-
Flavonoids are a class of specialized metabolites widely distributed throughout the plant kingdom, serving an essential role in both plant defense and human health owing to their distinctive chemical structures and diverse biological activities[1,2]. In plants, flavonoids provide protection against a broad range of biotic and abiotic stresses, including ultraviolet radiation, low temperature, drought, salinity, and pest attack[3−9]. In addition to their ecological functions, flavonoids exhibit notable pharmacological properties, such as antioxidant, antimicrobial, cardioprotective, and neuroprotective activities[10]. The flavonoid biosynthetic pathway responsible for flavonoid biosynthesis is evolutionarily conserved across plant species and relies on the coordinated action of several key enzymes, including chalcone synthase (CHS), chalcone isomerase (CHI), and flavonol synthase (FLS)[11]. Among these, CHS acts as the first rate-limiting enzyme and has therefore been extensively investigated in diverse plant taxa, such as maize, sorghum, petunia, orchids, and A. thaliana[12]. Functional studies in non-model plants have further validated the central role of CHS in flavonoid accumulation. For instance, heterologous expression of ScCHS1 from Stenoloma chusanum partially complemented the anthocyanin-deficient phenotype of the A. thaliana tt4 mutant[13]. Similarly, overexpression of CHS in Glycyrrhiza uralensis and Camellia sinensis resulted in significant increases in flavonoid content[14,15]. Conversely, suppression of CHS expression via RNA interference and related approaches markedly impairs flavonoid biosynthesis and leads to visible pigmentation changes. In Chinese cabbage, silencing BrCHS4 induced a transition from purple to green leaves and was accompanied by substantial reductions in flavonoid and anthocyanin levels[16]. Collectively, these findings highlight the indispensable role of CHS in flavonoid biosynthesis and provide a theoretical basis for the genetic manipulation of plant secondary metabolism.
Despite these advances, several challenges remain. Although considerable progress has been made in deciphering the transcriptional regulation of CHS genes, particularly through the identification of transcription factors such as MYB and bHLH that directly activate CHS expression, the regulatory roles of non-coding RNAs remain largely unexplored[17−19]. Recent evidence indicates that 22-nt small interfering RNAs (siRNAs) in soybean regulate seed coat color by targeting CHS genes[20]. However, the involvement of long non-coding RNAs (lncRNAs) or other non-coding RNA types in this process remains unclear. Furthermore, accumulating evidence suggests that CHS exhibits functional versatility beyond flavonoid biosynthesis. In addition to its metabolic role, CHS has been reported to negatively regulate endogenous auxin transport and to influence developmental processes such as floral organ morphogenesis[21,22]. Collectively, these observations imply that CHS may participate in a diverse physiological process through complex regulatory circuits, highlighting the necessity for a more comprehensive investigation of its regulatory networks and functional breadth.
The regulatory functions of lncRNAs in flavonoid biosynthesis are gradually being elucidated, though existing studies have mainly centered on indirect regulatory mechanisms. As an illustration, studies in apple have shown that the light-induced transcription factor MdWRKY1 binds to the promoter of MdLNC409 and subsequently modulates MdERF109 via this lncRNA to enhance anthocyanin biosynthesis[23]. Similarly, another light-induced lncRNA, MdLNC610, enhances the promoter activity of MdACO1, leading to increased ethylene production and elevated anthocyanin accumulation[24]. A particularly prevalent regulatory paradigm involves endogenous target mimics, in which lncRNAs function as competitive decoys for microRNAs (miRNAs), indirectly modulating the expression of downstream target genes. In sea buckthorn, for example, LNC1 and LNC2 act as endogenous target mimics by sequestering miR156a and miR828a, respectively[25]. A comparable mechanism has been described in apple, where the lncRNAs MLNC3.2 and MLNC4.6 bind miR156a, thereby blocking its cleavage of the target genes SPL2 and SPL33. The resulting SPL2/33 proteins subsequently form a transcriptional activation complex with the key anthocyanin regulator MdMYB1, which binds to the MdCHS promoter and activates the expression of downstream structural genes such as MdDFR and MdUFGT, ultimately enhancing light-induced anthocyanin accumulation[26]. By contrast, direct regulatory roles of lncRNA in flavonoid biosynthesis appear to be rare. One notable example is the intergenic LINC15957 in radish, which has been proposed to promote anthocyanin accumulation through the regulation of transcription factors; nevertheless, its precise target genes and molecular mechanism remain unresolved[27]. Taken together, existing studies predominantly demonstrated that intergenic lncRNAs regulate flavonoid biosynthesis indirectly, most commonly via endogenous target mimicry or transcriptional modulation. Whether antisense lncRNAs (lncNATs) can directly regulate flavonoid biosynthesis by physically interacting with structural genes such as CHS has not yet been reported.
As a living fossil plant with a distinctive evolutionary history, G. biloba also possesses substantial economic value. Extracts originating from G. biloba leaves are widely used in the prevention and treatment of cerebrovascular disorders and also show therapeutic potential in the clinical management of neurodegenerative disorders[28]. Among the bioactive constituents of G. biloba extracts, flavonoids represent the most abundant and pharmacologically valuable components. Consequently, characterizing the core genes that govern flavonoid biosynthesis in G. biloba is critically important for enhancing flavonoid accumulation and improving resource utilization. To date, numerous structural enzyme genes involved in flavonoid biosynthesis, as well as key regulatory factors such as MYB, bHLH, and bZIP transcription factors, have been identified in G. biloba through high-throughput sequencing[29−32]. However, the functional validation of these genes and the demonstration of their precise regulatory mechanisms in flavonoid biosynthesis remain inadequately explored. Furthermore, transcriptomic analyses of G. biloba leaves have suggested that lncRNAs may influence flavonoid biosynthesis by modulating the expression of transcription factors or flavonoid biosynthesis genes[33]. More recently, an integrative analysis combining PacBio third-generation and Illumina second-generation sequencing across eight G. biloba tissues predicted that five lncRNAs are potentially involved in the regulation of flavonoid biosynthesis[34]. However, direct experimental evidence demonstrating whether these lncRNAs can physically interact with flavonoid biosynthetic enzyme genes and directly modulate their function remains lacking.
In this present study, in order to identify key enzyme genes and lncRNAs associated with flavonoid biosynthesis in G. biloba, the study surveyed 26 cultivars and screened for individuals exhibiting high and low flavonoid contents. Leaves from three cultivars displaying pronounced differences in flavonoid accumulation were subsequently selected for integrated metabolomic and strand-specific RNA sequencing (ssRNA-seq) analyses. Through comprehensive data integration, a flavonoid-related LncNAT1-GbCHS regulatory module was identified. LncNAT1 is a type of lncNAT that partially overlaps with the GbCHS gene and suppresses its expression through sequence-specific binding, thereby negatively regulating flavonoid biosynthesis. In addition, the results demonstrate that GbCHS functions not only as a key enzyme in flavonoid biosynthesis but also plays broader roles in plant growth and development, including the regulation of lateral root formation, branching architecture, and leaf production. Collectively, this study reveals a previously unrecognized mechanism by which an lncNAT regulates flavonoid biosynthesis through its interaction with a structural enzyme gene and uncovers a novel developmental function of GbCHS. These findings provide valuable genetic insights for the improvement of G. biloba leaf yield and flavonoid accumulation in future breeding programs.
-
Plant materials were obtained from a G. biloba germplasm resource nursery located in Pizhou, Jiangsu Province, China. Leaves from 26 G. biloba cultivars of comparable age and grown under uniform environmental conditions were collected from the nursery. Detailed information for each cultivar is provided in Supplementary Table S1. Among these, leaves from the cultivars designated 'DHS', 'DTH' and 'HBAL' were selected for subsequent metabolomic profiling and high-throughput sequencing analyses.
Determination of flavonoids and flavonols
-
Flavonol compounds were quantified using high-performance liquid chromatography (HPLC) following a previously described protocol[33]. In addition, total flavonoid content was determined using a commercial plant flavonoid assay kit.
Widely targeted metabolic profiling
-
Freeze-dried leaf samples were ground into a fine powder, and 100 mg of each sample was extracted with 70% (v/v) aqueous methanol. The extracts were incubated at 4 °C overnight, with intermittent vortexing to enhance extraction efficiency. Widely targeted metabolomic profiling was performed using ultra-performance liquid chromatography coupled with tandem mass spectrometry (UPLC-MS/MS), as described previously[35]. Differentially accumulated metabolites (DAMs) were identified based on a variable importance in projection (VIP) ≥ 1 and a fold change ≥ 2 or ≤ 0.5.
lncRNA identification and ssRNA-seq
-
Total RNA was extracted from leaf tissues of the three selected cultivars ('DHS', 'DTH', and 'HBAL') using the RNAprep Pure Plant Kit following the manufacturer’s protocol. A total of nine strand-specific cDNA libraries were constructed and subjected to high-throughput sequencing. Clean reads obtained from each library were aligned to the G. biloba genome using Hisat2, followed by transcript assembly with StringTie[36]. LncRNAs were identified using an established pipeline[33], which included filtering based on transcript length, annotation status, and coding potential. This potential was assessed using CPC2, Pfam, and CNCI[37,38].
Differential expression analysis of lncRNAs and mRNAs
-
Differentially expressed lncRNAs (DELs) and mRNAs (DEGs) were identified using the DESeq2 package[39], with an adjusted p-value < 0.05 and an absolute log2 fold change > 1 set as the significance thresholds. Functional enrichment analysis was subsequently conducted, with GO enrichment performed using the GOseq tool[40], while KEGG pathway enrichment was assessed using the clusterProfiler 4.0 software[41,42].
