-
Due to continuous monoculture and intensive agricultural practices, continuous cropping obstacle (CCO) in the flower production system has long been recognized as a serious threat to the quality production of cut flowers[1]. The occurrence of CCO normally results in plant growth reduction, yield decline, and replant failure, which are typical outcomes of negative plant-soil feedback[2]. Plant-soil feedback (PSF) describes a loop in which soil properties conditioned by the legacy of previous abiotic and biotic processes, and current living plants consequently influence subsequent plants and soil biota[3,4]. PSF can be both positive and negative, depending on the direction of soil condition[5,6]. Negative PSF may occur through depletion of resources, accumulation of natural enemies, and excretion of allelopathic or autotoxic compounds[7]. These mechanisms do not act in isolation but are often interdependent and species-specific[4,8−10]. As such, resolving CCO in the flower production system with effective means requires fully understanding the interactive mechanisms within the species-specific context.
Lisianthus (Eustoma grandiflorum) is an important cut flower worldwide with enormous rose-like blossoms and an extended vase life[11]. In Yunnan Province of China, lisianthus has ranked the top best-selling cut flower over the last few decades with the production value reaching CNY ¥354 billion, and a cultivation area up to 500 ha[1]. However, the quality production of lisianthus is easily and increasingly impeded by the continuous cropping obstacle. For example, lisianthus grown in the closed greenhouse with either soilless substrate or soil, showed sickness right after one year of production. Our previous investigation revealed that soil metabolite alteration, fungal structural change and pathogen structural change were the top three important factors in explaining the CCO development in lisianthus, covering 24.55%, 22.68%, and 21.41% of the explainable variations, respectively[1]. In addition, the observed accumulation of some potential phytotoxic compounds such as cynaropicrin and nicotine further implied that lisianthus plants in the continuous cropping soil may suffer from autotoxicity induced by the accumulation of toxic compounds emitted from root exudates[1]. Nevertheless, allelopathic compounds are known to be rapidly degraded by soil microbes, and only showed a general toxicity without species-specific effect. Thus, such phytotoxic compounds may be insufficient to fully explain species-specific negative PSF due to their short persistence and lack of specificity.
Extracellular self-DNA fragments (50~2,000 bp) released from litter decomposition has been proposed as another important contributor to negative PSF[12]. Unlike those allelochemicals, extracellular DNA can persist longer in the soil due to its chemical stability and its protection against enzymatic degradation by absorption and binding onto soil minerals and organic matter components[13]. In addition, fragmented extracellular self-DNA (sDNA) showed species-specific inhibitory effects, which have been described as a general biological process on a wide range of taxa including microbe (i.e., Trichoderma harzianum), algae (i.e., Scenedesmus obliquus), plant (i.e., Arabidopsis thaliana), and animal (i.e., Sarcophaga carnaria)[14]. The inhibitory effects involve complex and different cascades of events in organisms exposed to sDNA such as growth inhibition, reactive oxygen species (ROS) production[15−22], cell cycle arrest, root apex necrosis[23−25], and increased genomic methylation[26,27]. This generality of observed toxic effects of sDNA is highly coherent with the autotoxicity hypothesis which accounts for both long persistence and species-specificity. Evidence has shown that both damage-associated molecular pattern (DAMP) and DNA damage response (DDR) signaling function as possible underlying mechanisms in the plant's defensive response to sDNA[16,23,25]. However, the scope of these investigations has been limited both in duration (typically several hours to a few days) and in the focal species, which have predominantly been model species such as Arabidopsis thaliana[18,23,28−30]. The fundamental role of sDNA in the negative PSF under longer, more environmentally realistic exposure durations remains difficult to quantify.
In this study, we used E. grandiflorum as our test organism and first examined the species-specific toxicity of fragmented extracellular DNA from various species (close and distant phylogenetic related), in pot trials. The effective concentration of sDNA was subsequently determined in an ecotoxicity test. We later analyzed the physiological and molecular responses of lisianthus plants after one month exposure to sDNA in the Murashige and Skoog (MS) solid medium by phenotypic change observation, root microscopy analysis, and plant transcriptome profiling. The possible pathway involving sDNA in the development of negative PSF was further validated by short-time exposure of sDNA with subsequent inoculation of a fungal pathogen.
-
Four plant species were selected according to their phylogenetic closeness with lisianthus (E. grandiflorum), including two closely related species (Gentiana scabra and Gentiana rigescens) and two distantly related species (Dianthus caryophyllus and Lactuca sativa). Genomic DNA was extracted from fresh leaf tissue of the above five species using a modified cetyltrimethylammonium bromide (CTAB) method[20]. Briefly, approximately 10 g of tissue was pulverized in liquid nitrogen and lysed in a larger volume of pre-warmed 2 × CTAB lysis buffer (using 50 mL centrifuge tubes). The subsequent steps of chloroform-isoamyl alcohol extraction and isopropanol precipitation were proportionally scaled. The detailed DNA extraction procedure is described in Supplementary Text 1. To obtain fragmented DNA, samples were ultrasonicated using an ultrasonic cell disruptor (JY92-IIDN, Scientz). Following an optimization process (Supplementary Table S1), the final parameters were set to 30% power with 30 s ON/10 s OFF cycles for 8 min. This treatment produced DNA fragments in the target size range of 50–2,000 bp, as verified by 1.5% agarose gel electrophoresis (Supplementary Fig. S1).
Plant material and general growth conditions
-
Lisianthus seeds were surface-sterilized with 2% NaClO solution for 8 min, rinsed thoroughly with sterile water, and then inoculated onto 1/2 MS medium. They were stratified at 4 °C for 7 d before being transferred to a light culture room for germination. Uniform three-week-old seedlings were used for all experiments. The basal culture medium consisted of MS salts supplemented with sucrose (10 g/L), agar (8 g/L), and cephalosporin (100 μg/mL) at pH 5.8–6.0. Unless otherwise stated, seedlings were cultured under a 16-h light/8-h dark cycle.
Long-term exposure of extracellular fragmented DNA from various species
-
Fragmented DNA solutions from E. grandiflorum (EG), G. scabra (GS), G. rigescens (GR), D. caryophyllus (DC), and L. sativa (LS) were added under sterile conditions in a laminar flow hood to achieve a final DNA concentration of 200 μg/mL. This concentration was normally regarded as effective in similar studies[14−16,23,26,31]. A control group (CK) received sterile water. Seedlings were transplanted into culture tubes (12 replicates per group) and grown at 25 °C/23 °C (day/night) for 30 d. After 30 d of growth, lisianthus plants were harvested and growth parameters such as plant height, fresh weight, root length, and number of root branches were measured.
