-
Flowering plants have evolved the capacity to sense favorable seasons to accurately initiate the seed dormancy-to-germination transition during their life cycles, leading to their successful adaptation and high suitability to an ever-changing environment[1,2]. Seeds usually become dormant during maturation in their mother plants. After the completion of seed development, the seeds perceive multiple exogenous cues, such as light, temperature, soil nitrate levels, and smoke, which provide information about soil status and time of year, facilitating them to determine whether to terminate seed dormancy and initiate germination[3−6]. Numerous studies have demonstrated that the phytohormones, mainly abscisic acid (ABA) and gibberellin (GA), function as key integrators between endogenous developmental cues and environmental conditions, to modulate gene transcription during seed dormancy-to-germination transition. Specifically, the dormancy and germination status of seeds are primarily determined by the dynamic balance between ABA and GA levels or sensitivity[7−9].
As the primary phytohormone in controlling seed dormancy and germination, ABA is perceived by the well-known receptors PYRABACTIN RESISTANCE (PYR)/PYR1-LIKE (PYL)/REGULATORY COMPONENT OF ABSCISIC ACID RECEPTOR (RCAR), which leads to the formation of PYR/RCAR–PROTEIN PHOSPHATASE 2C (PP2C) complex, and to their subsequent repression[10,11]. The inhibition of PP2Cs releases SNF1-related kinases 2 (SnRK2s) which phosphorylate downstream transcription factors (TFs), thereby initiating the ABA-responsive signaling cascade[12−16]. As a master transcription factor in ABA signaling, ABI5 (ABSCISIC ACID INSENSITIVE5) was revealed to directly bind and activate the promoters of downstream ABA-responsive genes like EM1 (EARLY METHIONINE-LABELED 1) and EM6, ultimately leading to the inhibition of seed germination[17]. Dissecting novel ABA signaling-associated components, especially critical transcription modulators, may contribute to the explanation of the molecular basis of ABA signaling or its crosstalk with other signaling pathways during seed germination.
The bHLH transcription factors are regulatory proteins that constitute a superfamily in plants. In tobacco, the bHLH transcription superfamily comprises over 190 members that can be divided into 23 subfamilies based on their protein sequence similarity[18]. Although much progress in bHLH transcription factors' functional research has been obtained in the past decades, especially in the model plant Arabidopsis thaliana, the functional elucidation of most bHLH genes in tobacco represent a major challenge. A body of genetic and molecular evidence has revealed that bHLH transcription factors perform crucial functions in plant development as well as in responses to both biotic and abiotic stresses[19,20]. Among them, the bHLH transcription factor MYC2 was revealed to function as the master regulator of the JA signaling cascade and participates in the regulation of various plant physiological processes, including JA-induced leaf senescence, root growth, and both biotic and abiotic stresses[21,22]. Furthermore, MYC2 can also participate in ethylene, salicylic acid, GA, and ABA signaling to mediate various plant developmental processes and defense responses[23]. In Nicotiana species, MYC2 transcription factors, including NtMYC2a/b/c from tobacco and their homolog NbbHLH2 from N. benthamiana, function as key regulators to control nicotine biosynthesis possibly through directly binding to the target G-box sequences present in the promoters of nicotine biosynthetic pathway genes, such as NtPMT1a/2, NbPMT, NtQPT2, NtA622, and NtMATE1[24−27]. One recent study also demonstrated that NtMYC2a participates in pollen development through modulating carbohydrate metabolism in tobacco[28]. Interestingly, their possible roles in seed germination in tobacco and the mechanisms involved still remain unclear.
Here, this study employed diverse molecular approaches to investigate the role of NtMYC2a and NtMYC2b in the regulation of ABA-mediated seed germination. We found that both mutation and overexpression of NtMYC2a and NtMYC2b can affect ABA-mediated seed germination. Furthermore, NtMYC2a and NtMYC2b can interact with each other to form a heterodimer and synergistically activate NtABI5 expression, leading to the transmission of ABA signaling during seed germination. Our results thus provide evidence that both NtMYC2a and NtMYC2b function as positive regulators to modulate ABA responses during seed germination in tobacco.
-
The seeds used in this study were all derived from the genetic background Nicotiana tabacum (cv. Coker176). Both the NtMYC2a/2b mutants and overexpression lines were kindly gifted by Yulong Gao (Yunnan Academy of Tobacco Agricultural Sciences). Apart from ABA (Sigma Co. Ltd.), all other chemicals were sourced from Shanghai Sangon Biotechnology Co. Ltd. (Shanghai, China).
Treatment of ABA
-
ABA working solution (100 μM) was prepared by diluting the 100 mM ABA stock solution, and six-week-old seedlings were used in this study. Samples were collected at 0, 1, 2, 4, 8, 12, and 24 h after treatment with H2O or ABA. For each time point, three biological replicates were set, with each replicate consisting of pooled tissues from at least three individual seedlings. All samples were immediately frozen in liquid nitrogen after collection, and stored at −80 °C until RNA extraction.
RT-qPCR analysis
-
Total RNA was isolated with Trizol reagent according to the manufacturer's protocol. Total RNA (1 μg) pre-treated with DNase1 was reverse-transcribed using Superscript II. Then, reverse transcription-quantitative polymerase chain reaction (RT-qPCR) was performed on a Roche Light Cycler 480 real-time PCR machine. All gene-specific primers used for RT-qPCR are listed in Supplementary Table S1.
Western blotting
-
Seeds of 35S:MYC-NtMYC2a/2b transgenic plants were harvested after treatment with ABA at the indicated time, and total proteins were extracted using RIPA lysis buffer (50 mM Tris-HCl [pH 8.0], 0.1% Nonidet P-40, 150 mM NaCl, 1% SDS, 0.5% sodium deoxycholate, and a protease inhibitor cocktail [CWBIO]). The anti-MYC (Abmart) and anti-ACTIN (Abiocode) were used to detect MYC-NtMYC2a/2b and Actin, respectively.
Measurement of germination or greening rates
-
Seeds of WT and NtMYC2a/2b mutants or overexpression lines were harvested at the same time and dried for 2 weeks. Seeds were stratified for 3 d at 4 °C and then germinated at 24 °C under a 16 h light/8 h dark photoperiod. Seeds were plated on 1/2 MS medium supplemented with different ABA concentrations (0, 0.5, or 1 µM) to assess ABA sensitivity of seeds. Germination was defined by the appearance of visible radicles, while seedling greening was determined based on the emergence of green cotyledons.
EMSA
-
The full-length CDSs of NtMYC2a/2b were inserted into the expression vector pET-28a to generate pET-28a-NtMYC2a/2b. His-NtMYC2a/2b recombinant proteins were purified using the His PurNi-NTA resin. The biotin-labeled WT or mutant NtABI5 promoter fragments were used as probes, and the biotin-unlabeled WT fragments were used as competitors. Electrophoresis mobility shift assays were conducted using a Chemiluminescent EMSA Kit following the manufacturer's protocol.