Gene cloning, vector construction, and GbCHS enzyme activity assays
-
The full-length sequences of LncNAT1 and the coding sequence (CDS) of GbCHS were amplified and cloned into the overexpression vector pRI101 through the ClonExpress II One Step Cloning Kit. Using the same strategy, GbCHS was also inserted into the pACT2-LUC and pET-32a vectors. For enzymatic characterization, the pET-32a-GbCHS construct was transformed into Escherichia coli BL-21 (DE3) cells, and recombinant protein expression was induced with 0.2 mM IPTG at 16 °C for 16 h. The resulting fusion protein was purified following a reported protocol[43] and verified by SDS-PAGE. Enzyme activity assays were conducted by incubating purified GbCHS in a reaction mixture comprising 160 μM p-coumaroyl-CoA, 480 μM malonyl-CoA, and 0.1 M potassium phosphate buffer. Three different amounts of purified protein (120, 150, and 180 μg) were assessed individually at 30 °C for 1 h. After incubation, reaction mixtures were extracted twice with ethyl acetate, centrifuged, evaporated under nitrogen gas, and resuspended in methanol. Heat-inactivated pET-32a-GbCHS protein was used as a negative control.
qRT-PCR analysis
-
For qRT-PCR analysis, GAPDH served as an internal control gene[30]. Gene-specific primers were designed with Primer Premier 5.0 and are listed in the Supplementary Table S2. Amplification was performed on a CFX96™ Real-Time PCR System using ChamQ SYBR qPCR Master Mix. Relative expression levels were calculated using the 2−ΔΔCᴛ method[44].
Genetic transformation of A. thaliana plants and G. biloba calli
-
The 35S::GbCHS construct was first introduced into Agrobacterium tumefaciens strain GV3101 and used to transform A. thaliana via the floral dip method. Seeds harvested from transformed plants were germinated on kanamycin-containing medium, and PCR analysis was performed to confirm transgene integration. PCR-positive transgenic plants were selected for subsequent analyses, including gene expression assessment, phenotypic observation, and determination of flavonoid content. Additionally, the 35S::LncNAT1 and 35S::GbCHS vectors were transformed into G. biloba calli following a previously described method[45]. Transgenic calli were used for gene expression analysis and flavonoid quantification.
Genetic transformation of G. biloba calli
-
The 35S::LncNAT1 and 35S::GbCHS constructs were introduced into A. tumefaciens strain EHA105 and subsequently used to infect G. biloba calli according to established protocols[29].
Virus-induced gene silencing (VIGS) transformation
-
Specific 300-bp fragments of the LncNAT1 and GbCHS sequences were amplified using designed primers (Supplementary Table S2) and cloned into the TRV vector. The resulting constructs were transformed into A. tumefaciens strain GV3101 and used for VIGS in seedlings following previously reported procedures[45].
Dual-luciferase reporter assay and RNase protection assay
-
For the dual-luciferase reporter (DLR) assays, protoplasts were isolated from poplar leaves as previously described[46]. Different plasmid combinations of effector and reporter plasmids, pRI101-GFP with GbCHS-LUC, or 35S::LncNAT1 with GbCHS-LUC, were co-transferred into these protoplasts. Firefly luciferase (LUC) and Renilla luciferase (REN) activities were quantified with a GLO-MAX® 20/20 Luminometer. In parallel, the same effector and reporter constructs were co-infiltrated into leaves of Nicotiana benthamiana, and luminescence signals were visualized using a Tanon-5200 in vivo plant imaging system.
RNase protection assay (RPA) was performed as previously described[47]. Briefly, total RNA extracted from G. biloba tissues was treated with RNase A in a digestion buffer to degrade single-stranded RNA. The remaining RNA was subjected to RT-PCR and agarose gel electrophoresis to evaluate the formation of the LncNAT1-GbCHS duplex and its protective effect against RNase degradation.
Measurement of photosynthetic rate
-
Photosynthetic parameters of rosette leaves from A. thaliana plants were measured using a portable gas-exchange system. Measurement conditions were set to 24 °C, a photosynthetically active radiation of 1,100 μmol photons/(m2·s), an ambient CO2 of 400 μmol/mol, and a relative humidity of 65%. Five biological replicates (one plant per replicate) were analyzed for each genotype.
-
Quantitative analysis of leaf samples from 26 G. biloba cultivars revealed substantial variation in total flavonoid contents (Fig. 1a). Among these cultivars, 'HBAL' exhibited the highest accumulation (36.46 mg/g), followed by 'DTH' (29.88 mg/g) and 'ZJCX' (23.14 mg/g). In contrast, 'YL' showed the lowest accumulation (10.15 mg/g), with 'DHS' (10.59 mg/g) and 'HBAL-3' (11.19 mg/g) also displaying relatively low levels. Overall, the total flavonoid content in 'HBAL' was approximately 3.45-fold higher than that in 'YL' (Fig. 1a).
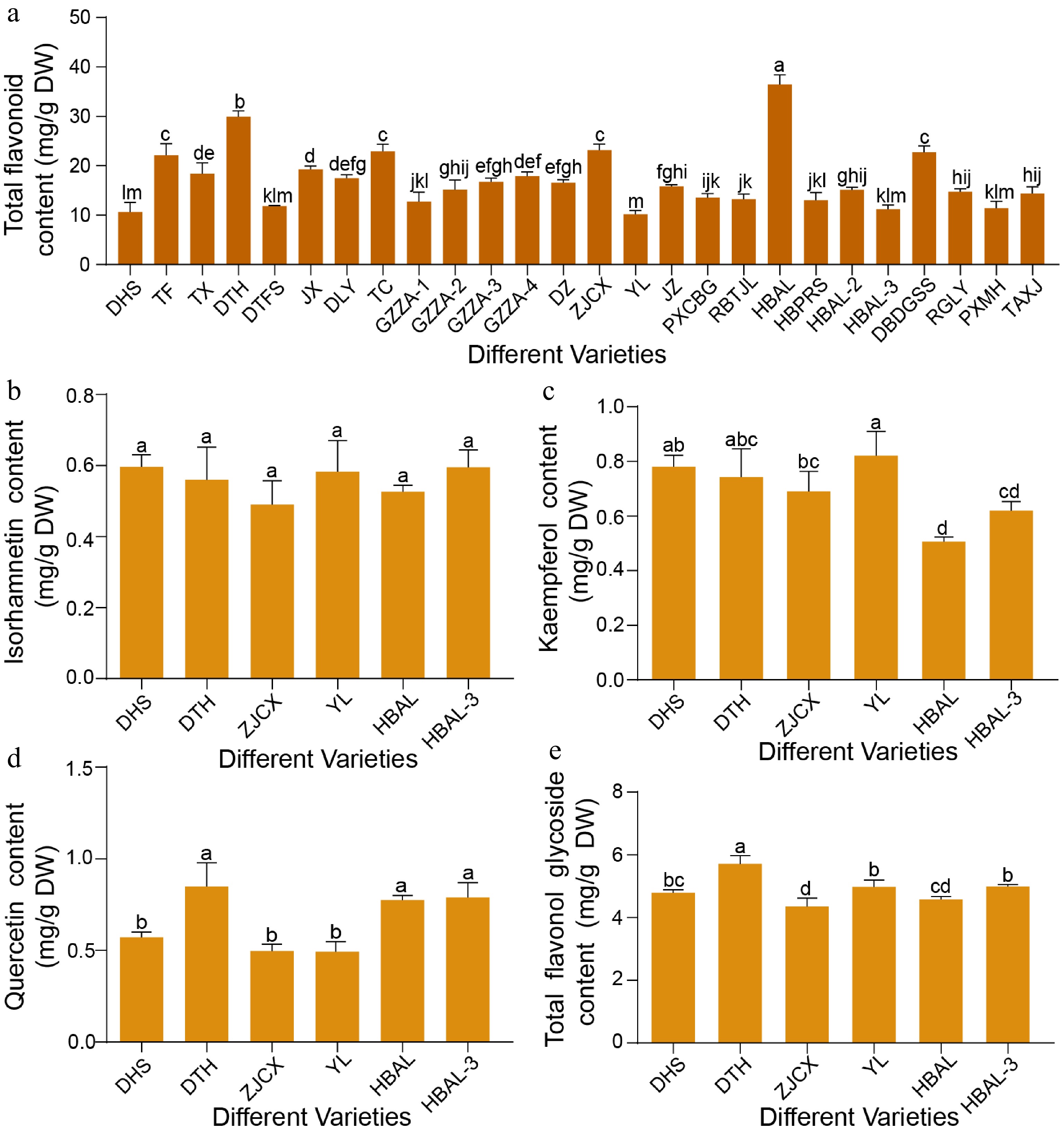
Figure 1.
Variation in flavonoid content among different G. biloba cultivars. (a) Total flavonoid levels in leaves of 26 G. biloba cultivars. (b) Isorhamnetin, (c) kaempferol, (d) quercetin, and (e) total flavonol glycoside contents in leaves of six selected cultivars. Statistical significance was determined by one-way ANOVA. Different lowercase letters indicate significant differences (p < 0.05). Data are presented as mean ± SD (n = 3 biological replicates).
Given that flavonol glycosides, including kaempferol, quercetin, and isorhamnetin, are the major bioactive constituents of G. biloba leaves, their contents were further quantified in six representative cultivars. Isorhamnetin content remained relatively stable across the six cultivars ('HBAL', 'DTH', 'ZJCX', 'YL', 'DHS', and 'HBAL-3') (Fig. 1b). In contrast, kaempferol content varied markedly, with 'YL' exhibiting the highest level (Fig. 1c). Quercetin content also differed significantly among cultivars, reaching a maximum of 0.85 mg/g in 'DTH' (Fig. 1d). Consistently, 'DTH' displayed the highest total flavonol glycoside content (5.71 mg/g). In contrast, the lowest quercetin content (0.57 mg/g) was observed in 'DHS', which also showed a comparatively low total flavonol glycoside level of 4.89 mg/g (Fig. 1e).