Assessment of the ecotoxicological effects of Lisianthus sDNA
-
To evaluate ecotoxicity, fragmented lisianthus sDNA solution was added to the basal medium to achieve final concentrations of 0, 50, 100, 200, and 400 μg/mL (30 replicates per treatment). Seedlings were cultured at a constant temperature of 25 °C for 4 weeks. At harvest, shoots and roots were separated, and their respective phenotypic traits were measured, including stem length, root length, fresh weight, and dry weight. The contents of malondialdehyde (MDA), hydrogen peroxide (H2O2), and superoxide anion (O2−) in roots were determined using commercial assay kits, strictly following the manufacturer's instructions. Root tip segments (approximately 1 cm in length) were collected from each treatment group and fixed in 70% FAA fixative at room temperature for 24 h. The fixed tissues were dehydrated, embedded in paraffin, and sectioned using standard histological procedures. Sections were stained with safranin and fast green and imaged using a 3DHISTECH Slide Scanner.
Transcriptome sequencing and analysis
-
Root samples of lisianthus from the control group (CK) and the 200 μg/mL (T-200) treatment group were selected for transcriptome sequencing, with three biological replicates per group. Total RNA was extracted from lisianthus roots treated with different concentrations of sDNA. Detailed information on the Illumina sequencing and data preprocessing is provided in supplementary material (Supplementary Text 1). After quality control checks, RNA sequencing was performed using an Illumina platform. Raw sequencing reads were quality-controlled and filtered using FASTP (v0.18.0) to obtain high-quality clean reads. Clean reads were aligned to the lisianthus reference genome using HISAT2 (v2.2.4). Differential expression gene (DEG) analysis was performed using DESeq2 software, with screening criteria set at |log2 (Fold Change)| ≥ 1 and false discovery rate (FDR) < 0.05. Gene Ontology (GO) functional enrichment analysis was conducted on the identified DEGs. Additionally, Gene Set Enrichment Analysis (GSEA) was employed to analyze differences at the pathway level, using significance thresholds of |Normalized Enrichment Score |(NES)| > 1, p < 0.05, and FDR < 0.25.
Validation of the synergistic effect of sDNA and fungal pathogen
Fungal culture and inoculum preparation
-
Fusarium oxysporum was selected as the representative fungal pathogen for lisianthus due to its severity in the practical cultivation[32]. The fungal strain F. oxysporum (strain PJZ1166, Shanghai BioHub Biotechnology, Shanghai, China) was cultured on potato dextrose agar (PDA) plates at 28 °C for 7 d. Conidia were harvested and further incubated in potato dextrose broth (PDB) at 28 °C, 170 rpm for 7 d. The conidia were then collected, resuspended in sterile water, and adjusted to a final concentration of 1 × 105 CFU/mL.
sDNA treatment and pathogen inoculation
-
Lisianthus sDNA was fragmented and diluted to concentrations of 50, 100, 200, and 400 μg/mL as above-mentioned. 3-week-old seedlings with similar sizes as mentioned above, were exposed in the sDNA with their roots immersed in the respective sDNA solutions for 24 h, with a control group (CK) treated with distilled water. Each of the five groups included 10 replicates. Seedlings were then transplanted into sterilized cups (height: 5 cm; volume: 200 mL) filled with sterilized Pindstrup peat (0–10 mm particle size, pH 5.5–6.0; Pindstrup Mosebrug A/S, Denmark). The seedlings from all groups were treated with 5 mL of the spore suspension after 3 d acclimation.
Soil DNA extraction and purification
-
Lisianthus plants were cultivated for 30 d before harvest. The accumulation of sDNA in the rhizosphere along with the population buildup of F. oxysporum were quantified as follows: Extracellular DNA (exDNA) from soil was extracted following a method adapted from Foscari et al.[33]. Briefly, 5 g of fresh soil sample was mixed with 5 mL of 0.12 M Na2HPO4 solution (pH 8.0) and shaken horizontally at 80 rpm for 30 min, followed by centrifugation at 7,500 g, 4 °C for 30 min to collect the supernatant. The extraction was repeated once, and the supernatants were combined. The combined supernatant was purified using a DNeasy PowerMax Soil Kit (omitting the cell lysis step), and the final DNA was eluted in 5 mL of 10 mM Tris-HCl (pH 8.0). DNA concentration was measured using a NanoDrop spectrophotometer.
Quantification of pathogen load in soil
-
To quantify the pathogen load in soil, a spore concentration gradient was prepared: the spore suspension was serially diluted 10-fold (from 1.6 × 102 to 1.6 × 107 spores/mL), and 5 mL of each dilution was mixed with 8 g of sterilized soil to create soil samples with pathogen loads ranging from 1.0 × 102 to 1.0 × 107 spores/g. Total DNA was extracted from each gradient soil sample using an Ezup Column Soil DNA Kit. Specific primers FaeF2 (5'-GGCATTTACTCCGCCACTTG-3')/FaeR2 (5'-AGCTCAGCGGCTTCCTATTG-3'), targeting the F. oxysporum elongation factor 1α (EF-1α) gene sequence, were used for absolute quantification by qRT-PCR. The reaction mixture (20 μL) contained 2 μL template DNA, 1 μL each of forward and reverse primer (10 μmol/L), 10 μL SYBR Green Mix, and topped up with ddH2O. The thermal cycling protocol was: 95 °C for 3 min; followed by 39 cycles of 95 °C for 5 s and 62 °C for 20 s. A standard curve was generated by plotting the log10 values of spore numbers against the obtained Cq values. Soil total DNA samples were subjected to qRT-PCR under identical conditions, and the pathogen spore quantity in soil samples was calculated based on the standard curve.
Statistical analysis
-
All measured data was obtained from at least 10 independent biological replicates. The normality of the data was verified using the Shapiro-Wilk test, and homogeneity of variances was assessed using Levene's test. For data meeting the assumptions of parametric tests, one-way analysis of variance (ANOVA) was used to compare significant differences among treatment groups, followed by Duncan's multiple range test for post-hoc comparisons. For data not meeting parametric assumptions, the non-parametric Kruskal-Wallis H test was employed. Growth indicators were converted to inhibition rates. The rates were then normalized to a 0–1 range using min–max normalization with negative values set to 0 to ensure biological plausibility. Dose-response curves were fitted using the four-parameter log-logistic model from the drc package (R version 4.5.1). Goodness-of-fit was evaluated using the coefficient of determination (R2), and the EC50 value was directly extracted from the fitted model. The statistical significance level was set at p < 0.05. All statistical analyses were performed using R version 4.5.1[34].
-
In general, the inhibitory effect of sDNA on lisianthus growth is species-specific and its intensity correlates positively with phylogenetic relatedness. The most severe growth inhibition on lisianthus seedlings was observed in response to sDNA (from lisianthus itself: EG group), followed by exDNA obtained from closely related species such as G. scabra (GS group) and G. rigescens (GR group). Exposure to sDNA significantly suppressed all measured growth parameters (p < 0.05), with the most pronounced effects on root architecture (Fig. 1). Root development in the EG group was nearly arrested, characterized by drastically shortened primary roots and a complete absence of lateral roots (Fig. 1b). The exDNA from G. scabra and G. rigescens also exhibited significant inhibitory effects, though to a lesser extent than sDNA. Notably, the inhibition by GS exDNA was consistently stronger than that by GR across multiple metrics, including plant height, root length, and total fresh weight (Fig. 1c−f). For instance, root systems in the GS group showed more severe suppression of lateral root formation and overall elongation compared to the GR group. In contrast, lisianthus plants treated with exDNA from distantly related species (D. caryophyllus and L. sativa) showed no such inhibitory effects. Their growth was as vigorous as that of the control (CK) group, with well developed root systems (Fig. 1a, b).