Molecular interaction simulation
-
The amino acid sequences of the two proteins were submitted to AlphaFold3 to predict the structure of the protein complex. The visualization of the interacting structures was achieved in PyMOL by uploading the protein structure. The image of molecular interaction was exported from PyMOL as png file.
Pull-down assay
-
The full-length cDNA of NtMYC2a was fused to pGEX-4T-1, and the full-length cDNA of NtMYC2b was cloned into pET-28a. Then these two plasmids were transformed into Escherichia coli BL21 cells, and the recombinant proteins were induced with isopropyl-beta-thiogalactopyranoside. Crude NtMYC2a-GST and GST recombinant protein extracts were obtained and then immobilized to glutathione affinity resin. NtMYC2b-His protein was co-incubated with the immobilized NtMYC2a-GST and GST fusion proteins for 4 h at 4 °C. Co-incubated proteins were eluted with glutathione elution buffer, and the eluted proteins were then subjected to western blot analysis. The membranes were probed respectively with anti-His and anti-GST antibodies to determine protein interaction. The primers used for pull-down are listed in Supplementary Table S2.
Split luciferase complementation assay
-
To confirm the interactions between NtMYC2a and NtMYC2b, their full-length CDS were fused to pCAMBIA1300-cLUC or pCAMBIA1300-nLUC to generate cLUC-MYC2b and nLUC-MYC2a respectively. All plasmids were introduced into Agrobacterium tumefaciens strain EHA105, and then infiltration of N. benthamiana leaves was performed as described in Zhang et al[29]. Infiltrated plants were kept under standard growth conditions for 48 to 72 h to allow optimal protein expression and potential interaction. Luciferase imaging was performed using an in vivo imaging system. The leaves were sprayed uniformly with a 1 mM D-luciferin substrate solution (in 0.1% Triton X-100), and placed in darkness for 5 min to minimize background luminescence. Bioluminescence signals were then captured using a low-light, cooled CCD camera imaging apparatus (Tanon-5200, China) with an exposure time optimized to avoid signal saturation. The split luciferase complementation assays were carried out with at least three independent biological replicates, with similar results. The primers used for this study are listed in Supplementary Table S2.
Transient expression assay
-
The native promoters of NtABI5 were amplified and cloned into the pGreenII 0800-LUC vector as a reporter. Full-length CDSs of NtMYC2a, NtMYC2b, and GFP were amplified and cloned into the pGreenII 62-SK vector as effectors. Agrobacterium tumefaciens EHA105 (pSoup) harboring the above constructs was infiltrated into five-week-old N. benthamiana leaves using a needleless syringe for transactivation analyses. The low-light cooled CCD imaging apparatus was used to capture the LUC image and count luminescence intensity. The transient expression assays were carried out with at least three independent biological replicates with similar results. The primers used for this study are listed in Supplementary Table S2.
-
Previous studies have demonstrated that NtMYC2s function as critical regulators in the synthesis and transport of nicotine and also participate in the regulation of carbohydrate metabolism during pollen development[22,27,28]. However, their distinctive roles in various physiological processes and stress responses in tobacco are still poorly characterized. This study performs experiments to determine their possible roles in ABA-mediated seed germination. To determine their roles in ABA signaling, we first conduct RT-qPCR analysis to explore their expression upon ABA treatment. As shown in Fig. 1a, b, the expression of both NtMYC2a and NtMYC2b was inhibited upon ABA treatment compared with the control treatment. To further reveal the expression patterns of NtMYC2a and NtMYC2b, the stabilities of both NtMYC2a and NtMYC2b proteins during ABA treatment was determined using 35S:MYC-NtMYC2a/2b transgenic plants. The western blotting analysis showed that both NtMYC2a and NtMYC2b protein accumulations were decreased upon ABA treatment (Fig. 1c, d). These results indicated that NtMYC2a and NtMYC2b may participate in the regulation of ABA signaling.
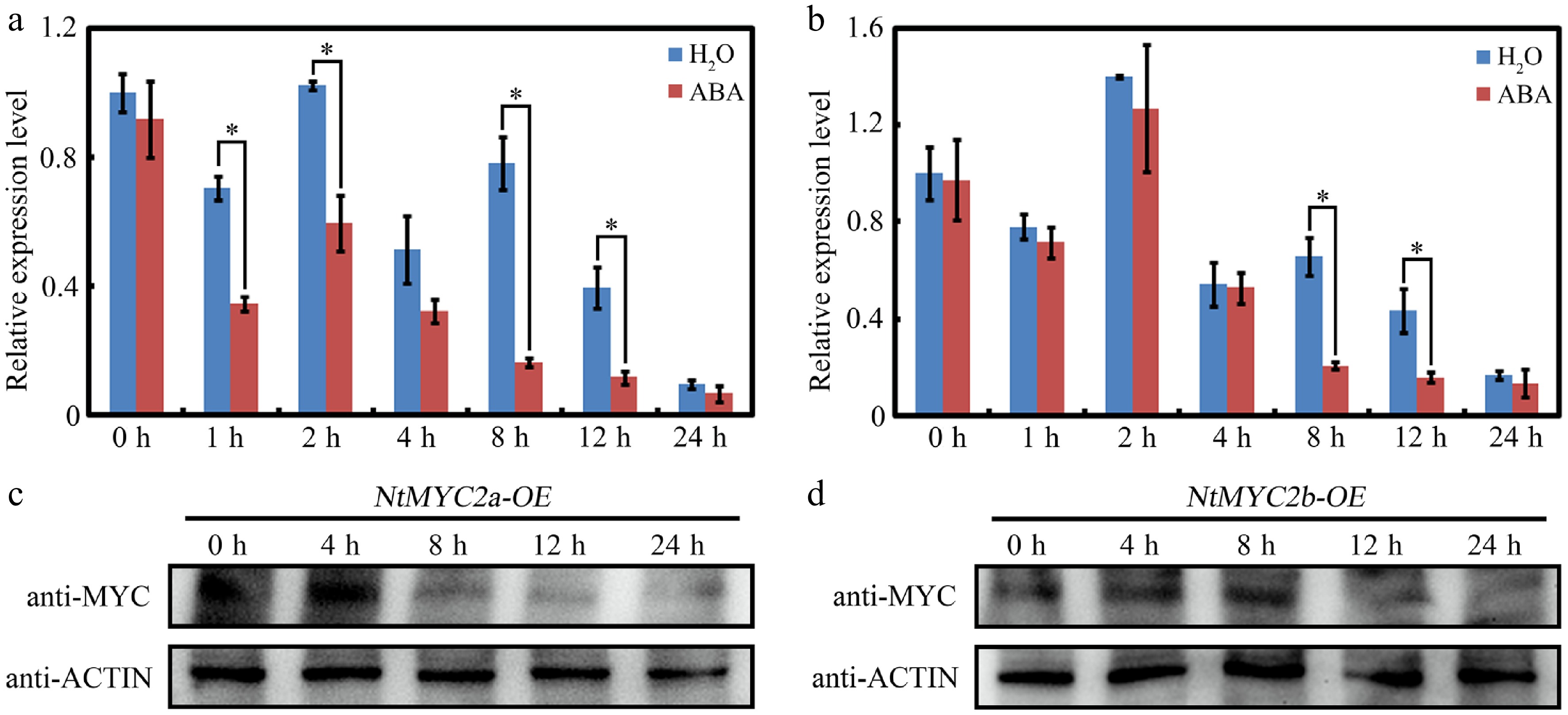
Figure 1.