Metabolomic analysis reveals distinct flavonoid profiles and pathway enrichment in G. biloba cultivars
-
To explore differences in flavonoid composition, leaves from three cultivars with distinct flavonoid profiles were selected: 'HBAL', which exhibited the highest total flavonoid content, 'DTH', which showed the highest flavonol glycoside accumulation, and 'DHS', which displays the lowest levels of both total flavonoids and flavonol glycosides. Totally, the analysis detected 976 metabolites, predominantly flavonoids (22.8%), phenolic acids (18.2%), lipids (17.1%), amino acids and their derivatives (9.2%), organic acids (7.4%), and other compound classes (8.4%) (Supplementary Table S3). DAM analysis identified 226 DAMs between 'DTH' and 'DHS', including 168 up-accumulated and 58 down-accumulated metabolites; 315 DAMs between 'HBAL' and 'DHS' (206 up- and 109 down-accumulated); and 181 DAMs between 'HBAL' and 'DTH' (97 up, 84 down) (Supplementary Fig. S1).
KEGG pathway enrichment analysis of the DAMs revealed that metabolites showing differential accumulation in all three pairwise comparisons ('DTH' vs 'DHS', 'HBAL' vs 'DHS', and 'DTH' vs 'HBAL') were predominantly enriched in pathways including flavonoid biosynthesis as well as flavone and flavonol biosynthesis pathways (Fig. 2a−c). Accordingly, further focus was placed on flavonoid-related DAMs, and a total of 125 differentially accumulated flavonoid metabolites were identified and classified into various categories: isoflavones, flavonols, flavones, dihydroflavonols, dihydroflavones, chalcones, biflavones, and anthocyanins. With the exception of anthocyanins, the majority of flavonoid subclasses exhibited higher accumulation in 'HBAL' or 'DTH' compared with 'DHS' (100%, 3/3), flavonols (73%, 27/37), flavones (69%, 31/45), flavanols (75%, 3/4), dihydroflavonols (100%, 4/4), dihydroflavones (100%, 6/6), chalcones (100%, 3/3), and biflavones (100%, 15/15) were higher in the 'HBAL' or 'DTH' cultivars compared to the 'DHS' cultivar. In contrast, 88% (7/8) of anthocyanin metabolites accumulated at higher levels in 'DHS' than in both 'DTH' and 'HBAL' (Fig. 2d, Supplementary Fig. S2). Correlation analysis further supported this divergence in flavonoid composition. Flavonols, isoflavones, flavones, dihydroflavonols, dihydroflavones, chalcones, and biflavones exhibited strong positive correlations with one another, whereas anthocyanins showed a negative correlation with these flavonoid subclasses (Fig. 2e).
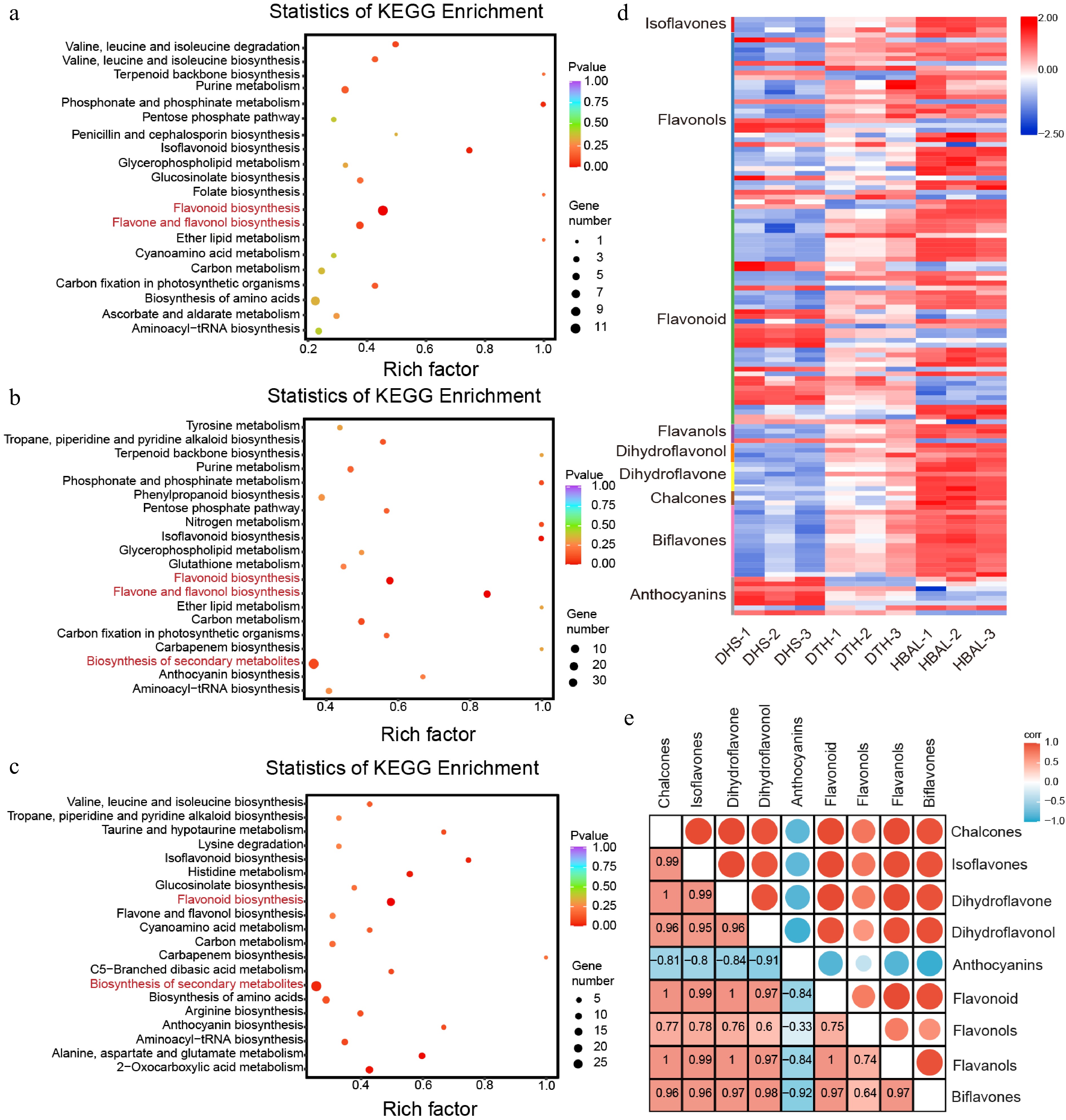
Figure 2.
Metabolomic analysis of three G. biloba cultivars. KEGG pathway enrichment analysis of DAMs in the comparisons between (a) 'DTH' and 'DHS', (b) 'HBAL' and 'DHS', and (c) 'DTH' and 'HBAL'. The rich factor represents the proportion of DAMs mapped to a given pathway relative to the total number of annotated metabolites in that pathway, and the circle size indicates the number of DAMs. (d) Distribution of differentially accumulated flavonoid metabolites among cultivars. (e) Correlation analysis between anthocyanins and other flavonoid subclasses.
Integration of transcriptome and metabolome identifies key genes in flavonoid biosynthesis of G. biloba
-
ssRNA-seq was conducted on leaf tissues from the three cultivars exhibiting marked differences in flavonoid accumulation. In total, 3,090 DEGs were identified across all pairwise comparisons. Specifically, 689 DEGs (318 upregulated, 371 downregulated) were identified in the 'DTH' vs 'DHS' comparison, while 'HBAL' vs 'DHS' showed 2,195 DEGs (1,418 upregulated, 777 downregulated), and 'DTH' vs 'HBAL' contained 1,545 DEGs (485 upregulated, 1,060 downregulated) (Supplementary Fig. S3). Additionally, there were 249 shared DEGs between the 'DTH' vs 'DHS' and 'DTH' vs 'HBAL' comparison groups, 300 shared DEGs between the 'DTH' vs 'DHS' and 'HBAL' vs 'DHS' comparison groups, 834 shared DEGs between the 'DTH' vs 'HBAL' and 'HBAL' vs 'DHS' comparison groups, and 44 DEGs consistently present across all three comparison sets (Supplementary Fig. S4). Integrative analysis of DEGs and DAMs revealed significant enrichment in pathways related to flavonoid biosynthesis as well as flavone and flavonol biosynthesis pathways (Supplementary Figs S5−S7).
Further examination identified 15 DEGs encoding key enzymes in the flavonoid biosynthesis pathway, including one GbCHS gene (Gb_19002), two GbDFR genes (Gb_26470, Gb_26459), two GbANR genes (Gb_09086, Gb_09087), two GbF3'H genes (Gb_32737, Gb_02188), four GbFLS genes (Gb_24242, Gb_14030, Gb_14029, Gb_14031), one GbOMT gene (Gb_18171), and three GbANS genes (Gb_06948, Gb_21870, Gb_33402). Except for one GbANS gene (Gb_33402) and two GbANR genes, the remaining genes exhibited higher expression levels in the high-flavonoid cultivar 'HBAL' or the high-flavonol cultivar 'DTH' (Fig. 3a). Transcript-metabolite correlation analysis further demonstrated that the expression profiles of GbCHS (Gb_19002), GbF3'H (Gb_32737, Gb_02188), and GbFLS (Gb_24242, Gb_14030, Gb_14029, Gb_14031) were closely aligned with the accumulation patterns of their corresponding flavonoid products (Fig. 3a).