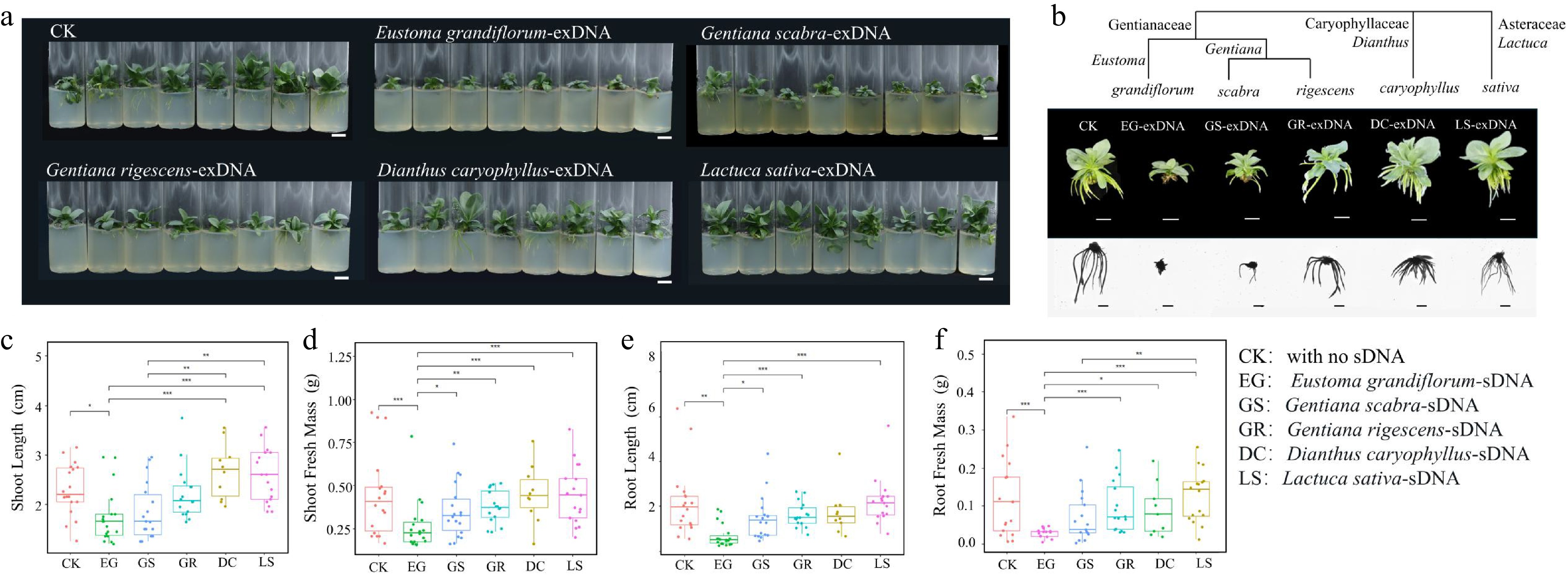
Figure 1.
Examination of species-specific inhibition of extracellular fragmented DNA (exDNA) in lisianthus plants. (a) Overall growth of lisianthus plants after 3 week exposure with 200 μg/mL exDNA from different species: Control with no exDNA addition (CK), Eustoma grandiflorum (EG), Gentiana scabra (GS), Gentiana rigescens (GR), Dianthus caryophyllus (DC), and Lactuca sativa (LS). (b)Phenotypic differences of individual plants from six groups along with the species phylogenetic relatedness. And morphological comparisons (mean ± SE, n = 12) in above- and below-ground traits among the six groups. (c) Shoot length. (d) Shoot fresh mass. (e) Root length. (f) Root fresh mass. Different symbols above the error bars indicated significant differences among the six groups based on Tukey multiple comparison test (n = 10, p < 0.05). * Indicated p-value < 0.05, ** indicated p-value < 0.01, *** indicated p-value < 0.001.
Ecotoxicological effects of sDNA in lisianthus plants
-
The inhibitory effect of sDNA on lisianthus plants exhibited a clear dose-dependent response (Fig. 2). Exposure to 50 μg/mL of sDNA resulted in significant reductions in plant height, shoot fresh weight, and root fresh weight (p < 0.05). The inhibitory effect intensified at 100 μg/mL and peaked at 200 μg/mL, with no statistically significant difference observed between the 200 and 400 μg/mL treatments (Fig. 2a−e). Exposure to sDNA significantly induced reactive oxygen species (ROS) accumulation and membrane lipid peroxidation in lisianthus plants (Fig. 2f−h). The contents of H2O2 and O2− increased steadily with rising concentrations (Fig. 2f, g). A similar dose-dependent pattern was observed for malondialdehyde (MDA) (Fig. 2h). The relationship between growth inhibition and sDNA concentration was fitted to a dose-response model (Fig. 2i). The data were well described by a four-parameter log-logistic model (R2 = 0.816). The estimated EC50 was 51.0 µg/mL, corresponding to the concentration that produced 50 % of the maximal inhibition.
