexpression caused delayed flowering, elevated levels of hormones (GA and ABA), and specific accumulation of (E)-β-ocimene and linalool, with activation of genes like NtDXS. Collectively, the findings of this study indicate that LiCMK enhances floral fragrance synthesis in Lilium 'Sorbonne' by activating MEP pathway genes. This study provides molecular insights into the mechanism of fragrance biosynthesis in lilies and will facilitate future breeding and application-oriented research.
-
Key words:
- Lilium 'Sorbonne' /
- Expression analysis /
- LiCMK /
- Floral monoterpene /
- Function verification