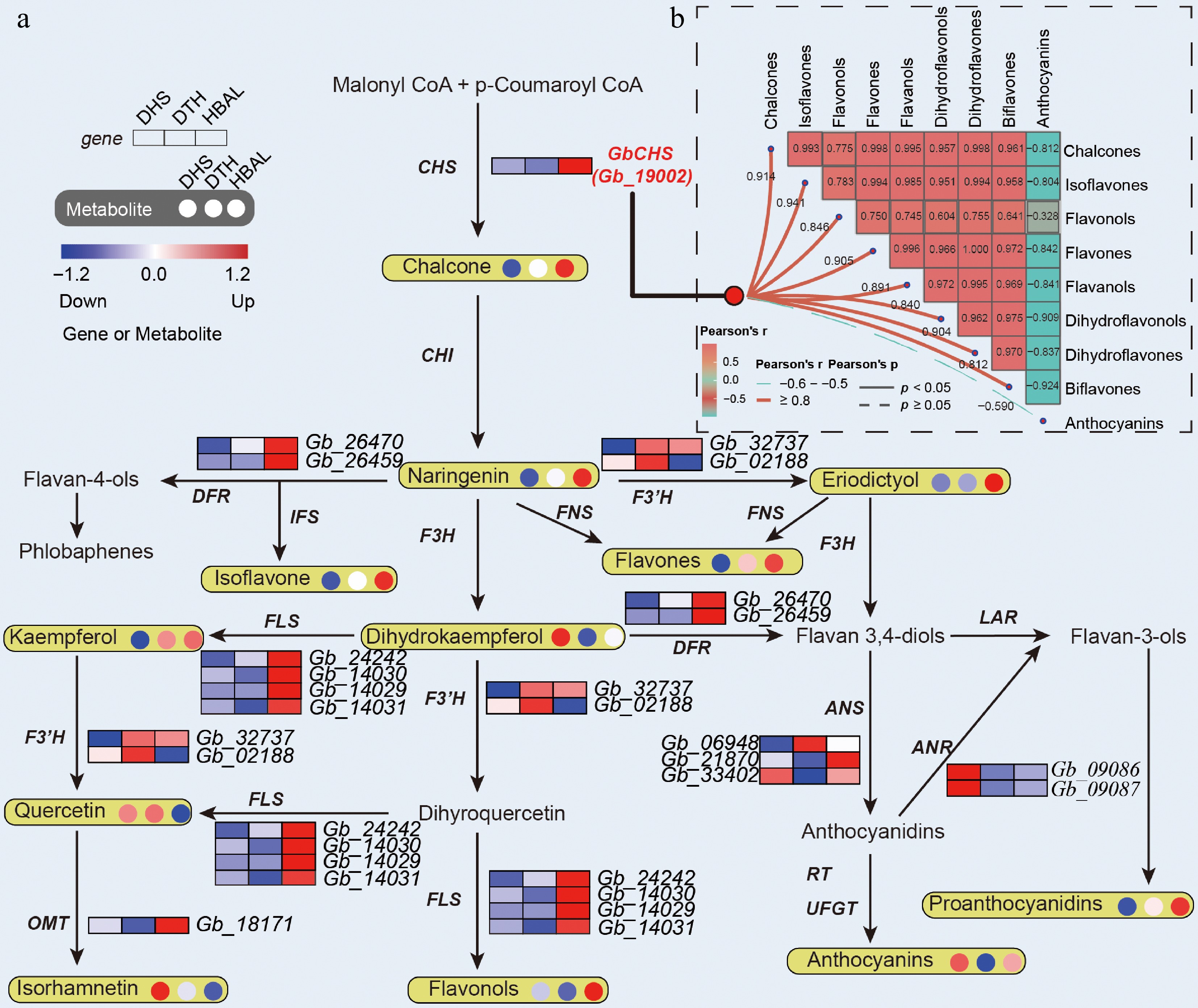
Figure 3.
Integrated transcriptomic and metabolomic profiling of flavonoid biosynthesis in G. biloba. (a) Expression patterns of DEGs and accumulation levels of DAMs associated with flavonoid biosynthesis. (b) Correlation network illustrating relationships between GbCHS expression and different flavonoid subclasses.
Given that CHS is the first rate-limiting enzyme in the flavonoid biosynthesis pathway, further focus was placed on the CHS gene family in G. biloba. A total of eight GbCHS genes were identified in the G. biloba genome, and multiple sequence alignment revealed a high degree of conservation among these genes (Supplementary Fig. S8). Expression analysis showed that only three GbCHS genes (Gb_20355, Gb_19002, and Gb_01519) were expressed at relatively high levels, whereas the remaining genes exhibited low transcript abundance (Supplementary Fig. S9). Notably, among these three, only Gb_19002 (GbCHS) was significantly differentially expressed. Its transcript level in the high-flavonoid cultivar 'HBAL' was approximately 15.5-fold higher than that in the low-flavonoid cultivar 'DHS', prompting its selection for subsequent functional analyses. Correlation analysis between GbCHS expression and flavonoid metabolites further revealed strong positive associations with chalcones, isoflavones, dihydroflavones, dihydroflavonols, flavones, flavonols, and biflavonoids, whereas a negative correlation was observed with anthocyanins (Fig. 3b). Collectively, these results indicate that GbCHS (Gb_19002) is a central regulatory gene influencing the biosynthesis of multiple flavonoid subclasses in G. biloba.
GbCHS serves as a pivotal structural gene for flavonoid biosynthesis
-
The GbCHS gene, containing a CDS of 1,191 bp, was successfully cloned. In order to investigate the catalytic properties of the GbCHS enzyme, a 6×His tag was added to its N-terminus, and the recombinant protein was heterologously expressed in E. coli (Fig. 4a). SDS–PAGE and Western blot analyses confirmed the successful expression and purification of GbCHS, yielding a protein with an apparent molecular weight of approximately 60 kDa (Fig. 4b, c). The catalytic activity of GbCHS was evaluated using purified recombinant protein in the presence of p-coumaroyl-CoA and malonyl-CoA. The resulting reaction products were analyzed by HPLC. As chalcone products are known to undergo spontaneous cyclization to naringenin during the in vitro reaction process, naringenin was detected in the reaction product. Compared with the heat-inactivated control, a distinct peak corresponding to naringenin appeared at a retention time of 4.57 min in reactions containing active GbCHS. Moreover, naringenin accumulation increased proportionally with the increased amount of GbCHS protein (Fig. 4d), confirming that GbCHS is a catalytically active chalcone synthase.
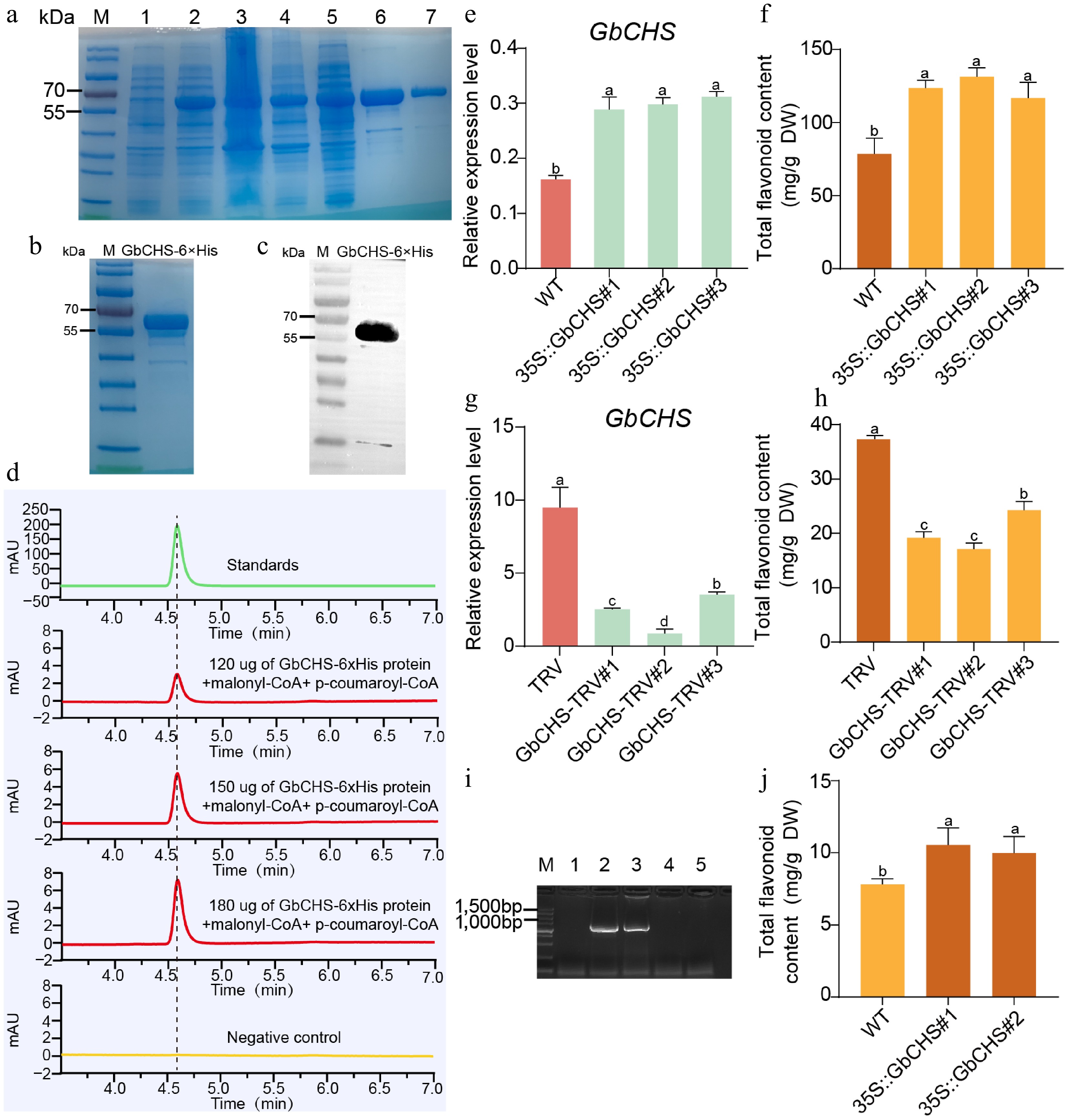
Figure 4.