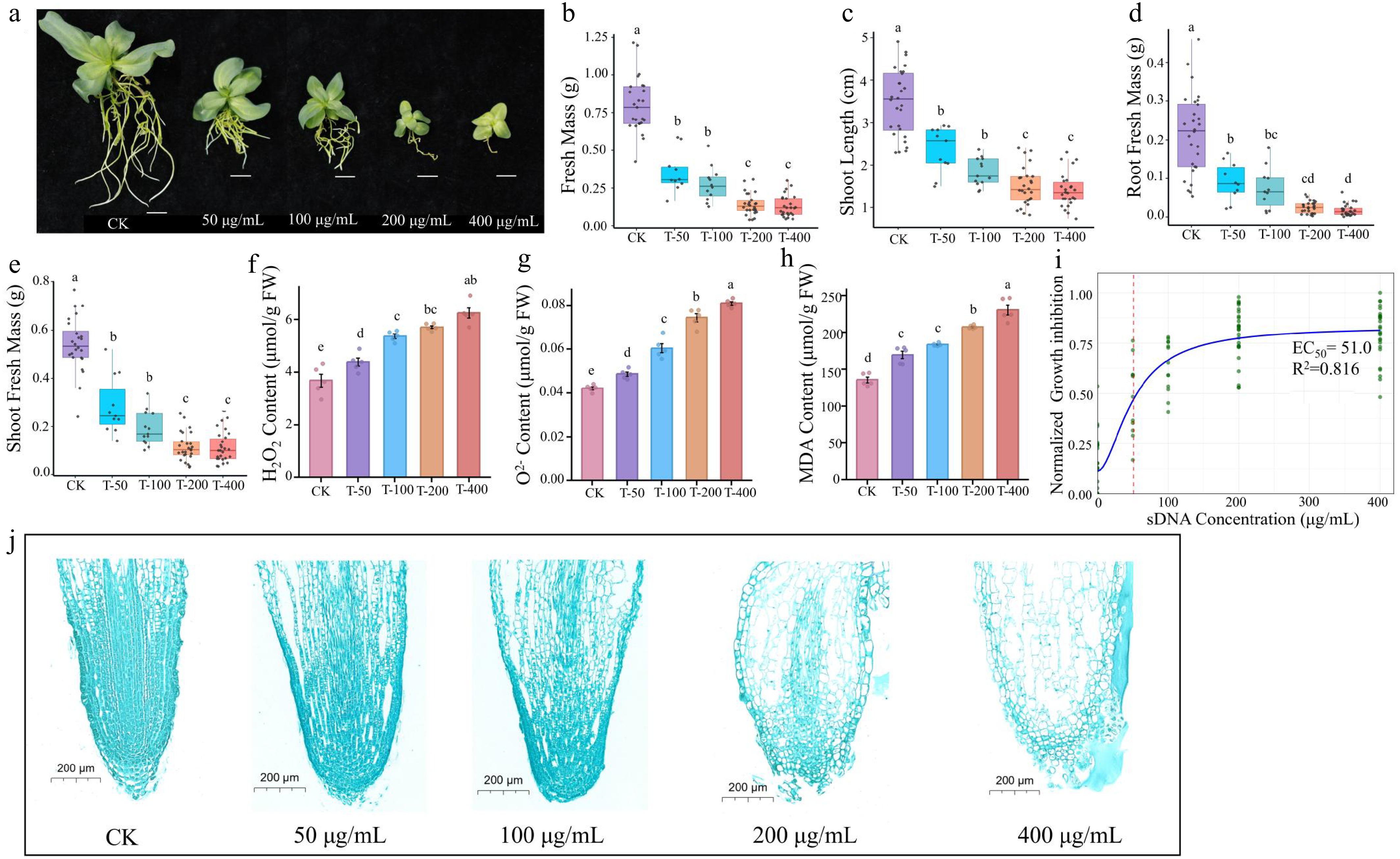
Figure 2.
Ecotoxicity effect of sDNA ranging from 0, 50,100 to 200 and 400 μg/mL on lisianthus. (a) Phenotypic images of seedlings under different sDNA treatments. From left to right: CK, T-50, T-100, T-200, T-400 treatment groups (scale bar = 1 cm). The growth inhibition and physiological responses with increasing concentration included (b) shoot length, (c) shoot fresh biomass, (d) root fresh biomass, (e) total fresh mass, (f) hydrogen peroxide (H2O2) content, (g) superoxide anion (O2−) concentration, and (h) malondialdehyde (MDA) accumulation. Different lowercase letters (a, b, c, d) above bars indicate significant differences among treatment groups (n = 10, p < 0.05, one-way ANOVA followed by Duncan's multiple range test). 'CK' represents the control group without sDNA treatment; 'T-50', 'T-100', 'T-200', and 'T-400' denote treatment groups with final sDNA concentrations of 50, 100, 200, and 400 μg/mL, respectively. (i) Dose-response curve of sDNA on growth inhibition. The y-axis indicates the normalized growth inhibition (0–1 range, processed using min-max normalization with negative values set to 0). Green dots represent normalized experimental data points, and the blue solid line denotes the fitted curve. The red dashed line marks the EC50 position (51.0 µg/mL). Microstructure of root tip stained with safranin and fast green (scale bar = 200 μm) from different exposure concentrations were shown in (j).
Microscopic analysis revealed corresponding concentration-dependent disruptions to root tip cellular architecture (Fig. 2j). Specifically, root tip cells in the control group (CK) exhibited an intact and orderly arrangement structure, with a clear demarcation between the meristematic and elongation zones. As the concentration of sDNA increased, cellular organization of the root tip became progressively disrupted. In the 50 μg/mL treatment group, slightly enlarged intercellular spaces were observed. Cellular deformation and reduced arrangement regularity became apparent at 100 μg/mL, while signs of compromised cell viability (such as blurred cell walls and cytoplasmic condensation) were evident at 200 μg/mL. The highest concentration (400 μg/mL) induced severe structural deterioration, characterized by indistinct cell boundaries, tissue fragmentation, and noticeable cavitation.
Transcriptomic responses of lisianthus root to sDNA exposure
-
Gene Set Enrichment Analysis (GSEA) confirms at the pathway level that lisianthus roots systemically respond to sDNA stress by coordinately activating key biological processes, including immune defense, redox regulation, and metabolic reorganization. GO enrichment analysis revealed broad transcriptional reprogramming upon sDNA treatment, with a general downregulation of genes involved in basic cellular and metabolic processes, alongside the activation of stress-responsive terms (Supplementary Fig. S3, Supplementary Text 2). To systematically decipher the underlying molecular mechanisms, GSEA was performed comparing the T-200 treatment and control (CK). As shown in Fig. 3a, the GSEA plot shows that gene sets such as 'response to chitin', 'lignin catabolic process', and 'hydroquinone: oxygen oxidoreductase activity' were significantly enriched in the T-200 group (NES > 1, FDR < 0.05). The coordinated upregulation of these pathways suggests that lisianthus roots may recognize sDNA as a danger signal analogous to pathogen infection, thereby activating classic immune defense pathways like the chitin response and lignin metabolism. Concurrently, the enrichment of genes related to oxidoreductase activity correlates with the accumulation of reactive oxygen species observed in prior physiological experiments, collectively indicating that sDNA may exacerbate cellular damage by inducing oxidative stress. Furthermore, other significantly enriched gene sets in the plot (e.g., cellular response to fungus, aerobic respiration) further corroborate the immune response and energy metabolic reprogramming triggered by sDNA stress. All significantly enriched gene sets had a Normalized Enrichment Score (NES) greater than 1.5 and a False Discovery Rate (FDR) less than 0.05, confirming the statistical reliability of the results.