Expression of NtMYC2a and NtMYC2b upon ABA treatment. (a) and (b) RT-qPCR analysis of transcript levels of NtMYC2a and NtMYC2b after ABA treatment. Samples were collected after ABA treatment for the indicated time. All data are presented as the mean ± SD from three independent biological replicates. Asterisks indicate Student's t-test significant differences as compared to controls, * p < 0.05. (c) and (d) Western blot analysis of NtMYC2a/2b protein accumulation in transgenic 35S:MYC-NtMYC2a/2b plants after ABA treatment for the indicated time. An anti-MYC antibody was used to detect MYC-NtMYC2a/2b. Actin was used as an internal control.
Knock out of NtMYC2a and NtMYC2b renders the seeds more insensitive to ABA
-
Based on the findings that both NtMYC2a and NtMYC2b are downregulated by ABA, we speculated that they may be involved in the regulation of ABA associated responses, such as ABA-mediated seed germination. Then, the phenotypes of the wild type, Ntmyc2a and Ntmyc2b single and double mutants were compared, in response to ABA during seed germination[27] (Supplementary Figs. S1, S2). No notable phenotypic differences were observed among the WT, Ntmyc2a and Ntmyc2b single mutants, and the double mutants on half-strength MS medium (Fig. 2a−c). However, both the Ntmyc2a and Ntmyc2b single mutant seeds exhibited a slight but consistent reduction in ABA sensitivity during seed germination and early seedling growth compared to the WT, especially at a concentration of 0.5 µM ABA. Furthermore, the Ntmyc2a/2b double mutant seeds were more insensitive to ABA compared with that of WT, and their single mutants, suggesting a redundant function of NtMYC2a and NtMYC2b during ABA-mediated seed germination (Fig.2a−c). Thus, our results implied that NtMYC2a/2b may act as positive regulators in ABA mediated seed germination and early seedling growth.
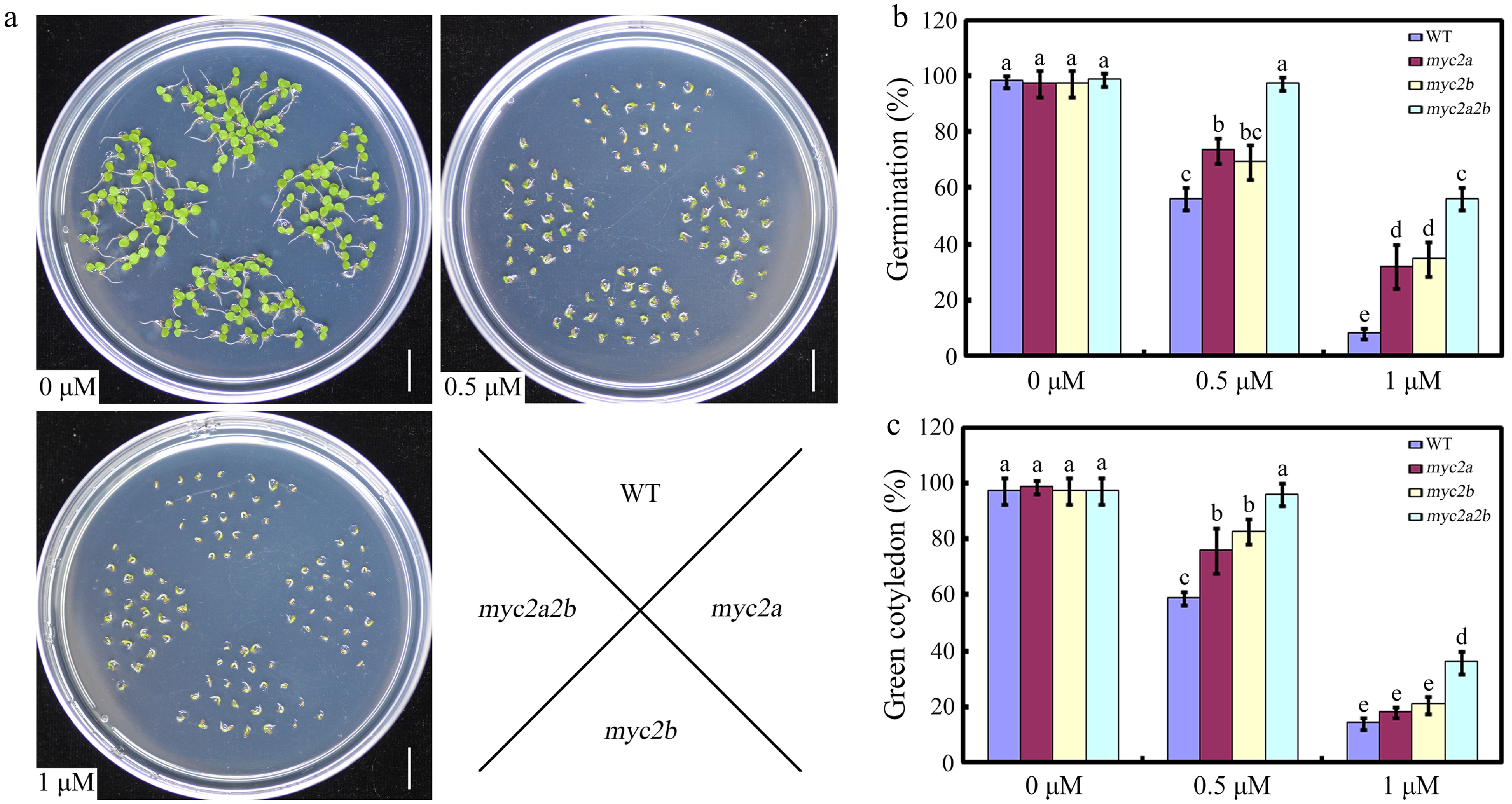
Figure 2.
The effect of NtMYC2a and NtMYC2b mutation on seed germination. (a) Seedlings of the indicated genotypes observed 7 d after germination on medium supplemented with 0, 0.5 or 1 µM ABA. (b) Statistic analysis of germination rate in the indicated genotypes. The seeds were scored 5 d after stratification on medium containing different concentrations of ABA. Germination is defined by radical emergence. (c) Statistic analysis of the indicated genotypes for cotyledon greening. Cotyledon greening was scored 7 d after stratification on medium containing different concentrations of ABA. All data are presented as the mean ± SD from three independent biological replicates. These data were analyzed by a two-way ANOVA using Tukey's HSD test. Bars with different letters are significantly different from each other (p < 0.05).