GbCHS promotes the accumulation of total flavonoids. (a) Expression and purification of the recombinant GbCHS protein. M: Protein marker; 1: non-induced sample; 2: induced sample; 3: supernatant after induction and lysis; 4: pellet after induction and lysis; 5: flow-through during purification; 6−7: eluted GbCHS recombinant protein. (b) Buffer exchange and quality assessment of purified GbCHS protein. (c) Western blot analysis of recombinant GbCHS protein. (d) Enzymatic activity assay of GbCHS based on naringenin production. (e) Relative expression levels of GbCHS in overexpressing G. biloba calli. (f) Total flavonoid content in GbCHS-overexpressing G. biloba calli. (g) GbCHS expression levels in VIGS-mediated silenced G. biloba plants. (h) Total flavonoid content in GbCHS-silenced G. biloba plants. (i) Positive detection of GbCHS-overexpressing A. thaliana lines. M: DNA molecular weight marker; 1: wild-type A. thaliana (WT); 2−3: GbCHS-overexpressing A. thaliana; 4−5: negative control. (j) Total flavonoid content in GbCHS-overexpressing A. thaliana plants. Different lowercase letters indicate significant differences (p < 0.05). Data are presented as mean ± SD of three biological replicates (n = 3).
In order to further assess the biological function of GbCHS, A. tumefaciens, carrying the 35S::GbCHS vector, was transformed into G. biloba calli. Transcript levels were quantified using qRT-PCR. Overexpression of GbCHS led to a marked increase in total flavonoid content in the G. biloba calli, which rose by 49% to 68% relative to the control group (Fig. 4e, f). Conversely, VIGS of GbCHS in G. biloba seedlings led to a pronounced reduction in flavonoid accumulation, with silenced lines showing 35%-55% lower total flavonoid content compared with plants transformed with the empty TRV vector (Fig. 4g, h). To further validate these findings in a heterologous system, this study generated two independent A. thaliana transgenic lines expressing 35S::GbCHS (Fig. 4i, Supplementary Fig. S10). Consistent with the results observed in G. biloba, total flavonoid content in these transgenic A. thaliana lines increased significantly by 36% and 28%, respectively, relative to wild-type plants (Fig. 4j).
GbCHS significantly promotes plant growth and development
-
Silencing of GbCHS via VIGS markedly impaired the growth of G. biloba. Compared with plants transformed with the TRV empty vector, GbCHS-silenced G. biloba exhibited a 30% reduction in plant height, a 20% decrease in leaf number, and a 13% reduction in fresh weight (Fig. 5a−d). Consistently, heterologous overexpression of GbCHS in A. thaliana significantly promoted root system development. Both transgenic lines overexpressing GbCHS produced substantially more lateral roots than wild-type plants (Fig. 5e). Quantitative analysis revealed that GbCHS#1 and GbCHS#2 formed an average of 14.78 and 16.00 lateral roots, respectively, representing a 20%-30% increase compared with WT of 12.33 (Fig. 5f). In addition, primary root length was dramatically enhanced in GbCHS-overexpressing plants, reaching 7.51 cm in GbCHS#1 and 7.60 cm in GbCHS#2, more than twice the length observed in WT plants (3.18 cm) (Fig. 5f). Aboveground growth was also strongly stimulated by GbCHS overexpression. The number of rosette leaves increased by 58% and 44% in the two GbCHS-overexpression A. thaliana lines, respectively, relative to WT (Fig. 5g−j), while plant height increased by 19.0% and 19.4% (Fig. 5k). Moreover, net photosynthetic rate was elevated 3.2-fold and 2.9-fold (Fig. 5l). Correspondingly, aboveground biomass accumulation was significantly enhanced, with fresh weight increasing by 38% and 22% and dry weight by 49% and 24% in the two overexpression lines (Fig. 5m, n).
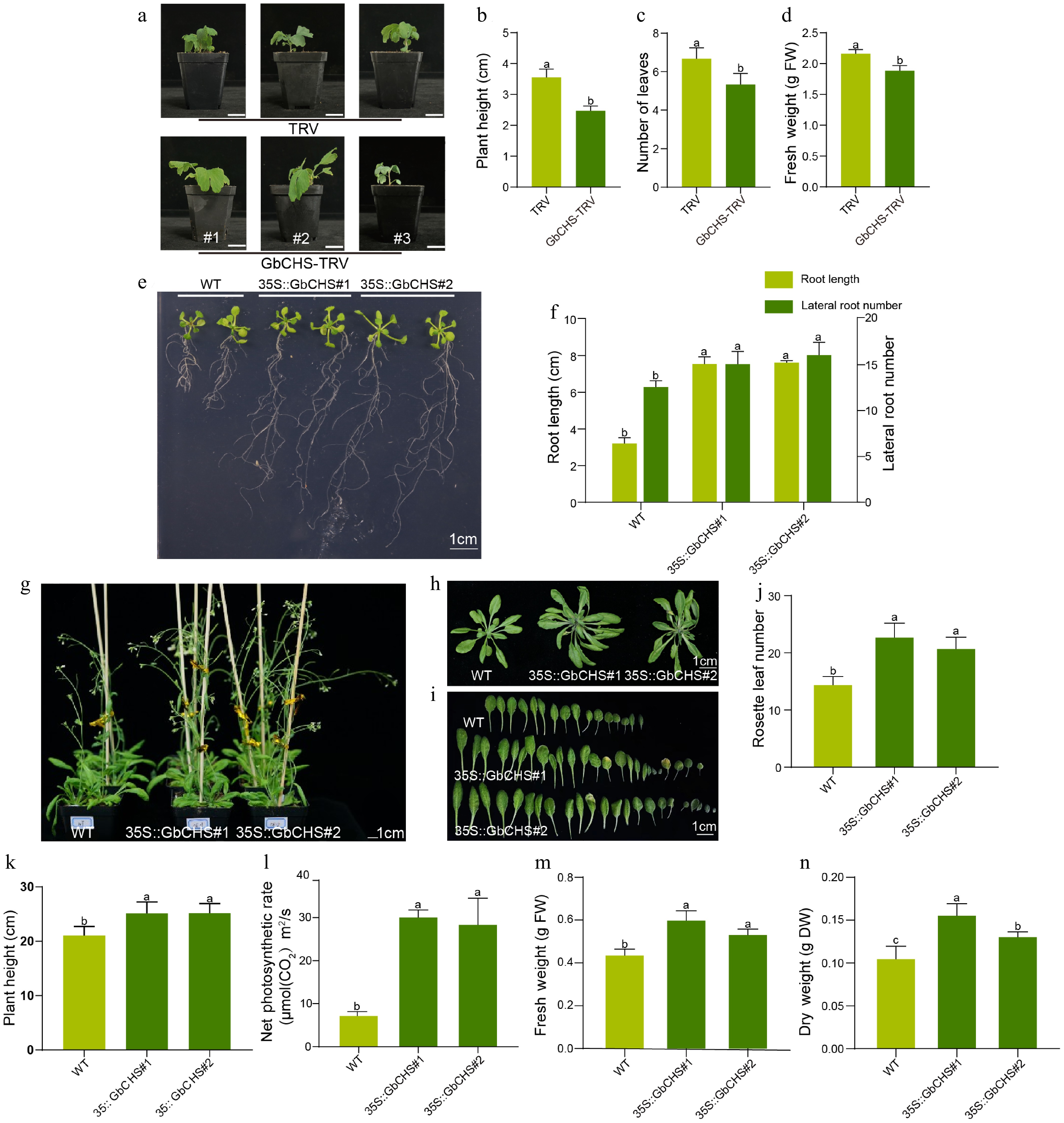
Figure 5.
Phenotypic effects of GbCHS silenced in G. biloba and GbCHS overexpression in A. thaliana plants. (a), (b) Comparison of plant height. (c) Comparison of the number of leaves. (d) Comparison of fresh weight. Plant height was measured from the soil surface to the apical meristem. Scale bar = 3 cm. (e), (f) Comparison of root traits. (g)−(j) Comparison of rosette leaf number. (k) Comparison of plant height. (l) Comparison of net photosynthetic rate. (m) Comparison of aboveground fresh weight (FW). (n) Comparison of aboveground dry weight (DW). Statistical significance was determined by ANOVA. Different lowercase letters indicate significant differences at p < 0.05. All data are shown as the mean ± SD (sample sizes: n = 3 for panels 5B−5F; n = 5 for panels 5H and 5K−5P).
Identification of lncRNAs predicts the potential LncNAT-GbCHS regulatory module in G. biloba
-
In order to systematically identify lncRNAs in G. biloba, this study assessed the protein-coding potential of newly assembled transcripts using CPC, PFAM, and CNCI. This analysis identified a total of 169,114 lncRNAs (Fig. 6a), comprising 156,761 lincRNAs (92.7%), 9,973 lncNATs (5.9%), and 2,380 sense-overlapping lncRNAs (1.4%) (Fig. 6b). Differential expression analysis revealed 3,448 DELs among the three cultivar comparisons. Specifically, 863 DELs were identified in 'DTH' vs 'DHS' (365 upregulated and 498 downregulated); 2,486 DELs in 'HBAL' vs 'DHS' (628 upregulated and 1,858 downregulated); and 1,330 DELs in 'DTH' vs 'HBAL' (759 upregulated and 571 downregulated) (Fig. 6c).