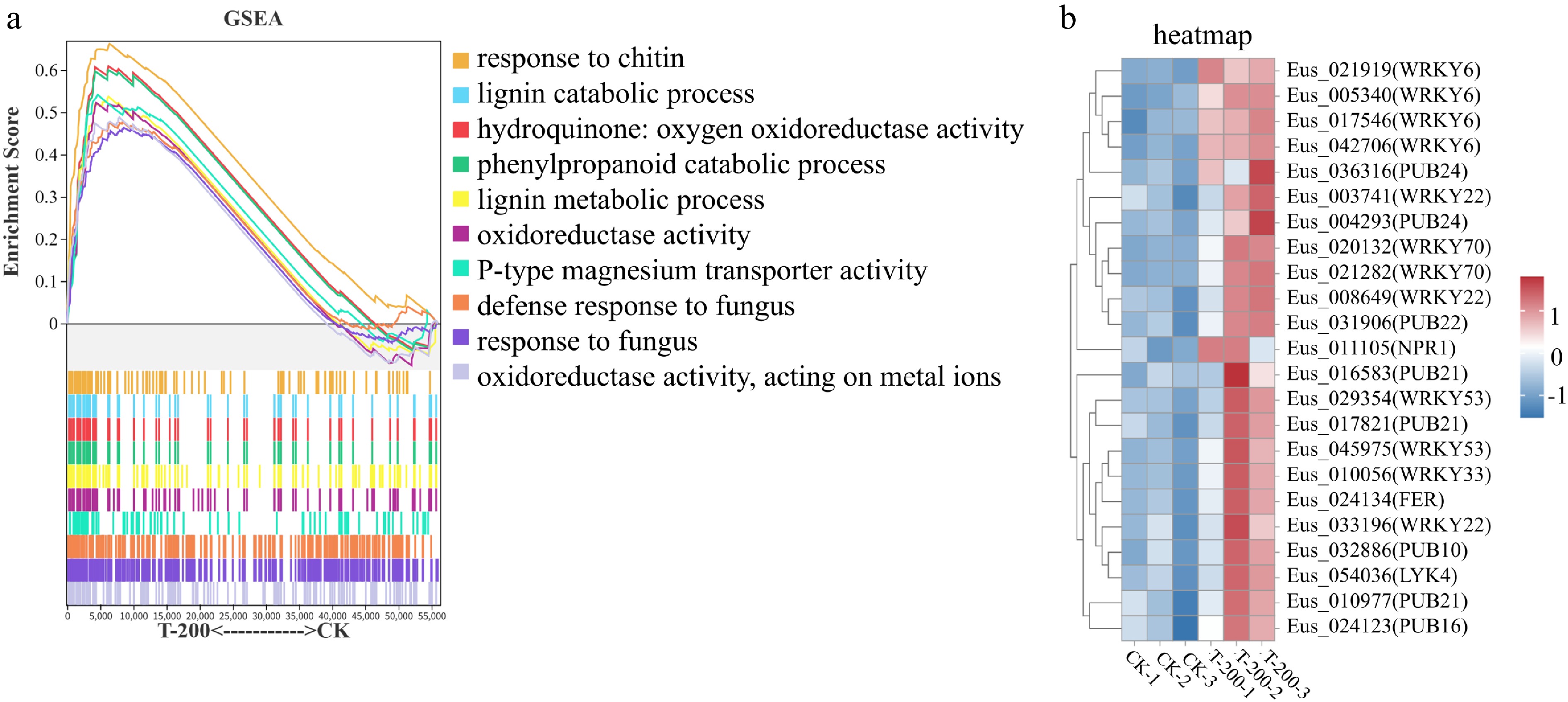

Figure 3.
Gene Set Enrichment Analysis (GSEA) and expression analysis of defense-related genes in response to sDNA treatment. (a) GSEA plot of significantly enriched up-regulated gene sets in the T-200 treatment group compared to the control group. Significantly enriched gene sets were identified with the criteria: |Normalized Enrichment Score (NES)| > 1, number of genes in the set > 50, nominal p-value < 0.05, and FDR < 0.25. The top ten significantly enriched gene sets are shown. The upper part of the plot shows the enrichment score profile for each gene set (y-axis: Enrichment Score, 0−0.6; x-axis: rank in ordered gene list). The lower part presents a heatmap indicating the position of each member gene within the ranked list. Curve colors correspond to the listed gene set names. (b) Heatmap of the expression patterns for key defense-related genes identified from the enriched GSEA pathways. The expression levels of selected WRKY transcription factors and PUB E3 ubiquitin ligase genes are shown, highlighting their concerted upregulation under sDNA treatment, which underpins the activated immune response observed in the GSEA.
Consistent with the pathway-level analysis, the heatmap analysis of specific defense-related genes (Fig. 3b) demonstrated significant upregulation of core immune regulators in the sDNA treatment group. These genes primarily involve the WRKY transcription factor family (such as WRKY6, WRKY22, WRKY33, WRKY53, WRKY70), and U-box E3 ubiquitin ligases (such as PUB21, PUB22, PUB24). It is noteworthy that most of the identified WRKY and PUB genes showed significant upregulation in the T-200 treatment. WRKY transcription factors are key regulatory nodes in plant immune signaling pathways, among which WRKY22 and WRKY33 are typically involved in pathogen defense responses downstream of the MAPK cascade[35−39], WRKY53 and WRKY70 function in SA-mediated systemic acquired resistance[40−43], while WRKY6 is associated with regulating ROS homeostasis and programmed cell death[44−46]. The coordinated upregulation of U-box E3 ubiquitin ligases (e.g., PUB21, PUB22) genes indicate widespread activation of protein ubiquitination in response to sDNA, a process crucial for fine-tuning immune responses[47−53]. This expression profile strongly supports the notion that sDNA mimics biotic stress signals and activates the plant's basal immune system.
Short-term exposure to sDNA increased pathogen infection
-
The plant-pathogen interaction experiment revealed that sDNA pretreatment significantly exacerbated F. oxysporum infection in a concentration-dependent manner. At 30 d post-inoculation, disease symptoms such as chlorosis and wilting were more severe in sDNA-pretreated plants (Fig. 4a). This phenotypic observation was corroborated by a quantitative increase in pathogen load in the rhizosphere, with F. oxysporum conidial abundance reaching 4.30 × 105 CFU/g soil and 7.34 × 105 CFU/g soil in the 100 and 400 μg/mL treatment groups, respectively. These values represent approximately a 2-fold and a 3-fold increase compared to the control (Fig. 4c). Furthermore, the concentration of sDNA in the rhizosphere also increased correspondingly with the pretreatment concentration (Fig. 4b).