Overexpression of NtMYC2a or NtMYC2b renders the seeds more sensitive to ABA
-
To further characterize the role of NtMYC2a and NtMYC2b in ABA-mediated seed germination, we also obtained transgenic tobacco plants constitutively expressing NtMYC2a or NtMYC2b under the control of the cauliflower mosaic virus (CaMV) 35S promoter. In contrast to the Ntmyc2a/2b mutants, seeds from NtMYC2a/2b overexpression lines exhibited significantly reduced germination and cotyledon greening relative to the WT (Fig. 3a−c). Thus, constitutive overexpression of NtMYC2a/2b led to enhanced sensitivity to ABA during seed germination and post-germinative growth, and further confirm that NtMYC2a and NtMYC2b function as positive regulators in ABA-mediated seed germination.
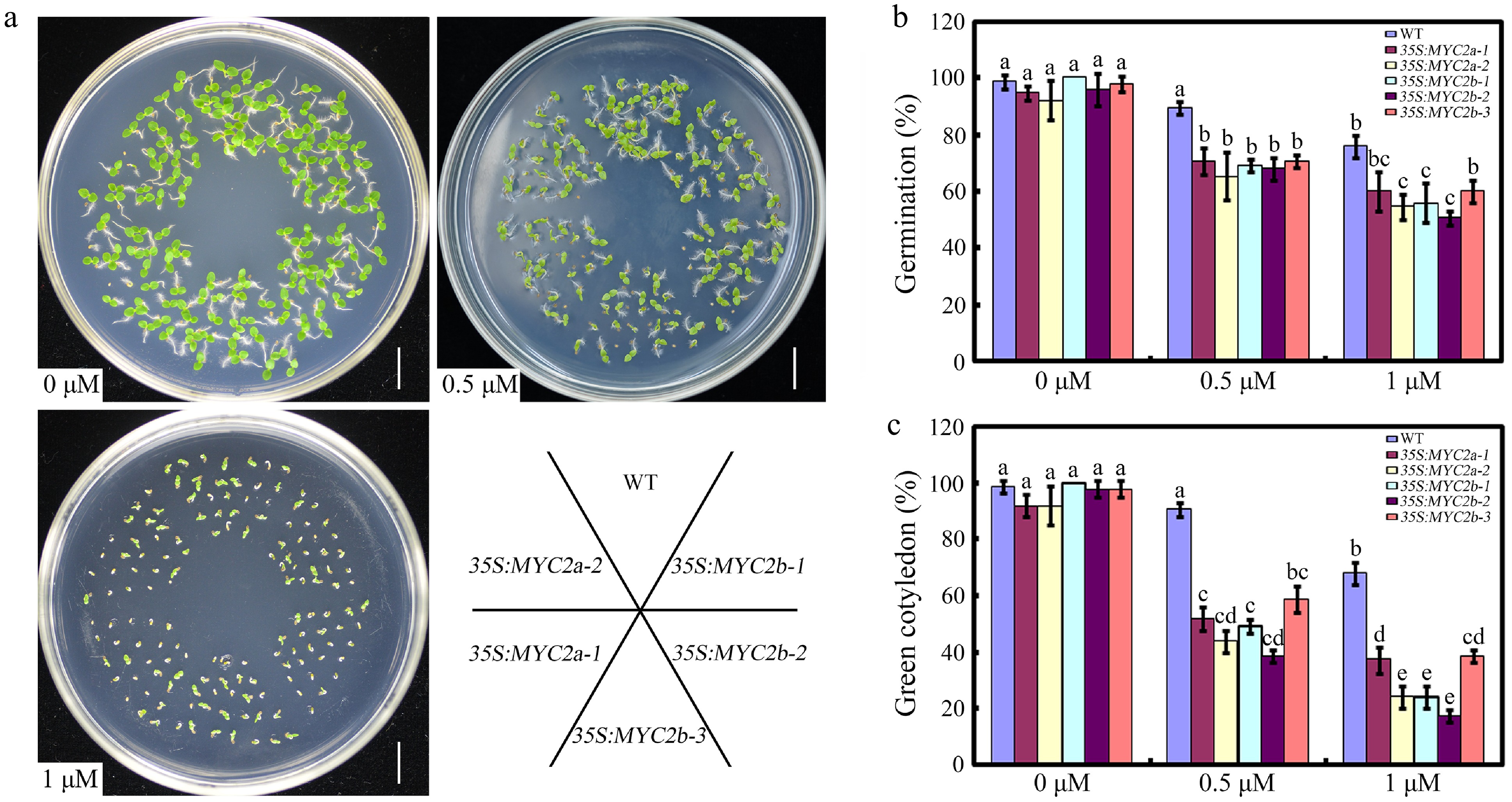
Figure 3.
The effect of NtMYC2a or NtMYC2b overexpression on seed germination. (a) Seedlings of the indicated genotypes observed 10 d after germination on medium supplemented with 0, 0.5 or 1 µM ABA. (b) Statistic analysis of germination rate in the indicated genotypes. The seeds were scored 8 d after stratification on medium containing different concentrations of ABA. Germination is defined by radical emergence. (c) Statistic analysis of the indicated genotypes for cotyledon greening. Cotyledon greening was scored 10 d after stratification on medium containing different concentrations of ABA. Error bars (mean ± SD) are derived from three biological replicates. Data were analyzed by a two-way ANOVA using Tukey's HSD test. Bars with different letters are significantly different from each other (p < 0.05).
NtMYC2a and NtMYC2b affect the expression of NtABI3 and NtABI5
-
Seeds of the Ntmyc2a/2b mutant or constitutive overexpression of NtMYC2a/2b showed contrasting phenotypes with respect to ABA mediated seed germination. This may indicate that MYC2a/2b may participate in this process by affecting the expression of germination associated genes. To test this possibility, we analyzed the expression of several ABA-related genes in NtMYC2a/2b mutant seeds or seeds of constitutive overexpression of NtMYC2a/2b by RT-qPCR. The results demonstrated that the expression levels of ABA signaling genes, including NtABI3 and NtABI5, decreased in Ntmyc2a/2b to different degrees compared with the WT, while enhanced in seeds that constitutively expressing NtMYC2a or NtMYC2b (Fig. 4). These results demonstrated that NtMYC2a/2b may participate in ABA mediated seed germination by affecting the expression of ABA signal transduction-related genes.