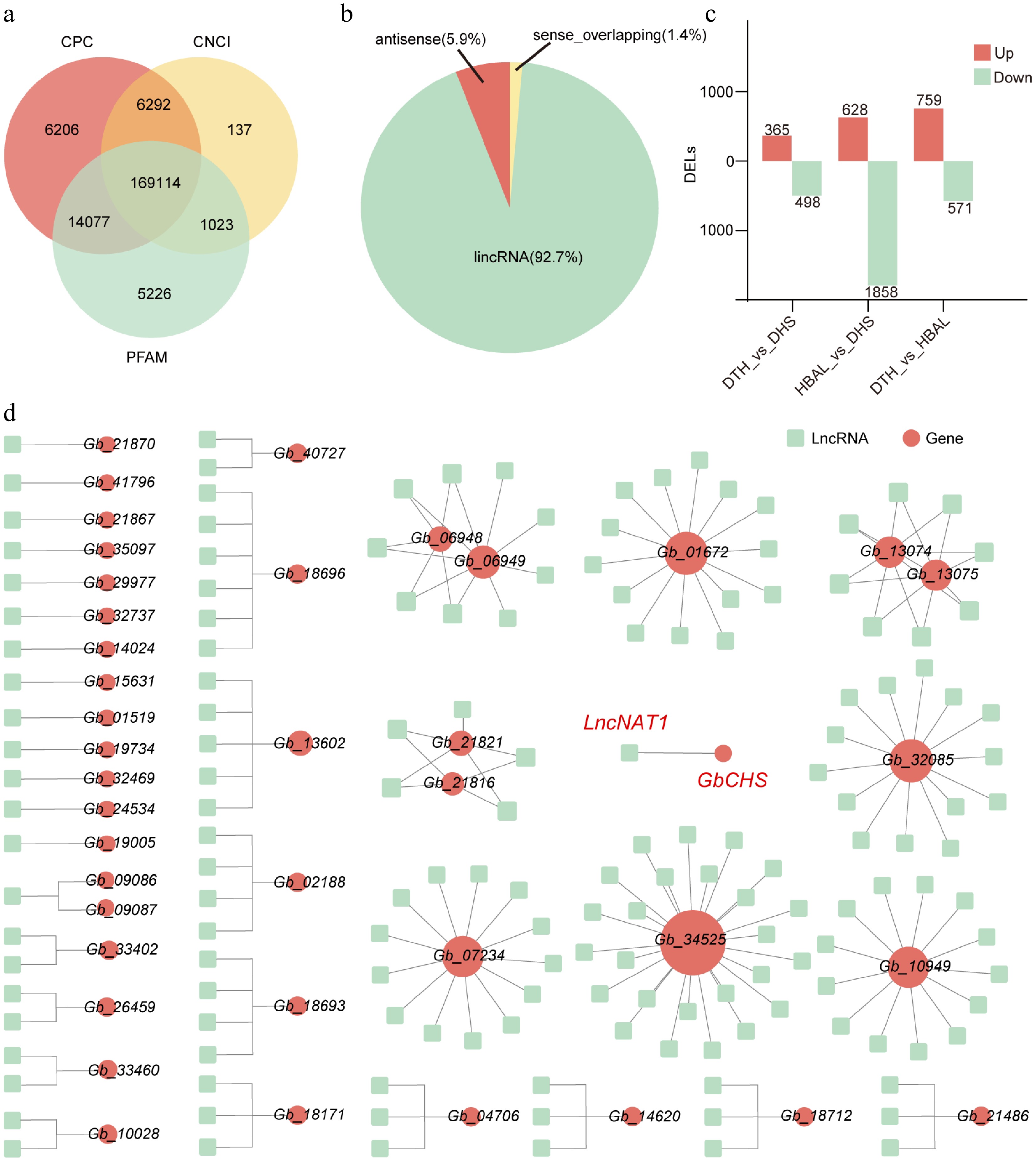
Figure 6.
Genome-wide identification and regulatory network analysis of lncRNAs in three G. biloba cultivars. (a) Identification pipeline and number of lncRNAs detected. (b) Classification of identified lncRNAs into lincRNAs. (c) Numbers of DELs among the three cultivar comparisons. (d) DELs-flavonoid biosynthesis gene networks.
Prediction of cis-regulatory targets of DELs indicated that 4,953 DELs may regulate 3,250 genes. Functional enrichment analysis showed that cis target genes of DELs in 'DTH' vs 'DHS' and 'DTH' vs 'HBAL' were enriched in the 'Metabolic pathways' and 'Biosynthesis of secondary metabolites' pathways. Notably, cis targets in the 'HBAL' vs 'DHS' were additionally enriched in the 'Flavonoid biosynthesis' pathway (Supplementary Figs S11−S13). Construction of DEL-flavonoid biosynthesis gene regulatory networks further identified 178 DELs as potential cis-regulators of 41 flavonoid biosynthesis genes, including the putative regulation of GbCHS by LncNAT1 (Fig. 6d).
LncNAT1 negatively regulates plant development and flavonoid biosynthesis by repressing GbCHS
-
Genomic locus analysis revealed that LncNAT1 is transcribed antisense to GbCHS (Gb_19002), with partial sequence complementarity between the two transcripts (Fig. 7a, Supplementary Fig. S14). Sequence alignment further showed that LncNAT1 shares only limited and fragmented similarity with other members of the CHS gene family (Supplementary Fig. S15). In contrast to GbCHS (Gb_19002), LncNAT1 displayed its highest expression in the low-flavonoid cultivar 'DHS' and the lowest expression in the high-flavonoid cultivar 'HBAL' (Supplementary Fig. S16). To investigate the biological function of LncNAT1, 35S::LncNAT1 was overexpressed in G. biloba calli, and qRT-PCR confirmed strong transgene expression (Fig. 7b) accompanied by a dramatic reduction (92%−94%) in total flavonoid content relative to the empty vector control (Fig. 7c). Conversely, VIGS-mediated silencing of LncNAT1 significantly decreased its transcript abundance and resulted in an 11%-14% increase in total flavonoids compared with the TRV control (Fig. 7d−f). In addition to its metabolic effects, LncNAT1 silencing markedly promoted vegetative growth, with plant height and fresh weight increasing by 65% and 14%, respectively (Fig. 7g, h).
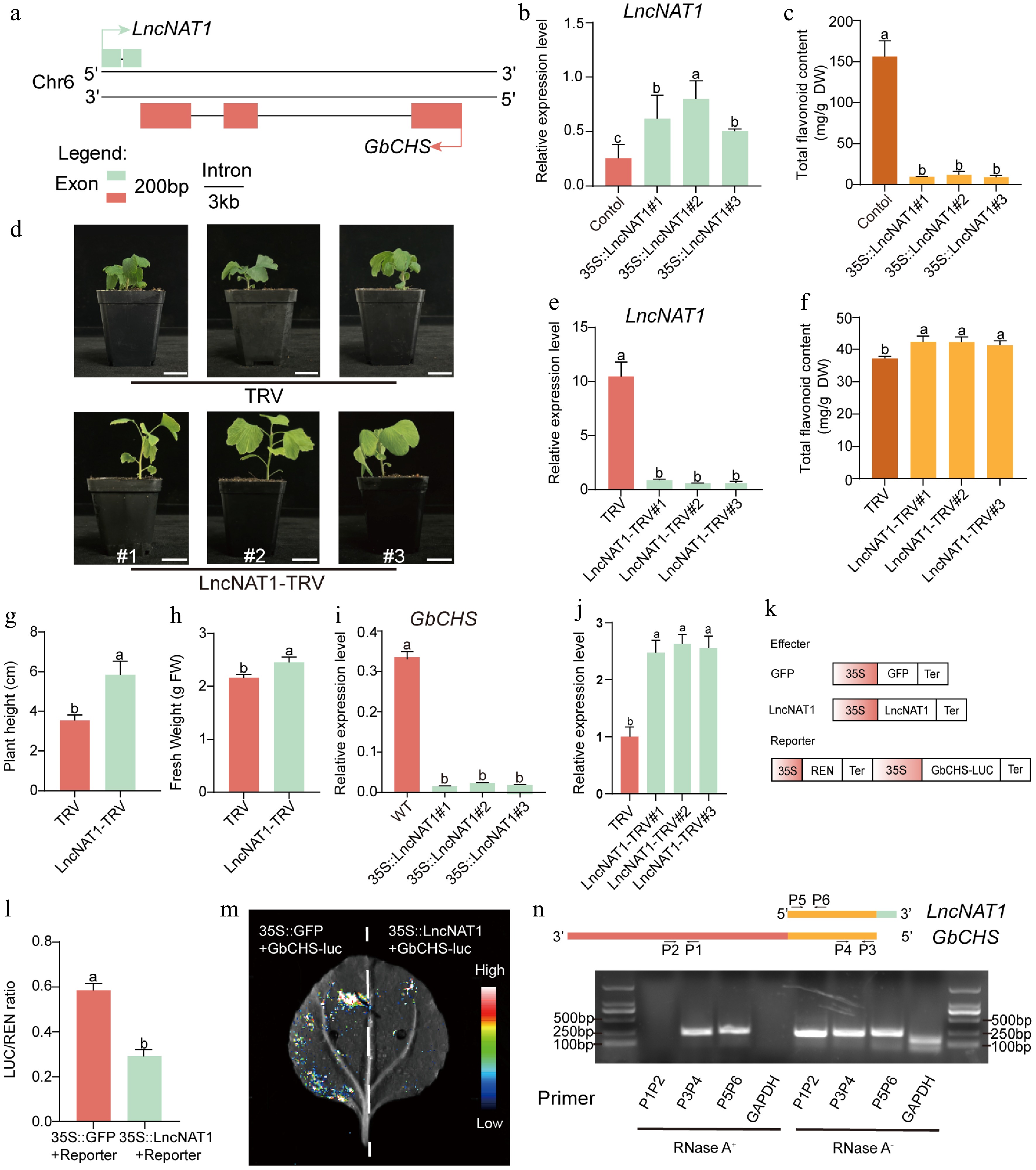
Figure 7.