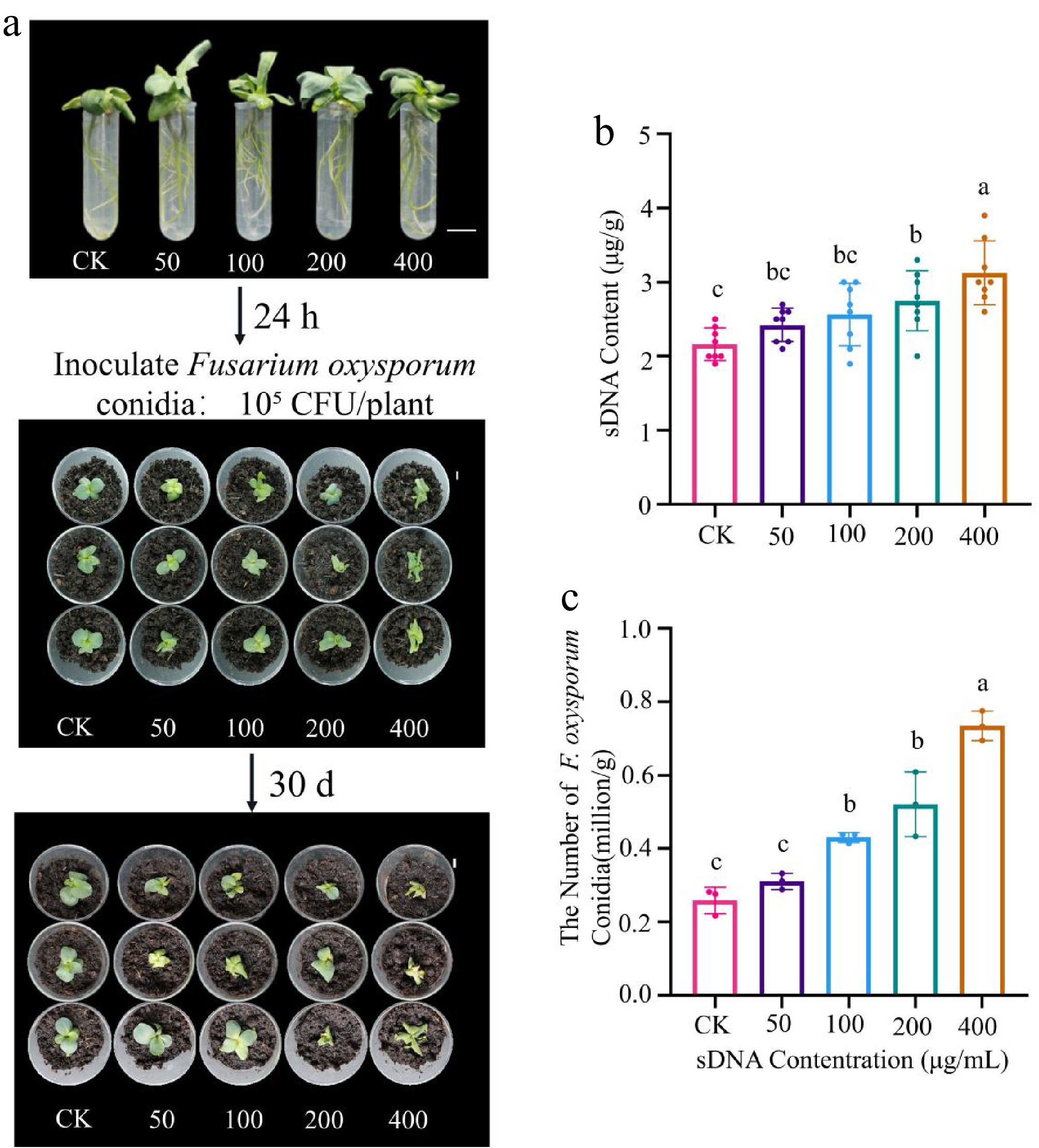
Figure 4.
Effect of sDNA pretreatment on the lisianthus - Fusarium oxysporum interaction. (a) Lisianthus seedlings were treated with sDNA (0, 50, 100, 200, 400 μg/mL); after 24 h, they were transplanted into a peat-based substrate and inoculated with F. oxysporum conidia (105 CFU/plant); the photograph shows representative disease symptoms across treatment groups (CK, T-50, T-100, T-200, T-400) at 30 d post-inoculation. The numbers 0 to 4 above the samples indicate the treatment groups corresponding to sDNA concentrations of 0 (CK), 50, 100, 200, and 400 μg/mL, respectively (scale bar = 1 cm). (b) Detection of extracellular DNA content in rhizosphere soil (n = 3). Different lowercase letters indicate significant differences between groups (p < 0.05, Duncan's test). (c) Quantitative analysis of F. oxysporum conidial abundance in rhizosphere soil (n = 8). The T-100 and T-400 treatments showed a 2-fold and 3-fold increase, respectively, compared to the control group.
-
The species-specific inhibitory effect of extracellular self-DNA has been discovered for over 10 years and provided new perspectives for the explanation of negative PSFs. However, few efforts ever since have been made to comprehensively investigate how extracellular self-DNA contributes to the build-up of negative PSF. A recent study proposed that self-DNA should be considered as the triggering mechanism explaining the species-specific negative PSF, in which the accumulation of self-DNA firstly causes root damage and weakens the plant, subsequently increasing its susceptibility to pathogens[28]. Nevertheless, self-DNA induced defensive responses seems contradictory with this hypothesis as self-DNA can function as Damage-Associated Molecular Pattern (DAMP) to increase its resistance against pathogens (e.g., Hyaloperonospora arabidopsidis, Pseudomonas syringae, Botrytis cinerea, and Pseudomonas syringae), and herbivores such as aphids[18,22,23,26]. This study has provided evidence suggesting that extracellular self-DNA (sDNA) may act as a key driver of negative plant-soil feedback in Eustoma grandiflorum by inducing root tip damage and facilitating pathogen infection.
Chronic toxicity of sDNA is more likely to occur in the field condition
-
There is a growing body of literature positioning self-DNA as a universal biological signal for conspecific inhibition, where the degree of inhibition correlates with taxonomic distance[12,14,54]. Consistent with this self-DNA autotoxicity phenomenon, we observed a strong phylogenetic gradient toxicity of sDNA in growth suppression of lisianthus seedlings. Lisianthus conspecific DNA exerted the most severe inhibition on lisianthus seedlings, followed by exDNA from closely related Gentianaceae species (G. scabra and G. rigescens), while exDNA from distantly related species had negligible effects. sDNA can accumulate gradually in the rhizosphere through root/litter decomposition and persist in the environment[19], thus creating a chronic inhibitory pressure which has been rarely considered in previous studies. For instance, sDNA can accumulate in bulk soil from 2.04 ± 0.11 μg/g after 1 year production to 4.04 ± 0.11 μg/g after 4 years of production in the lisianthus-grown field (Supplementary Fig. S5, data not published). Prior research has predominantly focused on the acute, early responses of self-DNA exposure within hours to a few days, such as calcium influx, MAPK activation, and transient ROS bursts[16,21,26,31,55,56]. In contrast, our work examines the long-term effects of chronic sDNA exposure, a scenario that mimics the sustained stress in agricultural systems and carries direct ecological and agronomic significance. The present results indicate that sustained sDNA exposure significantly suppressed growth, disrupted root architecture, and escalated oxidative stress after one month of exposure. The EC50 value derived from the fitted dose-response curve suggests that sDNA concentration as low as 51 μg/mL (R2 = 0.86) may be sufficient to induce significant growth suppression under long-term exposure. Notably, the observations of progressively disrupted root tip structure suggested that the root tip is particularly vulnerable to sDNA-induced damage. The cumulative damage to the root tip by the prolonged sDNA exposure can impair nutrient uptake and disrupt metabolic processes, thereby adversely affecting overall plant development[25]. This further suggests that in field conditions, where exposure lasts for an entire growing season or longer, the chronic toxicity of sDNA may be exacerbated, with even lower concentrations, potentially causing significant harm.