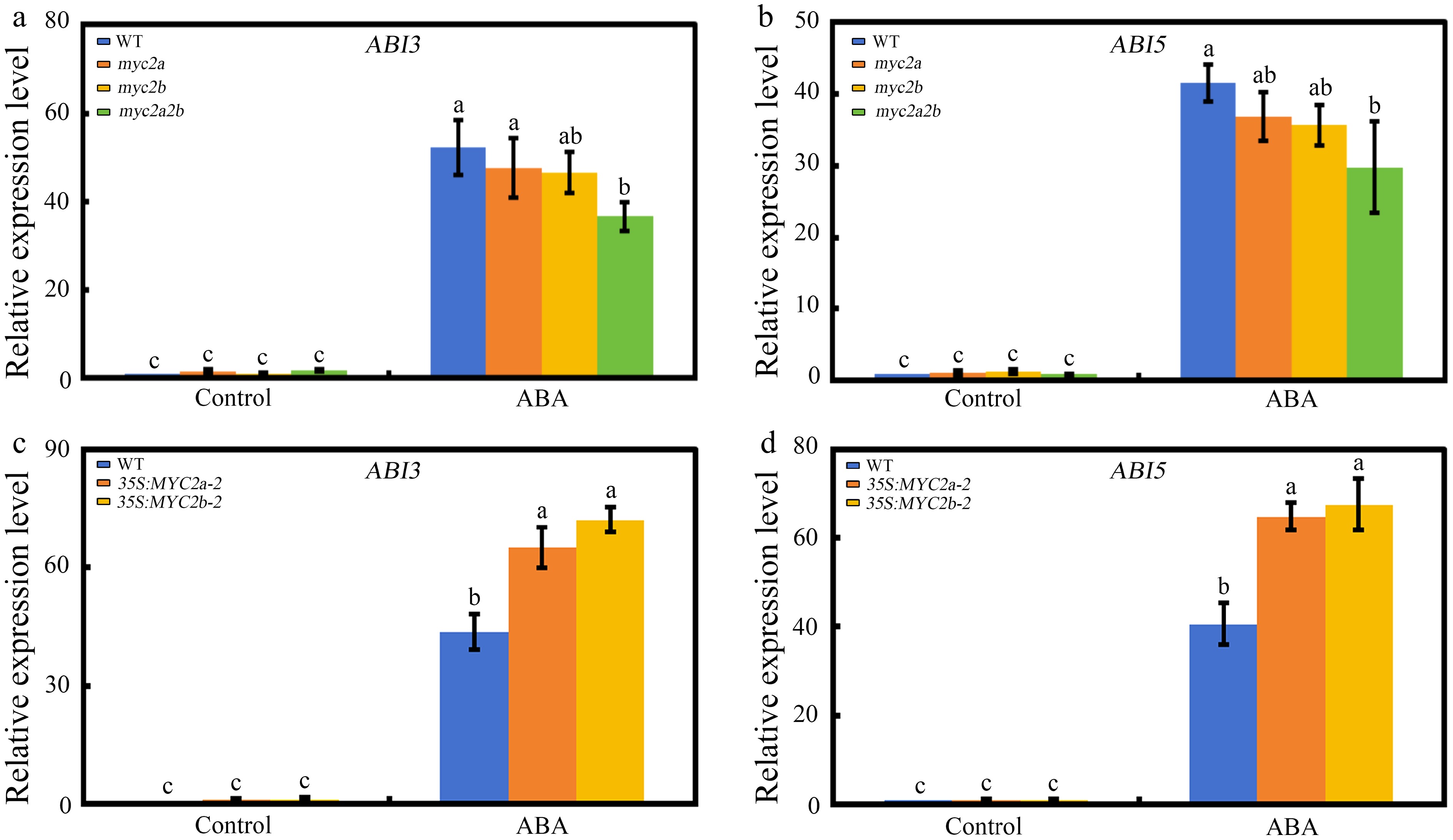
Figure 4.
The effect of NtMYC2a/2b mutation or overexpression on NtABI3 and NtABI5 expression. (a) and (b) Expression levels of NtABI3 and NtABI5 in Ntmyc2a and Ntmyc2b single mutant, Ntmyc2a2b double mutant and wild-type tobacco seeds grown on medium containing 1 µM ABA or not for 3 d. (c) and (d) Expression levels of NtABI3 and NtABI5 in NtMYC2a or NtMYC2b overexpression line and wild-type tobacco seeds grown on medium containing 1 µM ABA or not for 3 d. The data is shown as mean ± SD, error bars represent SD from three independent biological replicates, which were analyzed by a two-way ANOVA using Tukey's HSD test. Bars with different letters are significantly different from each other (p < 0.05).
NtMYC2a and NtMYC2b bind the promoter of NtABI5 to activate its expression
-
MYC2, as a bHLH transcription factor, was revealed to perform its biological functions through directly binding to the G-box element. Interestingly, one G-box element was found in the promoter of NtABI5, implying that NtMYC2a and NtMYC2b may bind to the G-box element to regulate NtABI5 expression. EMSA was performed to test whether NtMYC2a and NtMYC2b can directly bind to the G-box element. Incubation of NtMYC2a/2b proteins and the labeled probes resulted in a band shift, whereas inclusion of the unlabeled competitor DNA reduced the electrophoretic mobility shift in a dosage-dependent manner. Meanwhile, mutation of the G-box element in the probe abolished the band shift (Fig. 5a), confirming that NtMYC2a and NtMYC2b bind directly and specifically to the G-box element in the NtABI5 promoter.
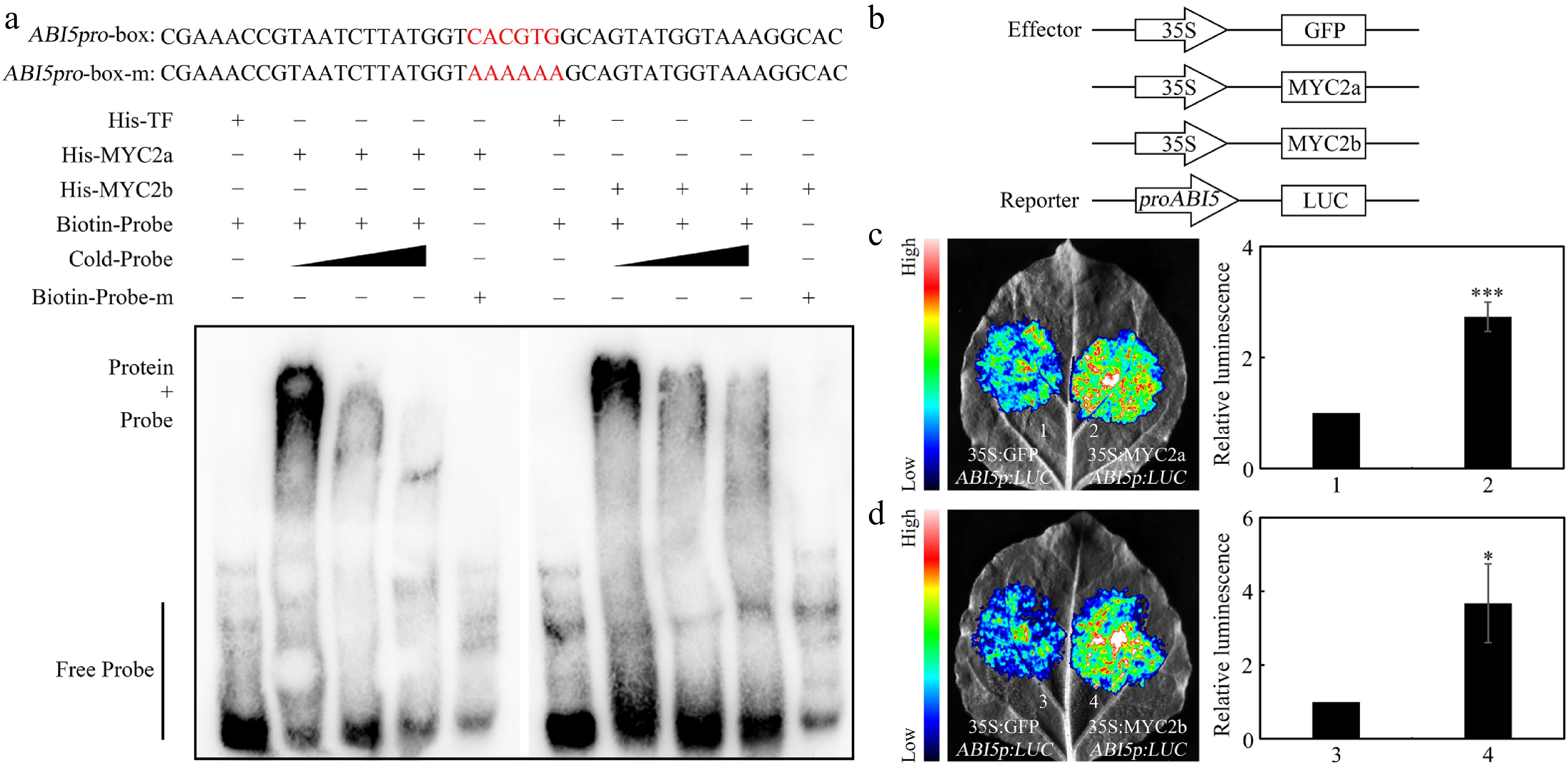
Figure 5.