LncNAT1 negatively regulates flavonoid biosynthesis and plant development by inhibiting GbCHS. (a) Genomic organization showing the antisense relationship between LncNAT1 and GbCHS. (b) LncNAT1 transcript levels, and (c) total flavonoid content in LncNAT1-overexpressing G. biloba calli. (d) Phenotypes of LncNAT1-silenced G. biloba plants. Scale bars = 3 cm. (e) LncNAT1 expression levels (f) and total flavonoid accumulation in LncNAT1-silenced G. biloba plants. (g), (h) Comparison of plant height and fresh weight between LncNAT1-silenced plants and TRV control. GbCHS transcript levels in (i) LncNAT1-overexpressing calli and (j) LncNAT1-silenced plants. (k) Schematic of DLR vector. (l) DLR assay showing repression of GbCHS activity by LncNAT1. (m) Luciferase imaging assay verifies the regulatory relationship between LncNAT1 and GbCHS. (n) RPA analysis of LncNAT1-GbCHS binding interaction. Different lowercase letters indicate significant differences (p < 0.05). Data represent mean ± SD of three biological replicates (n = 3).
This study further examined whether LncNAT1 regulates flavonoid biosynthesis by modulating GbCHS expression. It is revealed that overexpression of LncNAT1 significantly suppressed GbCHS transcript levels, whereas GbCHS expression was markedly enhanced upon LncNAT1 silencing (Fig. 7i, j). Furthermore, DLR assays demonstrated that co-expression of LncNAT1 significantly reduced the LUC/REN ratio of GbCHS (Fig. 7k, l). These results demonstrate the inhibitory effect of LncNAT1 on GbCHS at both the transcriptional and translational levels.
Given that LncNAT1 is a type of lncNAT, it was tested to determine if it functions by duplex formation with GbCHS through RPA. RPA demonstrated that complementary regions of LncNAT1 and GbCHS (P3/P4, P5/P6) remained resistant to RNase A digestion, whereas non-complementary regions (P1/P2) and the GAPDH control were almost completely degraded (Fig. 7m). These results indicate that LncNAT1 represses GbCHS expression by forming a stable LncNAT1-GbCHS RNA duplex. Finally, expression analysis of all other GbCHS family members in the LncNAT1-silenced plant revealed that only GbCHS (Gb_19002) was significantly upregulated, while other paralogs remained unchanged (Supplementary Fig. S17). This specificity demonstrates that LncNAT1 selectively targets GbCHS (Gb_19002) rather than broadly regulating the CHS gene family.
-
Flavonoids constitute a major class of polyphenolic secondary metabolites that are widely distributed in plants[10]. G. biloba is recognized as an important medicinal species owing to its abundant flavonoid constituents[28], which have been clinically validated for the treatment of cardiovascular and coronary heart diseases as well as neurodegenerative disorders[48]. To date, approximately 110 flavonoid compounds have been identified in G. biloba leaves[49]. More recent investigations have further revealed novel flavonoids such as biginkgosides A, B, C, and D[50]. Despite this chemical richness, the long juvenile period of G. biloba trees severely constrains conventional breeding programs aimed at improving flavonoid yield, resulting in a limited availability of elite, high-quality cultivars in the market. This study revealed a 3.45-fold variation in total flavonoid content among 26 G. biloba cultivars. 'HBAL' and 'DTH' accumulated high levels of eight major flavonoid subclasses, including biflavonoids and flavonols, whereas 'DHS' was characterized by elevated anthocyanin accumulation. These contrasting metabolic profiles suggest a competitive allocation of metabolic flux between the flavonoid and anthocyanin branches of the phenylpropanoid pathway, a phenomenon previously reported in other plant species where shared chalcone intermediates are differentially channeled toward distinct downstream products[50]. Together, these findings provide a scientific basis for the targeted selection of specialized G. biloba cultivars, such as high-flavonoid leaf-use types and high-anthocyanin ornamental types.
GbCHS functions in flavonoid biosynthesis and plant development
-
CHS is the first committed and rate-limiting enzyme in the flavonoid biosynthetic pathway and is highly conserved across angiosperms[51,52]. For example, silencing CiCHS in citrus or knocking out MdCHS in apple leads to a marked reduction in flavonoid content, while in A. thaliana, CHS protein abundance is dynamically regulated through the ubiquitin-proteasome system, thereby modulating anthocyanin accumulation[53−55]. The result establishes GbCHS as a critical structural gene in the flavonoid biosynthetic pathway of G. biloba. The recombinant enzyme catalyzes the formation of naringenin chalcone from p-coumaroyl-CoA and malonyl-CoA in vitro, confirming its catalytic competence. In vivo, overexpression of GbCHS in G. biloba calli increased total flavonoid content by more than 49%, whereas VIGS-based silencing resulted in a reduction exceeding 35%. Beyond its metabolic function, accumulating evidence indicates that CHS exerts broad regulatory effects on plant growth and development. In petunia, maize, and tomato, CHS mutations or silencing disrupt flavonol accumulation in pollen, leading to defective pollen tube growth, male sterility, or abnormal fruit development[56−58]. More recently, silencing CHS in apple was shown to cause not only a near-complete loss of flavonoids but also pronounced vegetative defects, including shortened internodes, reduced leaf size, and diminished growth rates, which were associated with altered auxin transport dynamics[54]. This study extends these findings to G. biloba. Silencing of GbCHS resulted in reduced plant height, leaf number, and biomass, whereas heterologous overexpression of GbCHS in A. thaliana significantly enhanced primary root length, lateral root number, rosette leaf number, and biomass. These phenotypes are consistent with previous observations in A. thaliana flavonol-deficient synthesis mutants, where impaired flavonol biosynthesis disrupts root elongation, regulates inflorescence branching, and alters stomatal aperture[59]. Conversely, the enhanced growth traits observed in GbCHS-overexpressing A. thaliana plants further support the notion that flavonoids act as key modulators of developmental processes. Collectively, these results demonstrate that GbCHS functions not only as a central enzymatic hub in flavonoid biosynthesis but also as an important metabolic regulator coordinating flavonoid-dependent control of plant growth and development in G. biloba and beyond.
Regulatory mechanism of the LncNAT1-GbCHS module: functional conservation and specificity of plant lncNATs
-
Natural antisense lncNATs represent an important class of regulatory RNAs in plants and typically exert their functions through sequence complementarity with their sense transcripts, thereby modulating gene expression at transcriptional and/or translational levels[60,61]. Several well-characterized lncNATs exemplify the mechanistic diversity of this regulatory mode. In petunia, the lncNAT SHO promotes the formation of double-stranded RNA and siRNA formation, leading to RNA interference-mediated degradation of the cytokinin biosynthesis gene SHO[62]. In rice, cis-NATPHO1;2 associates with polysomal to enhance the translation efficiency of PHO1;2[63]. Meanwhile, in A. thaliana, the lncNAT SVALKA interferes with RNA polymerase II elongation, thereby repressing transcription of the cold-responsive gene CBF1 through transcriptional collision[64,65]. Collectively, these studies highlight that plant lncNATs employ diverse regulatory strategies, including RNA interference, translational control, transcriptional interference, and chromatin-associated mechanisms.
The LncNAT1-GbCHS module identified in this study represents a distinct example of an antisense lncNAT directly targeting a structural gene. LncNAT1 forms an RNA duplex with GbCHS, thereby repressing its expression and modulating both flavonoid biosynthesis and plant growth. Unlike most previously characterized plant lncNATs, which primarily regulate transcription factors or hormone-related genes, LncNAT1 directly targets a core enzymatic gene in the flavonoid pathway, underscoring the specificity and precision of metabolic regulation in G. biloba. Furthermore, the RNA duplex-mediated mechanism LncNAT1 differs fundamentally from that of rice TL, which regulates OsMYB60 through histone modifications[66], and A. thaliana asHSFB2a, which represses transcription factor expression[67]. Together, these differences highlight the species- and target-dependent functional diversity of plant lncNATs. This variety suggests that lncNAT-mediated regulatory networks are substantially more complex than currently appreciated, warranting systematic investigation across diverse species and metabolic contexts.
Study limitations and future perspectives
-
Although this study clarifies the function and regulatory mechanism of the LncNAT-GbCHS module, several limitations remain. First, functional validation was primarily achieved through transient overexpression in G. biloba calli, VIGS-based gene silencing, and heterologous expression in A. thaliana, due to the lack of a stable genetic transformation system for G. biloba. Second, although the results demonstrate that GbCHS profoundly influences plant growth and development, the downstream molecular mechanisms, particularly the interplay between flavonoid accumulation and hormone signaling pathways such as auxin, need to be fully understood. Future research should therefore focus on two major directions. On the one hand, the establishment of a stable and efficient genetic transformation system for G. biloba will be essential for definitive functional validation of the LncNAT1-GbCHS module. On the other hand, integrative approaches combining transcriptomics, hormone profiling, and protein-protein or protein-metabolite interaction analyses will also play a critical role.
-
This study reveals substantial natural variation in flavonoid accumulation among G. biloba cultivars, with 'HBAL' exhibiting the highest total flavonoid content. Integrated metabolomic and transcriptomic analyses identified 976 metabolites and 3,090 DEGs. Among them, GbCHS (Gb_19002) genes show a strong positive correlation with flavonoid accumulation. Functional assays confirmed that GbCHS catalyzes naringenin chalcone synthesis, and its overexpression significantly enhanced flavonoid accumulation in G. biloba calli and A. thaliana, whereas VIGS-mediated silencing caused a marked reduction. Moreover, overexpressing GbCHS positively regulates plant growth and development. Additionally, a potential LncNAT-GbCHS regulatory module was identified. LncNAT1, an lncNAT, negatively regulates flavonoid biosynthesis and plant development by directly inhibiting GbCHS through RNA duplex formation (Fig. 8). Collectively, these results uncover a novel lncNAT-mediated regulatory mechanism controlling flavonoid metabolism and growth in G. biloba plants and offer promising genetic targets for the coordinated improvement of flavonoid yield and growth traits in G. biloba breeding programs.