Long-term sDNA exposure mimics a pathogen attack
-
Results from the transcriptomic analysis revealed that chronic stress of sDNA is not merely an amplified acute toxic response, where sDNA functions predominantly as a damage-associated molecular patterns (DAMPs). In general, research on the effects of sDNA has predominantly focused on aboveground plant organs involving leaves[16,18,22,31,57,58], and fruits[55,56,59] with its role in modulating immunity against pathogens and herbivores. In contrast, investigations into sDNA effects on roots remain limited[21,23,25,60], despite the root system being a primary site for both sDNA accumulation and soil-borne pathogen interaction in continuous cropping systems. In this study, long-term (1 month) exposure to sDNA elicited responses more analogous to pathogen-associated molecular patterns (PAMPs). Despite the sterile culture conditions, long-term exposure to sDNA upregulated a suite of genes classically associated with pathogen defense as indicated by GSEA results, including 'response to chitin', 'response to fungus', and 'defense response to fungus'. Specifically, we observed a concerted upregulation of WRKY transcription factors (e.g., WRKY22, WRKY33, WRKY70), and U-box E3 ubiquitin ligases (e.g., PUB21, PUB22). These genes are pivotal in mediating PAMP-triggered immunity (PTI) and systemic acquired resistance[35−46].
Plants detect invasion and damage through immune receptors that recognize both 'non-self' signals from colonizers and 'modified-self' signals from pathological alterations[61]. Conserved epitopes, or 'molecular patterns' (MPs), released into the apoplast during interactions with colonizers or via mechanical damage, are detected by cell-surface pattern recognition receptors (PRRs) to activate pattern-triggered immunity (PTI). DAMPs are endogenous molecules produced inside the cell and released to the apoplast under pathogen or insect attack, while PAMPS refer to the exogenous ones that are structurally conserved and unique to microbes such as fungal chitin or bacterial lipopolysaccharides, peptidoglycans, quorum-sensing factors, and flagellin[62−64]. In most cases, injury to the outer layers of an organism (e.g., the epidermis of leaves, and roots of plants) inevitably leads to infection[16]. Thus, mechanical damage and subsequent pathogen invasion are temporally linked events in natural environments. Here in this study, the findings suggest that lisianthus responds to long-term sDNA stress in a manner similar to fungal infections (PAMPs) rather than mere mechanical damage (DAMPs). We propose that this response may not be a case of mistaken identity, but rather a possible adaptation to a predictable sequence of events in nature. The persistent presence of sDNA could serve as a potential predictive danger signal, indicating a high probability of imminent microbial attack. Therefore, the immune system may have evolved to use this 'damage signature' to pre-emptively activate broad-spectrum, PAMP-triggered immunity, which could lead to a state of heightened alert against likely subsequent microbial attack. This is mechanistically plausible, as PRRs are known to mediate responses to both exogenous PAMPs and endogenous DAMPs[62,63,65,66], and recent perspectives in immunology suggest that effective immune activation often requires a concerted 'danger code' comprising both microbe-derived patterns (PAMPs) and host-derived damage signals (DAMPs)[67−69]. Consequently, resources are continuously diverted into defense preparedness at the expense of growth and metabolism, creating a growth-defense trade-off[70−72]. Consistent with this paradigm, chronic sDNA exposure triggered a broad downregulation of genes governing primary metabolism (e.g., 'cellular process' and 'metabolic process'). In this case, the chronic exposure of sDNA-induced prolonged defensive responses may force a prolonged reallocation of energy from growth to defense, resulting in the observed growth inhibition of lisianthus plants.
sDNA-induced root damage can outweigh the pathogen defensive responses
-
Results from our pathogen challenge experiment provided critical insights into the paradoxical relationship between sDNA-induced defense activation and actual disease susceptibility. Contrary to the typical priming effect observed in other studies typically investigated on above-ground organs such as leaves and fruit[16,18,22,26,55,56], 24 h root pretreatment with sDNA failed to induce effective resistance in lisianthus. Instead, it significantly increased host susceptibility to F. oxysporum, elevating the rhizosphere pathogen load by 2- to 3-fold. This supports the hypothesis proposed by Idbella et al.[28] that sDNA may first trigger the inhibitory effect, and subsequently contribute to the changes in the soil microbiome.
Extracellular fragmented DNA (exDNA) in the plant rhizosphere can originate from two fundamentally different processes with opposing consequences for plant health. On one hand, active and localized secretion of exDNA (size ranging from 150 bp to 5 kb) from root tips is a documented component of innate immunity, serving as root extracellular traps (RETs) that directly neutralize pathogens[73−77]. This represents an active, targeted deployment of exDNA as a defense effector. In monoculture systems such as lisianthus cultivation, however, the continuous accumulation of exDNA from litter and root degradation results in a passive, uncontrolled release of fragmented self-DNA at high concentrations into the rhizosphere. This pool of self-DNA can adsorb to root surfaces[23], reducing membrane permeability and impairing water and nutrient uptake, a phenomenon consistent with the dose-dependent disruption of root tip structure observed in our study (Fig. 2). Furthermore, the persistent DAMP/PAMP-like signal from accumulated sDNA may overstimulate and exhaust the plant's defense capacity, leading to the broad suppression of primary metabolism and growth. Hence, the theoretical protective benefits are likely outweighed by the actual physical and physiological damage, such as compromised nutrient uptake, diminished structural integrity, and depleted energy reserves, which could collectively increase overall vulnerability to disease. Consequently, when a pathogen such as F. oxysporum arrives, the plant is in a pre-weakened state that fails to support effective defense. Collectively, the results of this study provide the first evidence that chronic exposure to sDNA facilitates pathogen colonization, which in turn amplifies root damage and further sDNA release. This self-reinforcing loop (Fig. 5) simultaneously drives the accumulation of phytotoxic sDNA and the build-up of soil-borne pathogens, ultimately accelerating the negative plant–soil feedback observed in continuous cropping systems.