NtMYC2a/2b bind the promoter of NtABI5 to activate its expression. (a) EMSA results revealed that NtMYC2a/2b can bind to the promoter of NtABI5. The probes were synthesized according to the G-box element within the NtABI5 promoter. The red sequence signifies G-box sequence and the mutated base in the G-box element. His, His-NtMYC2a/2b, biotin-probe, biotin-probe-m and cold probes were present (+), or absent (−) in each reaction. (b) Schematic of the NtABI5p:LUC reporter and NtMYC2a/2b and GFP effectors. (c) and (d) Transient transcriptional activity assays. NtMYC2a/2b could activate NtABI5 expression in N. benthamiana leaves. Both representative leaf images and quantification of corresponding relative luminescence intensities were presented. Error bars represent ± SD. Asterisks indicate student's t-test significant differences as compared to controls, * p < 0.05, *** p < 0.001.
Based on the above results that NtMYC2a/2b binds directly to the promoter of NtABI5, the next steps examined whether NtMYC2a/2b could directly modulate NtABI5 expression through transient expression analysis in N. benthamiana leaves using constructs in which the NtABI5 promoter sequence was fused to LUCIFERASE (LUC) as a reporter. LUC signals were significantly enhanced when NtMYC2a or NtMYC2b was co-expressed with the construct (Fig. 5b−d), further implying that both NtMYC2a and NtMYC2b can act as transcriptional activators to directly enhance NtABI5 expression during seed germination.
NtMYC2a interacts with NtMYC2b
-
Because NtMYC2a and NtMYC2b function redundantly during ABA mediated seed germination, and both of them can directly activate NtABI5 expression during this process, we speculate that they may interact with each other to synergistically modulate NtABI5 expression. In order to determine our prediction, we first employed AlphaFold and PyMOL to analyse the potential interaction between NtMYC2a and NtMYC2b (Fig. 6a). Then, the in vitro pull-down assay was performed to detect their interaction. The pull-down assay demonstrated that GST-NtMYC2a can pull down NtMYC2b-His, while the GST control did not (Fig. 6b). Interaction of NtMYC2a and NtMYC2b was further corroborated by split luciferase complementation (split-LUC) assays. NtMYC2a and NtMYC2b were fused to the N- and C-terminal halves of the firefly luciferase, respectively (nLUC-NtMYC2a and cLUC-NtMYC2b). When fused nLUC-NtMYC2a was co-infiltrated with cLUC-NtMYC2b into N. benthamiana leaves, strong luminescence was observed (Fig. 6b). However, no luminescence was detected in all negative controls. Taken together, this data demonstrates a physical interaction between NtMYC2a and NtMYC2b, suggesting their possible synergistic action as a heterodimer in diverse stress responses.
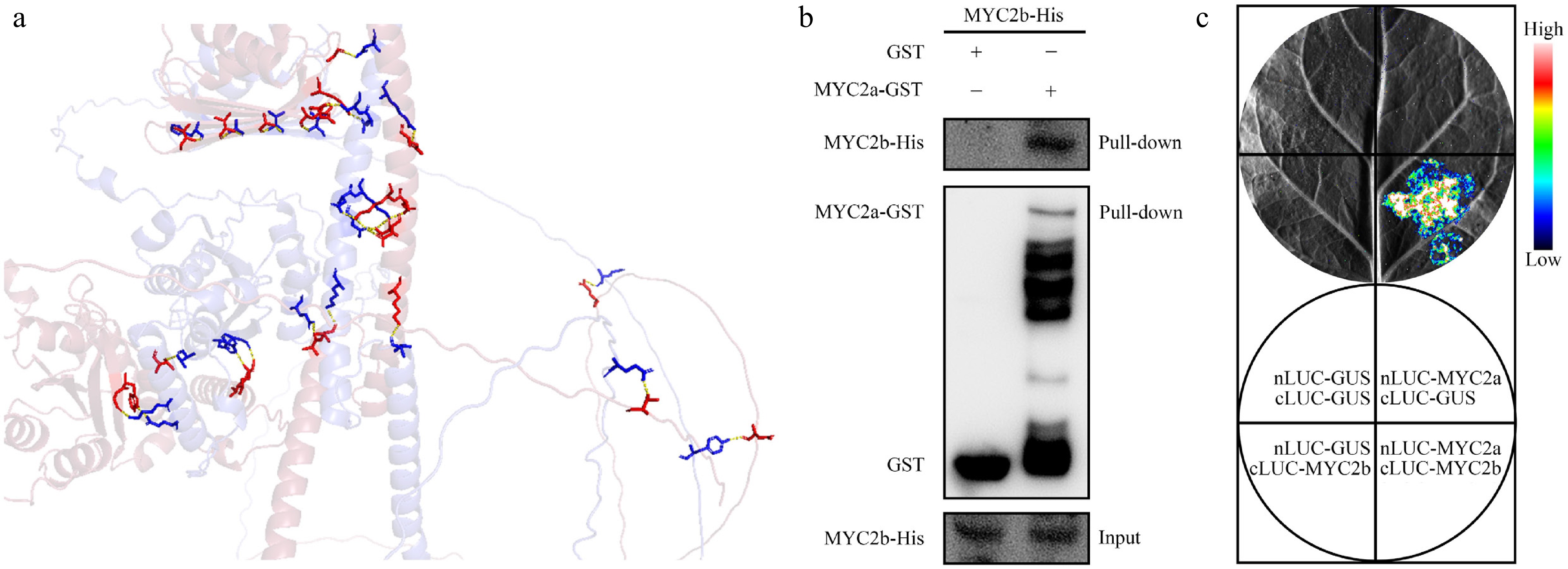
Figure 6.