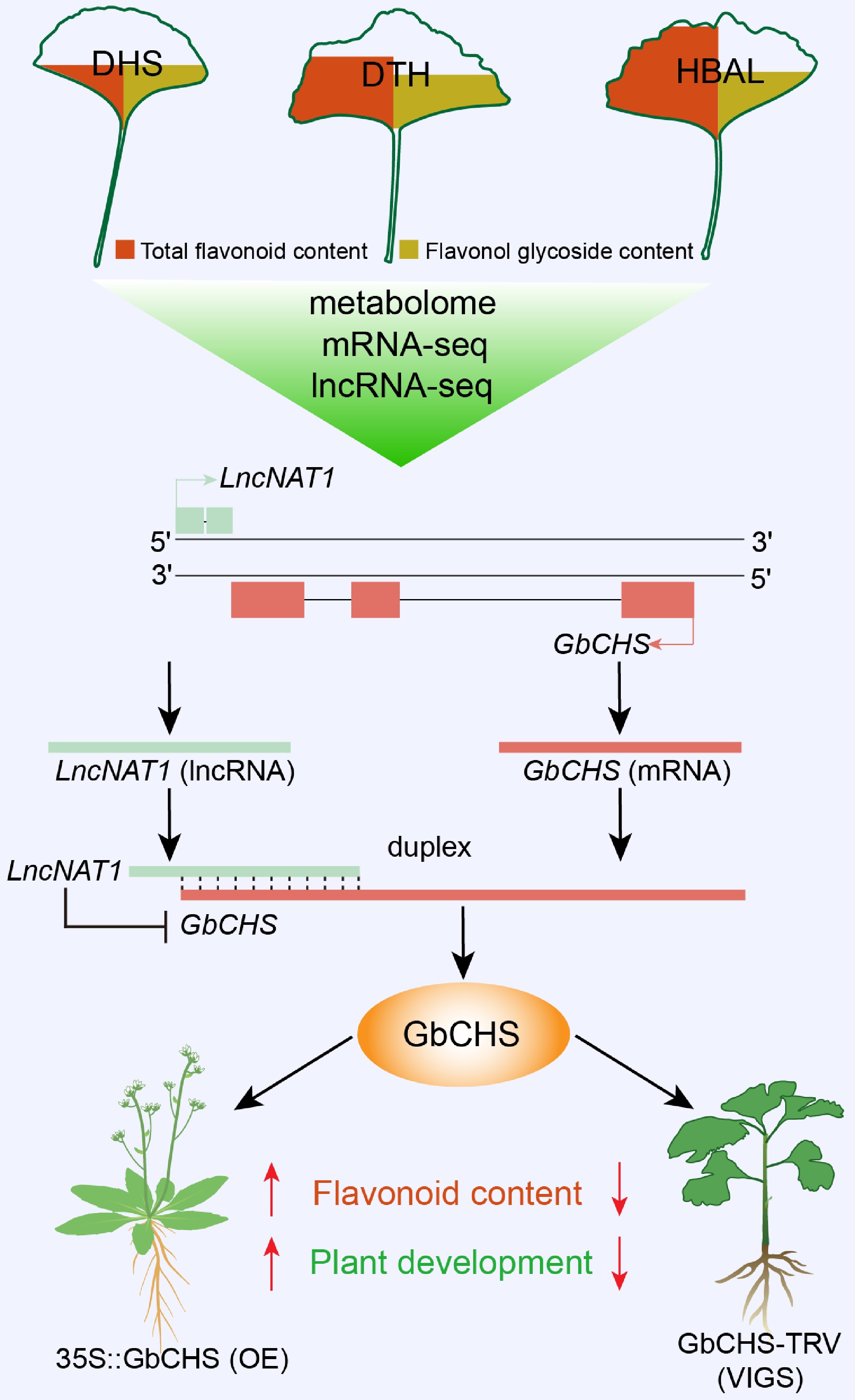
Figure 8.
Working model of the LncNAT1-GbCHS regulatory module in flavonoid biosynthesis and plant development. Integrated analysis of metabolomic, RNA-seq, and lncRNA-seq datasets identified a DEL, LncNAT1, along with its potential target gene, GbCHS. Functional validation using dual-luciferase assays, RPA, and genetic transformation experiments demonstrated that LncNAT1 directly binds to the GbCHS transcript to form an RNA duplex, thereby repressing GbCHS expression. This inhibition thereby negatively regulates flavonoid biosynthesis and plant growth and development in G. biloba.
-
The authors confirm contribution to the paper as follows: study conception and design: Wang L, Liu S; experimental work and data analysis: Shuai Y, Wan P, Cao M, Zhang H; draft manuscript preparation: Liu S, Wan P; manuscript revision: Wang L, Jin B, Zhang C, Zhang X, Ma L. All authors reviewed the results and approved the final version of the manuscript.
-
The ssRNA-seq data are available at the NCBI SRA database under accession number PRJNA1133634.
-
This project was supported by grants from the National Natural Science Foundation of China (Grant Nos 32171838 and 32001341), the Jiangsu Province Key Research and Development Plan (modern agriculture) (Grant No. BE2021367), and the Qing Lan Project of Yangzhou University.
-
The authors declare that they have no conflict of interest.
- Supplementary Table S1 Names and abbreviations of the Ginkgo biloba cultivars used in this study.
- Supplementary Table S2 Primer sequences used in the experiment.
- Supplementary Table S3 Summary statistics of metabolite classes and numbers identified by widely targeted metabolomics.
- Supplementary Fig. S1 Volcano plot of differentially accumulated metabolites (DAMs) in G. biloba leaves among different cultivar comparisons: (A)'DTH' vs 'DHS', (B) 'HBAL' vs 'DHS', and (C) 'DTH' vs 'HBAL'.
- Supplementary Fig. S2 Relative content of anthocyanins in the leaves of the three selected cultivars.
- Supplementary Fig. S3 Numbers of differentially expressed genes (DEGs).
- Supplementary Fig. S4 Venn diagram showing shared and unique DEGs across different comparison groups.
- Supplementary Fig. S5 KEGG enrichment analysis of DAMs and DEGs in 'DTH' vs 'DHS' comparison.
- Supplementary Fig. S6 KEGG enrichment analysis of DAMs and DEGs in 'DTH' vs 'DHS' comparison.
- Supplementary Fig. S7 KEGG enrichment analysis of DAMs and DEGs in 'DTH' vs 'HBAL'.
- Supplementary Fig. S8 Multiple sequence alignment of GbCHS (Gb_19002) protein with other CHS family members in G. biloba.
- Supplementary Fig. S9 Expression profiles of all eight CHS genes in three G. biloba cultivars with contrasting flavonoid contents.
- Supplementary Fig. S10 Expression levels of GbCHS in the GbCHS-overexpressing A. thaliana lines.
- Supplementary Fig. S11 KEGG enrichment analysis of cis-regulated target genes of DELs in 'DTH' vs 'DHS' comparison.
- Supplementary Fig. S12 KEGG enrichment analysis of cis-regulated target genes of DELs in 'DTH' vs 'HBAL'.
- Supplementary Fig. S13 KEGG enrichment analysis of cis-regulated target genes of DELs in 'HBAL' vs 'DHS' comparison.
- Supplementary Fig. S14 Sequence alignment between the reverse-complement strand of LncNAT1 and GbCHS.
- Supplementary Fig. S15 Sequence alignment of the LncNAT1 reverse-complement strand with other G. biloba CHS genes (excluding Gb_19002).
- Supplementary Fig. S16 Expression patterns of LncNAT1 and GbCHS across different cultivars.
- Supplementary Fig. S17 Relative expression levels of other CHS family genes (excluding Gb_19002) in LncNAT1-VIGS G. biloba seedlings.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Liu S, Shuai Y, Wan P, Cao M, Zhang H, et al. 2026. Unraveling the multifaceted roles of the LncNAT1-GbCHS module in Ginkgo biloba for flavonoid biosynthesis and plant development. Forestry Research 6: e006 doi: 10.48130/forres-0026-0006
Unraveling the multifaceted roles of the LncNAT1-GbCHS module in Ginkgo biloba for flavonoid biosynthesis and plant development
- Received: 25 June 2025
- Revised: 04 February 2026
- Accepted: 11 February 2026
- Published online: 25 March 2026
Abstract: Flavonoids are the primary bioactive compounds in Ginkgo biloba leaves. However, the regulatory mechanisms underlying their biosynthesis remain incompletely understood. In this study, three G. biloba exhibiting significant differences in flavonoid content were selected from a total of 26 cultivars. Integrated metabolomic, lncRNA, and mRNA sequencing analyses identified GbCHS as a core gene involved in flavonoid biosynthesis. Overexpression of GbCHS markedly enhanced flavonoid accumulation, whereas virus-induced gene silencing (VIGS) of GbCHS resulted in a significant reduction in total flavonoid content, confirming its essential role in flavonoid biosynthesis. Notably, GbCHS-silenced G. biloba plants also exhibited a significant decrease in plant height, leaf weight, root length, and lateral root number. In contrast, Arabidopsis thaliana plants overexpressing GbCHS showed significant increases in plant height, leaf weight, root length, and lateral root number, indicating that this gene also promotes plant development. Furthermore, GbCHS was found to be partially complementary to the lncRNA LncNAT1. Overexpression of LncNAT1 in G. biloba calli significantly suppressed GbCHS expression and reduced total flavonoid content, whereas silencing LncNAT1 led to increased GbCHS expression, flavonoid accumulation, plant height, and leaf weight. Mechanistically, LncNAT1 represses GbCHS expression through the formation of an LncNAT1-GbCHS RNA duplex. Collectively, these findings reveal the multidimensional regulatory functions of the novel LncNAT1-GbCHS module in flavonoid biosynthesis and plant development in G. biloba.
-
Key words:
- Flavonoids /
- Chalcone synthase /
- Ginkgo biloba /
- LncNAT1-GbCHS module /
- Plant development