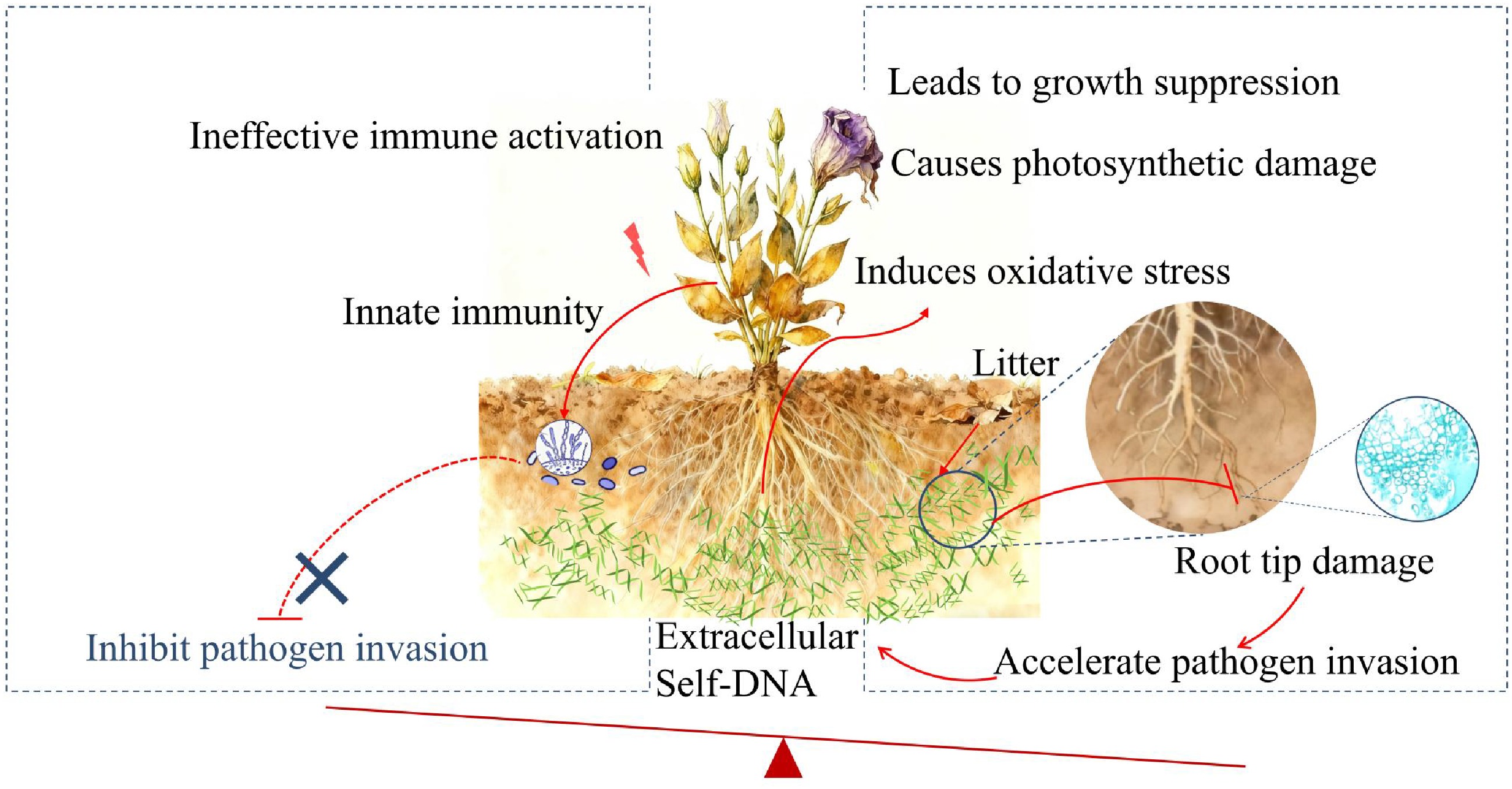
Figure 5.
Schematic diagram depicting the proposed mechanism of sDNA-mediated negative plant-soil feedback in lisianthus. Self-DNA released from accumulated root litter in the soil acts as a danger signal, activating the innate immune system and potentially enhancing pathogen resistance. Simultaneously, it induces oxidative stress and inhibits basic metabolic processes like photosynthesis, leading to root damage and reduced plant fitness. Crucially, the beneficial effects of immune activation are likely overridden by the growth-inhibitory and damaging effects, ultimately predisposing the plant to accelerate pathogen invasion and establishing a vicious cycle characteristic of soil sickness.
-
This study offers insights into the apparent paradox of sDNA-induced defense and heightened disease susceptibility, revealing a self-reinforcing loop in which sDNA accumulation and pathogen proliferation reciprocally accelerates, potentially underpinning the persistent soil sickness (negative PSF) in continuous cropping systems. Under the chronic exposure conditions of monocultures, sDNA appears to function as a dual-edged agent that directly impairs root architecture and primary metabolism while concurrently activating a maladaptive, PAMP-like defense program. Crucially, this sustained response likely compromises root integrity and exhausts plant resources, ultimately predisposing plants to severe infection by soil-borne pathogens such as F. oxysporum. Based on these findings, we propose that extracellular sDNA could be monitored as a novel biochemical indicator of soil sickness, enabling early detection and targeted management before visible crop decline. However, the mechanisms underlying self-DNA inhibition are still unclear regarding the specific DNA receptors and molecular processes[78]. The interplay between rhizosphere sDNA accumulation, plant root response, and the soil microbiome pathogenicity remains largely unexplored under ecologically relevant scenarios. By unraveling the mechanistic basis of its action and exploring targeted interventions, it may be possible to advance toward more resilient and sustainable horticultural systems that break the cycle of continuous cropping obstacle.
This study was financially supported by Provincial General Program of Department of Science and Technology of Yunnan Province (Grant No. 202501AT070091), and Innovative Talent-Led Demonstration Project for High-Quality Development of the Flower Industry by Yunnan Provincial Party Committee Organization Department.
-
The authors confirm their contributions to the paper as follows: study conception and design: Zhang P, He D; experiment organization and execution, data collection and analysis: He D, Li H, Yang Y; writing − draft manuscript preparation: He D, Zhang P; writing − revision: Zhang P, Li S. All authors reviewed the results and approved the final version of manuscript.
-
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
-
The authors declare that they have no conflict of interest.
- The supplementary files: Supplementary materials to this study.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
He D, Li H, Yang Y, Li S, Zhang P. 2026. Self-DNA as a chronic stress signal drives soil sickness in lisianthus by disrupting root integrity and facilitating pathogen invasion. Ornamental Plant Research 6: e008 doi: 10.48130/opr-0026-0005
Self-DNA as a chronic stress signal drives soil sickness in lisianthus by disrupting root integrity and facilitating pathogen invasion
- Received: 05 January 2026
- Revised: 25 January 2026
- Accepted: 03 February 2026
- Published online: 27 February 2026
Abstract: Continuous cropping obstacle (CCO) severely limits the sustainable production of high-value cut flowers like lisianthus (Eustoma grandiflorum), primarily driven by species-specific negative plant-soil feedback (PSF). Although autotoxic compounds are implicated, the mechanisms behind this specificity remain elusive. This study investigated the role of extracellular self-DNA (sDNA) as a species-specific autotoxic stress in mediating negative PSF under a prolonged exposure scenario. This study first demonstrated that the growth-inhibitory effect of fragmented extracellular DNA is phylogenetically dependent, with conspecific DNA causing the strongest suppression, followed by DNA from closely related Gentiana species. Chronic exposure to lisianthus sDNA induced a concentration-dependent ecotoxic response, including significant reactive oxygen species (ROS) accumulation, disruption of root cellular architecture, and overall growth inhibition, with a half-maximal effective concentration (EC50) of 51.0 μg/mL (R2 = 0.816). Transcriptomic profiling further indicated that long-term sDNA exposure activated defense pathways akin to pathogen-associated molecular pattern (PAMP) responses (e.g., 'response to chitin', 'response to fungus', and 'defense response to fungus'), while concurrently repressing genes involved in primary metabolism. To investigate the paradox of sDNA acting as both a defense elicitor and a growth suppressor, a pathogen challenge assay was conducted. Pretreatment with sDNA for 24 h significantly enhanced the susceptibility of lisianthus to the soil-borne pathogen Fusarium oxysporum, suggesting that the sDNA-induced physiological damage may outweigh its potential defensive benefits. The present findings propose that sDNA drives negative PSF through a self-reinforcing cycle, in which monoculture-induced sDNA accumulation directly impairs root health and plant vigor, which in turn predisposes the plant to pathogen infection, further exacerbating root decay and sDNA release. This mechanistic cycle highlights a key pathway underpinning the persistent soil sickness in the continuous cropping system of lisianthus.