Interaction between NtMYC2a and NtMYC2b. (a) The predicted cartoon diagram showing the interaction between NtMYC2a and NtMYC2b, as analysed via AlphaFold and PyMOL. The hydrogen bonds were shown as yellow dashed lines, the protein complex structures were shown as colored sticks, blue represents NtMYC2a, red represents NtMYC2b. (b) Pull-down assay confirming the interaction between NtMYC2a-GST and NtMYC2b-His. GST alone served as a negative control. (c) Split-luciferase complementation (split-LUC) assay confirming NtMYC2a-NtMYC2b interaction in N. benthamiana. A representative image of leaf taken 48 h after infiltration is shown, with luminescence intensity indicated by the pseudocolor bar.
NtMYC2a and NtMYC2b synergistically activate NtABI5 expression
-
Since NtMYC2a and NtMYC2b function redundantly in ABA-mediated seed germination, and they can interact with each other to form a heterodimer, it can be speculated that they may function synergistically to activate NtABI5 expression. To assess this possibility, a LUC reporter assay was performed to examine the effects of NtMYC2a on the transcriptional activity of NtMYC2b in N. benthamiana leaves. As shown in Fig. 7, co-expression of NtMYC2a with NtMYC2b increased LUC expression driven by the NtABI5 promoter, compared with the expression of NtMYC2a or NtMYC2b alone. These results suggest that NtMYC2a and NtMYC2b may function synergistically to promote NtABI5 expression during ABA-mediated seed germination.
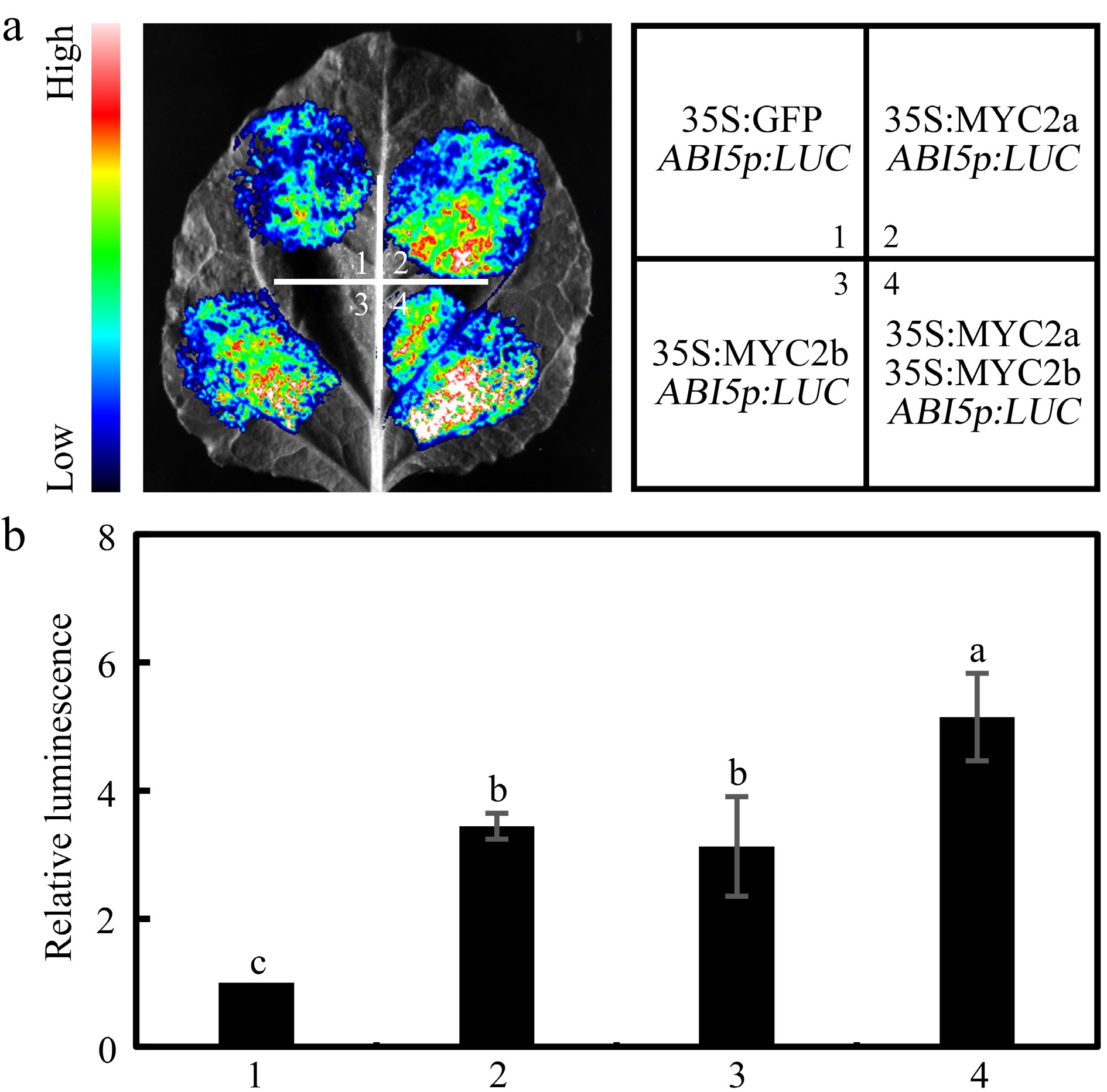
Figure 7.
NtMYC2a and NtMYC2b synergistically activate NtABI5 expression. Transient expression analysis in N. benthamiana leaves to show that NtMYC2a and NtMYC2b can synergistically activate NtABI5 expression. Both the (a) representative leaf image, and the (b) corresponding relative luminescence intensity, were presented. Three biological replicates data were analyzed by a one-way ANOVA using Tukey's HSD test. Bars with different letters are significantly different from each other (p < 0.05).
-
Seeds have evolved sophisticated gene-regulated networks to perceive favorable environmental conditions, such as moisture, temperature, and light quality, to determine timing of germination[3,5,6]. These cues then modulate the accumulation and perception of GA and ABA, leading to the promotion and repression of seed germination respectively[1]. Numerous studies have demonstrated that GA and ABA control seed germination in an antagonistic manner[30,31]. ABA, perceived by the PYR/RCAR–PP2C complex and transduced by ABA response transcription factors, accumulates during seed development and facilitates the seeds to enter a physiologically dormant state, which represents an adaptive strategy under unfavorable conditions. Identifying new components which function in ABA-mediated seed germination will help reveal the molecular basis of ABA signaling. In this study, we demonstrated that NtMYC2a/2b may function as critical components of ABA signaling during ABA-mediated seed germination in tobacco.
Numerous studies have demonstrated that MYC2 plays a critical role in diverse respects of plant growth and development, stress tolerance, and secondary metabolisms[23]. Despite its diverse roles, MYC2 mainly performs its biological functions by directly binding to the cis-acting G-box element (CACGTG), or G-box-like motifs (AACGTG or CATGTG) present in its target promoters[32−35]. In Tanacetum cinerariifolium, TcMYC2 binds to the E/G-box elements in the promoters of TcCHS, TcAOC, and TcGLIP and acts as a positive regulator of these pyrethrin biosynthesis genes upon methyl jasmonate (MeJA) treatment[36]. In Arabidopsis, AtMYC2 directly targets the jasmonate-induced LIPOXYGENASE2 to modulate jasmonate-induced anthocyanin accumulation and root growth inhibition[37]. Here, we also revealed that NtMYC2a/2b can directly target NtABI5 through the G-box in its promoter, and subsequently regulate ABA-mediated seed germination (Fig. 5). Based on the contrasting expression pattern of NtMYC2a/2b and NtABI5 in NtMYC2a/2b mutants and overexpression plants, together with the enhanced LUC intensity in transient expression assays, we concluded that NtMYC2a/2b act as positive regulators of NtABI5. Thus, our results implied that NtMYC2a and NtMYC2b may act upstream of NtABI5 to positively mediate the ABA-mediated germination process.
Protein-protein interaction represents an important mechanism for the functional performance of proteins. Upon receipt of various signals, plants usually amplify them through ordered interactions between proteins, thereby triggering various physiological responses[38]. Numerous studies have demonstrated that MYC2 can interact with various types of proteins which function as key components of various signaling pathways to mediate various plant developmental processes, and both biotic and abiotic stresses[23]. For example, NtMYC2a and NtMYC2b can interact with NtJAZ1 and modulate multiple jasmonate-inducible genes in the nicotine biosynthetic pathway[26]. Arabidopsis AtMYC2 physically associates with AtPHR1 to synergistically enhance Pi deficiency-activated jasmonate signaling[37]. Thus, it will be interesting to further determine whether certain proteins can interact and form complexes with MYC2 to modulate its DNA binding ability, or transcriptional activation, or repression ability. In this study, NtMYC2a and NtMYC2b was found to interact with each other to form a functional heterodimer and synergistically activate NtABI5 expression, suggesting that they may function together during ABA-mediated seed germination. Identification of their interacting partners can contribute to the elucidation of NtMYC2-mediated ABA responses during seed germination.
In this study, an interesting finding is that NtMYC2a and NtMYC2b can interact with each other to form a hetreodimer and synergistically activate NtABI5 expression during ABA-mediated seed germination. This finding may be contradictory to the inhibited expression patterns of NtMYC2a and NtMYC2b upon ABA treatment. Furthermore, ABA can also attenuate the interaction between NtMYC2a and NtMYC2b (Supplementary Fig. S3). These findings suggested that seeds may actively reduce the RNA and protein levels of NtMYC2a and NtMYC2b and also their interaction strength during seed germination, which may contribute to the avoidance of over accumulation of NtABI5 transcripts under high ABA levels. It may represent a feedback regulation of ABA signaling on the NtABI5 transcripts through the regulation of NtMYC2a and NtMYC2b expression and interaction. However, this hypothesis needs to be further determined in the future.
Seeds are irreplaceable genetic resources, and their rapid and synchronous germination and subsequent seedling establishment are the prerequisites of successful agricultural production and sustainable agriculture[39]. This study has revealed the molecular mechanisms underlying the regulation of ABA-mediated seed germination by NtMYC2a/2b in tobacco. Our results demonstrated that NtMYC2a/2b may act as critical components of the seed germination regulatory network and participates in the ABA-mediated seed germination process in tobacco. It will be interesting to further investigate how NtMYC2's transcription and translation is regulated during seed germination processes. These results are consistent with the role of AtMYC2 in ABA-mediated seed germination[40], implying that the function of MYC2 in different plant species may be conserved. Considering the role of NtMYC2a/2b in ABA-mediated seed germination, it will also be interesting to determine their possible involvement in plant abiotic stresses in the future. The molecular mechanisms revealed in this study contribute to the understanding of the regulatory mechanism of MYC2 in plants.
-
The authors confirm their contributions to the paper as follows: designed the research: Chen L, Yao H; performed experiments: Jing Y, Wang R, Zhao Z, Shi L, Wang B, Yang D, Pan W; analyzed data: Jing Y, Wang R, Chen L; wrote the paper: Chen L. All authors reviewed the results and approved the final version of the manuscript.
-
The original contributions presented in the study are included in the article or supplementary material, further inquiries can be directed to the corresponding author/s.
-
The authors declare that they have no conflicts of interest with the contents of this article.
-
# Authors contributed equally: Yifen Jing, Ruling Wang
- Supplementary Table S1 Primers used for RT-qPCR.
- Supplementary Table S2 Primers used for generating various clones.
- Supplementary Fig. S1 Generation of Ntmyc2a mutant using CRISPR-Cas9 system.
- Supplementary Fig. S2 Generation of Ntmyc2b mutant using CRISPR-Cas9 system.
- Supplementary Fig. S3 ABA suppresses the interaction between NtMYC2a and NtMYC2b.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of Hainan Yazhou Bay Seed Laboratory. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Jing Y, Wang R, Zhao Z, Shi L, Wang B, et al. 2026. The NtMYC2-NtABI5 module regulates abscisic acid mediated seed germination in tobacco. Seed Biology 5: e012 doi: 10.48130/seedbio-0026-0008
The NtMYC2-NtABI5 module regulates abscisic acid mediated seed germination in tobacco
- Received: 28 October 2025
- Revised: 04 January 2026
- Accepted: 21 January 2026
- Published online: 13 April 2026
Abstract: In tobacco, the basic helix-loop-helix (bHLH) transcription factor MYC2 contributes significantly to the biosynthesis and transport of nicotine, and also participates in the regulation of pollen development. However, their biological functions in seed germination remain largely unknown. Here, we examined the regulatory roles of NtMYC2a and NtMYC2b in abscisic acid (ABA) mediated seed germination. The expression of both NtMYC2a and NtMYC2b was repressed by ABA at both mRNA and protein levels. Simultaneous knock out of NtMYC2a and NtMYC2b reduced ABA signaling to promote seed germination, while overexpression of NtMYC2a or NtMYC2b resulted in the germinated seeds being more ABA-sensitive. Gene expression analysis showed that the expression of both NtABI3 and NtABI5 was reduced in Ntmyc2a/2b double mutants, but enhanced in transgenic plants overexpressing NtMYC2a or NtMYC2b. Both electrophoretic mobility shift assays (EMSA) and transient dual-luciferase reporter assays, revealed that both NtMYC2a and NtMYC2b could bind to the promoter of NtABI5 to activate its expression. Furthermore, NtMYC2a and NtMYC2b could interact with each other to form a heterodimer and function synergistically to activate the expression of NtABI5. Thus, these results provide evidence that the NtMYC2-NtABI5 module may function as a novel component in ABA signaling to positively modulate ABA-mediated seed germination in tobacco.
-
Key words:
- Tobacco /
- NtMYC2 /
- NtABI5 /
- Abscisic acid /
- Seed germination














